Abstract
Background
Liver cancer is a common threat to human health. Schizandrin A (SA) has certain therapeutic effect on liver cancer, but its related mechanism of action is not clear. Elucidating this mechanism is of great significance for the development of drugs to target liver cancer.
Objective
The objective is to study the effect and mechanism of SA on liver cancer.
Methods
The potential targets of SA in the treatment of liver cancer were screened based on network pharmacology, and the core targets were further screened by protein network interaction. Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis was performed based on the core targets to find the key pathways of action. The results obtained for key targets and pathways were verified in vitro by pharmacological experiments with HepG2 cells. Specifically, an Enzyme Linked Immunosorbent assay (ELISA) kit was used to detect the content of key indicators (interleukin-1 (IL-1), Tumor Necrosis Factor-alpha (TNF-α), interleukin-6 (IL-6), interleukin-10 (IL-10), superoxide dismutase [SOD], malondialdehyde [MDA], interferon (IFN), and Glutathione peroxidase (GSH-px)), Quantitative Real-time Polymerase Chain Reaction (qRT-PCR) was used to detect the key messenger Ribonucleic Acid (mRNA) (phosphatidylinositol 3 kinase (PI3K), protein kinase B (Akt), vascular endothelial growth factor A (VEGFA), and Transforming Growth Factor Beta-1 (TGF-β1)) after drug intervention, and Western blotting (WB) detected the expression of key target proteins (p-PI3K, PI3K, p-Akt, and Akt).
Results
The results showed that SA mainly acts on IL-1β, TNF-α, IL-6, IL-10, SOD, MDA, TGF-β1, IFN, and VEGFA, affecting the PI3K-Akt signaling pathway and playing an anti-inflammatory role in the treatment of liver cancer. The experimental results showed that SA significantly reduced the inflammation level of HepG2 cells; improved the oxidative stress state (decreasing SOD level and increasing MDA level of cells); downregulated TGF-β1, VEGFA, PI3K, and Akt mRNA levels; and inhibited the expression of the PI3K-Akt signaling pathway.
Conclusions
SA has a significant anti-HepG2 effect, which may be achieved by reducing inflammation levels, improving oxidative stress state, and inhibiting the PI3K-Akt pathway.
This is a visual representation of the abstract.
Introduction
As the second most prevalent cancer in the world, liver cancer is a serious threat to human physical and mental health. 1 According to epidemiological data, China has more than 4 million patients every year. Diagnosed with liver cancer, about 400 000 people die from the disease, and the incidence is increasing every year. 2 High heterogeneity leads to low clinical treatment efficiency, which is a serious concern to many doctors and patients.
Inflammation and oxidative stress are important factors affecting the occurrence and development of liver cancer, and some factors play an important role in its occurrence, such as the inflammatory cytokines TNF-α, IL-1β, IL-6, and IL-10, which can damage hepatic epithelial cells. Because the liver has a high regenerative capacity, this damage induces substantial cell proliferation. When the high rate of cell proliferation is coupled with DeoxyriboNucleic Acid (DNA) mutation, the incidence of malignant transformation increases. 3 Oxidative stress can induce hepatocyte injury, produce multiple cytokines and chemokines which promote the occurrence of fibrosis, and affect liver inflammation and cell apoptosis. Malondialdehyde (MDA) is the lipid peroxide formed by reactive oxygen species (ROS) attacking the lipid and oxidizing the lipid on the cell membrane surface. Superoxide dismutase (SOD) maintains a dynamic balance with oxygen free radicals under normal conditions and changes with changes in oxygen free radical levels and membrane lipid peroxidation under pathological conditions. 4 GSH-px can change lipid hydroperoxides and free hydrogen peroxide into their corresponding alcohols and water, thus protecting the organism from oxidative damage. 5 In addition to the above factors, other factors are also important to the occurrence of liver cancer. For example, VEGFA, as one of the most recognized and effective endothelial growth factors, mobilizes the endothelial progenitor cells to participate in tumor angiogenesis and is a key target for the treatment of liver cancer. 6 It is reported that aberrant TGF-β1 signaling probably has a pro-ontogenetic role in liver cancer occurrence and is a well-characterized inducer of epithelial-mesenchymal transition (EMT) in liver cancer. 7 In summary, the above factors are closely related to the occurrence of liver cancer and may be the key targets for the treatment of liver cancer.
