Abstract
Introduction
Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2)-associated symptoms and infectious respiratory illness are designated as coronavirus disease 19 (COVID-19). According to the World Health Organization (WHO) statistics, as of February 25, 2021, there have been nearly 112 million confirmed cases of COVID-19 and 2.49 million deaths worldwide. This has brought severe menace to the lives and health of people worldwide and has become a vital global public health emergency.1,2 SARS-CoV-2 is a novel coronavirus with respiratory droplet transmission as the main route of transmission. It can also be transmitted through contact. It has the features of fast and widespread transmission, high infectivity, and common susceptibility to all kinds of individuals. Patients with mild COVID-19 have symptoms such as fever, weakness, and tussiculation. Patients with severe infection may have symptoms such as respiratory distress syndrome, dyspnea, or septic shock. At present, there is no specific medicine for COVID-19 treatment.3–5 To facilitate early detection, reporting, isolation, and treatment of the disease, and to increase the cure rate and reduce the mortality rate, the General Office of the National Health and Wellness Commission of the People’s Republic of China and the Office of the State Administration of Traditional Chinese Medicine of the People's Republic of China recommended the use of traditional Chinese medicine (TCM) in the treatment of novel coronavirus pneumonia (Trial Version 7-9),6–8 based on the diagnosis and treatment experience of COVID-19 and the clinical treatment guidelines of the WHO and other countries. COVID-19 falls within the field of “epidemic” disease in TCM. Based on the disease situation, local climate features, and different body constitutions, Huo-Xiang-Zheng-Qi prescription (HXZQP) (including HXZQ capsules, HXZQ water, HXZQ dropping pills, and HXZQ oral liquid) can be used to cure COVID-19. Clinical observations have confirmed that based on syndrome differentiation, HXZQ, combined with western medicine, can alleviate the clinical symptoms, prevent the transition from mild to severe, and improve the clinical cure rate in patients with COVID-19. 9 A large sample prospective clinical study on HXZQ oral liquid and Jinhaojiere granules for the prophylactic intervention of COVID-19 in the community showed that the combined use of these 2 drugs effectively improved the protection rate of community residents against cold and other respiratory diseases. 10 Huang 11 used homemade HXZQ medicated soap for protection against COVID-19.
HXZQP originated from the “Prescription of Peaceful Benevolent Dispensary” during the Song Dynasty. It has a history of approximately 1000 years and has been widely used in the treatment of pathogenic dampness in febrile diseases. According to the record in the “Chinese Pharmacopoeia,” the raw materials of HXZQP are Atractylodis Rhizoma (Cangzhu), Citri Reticulatae Pericarpium (Chenpi), Magnoliae Officinalis Cortex (made from ginger) (Houpu), Angelicae Dahuricae Radix (Baizhi), Poria (Fuling), Arecae Pericarpium (Dafupi), Pinelliae Rhizoma (Banxia), Glycyrrhizae Radix et Rhizoma (Gancao), Pogostemonis Herba (Huoxiang), and Perillae Folium (Zisu). HXZQP functions to “jiebiaohuashi” and “liqihezhong,” it can be used for symptoms include headache and dizziness, vomiting and diarrhea, chest and diaphragm tightness, abdominal distension and pain, and cold of the gastrointestinal tract. HXZQP showed positive effects against COVID-19; however, the mechanism of action remains unclear.
As a new field of pharmacological research, network pharmacology emphasizes the concept of a “multi-component, multi-target treatment network” and aligns with the principle of TCM. This study offers a new approach to research on the multi-target mechanism of a TCM to prevent COVID-19. This study used network pharmacology to analyze the mechanism of HXZQP in the prevention and treatment of COVID-19. First, the effective components of HXZQP were screened from the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP) database, and effective targets were identified. Second, we used network pharmacology to analyze the compound–target interaction and found the compound–target network and the compound–target–COVID-19 network. Finally, we used bioinformatics analysis to expound on HXZQP's multitarget multi-pathway mechanism of action in the prevention and treatment of COVID-19.
