Abstract
Since the COVID-19 pandemic in 2020, many reports have highlighted several potential anti-SARS-CoV-2 drug candidates, including phenolic compounds. Therefore, this study aimed to evaluate the anti-SARS-CoV-2 activity of nine common phenolic compounds found in plants using the in vitro cellular infection model. The anti-SARS-CoV-2 activity of curcumin, quercetin, gallic acid, catechin, rutin, kaempferol, naringenin, coumaric acid and caffeic acid were evaluated on SARS-CoV-2-infected Vero E6 cells by using a cytopathic effect (CPE)-based assay. The anti-SARS-CoV-2 activity in human lung cells, A549 expressing human ACE2 and TMPRSS2, was evaluated by the RT-qPCR technique. S1-ACE2 interaction and 3CL protease activity assays were also performed for the potent compound. Of the nine phenolic compounds, only curcumin inhibited the SARS-CoV-2 induced CPE activity (EC50 of 13.63 µM) in Vero E6 cells, but with a low selective index (SI) value. Interestingly, curcumin exhibited potent anti-SARS-CoV-2 activity in A549 cells with an EC50 of 4.57 µM and an SI value of 7.96. S1-ACE2 interaction and 3CL protease inhibitory activities of curcumin were also observed. In conclusion, curcumin showed a moderate in vitro anti-SARS-CoV-2 activity. The true potential of curcumin as an anti-SARS-CoV-2 candidate could be further evaluated in a COVID-19 animal model.
Introduction
Since the start of the COVID-19 pandemic in 2020, many studies have been initiated to find a cure for the disease including the development of vaccines and antiviral drugs targeting the essential components for virus entry and replication. To date, mRNA-based, inactivated, and viral vector-based COVID-19 vaccines have been approved by regulatory authorities and used widely to reduce the disease severity.1–3 However, as the COVID-19 virus, SARS-CoV-2, frequently mutates its spike protein (S protein), the effectiveness of current vaccines was affected.4,5 Due to the waning of spike-antibodies in vaccine recipients over time, 6 vaccine booster shots are needed to maintain the optimum antibody levels against different SARS-CoV-2 variants. 7
COVID-19 drug treatments, especially the antivirals, have showed promising benefits in preventing severe disease8–10 and were reported to remain effective against different SARS-CoV-2 variants. 11 After remdesivir, two oral anti-SARS-CoV-2 drugs have been authorized for emergency use by the US Food and Drug Administration (FDA).8,10 These drugs are Paxlovid, a combination of nirmatrelvir tablet (3CLpro main protease or MPro inhibitor) with ritonavir tablet (to slow down nirmatrelvir breakdown), and molnupiravir, a small-molecule ribonucleoside prodrug of N-hydroxycytidine that induced RNA mutagenesis by viral RNA dependent RNA polymerase. 10
Apart from the FDA approved antiviral drugs, several other potential antiviral candidates have been highlighted by many studies.12–15 These candidates include the phenolic compounds commonly found in plants and dietary sources, which have been previously studied for their potential in inhibiting the essential viral infection16–19 and replication components, such as 3CLpro main protease,12,20–23 human type-2 transmembrane serine (TMPRSS2), angiotensin-converting enzyme-2 (ACE2) 24 and viral spike S1-protein. 25 In detail, several in silico studies have uncovered the potential of rutin in binding with the SARS-CoV-2 3CLpro main protease and helicase and the host ACE2 receptor. At higher concentration (EC50 > 100 µM), rutin was reported to inhibit several viruses such as influenza H5N1, hepatitis C, enterovirus and dengue virus. 17 Quercetin was also highlighted by many in silico studies for its potential in inhibiting the SARS-CoV-2 3CLpro main protease. In addition, the antiviral property of quercetin against live virus was previously shown by its antiviral activity against the SARS-CoV-1 entry into Vero E6 cells. 