Abstract
Cryptocarya densiflora Blume (Lauraceae) is an evergreen tree widely distributed throughout the hills and mountain forests up to 1500 m in Malaysia and Indonesia. The plant has been reported to contain phenanthroindolizidine-type of alkaloids. In the present work, a new phenanthroindolizidine alkaloid named (R)-13aα-densiindolizidine, was isolated from the dichloromethane (DCM) extract of the leaves. The structure of the alkaloid was established based on 1D and 2D nuclear magnetic resonance (NMR) and liquid chromatography mass spectrometry-ion trap-time of flight (LCMS-IT-TOF) analysis. (R)-13aα-densiindolizidine displayed binding interactions with crucial amino acid residues in the active sites of
Introduction
Cryptocarya genus comprises around 200 to 250 plant species distributed across South China and India to North Australia, Madagascar, and South America, in which 17 species can be found in Peninsular Malaysia, including Cryptocarya densiflora Blume. 1 The species C densiflora is a medium sized tree, up to 20 m in height and 135 cm in girth. 2 Although several Cryptocarya species are known for traditional medicine such as for women after childbirth and treating diarrhea, 3 however, none is known for C densiflora. Phytochemically, Cryptocarya species have been reported to contain flavonoids, 4 pyrones, 5 lignans, 6 chalcones, 7 and alkaloids. 8 Several phenanthroindolizidine alkaloids have been isolated from Crytocarya species including three from C densiflora from our previous work.9,10
Phenanthroindolizidine alkaloids are known to exhibit interesting pharmacological properties. 11 Apart from Cryptocarya genus, these alkaloids are present in a few species of Asclepiadaceae, Acanthaceae, and Moraceae families. 12 Since the first isolation of tylophorine in 1935, 13 phenanthroindolizidine alkaloids have attracted much attention because they exhibited antitumor and anticancer activity, as well as inhibitors of protein synthesis. 14 More than 100 natural phenanthroindolizidines have been reported to date. Antofine has been extensively studied towards the development of potent anticancer agent due to its excellent cytotoxic activity.14,15 In addition, several phenanthroindolizidine alkaloids such as tylophorine, tylophorinine and 7-methoxycryptopleurine have been reported to display potency against SARS-CoV-2 and transmissible gastroenteritis virus (TGEV) in vitro at low nanomolar level with high oral availability in rats. These suggested they could be used as potential therapeutic agents for coronavirus infections.16–18
This paper describes the isolation and characterization of a new phenanthroindolizidine alkaloid, (R)-13aα-densiindolizidine, from the DCM extract of the leaves of C densiflora. Inspired by the pharmacological properties of previously reported phenanthroindolizidines, we evaluated the binding interactions of (R)-13aα-densiindolizidine with important amino acid residues in the active sites of SARS-CoV-2 Mpro and RdRp in silico, as well its drug-likeness properties.
Results and Discussion
The new alkaloid (Figure 1) was obtained as an optically active dark brownish amorphous solid with

Molecular structure of (R)-13aα-densiindolizidine.

Key HMBC and COSY correlations of (R)-13aα-densiindolizidine.

NOESY correlations of (R)-13aα-densiindolizidine.
1H and 13C NMR Spectroscopic Assignments of (R)-13aα-Densiindolizidine in CDCl3.

Abbreviations: s, singlet; d, doublet; brs, broad singlet; brd, broad doublet; brt, broad triplet; m, multiplet.
Lipinski's Rule
The physicochemical properties for (R)-13aα-densiindolizidine were predicted using SwissADME. A total of eight descriptors were taken into consideration (Table 2). Based on the result, (R)-13aα-densiindolizidine can be considered as a potential lead compound that obeys Lipinski's rule with good pharmacokinetic properties. (R)-13aα-densiindolizidine passed ADMETsar criteria for druggability as indicated in Table 3.
Physicochemical Properties Prediction for (R)-13aα-Densiindolizidine.
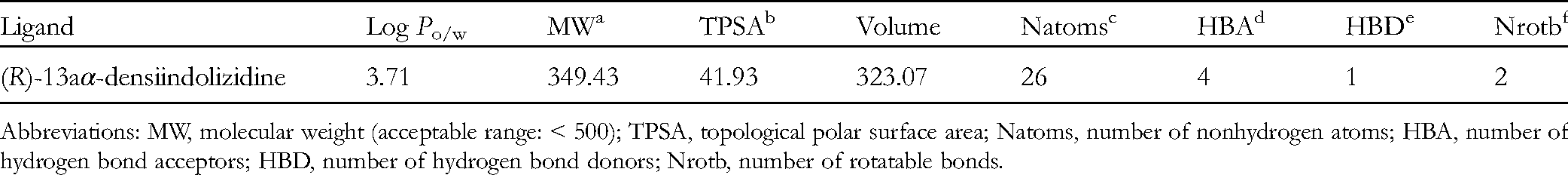