Results
Prediction of Active Ingredients and Targets of HXZQP
Using the TCMSP database, the targets of the active ingredients of HXZQP were identified. The UniProt database was used to obtain all the official gene symbols corresponding to the target information. Finally, we obtained HXZQP with 151 active ingredients that met the 2 screening conditions of oral bioavailability (OB) value ≥30% and drug-likeness (DL) value ≥0.18. The results show that Huoxiang has 8 active ingredients and 157 targets, Zisu has 13 active ingredients and 90 targets, Cangzhu has 4 active ingredients and 52 targets, Chenpi has 5 active ingredients and 61 targets, Houpu has 2 active ingredients and 23 targets, Baizhi has 20 active ingredients and 49 targets, Fuling has 6 active ingredients and 18 targets, Dafupi has 2 active ingredients and 10 targets, Banxia has 11 active ingredients and 88 targets, and Gancao has 92 active ingredients and 210 targets. In total, 250 targets were identified.
Mining Disease Targets of COVID-19
Using GeneCards, Online Mendelian Inheritance in Man (OMIM), Pharmacogenomics Knowledge Base (PharmGKB), Therapeutic Target Database (TTD), and DrugBank databases, with “2019-nCoV,” “COVID-19,” and “SARS-CoV-2” as keywords, COVID-19 human disease–related genes were identified. A total of 749 related genes were screened, including 637 GeneCard-related genes, 1 OMIM-related gene, 15 PharmGKB-related genes, 128 TTD-related genes, and 1 DrugBank-related gene. A Venn diagram of disease-related genes is shown in Figure 1.

Venn diagram of COVID-19 disease-related genes merged in each database.
Drug-Active Ingredient–Target Interaction Network Construction of HXZQP
For the 749 COVID-19-related targets, we took the intersection with 250 HXZQP action targets to obtain 67 intersection targets (Figure 2). The drug-active ingredient–target interaction network of HXZQP for the prevention of COVID-19 is shown in Figure 3.

Venn diagram of the intersection of HXZQP and COVID-19 disease-related genes.

The drug-active ingredient–target interaction network of HXZQP for the prevention and treatment of COVID-19.
Protein–Protein Interaction (PPI) Network Construction and Topological Analysis of HXZQP for the Prevention and Treatment of COVID-19
The 67 intersection genes of HXZQP and COVID-19 were entered into STRING online software to construct a PPI network, and the protein interaction network (.tsv) files and pictures were downloaded (Figure 4). The files were imported into Cytoscape 3.8 software, and 6 topological features of Betweenness Centrality (BC), Closeness Centrality (CC), Degree Centrality (DC), (Eigenvector Centrality) EC, (network centrality) NC, and (local average connectivity) LAC were selected using CytoNCA. The first circle candidate targets showed BC values > 20.978, CC > 0.625, DC > 26.5, EC > 0.112, LAC > 20.419, and NC > 22.691; whereas the second circle candidate targets showed BC values > 3.314, CC > 0.912, DC > 28, EC > 0.179, LAC > 24.571, and NC > 26.661 (see Figure 5). We identified 13 PPI network core proteins: MAPK1, MAPK3, MAPK8, MAPK14, STAT3, PTGS2, CXCL8, CASP3, CCL2, TP53, EGFR, VEGFA, and FOS.

PPI network of HXZQP for the prevention and treatment of COVID-19.

The core protein of the PPI network of HXZQP for the prevention and treatment of COVID-19.
Gene Ontology (GO) Enrichment Analysis of the Targets of HXZQP for the Prevention and Treatment of COVID-19
GO enrichment analysis was performed using R software and colorspace, stringi, ggplot2, DOSE, clusterProfiler, and enrichplot packages for GO enrichment analysis, including 3 parts: biological process (BP), molecular function (MF), and cell component (CC). Considering P < .05, q < .05, the result of the targets was related to 2103 items for BP, 46 items for CC, and 129 items for MF. Each category was sorted according to the number of enriched genes, and the top 10 items were displayed in the form of a bar plot and a bubble chart, as shown in Figure 6 and Figure 7. The results show that, in terms of the enrichment results of biological processes, the targets of HXZQP are mainly involved in defense response to the virus, modulation of host cellular process by the virus, regulation of defense response to virus by the host, cellular response to biotic stimulus, response to lipopolysaccharide, response to oxidative stress, response to molecule of bacterial origin, and cellular response to chemical stress. From the results of molecular function enrichment, HXZQP is mainly reflected in virus receptor activity, membrane raft, RNA polymerase II transcription regulator complex, protein kinase complex, plasma membrane raft, serine/threonine protein kinase complex, caveola, cyclin-dependent protein kinase holoenzyme complex, transcription regulator complex, membrane region, and membrane microdomain. From the results of the enrichment results of cellular components, the targets of HXZQP in the prevention and treatment of COVID-19 were identified to be mainly involved in cytokine receptor binding, protein serine/threonine kinase activity, death domain binding, BH domain binding, DNA-binding transcription factor binding, chemokine receptor binding, cytokine activity, RNA polymerase II-specific DNA-binding transcription factor binding, protein phosphatase binding, and phosphatase binding.