18 Naringenin was also considered a broad antiviral compound as many studies have reported its antiviral activities against dengue, hepatitis C, hepatitis B, zika, chikungunya, herpes simplex 1 and 2, rotavirus, yellow fever, and human immunodeficiency viruses. Furthermore, in silico studies on naringenin have reported its multi-protein target potential activities against the SARS-CoV-2 spike S1-protein, TMPRSS2, 3CLpro main protease and the host ACE2 protein receptor. 16 An anti-inflammatory compound, curcumin, was also a compound of interest during the COVID-19 pandemic. Curcumin was found to be effective in inhibiting the wild-type, D614G and Delta SARS-CoV-2 infections in Vero E6 cells.19,26 Other phenolic compounds, such as gallic acid, caffeic acid, catechin, kaempferol and coumaric acid, were also reported as potential anti-SARS-CoV-2 candidates by several in silico and enzyme target-based assay studies.20,21,27
Based on the available evidence, some of these compounds need further validation for their potential antiviral activity against live SARS-CoV-2 virus. Although some of these compounds have been previously demonstrated for their antiviral activities against live SARS-CoV-2 virus, the antiviral screenings were limited to either certain virus variants or host cells. Therefore, the aim of our study was to screen and validate the antiviral activity of nine phenolic compounds (curcumin, quercetin, gallic acid, rutin, caffeic acid, catechin, kaempferol, naringenin, and coumaric acid), together with the well-known antivirals as positive controls (andrographolide, remdesivir, and chloroquine),14,28 against the Malaysian clinical SARS-CoV-2 virus isolates by using cytopathic effect (CPE) based assay. The potential phenolic compound candidate was further screened for its anti-SARS-CoV-2 activity against alpha, delta and omicron SARS-CoV-2 variants. In addition, the potential phenolic compound candidate was also validated against the SARS-CoV-2 infection in human lung cells and studied for its potential targets against the essential viral infection components by performing the SARS-CoV-2 spike (S1)-human ACE2 interaction and 3CL protease or main protease (Mpro) assays.
Materials and Methods
Drugs and Compounds Preparation
All drugs and compounds (Curcumin, 95% [Alfa Aesar, Great Britain], quercetin, 95% [Sigma, USA], gallic acid, 98% [Sigma, USA], rutin hydrate, 94% [Sigma, USA], caffeic acid, 98% [Sigma, USA], andrographolide, 98% [Sigma, USA], catechin 98% [Santa Cruz Biotechnology, USA], kaempferol, 98% [ChemFaces, China], naringenin, 98% [ChemFaces, China], coumaric acid, 98% [Sigma, USA] and remdesivir [Cayman Chemical, USA]) were dissolved in 100% DMSO, except chloroquine diphosphate salt (Sigma, USA), which was dissolved in water. The stocks were prepared at 20 mM concentration and kept at −20 °C prior to use.
Cell Culture
African green monkey-derived epithelial kidney cells (Vero E6; ATCC-CRL-1586) and human lung carcinoma expressing human angiotensin I-converting enzyme-2 (hACE2) and transmembrane protease serine 2 (TMPRSS2) (A549-hACE2-TMPRSS2) were purchased from the American Type Cell Culture Collection (ATCC, USA) and InvivoGen (InvivoGen, USA), respectively. Vero cells were cultured in Dulbecco's modified Eagle's medium (DMEM) (Gibco, USA) supplemented with 10% fetal bovine serum (FBS) (Gibco, USA) and 100 units/mL penicillin–streptomycin (Gibco, USA). A549-hACE2-TMPRSS2 was cultured in DMEM (Gibco, USA) supplemented with 10% FBS (Gibco, USA), 100 µg/mL normocin (Invivogen, USA), 100 units/mL penicillin–streptomycin (Gibco, USA), 0.5 µg/mL puromycin (Invivogen, USA) and 300 µg/mL hygromycin (Invivogen, USA). All cell lines were grown at 37 °C in a 5% CO2 incubator.