Abbreviations: MW, molecular weight (acceptable range: < 500); TPSA, topological polar surface area; Natoms, number of nonhydrogen atoms; HBA, number of hydrogen bond acceptors; HBD, number of hydrogen bond donors; Nrotb, number of rotatable bonds.
Predicted ADMET Properties for (R)-13aα-Densiindolizidine.

Abbreviations: CYP2D6, cytochrome P450 2D6; HIA, human intestinal absorption; BBB, bloodbrain barrier permeability; H-HT, human hepatotoxicity; DILI, drug induced liver injury.
Molecular Docking Studies
The alkaloid (R)-13aα-densiindolizidine was subjected to docking studies against potential targets, SARS-CoV-2 protease Mpro (PDB ID: 6LU7), and RdRp (PDB ID: 7D4F). Based on the docking results, this alkaloid was able to fit in the substrate-binding pocket and binds to one of catalytic triad residue, Cys145, with docking interaction energy of −33.5 kcal/mol (Table 4). (R)-13aα-densiindolizidine was observed to form interactions with substrate binding sites (Glu166 and Thr190). The binding mode of (R)-13aα-densiindolizidine within the SARS-CoV-2 Mpro cavity is shown in Figure 4 whereby four hydrogen bonds were observed. Among these hydrogen bonds, a conventional hydrogen bond (O-H---N-H) between 6-hydroxyl group of the (R)-13aα-densiindolizidine with amino group (NH) from the residue Glu166 was the shortest distance (2.02 Å). The 3-methoxy group of (R)-13aα-densiindolizidine displayed the ability to form two carbon hydrogen bonds (C = O---H-C-O) with the oxygen from carboxylic acid group Thr190 residue (2.69 and 3.09 Å) and carbon-hydrogen bond interaction with amino acid residue Pro168 (2.73 Å) to strengthen the ligand-enzyme complex. The complex also participated in hydrophobic interaction between the sulfur of the Cys145 thiol group (SH) with one of the hydrogen at the E ring of (R)-13aα-densiindolizidine. This suggests that (R)-13aα-densiindolizidine is able to fit in the substrate-binding pocket and the interactions with the catalytic active residues are expected to enhance inhibition activity of the enzyme and prevent replication process of SARS-CoV-2.
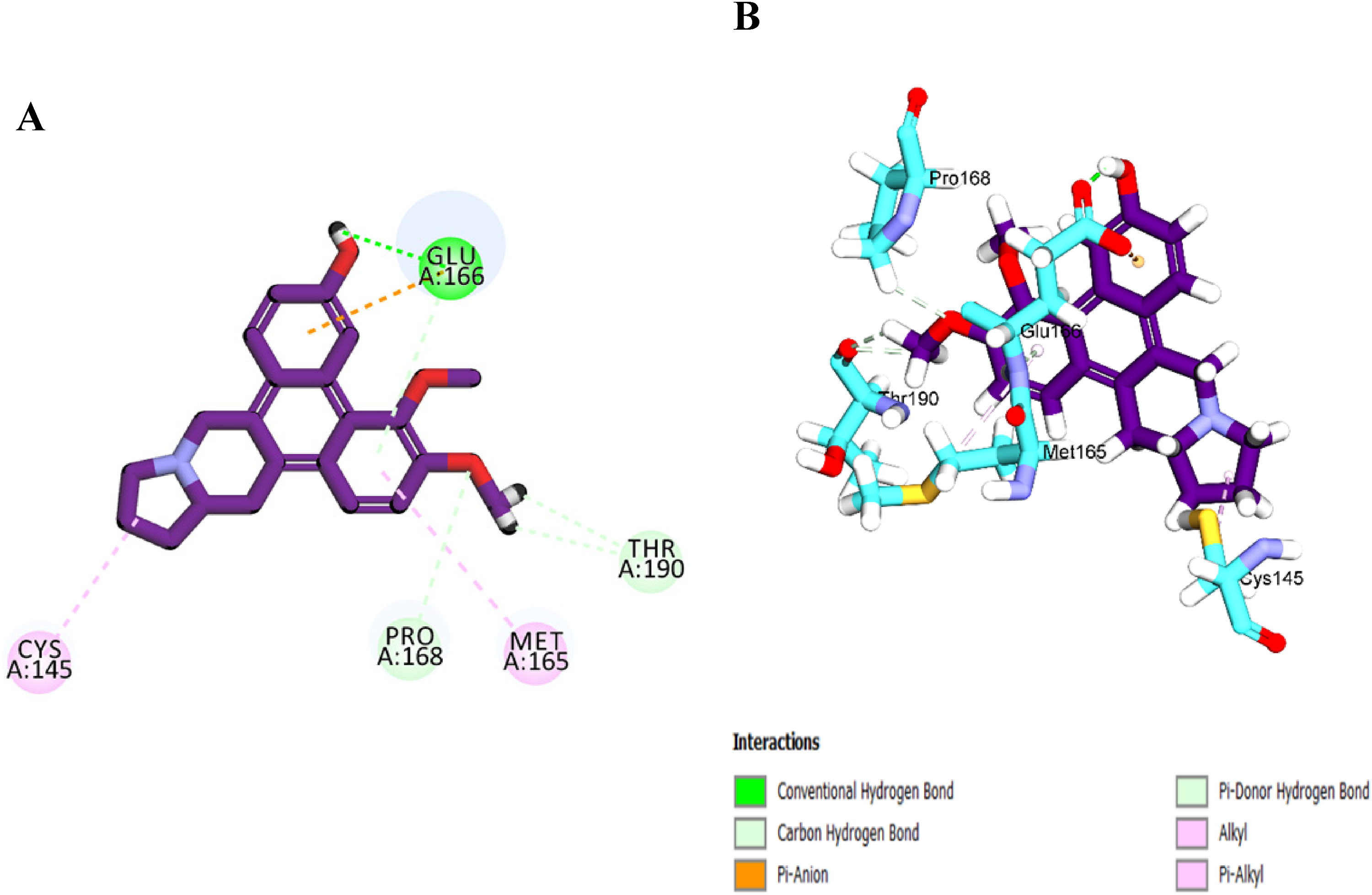
(A) 2D interaction of (R)-13aα-densiindolizidine with SARS-CoV-2 Mpro protein; (B) 3D interaction of (R)-13aα-densiindolizidine with SARS-CoV-2 Mpro protein.
Docking Interaction of (R)-13aα-Densiindolizidine Against SARS-CoV-2 Mpro and RdRp.