In recent years, Traditional Chinese Medicine (TCM) has been increasingly widely used in clinics due to its high efficacy, safety, and other comprehensive therapeutic effects. 8 The multilevel, multitarget, and multipathway therapeutic mechanism is the main way to exert its efficacy. However, how to treat diseases through the key target or pathway of TCM has been a difficult problem puzzling the modernization of TCM. With the development of network pharmacological technology, the efficiency of TCM has been greatly improved. 9 This technology can efficiently provide researchers with possible key targets for TCM treatment of diseases and point out the direction for pharmacological experiments. 10 At the same time, the “multiple targets–single ligand” docking mode of molecular docking technology can be matched according to space and power. This technology is accurate and low-cost, currently mainly used in drug design and elucidating biochemical pathways. 11 Therefore, network pharmacology–molecular docking lays a scientific and theoretical basis for research and development of new Chinese medicine.
schizandrin A (SA), a lignan found in Schisandra chinensis (Turcz.) Baill., has significant pharmacological activities such as anti-inflammation, antioxidation, antitumor, and antiviral. 12 It has been shown that SA has a significant effect on the treatment of liver cancer.13,14 However, the mechanism of SA in the treatment of liver cancer has not been reported. Therefore, based on the strategy of “network pharmacology–molecular docking–targeted validation,” this study effectively screened out the target and key pathway of SA in the treatment of liver cancer and verified its prediction in in vitro cell experiments to clarify its related mechanism. This study lays a certain foundation for the treatment of liver cancer and the development of target drugs and provides a scientific reference for the future of TCM pharmacological research.
Material and Methods
Instrument and Software
The instrument and software include a multifunctional enzyme marker (Molecular Devices; Catalog number: SpectraMax M5), gel imaging system (Bio-Rad Bole; Catalog number: ChemiDoc XRS+), ultrasonic crushing machine (Shanghai Zhixin; Catalog number: JYD-900), carbon dioxide incubator (Shanghai Yiheng; Catalog number: BPN-80RNP), electrometer (Beijing Liuyi; Catalog number: DYCZ-24K), membrane transfer instrument (Beijing Liuyi; Catalog number: DYCZ-40D), real-time Polymerase Chain Reaction (PCR) (Applied Biosystems™; Catalog number: 7500), Cytoscape 3.7.2 software, Open Babel 3.1.1 software, and AutoDockTools 1.5.7 software.
Drugs and Reagents
SA (purity: 97%, batch number: 111520-202105) was purchased from the National Institutes for Food and Drug Control, IL-1β (NeoBioscience; Catalog number: ERC007.48), TNF-α (NeoBioscience; Catalog number: EMC102a.96), IL-6 (NeoBioscience; Catalog number: ERC003.48), IL-10 (NeoBioscience; Catalog number: ERC004.96), and IFN kit (NeoBioscience; Catalog number: ERC101g.48) were purchased from Nanjing Kechuang Biotechnology Co, Ltd. SOD (Nanjing Jiancheng; Catalog number: A001-4-1) and MDA (Nanjing Jiancheng; Catalog number: A003-1-2) kits were purchased from Nanjing Jiancheng Biological Co, Ltd. Human hepatoma cell line HepG2 cells were provided by the School of Basic Medical Sciences, Zhengzhou University. Antibodies p-PI3K (CST; Catalog number: 17366), PI3K (CST; Catalog number: 4249), p-Akt (CST; Catalog number: 4060), Akt (CST; Catalog number: 9272), and glyceraldehyde-3-phosphatedehydrogenase (GAPDH) (CST; Catalog number: 2118) were purchased from Cell Signaling Technology, Inc.
Cells and Groups
HepG2 cells were stored in dulbecco's modified eagle medium (DMEM) medium supplemented with fetal bovine serum and double antibody and cultured in a humidified incubator at 37 °C and 5% CO2. The cells were divided into the control (DMEM) and SA low-dose (10 μmol/L), medium-dose (20 μmol/L), and high-dose (40 μmol/L) groups.