Virus Propagation
The clinical SARS-CoV-2 variants, Wuhan-Wild-Type (WT), Alpha, Delta and Omicron were isolated from COVID-19 patients’ diagnostic nasal swab samples by the Virology Unit, Institute for Medical Research, National Institutes of Health, Malaysia. The viral isolates were propagated in Vero E6 cells grown in 2% FBS supplemented DMEM media at 37 °C in a 5% CO2 incubator at a biosafety level 3 (BSL-3) lab facility, Institute for Medical Research, National Institutes of Health, Malaysia. The virus culture supernatant was collected once the viral-induced cytopathic effect (CPE) was observed and kept at −80 °C prior to use. Virus isolated from human diagnostic samples is not considered as a human biological specimen by the Medical Research and Ethics Committee (MREC), Ministry of Health, Malaysia. Therefore, informed consent was not obtained from the patients.
Virus Titer Determination
The virus titer was measured by the 50% tissue culture infective dose (TCID50) method. Vero E6 (2 × 104 cells/well) and A549-hACE2-TMPRSS2 (1 × 104 cells/well) cells were seeded in 96 well culture plates and grown overnight. The cells were exposed to serially diluted (10-fold) viruses and incubated for 72 hours at 37 °C in a 5% CO2 incubator. The CPE was observed microscopically. The viral-induced CPE was measured by ATP-based assay (ViralToxGlo®, Promega, USA). The luminescent signal intensity was measured by a multimode microplate reader (GloMax® Explorer, Promega, USA). The TCID50 values were determined by dose–response curve analysis using GraphPad Prism software (Version 7.0).
Cell Viability Assay
Vero E6 (2 × 104 cells/well) and A549-hACE2-TMPRSS2 (1 × 104 cells/well) cells were seeded in 96 well plates and incubated overnight in 2% FBS supplemented DMEM media at 37 °C in a 5% CO2 incubator. Then, the confluent cells were exposed to a serially diluted concentration of compounds or drugs and incubated for 72 hours at 37 °C in a 5% CO2 incubator. The healthy cell control was cells exposed to media containing 0.5% DMSO and the lysis cell control was cells exposed to media containing 0.05% Triton X 100. The cell viability was measured by either MTT assay or ATP-based assay. For the MTT assay, the MTT reagent solution was added and after 4 hours of incubation, the culture supernatant was discarded. The purple formazan was solubilized by addition of 100% DMSO. The intensity of the solubilized formazan was measured by a multimode microplate reader (FLUOstar Omega, BMG LABTECH, Germany) at 570 nm wavelength. 29 For the ATP-based cell viability assay, the ATP detection reagent (CellTiter-Glo®, Promega, USA) was added and incubated at room temperature for 10 minutes. The luminescent signal intensity was measured by a multimode microplate reader (GloMax® Explorer, Promega, USA). The half-maximal cytotoxic concentration value (CC50) was determined by dose–response curve analysis using GraphPad Prism software (Version 7.0).
Antiviral Screening in Monkey Kidney Cells by Cytopathic Effect (CPE) Inhibition Assay
Vero E6 cells were seeded at 2 × 104 cells/well in a 96 well plate and incubated overnight in 2% FBS supplemented DMEM media at 37 °C in a 5% CO2 incubator. Then, either compounds or drugs, which were serially diluted with 2% FBS supplemented DMEM media, were added to the confluent cells. In the BSL3 lab, SARS-CoV-2 virus isolate was diluted with 2% FBS supplemented DMEM media and added to the confluent cells in a 96 well plate (25TCID50 per well) containing serially diluted concentration of either compounds or drugs. The healthy cell control was cells exposed to media containing 0.5% DMSO only, and the CPE control was cells exposed to the virus only. The test plate was incubated for 72 hours at 37 °C in a 5% CO2 incubator. The viral cytopathic effect (CPE) was measured by ATP-based assay (ViralToxGlo®, Promega, USA). The luminescent signal intensity was measured by a multimode microplate reader (GloMax® Explorer, Promega, USA). The half-maximal effective concentration value (EC50) was determined by dose–response curve analysis using GraphPad Prism software (Version 7.0).