The bar plot of gene ontology enrichment analysis.

The bubble chart of gene ontology enrichment analysis.
Kyoto Encyclopedia of Genes and Genomes (KEGG) Enrichment Analysis of the Targets of HXZQP in the Prevention and Cure of COVID-19
Using R software and colorspace, stringi, ggplot2, DOSE, clusterProfiler, enrichplot, and pathview packages for KEGG enrichment analysis of the targets of HXZQP in the prevention and treatment of COVID-19, screening rules were set at P < .05, q < .05. The top 30 terms were visually displayed through bar plots and bubble charts (Figure 8 and Figure 9), and a total of 170 signaling pathways were screened. The main pathways of HXZQP in preventing COVID-19 include COVID-19, human papillomavirus infection, VEGF signaling pathway, PI3K-Akt signaling pathway, Epstein-Barr virus infection, human T-cell leukemia virus 1 infection, Th17 cell differentiation, influenza A, HIF-1 signaling pathway, TNF signaling pathway, IL-17 signaling pathway, apoptosis, toxoplasmosis, hepatitis C, measles, hepatitis B, human cytomegalovirus infection, and Kaposi sarcoma–associated herpesvirus infection. The COVID-19 pathway map was selected to draw the pathway map; the part marked in red is the target of HXZQP on this pathway with a total of 17 targets (Figure 10).

The bar plot of Kyoto Encyclopedia of Genes and Genomes enrichment analysis.

The bubble chart of Kyoto Encyclopedia of Genes and Genomes enrichment analysis.
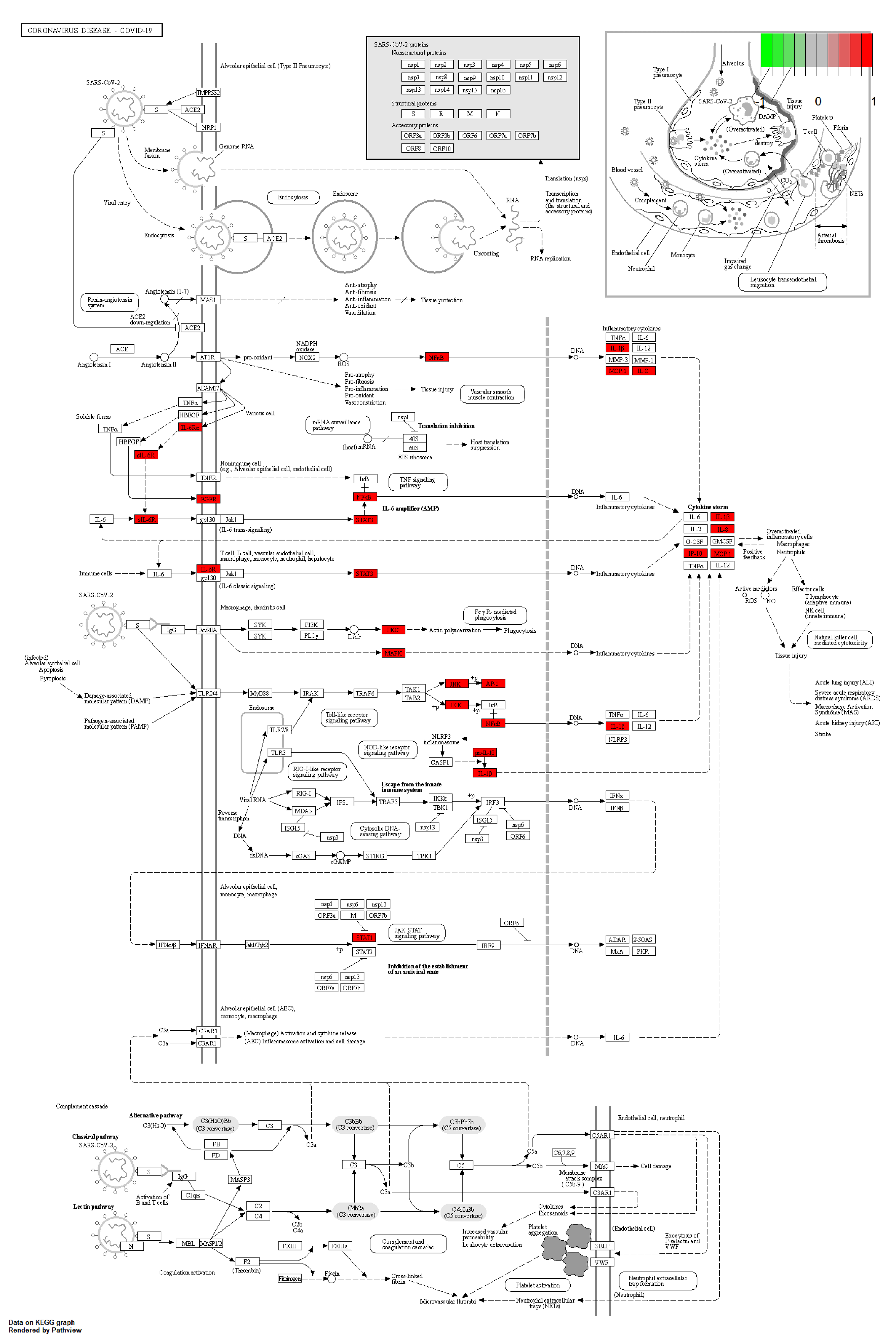
Target annotation mapping of active components of HXZQP in COVID-19 treatment.
Analysis of the Molecular Docking Results of Some Active Ingredients of HXZQP and Target Protein
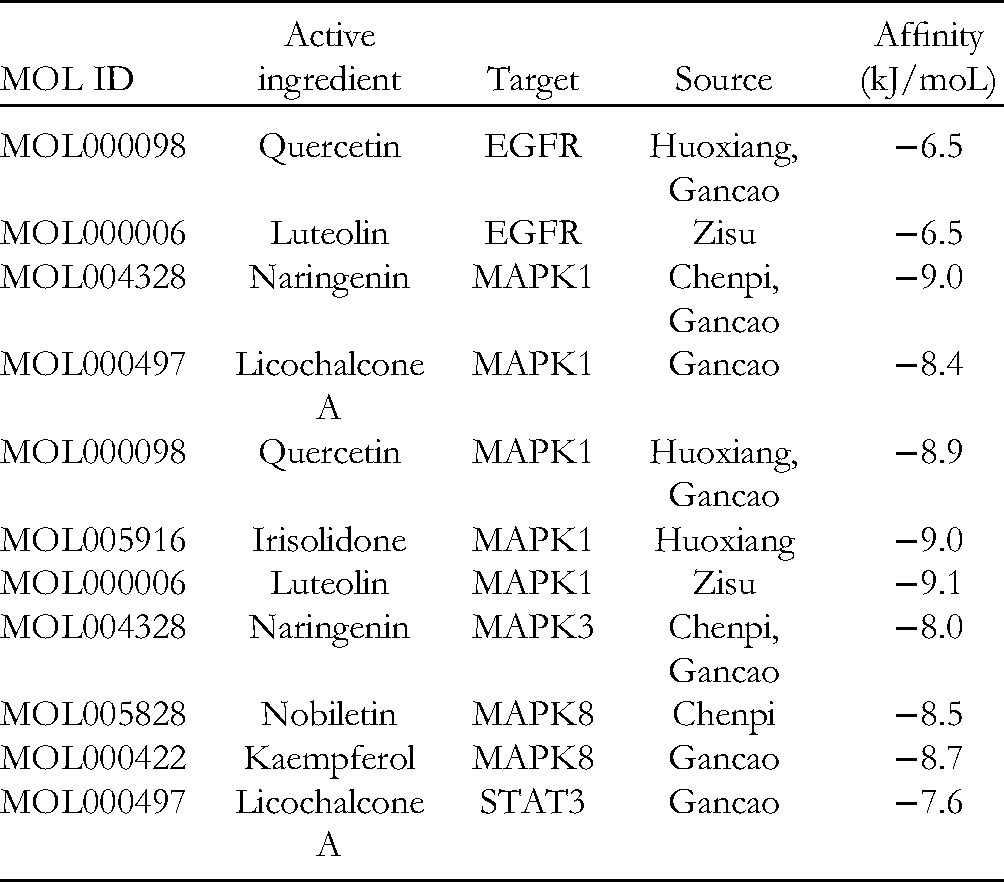
The key target proteins involved in the COVID-19 treatment pathway were selected, namely, EGFR (PDB ID: 2M0B), MAPK1 (PDB ID: 4FV5), MAPK3 (PDB ID: 2ZOQ), MAPK8 (PDB ID: 3ELJ), and STAT3 (PDB ID: 6TLC), to analyze the molecular docking results of the target protein and chemical components of HXZQP. The low energy stable conformation between the ligand and receptor suggests a large likelihood of interaction between them. In this study, AutoDock Vina (v4.2) docking software was used to perform molecular docking of the effective components of HXZQP (small-molecule ligands) and drug target proteins (receptors). The smaller the docking score, the stronger the binding affinity. The conformation with the highest score (the lowest affinity value) was selected as the docking conformation, and PyMol was used to visually analyze the docking conformation. Among the several binding conformations of the docking results for every active ingredient, we selected the conformation with the lowest binding energy. The smaller the binding energy, the higher the affinity between the receptor ligands (see Table 1). The molecular docking model of the representative core active ingredients of HXZQP and the key target proteins involved in the COVID-19 treatment are shown in Figure 11.