In the development of the drug targets for SARS-CoV-2, RdRp protein play an important role due to the nonsimilar enzyme in host cell homologs, fewer off-targets effects against human host proteins, and development of selective SARS-CoV-2 RdRp inhibitors. 23 This enzyme has two binding sites including RNA template strand and RNA primer strand (located near to RdRp catalytic site). Yin et al 24 reported that ligand suramin can inhibit the SARS-CoV-2 RdRp at these two binding sites. Residues Asn497, Lys500, Arg569, Gln573, Asn496, Lys577, Gly590, Leu576, Ala580, Ala685, Tyr689, and Leu758 were reported to interact with suramin in the RNA template strand binding site while residues Arg555, Lys551, Arg553, Arg836, Ala550, Lys551, Arg865, His439, Ile548, Ser549, Ala840, Ser861, and Leu862 are crucial amino acids for RNA primer strand. 24 Thus, any close interaction with these residues will disrupt the function of the RdRp protease. The result showed that the alkaloid (R)-13aα-densiindolizidine fits well in the RNA template and RNA primer strands binding sites with docking energy value of −29.9 and −27.1 kcal/mol, respectively.
In the case of RNA template strand binding site shown in Figure 5, alkaloid (R)-13aα-densiindolizidine displayed the ability to interact with active residues Lys500, Asn496, Leu576, Ala580, Lys577, and Ala685. Interaction with these residues is expected to block the binding site of RNA template strand, thus inhibiting RdRp activity. The ligand-enzyme complex was stabilized by a conventional hydrogen bond between oxygen from 6-hydroxyl group of (R)-13aα-densiindolizidine and amino group (NH) of the residue Lys500 (2.05 Å). Alkaloid (R)-13aα-densiindolizidine can be potentially stabilized by carbon-hydrogen bond interactions (C = O-----H-C) between oxygen from carbonyl group of Leu576 (2.39 and 2.68 Å) and hydrogen of the C-11 methylene of (R)-13aα-densiindolizidine. In addition, the hydrogen of the C-9 of (R)-13aα-densiindolizidine has the potential to form carbon-hydrogen bond interaction (C = O---H-R) with oxygen from the carbonyl functional group on residue Asn496. The alkaloid was further stabilized by hydrophobic interaction with residues Ala580 and Lys577 and π-alkyl interaction with residue Ala685.