Antiviral Screening in Human Lung Cells by RT-qPCR Technique
A549 cells expressing human ACE2 and TMPRSS2 were seeded at 1 × 104 cells/well in a 96 well plate and incubated overnight in 2% FBS supplemented DMEM media at 37 °C in a 5% CO2 incubator. Then, either compounds or drugs, which were serially diluted with 2% FBS supplemented DMEM media, were added to the confluent cells. In the BSL3 lab, SARS-CoV-2 virus isolate was diluted with 2% FBS supplemented DMEM media and added to the confluent cells in a 96 well plate (0.03 MOI) containing serially diluted concentration of either compounds or drugs. The healthy cell control was cells exposed to media containing 0.5% DMSO only, and the infection control was cells exposed to the virus only. The test plate was incubated for 72 hours at 37 °C in a 5% CO2 incubator. The viral RNA from the culture supernatant was automatically extracted by KingFischer Duo Prime System (ThermoFischer Scientific, USA) using a MagMaxTM Viral/Pathogen Nucleic Acid Isolation Kit (Applied BiosystemsTM, USA). The viral RNA copy numbers were quantified by using the Applied Biosystems 7500 thermocycler (Applied Biosystems, USA). Briefly, the primer set targeting the N gene region 30 was used for the viral RNA copy number quantification. The RT-qPCR condition was as recommended by the Quantitect SYBR Green RT-PCR Kit (Qiagen, Germany). Synthetic SARS-CoV-2 RNA (VR-3276SD) (ATCC, USA) with known copy number was used as a standard for this assay.
Spike S1 (SARS-CoV-2)-ACE2 Inhibitor Assay
The effect of curcumin on the spike S1-ACE2 interaction was determined by using the Spike S1 (SARS-CoV-2): ACE2 inhibitor screening colorimetric assay kit (BPS Bioscience, Canada). Briefly, curcumin (5, 10, and 20 µM) was added to the spike S1, Fc fusion, avi-tag (2 µg/mL) pre-coated 96 well plated together with the reaction mixture containing 1X immunobuffer and ACE2-Biotin (0.4 ng/µL). The positive control was the reaction mixture without the curcumin. The reaction mixture was incubated for 10 minutes at room temperature prior to 1 hour incubation with streptavidin-horse radish peroxidase (HRP). The signal intensity was measured at 450 nm by using a microplate reader ((FLUOstar Omega, BMG LABTECH, Germany) after 5 minutes incubation with HRP substrate and stopped by 1 M sulfuric acid.
3CL Protease (SARS-CoV-2) Assay
The effect of curcumin on 3CL protease (Mpro) activity was determined by using the 3CL protease, untagged (SARS-CoV-2) assay kit (BPS Bioscience, Canada). Briefly, the curcumin (5, 10, and 20 µM) was incubated with the reaction mixture containing assay buffer and 3CL protease (0.3 ng/µL). The positive control was the reaction mixture without the curcumin. The inhibitor control was GC376 (100 µM). The reaction mixture was preincubated for 20 minutes at room temperature with slow shaking. The reaction was started by adding 3CL protease substrate (40 µM) and incubated at room temperature for 4 hours with slow shaking. The fluorescence intensity was measured at an excitation wavelength of 350 nm and an emission wavelength of 460 nm by using a microplate reader ((FLUOstar Omega, BMG LABTECH, Germany).
Data Analysis
The dose–response curve was generated by nonlinear regression (curve fit) XY analysis using (inhibitor) versus normalized response (variable slope) algorithm (GraphPad Prism software, version 7.0). The CC50 and EC50 values were used to calculate the selectivity index value (CC50/EC50).
Results
Cytotoxic Activity of the Phenolic Compounds
Curcumin and gallic acid showed cytotoxic activities in Vero-E6 cells with CC50 values of 44.83 µM and 200 µM, respectively. Besides curcumin and gallic acid, none of the tested phenolic compounds (quercetin, rutin, caffeic acid, catechin, naringenin, kaempferol, and coumaric acid) showed any cytotoxic activity, even at the highest tested concentration of 200 µM (CC50 > 200 µM) (Figures 1 and 2). The standard antiviral compound, andrographolide, showed cytotoxic activity with a CC50 value of 40.19 µM (Figure 1). The standard antiviral drug, chloroquine, showed cytotoxic activity with a CC50 value of 54.08 µM, while remdesivir did not show any cytotoxic activity (CC50 > 200 µM) (Figure 2). Only the non-cytotoxic concentrations of the phenolic compounds, standard antiviral compound, and drugs were chosen for the SARS-CoV-2-induced CPE-based assay.