Molecular docking model of the representative core active ingredients of HXZQP and the key target proteins involved in the COVID-19 pathway.
Binding Energy of Representative Core Active Ingredients of HXZQP With Key Target Protein Molecules Involved in COVID-19 Treatment.
Discussion
According to the National Health Commission of the People's Republic of China, as of February 25, 2021, 101 778 patients with COVID-19 have been confirmed, and 96 361 patients have been cured and discharged in China. 12 TCM has shown distinctive advantages in epidemic prevention and control. The number of confirmed cases treated by TCM in China has exceeded 90%. HXZQP and other Chinese patented medicines have been recommended by the state and have played a positive role in the front line of the epidemic. However, owing to its complexity, the specific active components and pharmacological effects of HXZQP cannot be accurately clarified. Network pharmacology studies interactions among drugs, targets, and diseases from the standpoint of biomolecular networks. The interactions among drugs, active ingredients, targets, and related diseases are abstracted as a network model. The topological parameters of each node in the network are obtained. These parameters are used to appraise the significance of the nodes in the entire network. 13
In this study, with the aid of TCMSP, the OB and DL values for screening conditions were used to collect the targets of the active ingredients of all the drugs of HXZQP, and the UniProt database was used to obtain all official gene symbols of correspondence target information and 151 active ingredients of HXZQP. The results reflected that the most active ingredients in HXZQP were Glycyrrhizae Radix et Rhizoma, Angelicae Dahuricae Radix, Perillae Folium, Pinelliae Rhizoma, Pogostemonis Herba, Poria, Citri Reticulatae Pericarpium, Atractylodis Rhizoma, Magnoliae Officinalis Cortex, and Arecae Pericarpium. Thereafter, we explored the potential biological mechanisms of HXZQP in the prevention and treatment of COVID-19. We used GeneCards, OMIM, PharmGKB, TTD, and DrugBank databases and obtained 749 COVID-19-related targets and 67 intersection targets through the intersection with HXZQP targets. By constructing a PPI network with 67 intersection targets, we discovered that there were complicated connections between these proteins and not a single-line action. The proteins MAPK1, MAPK3, MAPK8, MAPK14, STAT3, PTGS2, CXCL8, CASP3, CCL2, TP53, EGFR, VEGFA, and FOS in the PPI core network could be the major direct targets of HXZQP in the prevention and treatment of COVID-19. Among them, MAPK1, MAPK3, MAPK8, and MAPK14 are members of the mitogen-activated protein kinase (MAPK) family, which plays a critical role in many cell multiplication-related signaling pathways. It is a key signaling pathway that participates in the physiological control of cell differentiation, growth, survival, apoptosis, and migration. Research has shown that the MAPK pathway is activated by many viral infections and is indispensable for viral proliferation. 