(A) 2D interaction of (R)-13aα-densiindolizidine at RNA template binding site of SARS-CoV-2 RdRp protein; (B) 3D interaction of (R)-13aα-densiindolizidine at RNA template binding site SARS-CoV-2 RdRp protein.
The alkaloid (R)-13aα-densiindolizidine was also predicted to be able to fit in the RNA primer strand binding site (Figure 6) through several hydrogen bonds and electrostatic interactions. This alkaloid binds to the enzyme through four conventional hydrogen bond interactions which include (1) oxygen of 3- and 4-methoxy groups (R-O---H-N) with the hydrogen of amino group Lys545 residue (2.05 Å); (2) oxygen of 4-methoxy group (N-H---O-R) with hydrogen from amino group (NH) of Arg555 residue; (3) 6-hydroxyl group (N-H---O-H) with hydrogen of amino group from the Arg555. This ligand-protease complex was further stabilized through electrostatic π-cation interaction of rings A, B, and C of the alkaloid with Arg555. Overall, (R)-13aα-densiindolizidine was observed to display potential binding interactions with the catalytic triad of SARS-CoV-2 Mpro and can potentially act as inhibitor of SARS-CoV-2 RdRp by blocking both crucial binding sites, the template and primer strands of RdRp.

(A) 2D interaction of (R)-13aα-densiindolizidine at RNA primer strand of SARS-CoV-2 RdRp protein; (B) 3D interaction of (R)-13aα-densiindolizidine at RNA primer strand SARS-CoV-2 RdRp protein.
Experimental
General
Analytical and preparative thin-layer chromatography (TLC) was carried out on Merck 60 F254 silica gel plates (absorbent thickness: 0.25 and 0.50 mm, respectively). Column chromatography (CC) was performed using silica gel (Merck 230-400 mesh, ASTM). Ultraviolet (UV) spectra were recorded using a Shimadzu UV-250 UV–Visible Spectrophotometer. IR spectra were recorded using a Perkin-Elmer Spectrum 400 FT-IR Spectrometer. NMR spectra were acquired in deuterated chloroform (CDCl3) (Merck) with tetramethylsilane (TMS) as the internal standard using the BRUKER Avance III 400 MHz NMR and BRUKER Avance III 600 MHz NMR spectrometers. Chemical shifts are given in the δ scale. LCMS-IT-TOF spectra were obtained using an Agilent Technologies 6530 AccurateMass Q-TOF LC/MS system. A Jasco P1020 polarimeter was used to measure optical rotation. All solvents were of analytical grade and were distilled prior to use.
Plant Materials
The leaves of C densiflora was collected from Hutan Simpan Tembat, Ulu Terengganu, Terengganu, Malaysia and were authenticated by a certified botanist, Teo Leong Eng, Department of Chemistry, Faculty of Science, University of Malaya. A voucher specimen (KL 5211) has been deposited with the University of Malaya herbarium.
Extraction and Isolation
Plant extraction was carried out by cold percolation. Dried grounded leaves of C densiflora (2.5 kg) was first defatted with hexane (15 L) for three days at room temperature. The resulting slurry was filtered, and the residual plant material was moistened with 25% ammonia solution (1 L) and left for two hours to aggregate the nitrogen-containing compounds in the plant. The basified residual plant material was then successively reextracted with DCM (15 L, 3×). The DCM extract was repeatedly extracted with a solution of 5% hydrochloric acid (0.5 L, 1×) until it gave a negative result for Mayer's test. It was next basified with 25% ammonia solution to about pH 11 and reextracted with DCM (3 L, 1×) to yield 13 g of extract. The DCM crude extract was subjected to exhaustive CC over silica gel and eluted with DCM which was gradually enriched with methanol (MeOH). The ratio of the solvent between DCM and MeOH were (100:0; 99:1; 98:2; 97:3; 96:4; 95:5; 94:6; 93:7; 92:8; 90:10; 85:15; 80:20, and 50:50). Fractions were collected every 100 mL and