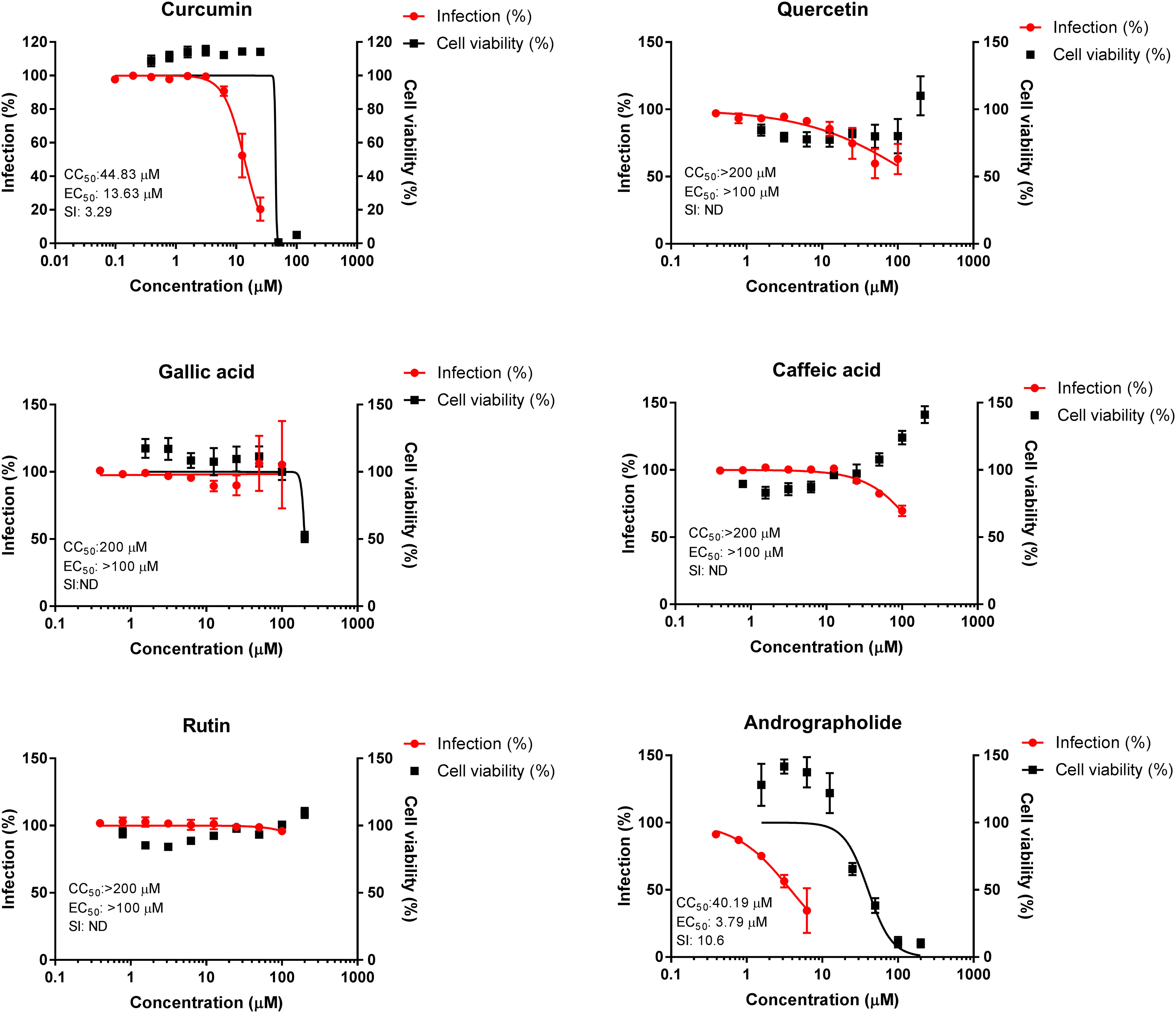
Anti-SARS-CoV-2 and cytotoxic activities of curcumin, quercetin, gallic acid, caffeic acid, rutin, and a standard compound, andrographolide (anti-SARS-CoV-2 plant derived compound). Antiviral and cytotoxic activities were determined by CPE inhibition assay and MTT assay, respectively after 72 hours’ exposure in a 37 °C CO2 incubator. The data were normalized to percentage of infection (left Y-axes: red) and percentage of cell viability (right Y-axes: black). Data points are mean values and bars are the standard error of mean of experiments performed in 2 to 3 independent tests.