14 Studies have confirmed 15 that SARS-CoV-2 is transmitted through the respiratory system, and the virus enters the circulatory system, causing an immune response in the blood and activating the MAPK signaling pathway to resist the virus. STAT3 mediates the expression of multiple genes involved in cytostimulation; therefore, it plays a critical role in many cellular processes, such as cell multiplication and apoptosis. In SARS-CoV-2-infected cells, a positive feedback loop established between STAT3 and plasminogen activator inhibitor-1 (PAI-1) may lead to an escalating cycle of activation, similar to the interdependent signaling networks affected in COVID-19. 16 Cytokine release syndrome is a manifestation of excessive activation of the immune system. Compared with other respiratory viral infections such as influenza, the main pathogenic mechanism involved in the severe clinical features of COVID-19 is an abnormal host immune response, which causes the excessive release of cytokines and chemokines, called “cytokine release syndrome (CRS).” 17 Although the pathophysiological mechanism of CRS is not yet clear, many studies have shown that the pathogenesis of CRS is closely related to the disorder of the interaction and regulation of various cytokines between cells and the imbalance between pro-inflammatory and anti-inflammatory responses. The results of the PPI core network in this study showed that targets belonging to the pro-inflammatory cytokines PTGS2 and CXCL8 are closely related to CRS. Cyclooxygenase-2 (COX-2) catalyzes the production of prostaglandin E2 (PGE2), which is widely involved in the inflammatory response in multiple tissues and organs and plays a very important role in the body's inflammatory response. CXCL8, also known as interleukin-8 (IL-8), is a glutamate (leucine) CXC chemokine. Studies have found that COVID-19 patients have higher levels of CXCL8, VEGFA, and VEGF in the peripheral blood than healthy individuals. 18 CASP3 is the main virus-induced apoptotic effector 19 and an important mediator of p53-induced apoptosis. 20 CCL2 is considered one of the earliest chemokines significantly upregulated in SARS-CoV-infected pulmonary epithelial cells, and its serum level correlated positively with the severity of SARS infection. 21
In addition, GO and KEGG enrichment analysis of the signaling pathways of 67 proteins revealed that the targets of HXZQP are involved in defense response to the virus, modulation of host cellular process by the virus, regulation of defense response to virus by the host, virus receptor activity, and other virus-related regulation. The main actions of HXZQP to prevent and cure COVID-19 include COVID-19, human papillomavirus infection, VEGF signaling pathway, PI3K-Akt signaling pathway, Epstein-Barr virus infection, human T-cell leukemia virus 1 infection, Th17 cell differentiation, influenza A, HIF-1 signaling