each fraction was tested with aluminum TLC plate for their alkaloids. The alkaloid spots were first detected by UV light (254 and 366 nm) and confirmed by spraying with Dragendorff's reagent. Fraction having spots with the same Rf values and stains were combined and treated as a group. The combined groups were purified with CC and preparative TLC. Isolation and purification (13 g) of alkaloid yielded 20 fractions. Further purification of fraction F7 by a preparative TLC using DCM: MeOH with 97:3; v/v, saturated with ammonium hydroxide (NH4OH); gave (R)-13aα-densiindolizidine.
(R)-13aα-densiindolizidine: dark brownish amorphous solid;
Molecular Descriptors Calculation
The molecular properties of (R)-13aα-densiindolizidine were predicted through SwissADME server (http://www.swissadme.ch/index.php). The descriptors include volume, log-P, topological polar surface area, number of OH or NH, molecular weight, number of rotatable bonds, number of atoms, number of O or N, drug-likeness including G protein-coupled receptors ligand, nuclear receptor ligand, a kinase inhibitor, ion channel modulator and the number of Lipinski's rule violations.
ADMET Prediction
The alkaloid (R)-13aα-densiindolizidine was subjected through ADMETlab (https://admet.scbdd.com/home/index/) for ADMET prediction. The prediction provides information about human intestinal absorption, atom-based logP (Alog P98), aqueous solubility, hepatotoxicity, blood–brain barrier, plasma protein binding, polar surface area, and cytochrome P450 2D6 (CYP2D6) descriptors.
Molecular Docking
In this work, SARS-CoV-2 RNA-dependent RNA-polymerase (RdRp) (PDB ID: 7D4F, 2.57 Å) 25 and main protease (Mpro) crystal structure complexed with inhibitor NS3 (PDB ID: 6LU7, 2.16 Å) 26 were retrieved from the Protein Data Bank (http://www.rcsb.org/). 27 Molecular docking was performed using CDOCKER module in Discovery Studio® (Accelrys). The co-crystallized ligands were first removed and then redocked with the protein binding site to validate the molecular docking protocol. Ligand binding pose were ranked based on their CDOCKER energy values.
Conclusions
Isolation, identification, and characterization of the compound isolated from the leaves of C densiflora yielded a new phenanthroindolizidine alkaloid named (R)-13aα-densiindolizidine. Since phenanthroindolizidine alkaloids are well known for their interesting pharmacological activities, thus the new alkaloid was subjected to in silico ADMET analyses. In silico findings suggest that the alkaloid could be a potent therapeutic lead as it fulfills Lipinski's rule criteria as well as the ability to properly bind and interact well with amino acid residues in the active site of SARS-CoV-2 Mpro and RdRp.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X221114227 - Supplemental material for (R)-13aα-Densiindolizidine, A New Phenanthroindolizidine Alkaloid From Cryptocarya densiflora Blume (Lauraceae) and Molecular Docking Against SARS-CoV-2
Supplemental material, sj-docx-1-npx-10.1177_1934578X221114227 for (R)-13aα-Densiindolizidine, A New Phenanthroindolizidine Alkaloid From Cryptocarya densiflora Blume (Lauraceae) and Molecular Docking Against SARS-CoV-2 by Wan N Nazneem Wan Othman, Fatimah Salim, Nor N Abdullah, Syahrul I Abu Bakar, Khalijah Awang, Lalith Jayasinghe and Nor H Ismail in Natural Product Communications
Footnotes
Acknowledgements
The authors would like to acknowledge the Department of Chemistry, Faculty of Science, the University of Malaya for the facilities and support of this work.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Univesriti Teknologi MARA, UiTM Dynamic Research (Grant No: 600-RMC/DINAMIK-POSTDOC 5/3-006/2020).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