Prediction of Potential Targets of SA
TCMSP (https://old.tcmsp-e.com/tcmsp.php), SwissTargetPrediction (http://swisstargetprediction.ch/), STITCH (http://stitch.embl.de/), and SEA (http://sea.edbc.org) were used to predict the SA targets, which were changed to the uniform target name in the UniProt database. 15
Prediction of Potential Targets for Liver Cancer
With “liver cancer” as the index word, GeneCards (https://www.genecards.org/), OMIM (http://www.omim.org/), CTD (http://ctdbase.org/), TTD (http://db.idrblab.net/ttd/), and DisGeNET (http://www.disgenet.org/home/) were used to search for the targets related to the disease. 16
Target Intersection and “Drug–Target” Network Construction
The intersection of SA and potential targets of liver cancer was imported into Cytoscape 3.7.2 software to construct a “drug–target” network diagram, analyze the topological properties of the network, and use a network analyzer plug-in for data processing. 17
Construction of Protein–Protein Interaction Network and Analysis of KEGG Pathway
Potential targets were imported into the string database, “multiple proteins” were checked, “Homo sapiens” was selected for the species, the protein–protein interaction (PPI) diagram was constructed, and further analysis was conducted to screen the core targets. The core targets were imported into the Database for Annotation,Visualization and Integrated Discovery (DAVID) database, with P < .05 considered as biological significance, and then, KEGG analysis was performed. Finally, the number and importance of the enriched targets were sorted successively, and the top 20 items were selected to draw the diagram. 18
Molecular Docking Analysis
First, three-dimensional (3D) protein structures associated with key targets were exported from the RCSB PDB online platform (http://www.rcsb.org/) in PDB format. Then, the 3D structures of active compounds were downloaded from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/) and saved in SDF format. Then OpenBabel 3.1.1 software was imported, and the 3D protein was converted into a format that AutoDock could recognize. Pymol was used to dewater and remove small molecules from the active compounds and protein structures, and AutoDockTools 1.5.7 software was imported to set the active compounds as ligand (add total hydrogen, and set the torskey for detection), and the protein structure was set as receptor (add total hydrogen) and then converted into “pdbqt” format. AutoDock4 (set docking box, docking parameters, calculation methods, etc) was used for molecular docking, and the docking results were visualized on Pymol. Finally, docking binding energy was used to evaluate the ability of the small molecules to bind to the disease target proteins. 19
Determination of Survival Rate of HepG2 Cells
HepG2 cells (5 × 103) were inoculated into each well of a 96-well plate and then cultured in a CO2 incubator for 24 h. The culture medium was discarded, and the cells were washed twice with phosphate buffer solution (PBS) before being treated with different concentrations of SA (0, 2.5, 5, 10, 20, 40, 80, and 160 μmol·L−1) for 24 and 48 h, respectively, and then, the medium was discarded. After cleaning with PBS twice, the Cell-Counting-Kit-8 (CCK-8) reagent (10 μL) was added to each well and then cultured in a CO2 incubator for 2 h. The wavelength was set at 450 nm, the optical density (OD) of each well was determined, and the survival rate of cells at different concentrations was calculated. 20
Determination of Cell Supernatant-Related Factors
About 5 × 105 cells from different experimental groups were inoculated on 6-well plates. Twenty-four hours later, the cells were collected and added to PBS ice for ultrasonic cell lysis. Then, the cells were placed in a centrifuge at 4 °C for 10 min at 12 000 g, and the supernatant was collected for detection. The contents of TNF-α, IL-6, IL-10, IL-1β, SOD, GSH-px, MDA, and interferon-γ (IFN-γ) in each group were detected and analyzed according to the instructions of the kit. 21
qRT-PCR Detection
According to the product description, the total cell Ribonucleic Acid (RNA) was extracted by TRIzol, and cDNA was synthesized by a Servicebio®RT First-Strand cDNA Synthesis Kit. Quantitative detection of PI3K, Akt, VEGFA, TGF-β1, and GAPDH genes was performed by 2× SYBR Green qPCR Master Mix. 22 The primary information is shown in Table 1.
Primary Information of qRT-PCR Detection.

Note: annealing temperature 54 °C.
Relative expression level = 2−(α−β), α = CTTarget gene after SA intervention−CTReference gene after SA intervention, β = CTTarget gene without SA intervention−CTReference gene without SA intervention.