Anti-SARS-CoV-2 and cytotoxic activities of coumaric acid, catechin, kaempferol, naringenin, and standard antiviral drugs, remdesivir and chloroquine. Antiviral and cytotoxic activities were determined by CPE inhibition assay and MTT assay, respectively after 72 hours’ exposure in a 37 °C CO2 incubator. The data were normalized to percentage of infection (left Y-axes: red) and percentage of cell viability (right Y-axes: black). Data points are mean values and bars are the standard error of mean of experiments performed in 2 to 3 independent tests.
Effect of the Phenolic Compounds on SARS-CoV-2 Induced CPE
Antiviral screening of the selected phenolic compounds against WT Wuhan SARS-CoV-2 showed that only curcumin inhibited the viral-induced CPE with an EC50 value of 13.63 µM (SI = 3.29) (Figures 1 and 2). The positive control compounds and drugs, such as andrographolide, chloroquine and remdesivir, inhibited the viral-induced CPE with EC50 values of less than 10 µM with a high selectivity index (SI ≥ 10) (Figures 1 and 2).
Antiviral Activities of Curcumin Against SARS-CoV-2 Variants of Concern (VOCs)
Curcumin also exhibited antiviral activities against SARS-CoV-2 VOCs (Figure 3). Exposure of curcumin to alpha, delta and omicron variants of SARS-CoV-2 resulted in a reduction of CPE with EC50 values of ∼25 µM (extrapolated data: 25.25 µM) (SI: 1.78), 20.64 µM (SI: 2.17), and ∼25 µM (extrapolated data: 25.36 µM) (SI: 1.77), respectively (Figure 3).

Anti-SARS-CoV-2 activities of curcumin against SARS-CoV-2 VOCs. Antiviral activities were determined by CPE inhibition assay after 72 hours’ exposure in a 37 °C CO2 incubator. The data were normalized to percentage of infection (Y-axes: red). Data points are mean values and bars are the standard error of mean of experiments performed in 3 independent tests.
Antiviral Activities of Curcumin Against SARS-CoV-2 Infection in Human Lung Cell Lines
To validate the in vitro anti-SARS-CoV-2 activity of curcumin in human lung cells, curcumin was further tested in WT Wuhan SARS-CoV-2-infected A549 lung carcinoma cells expressing the human ACE2 and TMPRSS2 genes (Figure 4). The curcumin treatment showed anti-SARS-CoV-2 activity with an EC50 value of 4.57 µM and a SI value of 7.96 (Figure 4).

Anti-SARS-CoV-2 activity of curcumin against SARS-CoV-2 infection in a human lung cell line (A549-hACE2-TMPRSS2 cells). Antiviral and cytotoxic activities of curcumin were determined by RT-qPCR and ATP-based assay, respectively, after 72 hours’ exposure in a 37 °C CO2 incubator. The data were normalized to percentage of infection (Y-axes: red) and percentage of cell viability (right Y-axes: black). Data points are mean values and bars are the standard error of mean of experiments performed in 3 independent tests.