pathway, TNF signaling pathway, IL-17 signaling pathway, apoptosis, toxoplasmosis, hepatitis C, measles, hepatitis B, human cytomegalovirus infection, Kaposi sarcoma–associated herpesvirus infection, and other signaling pathways that are closely related to COVID-19. In the COVID-19 pathway, SARS-CoV-2 infects alveolar epithelial cells (mainly alveolar epithelial type 2 (AEC2) cells) via angiotensin-converting enzyme 2 (ACE2) receptors. When ACE2 is occupied by SARS-CoV-2, the level of serum-free angiotensin II (Ang II) increases due to a decrease in ACE2-mediated degradation, which promotes the activation of the NF-κB pathway through Ang II type 1 receptor (AT1R), and then produces interleukin-6 (IL-6). SARS-CoV-2 activates the innate immune system. Macrophage stimulation causes excessive production of pro-inflammatory cytokines (including IL-6) and “cytokine storm,” leading to systemic inflammatory response syndrome and multiple organ failure. The synthetic effects of endothelial injury, neutrophil imbalance, complement activation, and a hypercoagulable state seem to be intertwined, leading to serious features of COVID-19. Targeting the IL-17 signaling pathway axis can be used as a new strategy for the treatment of lung injury. 22 TH17 type reaction participates in the cytokine storm in viral lung infections, including SARS-CoV-2, causing tissue damage and possibly promoting pulmonary edema. Targeting the TH17 pathway may benefit patients with dominant TH17 immunity. 23 The PI3K-Akt signaling pathway plays a major role in cell differentiation, proliferation, oxidative stress, and apoptosis. It inhibits autophagy in lung fibroblasts and reduces the formation of lung fibrosis. 24
There are 17 targets of HXZQP related to COVID-19 treatment: PRKCA, RELA, FOS, MAPK14, IL6R, CCL2, CXCL8, MAPK3, MAPK1, MAPK8, IKBKB, STAT1, STAT3, EGFR, IL1B, PRKCB, and CXCL10. Among these, EGFR, MAPK1, MAPK3, MAPK8, and STAT3 were the core targets. The core active components of EGFR, MAPK1, MAPK3, MAPK8, and STAT3 target proteins involved in HXZQP were MOL000098 (quercetin) and MOL000006 (luteolin), MOL004328 (naringenin), MOL005916 (irisolidone), MOL005828 (nobiletin), MOL000422 (kaempferol), and MOL000497 (licochalcone A). We used molecular docking technology to preliminarily explore the affinity of core active ingredients for EGFR (PDB ID: 2M0B), MAPK1 (PDB ID: 4FV5), MAPK3 (PDB ID: 2ZOQ), MAPK8 (PDB ID: 3ELJ), and STAT3 (PDB ID: 6TLC). The low-energy and stable conformation between the ligand and receptor indicates a greater likelihood of interaction between the 2. Generally, a binding energy ≤5 kJ/moL is used as the screening standard. The results revealed that after entering the human body, quercetin with EGFR and MAPK1 in Pogostemonis Herba and Glycyrrhizae Radix et Rhizoma, luteolin with EGFR and MAPK1 in Perillae Folium, naringenin with MAPK1 and MAPK1 in Citri Reticulatae Pericarpium and Glycyrrhizae Radix et Rhizoma, licochalcone a with MAPK1 and