Key Protein Expression Detected by Western Blotting
Five culture dishes were inoculated with about 1 × 106 HepG2 cells in each dish and divided into the control and SA low-dose (10 μmol/L), medium-dose (20 μmol/L), and high-dose (40 μmol/L) dose groups. Dishes were cultured for 24 h; then, the cells were collected and the proteins of each group extracted. A BCA protein quantitative detection kit was used to measure the protein concentration of each group, and polyacrylamide gel electrophoresis was performed according to the total protein loading amount (40 μg) of each group. The protein on the glue was transferred to a polyvinylidene difluoride (PVDF) membrane using a semidry membrane transferer/automatic fast protein transfer system and sealed with 5% skim milk powder at room temperature for 1 h. The protein bands of PI3K, p-PI3K, Akt, p-Akt, and GAPDH were cut down according to the molecular weight of the marker. After Tris-Buffered Saline and Tween (TBST) cleaning, the bands were put into zip-lock bags, and the corresponding primary antibody was added (the dilution ratio of primary antibody was 1:000) and incubated at 4 °C overnight. Each strip was added to the corresponding secondary antibody (the dilution ratio of secondary antibody was 1:1 × 104) and incubated at room temperature for 1 h. After TBST cleaning, an appropriate amount of luminescent liquid was added to the strip, which was developed and photographed in a professional luminescent chemical imaging system. The experiment was repeated 3 times. GAPDH was used as the internal reference, and ImageJ software was used to analyze the protein expression levels of each group. 23
Statistical Analysis
The experiment was independently repeated 3 times with 6 samples in each group. The results are represented by mean plus or minus standard deviation. The experimental data were statistically analyzed using Statistical Product and Service Solutions (SPSS) 21.0 and an independent sample t-test. P < .05 was considered statistically significant.
Results
Construction of Target Network of SA in the Treatment of Liver Cancer
In this study, 358 targets of SA and 2401 targets of liver cancer were screened, and 156 targets of SA directly acting on liver cancer were obtained after intersection; the drug–target network was constructed, as shown in Figure 1.

“Drug–target” network of schizandrin A (SA) in the treatment of liver cancer.
PPI Network
As shown in Figure 2, the network contains nodes (nodes represent targets) and edges (edges represent interactions between targets). The larger and darker the nodes in the figure are, the larger the degree value. The most significant core targets were PI3K, AKT, IL-1β, TNF-α, IL-6, IL-10, SOD, MDA, TGF-β1, IFN, and VEGFA.

Protein–protein interaction (PPI) network of schizandrin A (SA) in the treatment of liver cancer.
KEGG Pathway Analysis and Molecular Docking Results
As shown in Figure 3, after KEGG analysis of the core target, the PI3K-Akt signaling pathway ranked the highest, providing a scientific theoretical basis for further experimental verification. It is generally believed that a binding energy less than −5.0 kcal·mol−1 indicates that the ligand has a good binding activity to the receptor, while a binding energy less than −7.0 kcal·mol−1 indicates a strong binding activity. The results of molecular docking in this study showed that the binding energy of SA with key target proteins PI3K (−8.1 kcal·mol−1) and Akt (−7.2 kcal·mol−1) was low, and the hydrogen bonds between molecules and proteins were stable. SA can stably bind to the active pockets of PI3K and Akt and is “embedded” in the active pockets of the proteins, suggesting that the biological function of SA may be mediated by affecting these 2 proteins and their downstream signaling pathways. The 3D structure of SA and its docking with the active pocket molecules of PI3K and Akt are shown in Figure 4.

KEGG pathway enrichment analysis.

Molecular docking results of core targets.
Cell Proliferation Assay
Compared with the blank group, the survival rate of HepG2 cells in the experimental group decreased in a concentration- and time-dependent manner. When SA ≤ 40 μmol·L−1 was used for 24 h, the cell survival rate was more than 80%. When the SA concentration was less than 10 μmol·L−1 for 48 h, the survival rate of cells was more than 80%. In order to avoid subsequent experimental errors caused by SA cytotoxicity, this research selected the drug concentration with no obvious cytotoxicity as the administration concentration. Therefore, we selected 24 h as the dosing time, and 10.0, 20.0, and 40.0 μmol·L−1 as the dosing concentration. The results are shown in Table 2.