The Effect of Curcumin on SARS-CoV-2 Spike S1-ACE2 Interaction and 3CL Protease Activity
To confirm the potential targets of curcumin in preventing SARS-CoV-2 infection of the cells, the direct inhibitory effects of curcumin on the SARS-CoV-2 spike S1-ACE2 interaction and 3CL protease activity were measured. The effect of curcumin on the S1-ACE2 interaction was observed at 10 µM and 20 µM, which reduced the interaction activity to 85% and 67%, respectively, as compared to 100% activity in the positive control (no treatment) (Figure 5). An inhibitory effect of curcumin on 3CL protease activity was also observed at 10 µM and 20 µM, which reduced the activity to 81% and 60%, respectively, as compared to the total activity in the positive control without treatment (Figure 5). There was no reduction in activity for both S1-ACE2 interaction and 3CL protease by 5 µM of curcumin (Figure 5).

Inhibitory effect of curcumin on SARS-CoV-2 spike S1-ACE2 interaction and 3CL protease activity. Curcumin (5, 10, and 20 µM) was tested against the SARS-CoV-2 spike S1-ACE2 interaction (A) and 3CL protease (B) activities at room temperature for 1 and 4 hours, respectively. The data were normalized to percentage of activity. The bar charts are mean values with the standard deviation of each test in triplicates.
Discussion
The screening of anti-SARS-CoV-2 activity in our study has showed that out of the nine phenolic compounds tested, only curcumin inhibited the viral-induced CPE in Vero E6 cells against WT, alpha, delta and omicron variants. However, the SI was considered low (SI < 5) due to the cytotoxic effect of curcumin on the uninfected Vero E6 cells. Interestingly, the anti-SARS-CoV-2 activity of curcumin was found to be more potent in the human A549-hACE2-TMPRSS2 lung cells (EC50: 4.57 µM, SI: 7.96). In addition, our study has also highlighted the dual action potential of curcumin whereby the S1-ACE2 interaction and 3CL protease activities were inhibited by curcumin in a dose dependent manner.
The S protein of SARS-CoV-2 was one of the proposed target sites for curcumin by a computational study. 26 This could explain the less potent antiviral activities against alpha, delta and omicron variant infections in vero cells (EC50 values 20.64-25.36 µM), which could be due to viral S protein mutations that could possibly affect the binding affinity. A study conducted by Goc et al has highlighted the capability of curcumin to affect the binding and cellular entry of pseudo-typed SARS-CoV-2 virions by disrupting the interaction of spike protein with human angiotensin-converting enzyme-2 (hACE2) receptor, as well as inhibiting the essential proteases (TMPRSS2 and cathepsin L) for entry mechanisms. 31 The multiple targets of curcumin as demonstrated by previous studies could also explain the more potent anti-SARS-CoV-2 activity of this compound in the human lung cell line used in our study, whereby the cells overexpressed human ACE2 receptor and TMPRSS2. Besides that, an in silico study conducted by Teli et al showed that curcumin was able to covalently inhibit 3CL and Mpro with good pharmacokinetic parameters and passes Lipinski's rule of five. 32
Our in vitro findings were further supported by a study by Bormann et al, 26 which showed the anti-SARS-CoV-2 activity of curcumin in infected Vero-E6 cells, with an EC50 value of 7.9 µg/mL (21.44 µM). Furthermore, another study conducted by Marin-Palma et al 19 has demonstrated that curcumin reduced the viral titer of the D614G mutation SARS-CoV-2 strain at different points of treatments (pre-infection, post-infection treatments and co-treatment) with EC50 values ranging from 3.57 to 6.03 µg/mL (equivalent to 9.69-16.37 µM) and a SI value of less than 5. In contrast to our findings, the anti-SARS-CoV-2 activity of curcumin was more potent against the Delta variant with EC50 values ranging from 1.14 to 1.66 µg/mL (equivalent to 3.09-4.51 µM) and SI values of 9.94 to 14.5. 19 The differences in these data could be due to the different approaches used in each study. Despite its potential as an antiviral agent, there are concerns on the poor bioavailability of curcumin in humans (Cmax ≤ 150 ng/mL). Therefore, the use of curcumin as a single compound treatment is not advisable and further development on curcumin formulations and drug delivery systems to enhance its pharmacokinetic profile is warranted. 33 Indeed, several clinical trials of nano-curcumin (SinaCurcumin® Exir Nano, SinaCurcumin® Soft gel and curcumin-piperine combination) have positively affected the COVID-19 patient symptoms through its anti-inflammatory effects. 34