STAT3 in Glycyrrhizae Radix et Rhizoma, kaempferol with MAPK8 in Glycyrrhizae Radix et Rhizoma, irisolidone with MAPK1 in Pogostemonis Herba, and nobiletin with MAPK8 receptor in Citri Reticulatae Pericarpium showed strong binding ability. Quercetin is an expectorant and cough suppressant and has anti-inflammatory and anti-asthmatic effects and immunosuppressive effects; luteolin has various pharmacological activities, such as anti-inflammatory, anti-allergic, antibacterial, and antiviral. Naringenin has various pharmacological activities such as anti-inflammatory, antiviral, and anti-fibrosis. Kaempferol, as a common flavonoid, has anti-infective, anti-inflammatory effects, and anticancer effects. Licorice chalcone A (licochalcone a) has antibacterial, anti-inflammatory effects, and antioxidant effects. Nepalese irisolidone (irisolidone) has evident effects on dilating coronary arteries, cerebral arteries, peripheral blood vessels, and microvessels. It also reduces blood vessel resistance, cholesterol, and blood viscosity. Chuan Chen Pi (nobiletin) has anti-infective and antiviral therapeutic properties. The core active ingredients from the medicinal flavors of patchouli, licorice, chenpi, and perilla may be part of the basis for the treatment of COVID-19 in the HXZQ formula. This indicates that HXZQP can regulate the COVID-19 pathway and contribute to the prevention of COVID-19.
Conclusion
In conclusion, using network pharmacology, we found that HXZQP could contribute to the prevention and cure of COVID-19 through multiple compounds, targets, and pathways, providing theoretical evidence for the study of its active components and experimental research. Using network pharmacology and molecular docking to study the active ingredients and action targets of HXZQP may have some limitations; for example, the initial message comes from discordant experimental conditions, the limited number of drug action targets and small molecular compounds, and the information in the database tilts toward certain hotspots. Therefore, it is essential to thoroughly verify the outcomes of this study with the aid of future experimental studies, particularly the regulatory mechanism of HXZQP and its components in the COVID-19 signaling pathway.
Materials and Methods
Mining and Screening of Active Ingredients and Targets of HXZQP
The Huoxiang, Zisu, Cangzhu, Chenpi, Houpu, Baizhi, Fuling, Dafupi, Banxia, and Gancao in HXZQP were input into the TCMSP database (http://tcmspw.com/tcmsp.php) to search and obtain their respective active ingredients by screening and meeting the 2 conditions of OB value ≥30% and DL value ≥0.18. Thereafter, the targets of the active ingredients of all the drugs of HXZQP in the TCMSP database were identified using the UniProt database (https://www.uniprot.org/) to obtain all the official gene symbols corresponding to the target information, and this section of the information was applied for follow-up network pharmacology data analysis.