Effect of SA on HepG2 Cell Viability (x ± s) (n = 6).

Note: versus blank control.
*P < .05, **P < .01.
SA, schizandrin A.
Cytokine Detection Results
Results of inflammatory cytokines showed that, compared with the control group, the levels of TNF-α (t = 3.86, 4.94, 5.409), IL-1β (t = 4.269, 9.414, 16.24), IL-6 (t = 3.503, 7.696, 11.62), and IFN-γ (t = 5.629, 8.356, 13.38) were decreased; GSH-px (t = 4.319, 7.405, 12.18) and SOD (t = 3.418, 5.58, 9.654) contents were significantly decreased in the SA low-dose, medium-dose and high-dose groups (P < .05). The contents of IL-10 (t = 7.92, 12.73, 19.28) and MDA (t = 11.06, 13.31, 49.77) of the SA low-dose, medium-dose, and high-dose groups were significantly increased, with statistical significance (P < .05), as shown in Figure 5.
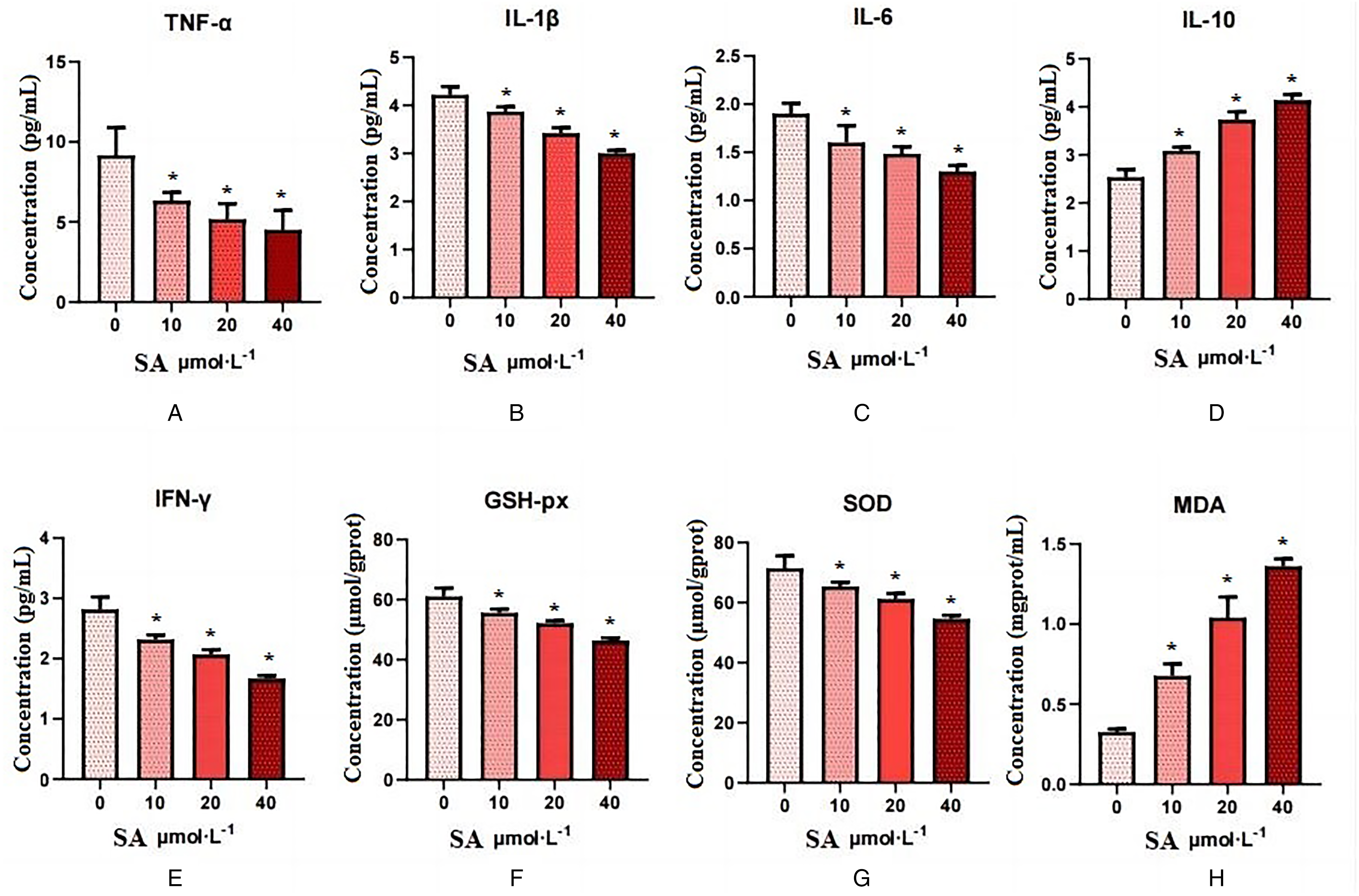
The expression levels of different cytokines in experimental groups.