Besides curcumin, other phenolic compounds, such as quercetin, rutin, kaempferol, naringenin, catechin, gallic acid, and caffeic acid, have been reported to inhibit the activity of SARS-CoV-2 3CLpro main protease.12,20–23 However, none of them inhibited the SARS-CoV-2-induced CPE in our study. This could be due to the unfavorable cellular bioavailability property, as previously showed in the study of rutin. 23 Another possible reason is the lack of viral-induced CPE inhibitory activity at the concentration of 100 µM and below as set in our study. Quercetin has been showed to significantly inhibit SARS-CoV-2 viral replication at high concentrations of 400 to 500 µM, 22 while gallic acid has been reported to exert anti-SARS-CoV-2 activity with an IC50 of 108 µg/mL, which is equivalent to 635 µM. 35 In the case of naringenin, a study by Abdallah et al have showed that naringenin exerted anti-SARS-CoV-2 activity with the EC50 value of 28.35 µg/mL. 12 This concentration (equivalent to 104 µM) is two times higher than the reported human serum concentration (Cmax about 50 µM) of healthy volunteers given 600 mg of naringenin. 36
As reported by other studies,14,28,37 in vitro anti-SARS-CoV-2 activity was similarly showed by the positive control antivirals (andrographolide, chloroquine and remdesivir) in our study. Andrographolide is an active compound found in the medicinal plant Andrographis paniculata (Burm. G.) Wall. ex Nees. The compound was previously reported to exert an anti-SARS-CoV-2 activity in Vero-E6 cells and human lung cells, Calu-3, with EC50 values of 6.58 and 1.68 µM, respectively.14,38 Initially, chloroquine and remdesivir were reported to inhibit SARS-CoV-2 virus replication in Vero E6 cells with EC50 values of 1.13 µM and 0.77 µM, respectively. However, chloroquine was later found not to inhibit the SARS-CoV-2 virus in human lung cells, Calu-3. 39 This is due to the lack of the endosomal-pH-dependent cysteine protease cathepsin L viral entry requirement in Calu-3 cells, which is a target for chloroquine. 39 Hence, this phenomenon emphasized the importance of validating the anti-SARS-CoV-2 activity of potential antiviral candidates in human lung cells, although Vero E6 cells were conveniently used for primary screening due to their measurable cytopathic effect.13,15 Therefore, our study has also demonstrated that curcumin anti-SARS-CoV-2 activity was more potent (lower EC50 value) in human lung cells, A549 expressing human ACE2 and TMPRSS2 infected with the WT Wuhan SARS-CoV-2 variant. Despite the different techniques or assays used, the higher EC50 values for some compounds in Vero E6 cells could also be due to the increase of compound efflux activity in the cells. Inhibition of compound efflux activity by the presence of P-glycoprotein efflux inhibitor in the antiviral assay has been shown to improve the efficacy of anti-SARS-CoV-2 drugs in Vero E6 cells. 11
Conclusions
Curcumin exhibited a moderate anti-SARS-CoV-2 activity in monkey kidney (Vero E6) and human lung (A549 expressing human ACE2 and TMPRSS2) cells. The dual targets of curcumin on S1-ACE2 interaction and 3CL protease activities were confirmed in our study. Further pre-clinical evaluation of curcumin anti-SARS-CoV-2 activity in a COVID-19 animal model is warranted. However, to make curcumin a clinically relevant candidate, the issue of bioavailability needs to be addressed.
Footnotes
Acknowledgments
The authors would like to thank the Director General of Health Malaysia for his permission to publish this article. They also would like to thank the staff of Bioassay and Virology Units for their supports and contributions in this study.
Additional Information
The data presented in this study are available on request from the corresponding author.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Ethical Approval is not applicable for this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by Ministry of Health Malaysia (NMRR-20-874-54715).
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Informed Consent
There are no human subjects in this article and informed consent is not applicable.