Mining Disease Targets of COVID-19
We used GeneCards (https://www.genecards.org/), OMIM (https://omim.org/), PharmGKB (https://www.pharmgkb.org/), TTD (http://db.idrblab.net/ttd/), and DrugBank (https://www.drugbank.ca/) database to identify disease targets of COVID-19 using “2019-nCoV,” “COVID-19,” and “SARS-CoV-2” as keywords to filter and merge COVID- 19 human disease–related genes.
Drug-Active Ingredient–Target Interaction Network Construction of HXZQP to Prevent and Treat COVID-19
Using Venny 2.1 (https://bioinfogp.cnb.csic.es/tools/venny/index.html) online software, we considered the intersection of HXZQP action targets and COVID-19 disease targets, and the intersection targets were considered the core targets of HXZQP to prevent and treat COVID-19. We used Cytoscape 3.8 software to construct a drug-active ingredient–target interaction network for HXZQP in the prevention and treatment of COVID-19.
PPI Construction and Topological Analysis of HXZQP to Prevent and Treat COVID-19
The online software STRING: functional protein association networks (https://www.string-db.org/), select multiple proteins, protein name: input the intersection gene of front HXZQP and COVID-19; Organism: Select Homo sapiens to build a protein-protein interaction (PPI) network. We used Cytoscape 3.8 software to obtain the core protein of the PPI network.
GO and KEGG Pathway Enrichment Analysis for the Targets of HXZQP
R software was used to carry out GO and KEGG pathway enrichment analysis for the prevention and treatment of COVID-19 by HXZQP. To elucidate the molecular mechanism of HXZQP in preventing COVID-19, we need to begin with 2 aspects of gene function and pathway analysis.
Component-Target Molecular Docking
The PPI network core protein was used as an example to analyze the molecular docking results of the target proteins and chemical components. First, we downloaded the 2D structure SDF format file of the small-molecule ligand (the active ingredient of Huoxiang Zhengqi Chengfang) from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/) and used ChemOffice software to convert its 2D structure into a 3D structure. The 3D structure PDB format file of the PPI network core protein was downloaded from the RSCB PDB database (https://www.rcsb.org/), and the PyMol software removed the water molecules and small-molecule ligands in the target protein and then saved it as a PDB format file. AutoDockTools software was used to convert the SDF format files into PDBQT format files and to determine the active pockets. Finally, AutoDock Vina was used for molecular docking of the active ingredients of traditional Chinese medicine (small-molecule ligands) and drug target proteins (receptors).
Supporting Information
This study was funded by the National Natural Science Foundation of China (Grant no. 81603412); Key R&D Projects of Hebei Province (Grant no. 18277731D); Scientific Research Project of Hebei Administration of Traditional Chinese Medicine (Grants nos. 2017163, 2019008, and 2020014); General Projects for Improving Scientific Research Capacity of Hebei College of Traditional Chinese Medicine (Grant no. KTY2019009); Hebei Key Laboratory of Chinese Medicine Research on Cardio-Cerebrovascular Disease; Key Laboratory of Integrated Traditional Chinese and Western Medicine Hepatonephrosis in Hebei Province (Grant no. A201902); and Hebei Province “Three Three Three Talent Project” funded project (Grant no. A202002008), Hebei Provincial Natural Science Foundation Grant (H2021423060).
Footnotes
Acknowledgments
The author is very grateful for the data provided by the TCMSP, GeneCards, OMIM, PharmGKB, TTD, and DrugBank database.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Hebei Province “Three Three Three Talent Project” funded project, National Natural Science Foundation of China, Key R&D Projects of Hebei Province, General Projects for Improving Scientific Research Capacity of Hebei College of Traditional Chinese Medicine, Scientific Research Project of Hebei Administration of Traditional Chinese Medicine, Hebei Key Laboratory of Chinese Medicine Research on Cardio-Cerebrovascular Disease; Key Laboratory of Integrated Traditional Chinese and Western Medicine Hepatonephrosis in Hebei Province, Hebei Provincial Natural Science Foundation Grant (grant number A202002008, 81603412, 18277731D, KTY2019009, 2017163, 2019008, 2020014, A201902 and H2021423060).