qRT-PCR Results
Compared with the control group, the relative expression levels of PI3K mRNA were 0.76, 0.69, and 0.57 (t = 10.05, t = 19.64, and t = 5.351; P < .05); the relative expression levels of Akt mRNA were 0.91, 0.83, and 0.71 (t = 12.41, t = 18.28, and t = 23.79; P < .05) at low, medium, and high concentrations of SA for 24 h, respectively. The relative expression levels of VEGFA mRNA were 0.89, 0.85, and 0.77 (t = 2.902, t = 3.293, and t = 6.627; P < .05); the mRNA relative expression levels of TGF-β1 were 0.86, 0.79, and 0.78 (t = 11.73, t = 14.77, and t = 14.27; P < .05) at low, medium, and high concentrations of SA for 24 h, respectively, as shown in Figure 6.

Determination of expression levels of key target genes in different experimental groups.
The Protein Expression of PI3K-Akt Signaling Pathway Detected by Western Blotting
Compared with the control group, the expression levels of p-PI3K/PI3K and p-Akt/Akt in the supernatant of the SA low-dose, medium-dose and high-dose groups were significantly decreased (Figure 7).

PI3K/Akt protein levels in different experimental groups.
Discussion
Liver cancer is a common malignant tumor. In recent years, its morbidity and mortality have increased quickly. The occurrence and development of liver cancer are closely related to inflammation and oxidative stress, which are also common ways of anti-liver cancer therapy. 24 As the active monomer component of TCM, SA has certain anti-inflammatory and antioxidant stress effects, and therefore, its antitumor possibility has gradually become a research focus.25,26 The results of network pharmacology in this research suggest that SA may relieve the inflammatory state of HepG2 cells by regulating TNF-α, IL-6, IL-1β, IFN, IL-10, IFN, and other inflammatory factors, so as to play a certain therapeutic effect. It has been reported that the levels of IL-6, TNF-α, and IL-1β in mouse serum with liver cancer are significantly higher than those in normal mice and are positively correlated with the degree of malignancy of the tumors.27,28 Therefore, they can be used as important indicators for screening and evaluating the prognosis of liver cancer. Cell experimental results in vitro showed that SA could significantly reduce the levels of inflammatory factors (TNF-α, IL-6, IL-1β, and IFN) in HepG2 cells and increase the levels of anti-inflammatory factors (IL-10), with statistical significance (P < .05), which was mutually supported by the results of network pharmacology. These results suggest that inhibiting inflammation may also be an important pathway for SA to play in its anti-liver cancer role.
Oxidative damage refers to the phenomenon of protein damage caused by excessive production of ROS and active nitrogen (Reactive Nitrogen Species (RNS)) in the body, which cannot be timely cleared by the body. 29 SOD and GSH-px are classic antioxidants, which can remove oxygen free radicals, protect cells from damage, and reduce lipid peroxidation, protein damage, and DNA mutation caused by oxidative stress, and are important indicators of cellular oxidative damage.30,31 In vivo, free radicals acting on lipids can cause peroxidation, and the end product of oxidation is MDA, which can cause cross-linked polymerization of proteins, nucleic acids, and other macromolecules of life, and has cytotoxicity. 32 Therefore, MDA content is an important parameter reflecting the potential antioxidant capacity of cells, which can reflect the rate and intensity of lipid peroxidation of cells and indirectly reflect the degree of tissue peroxidation damage. 33 The results of this study showed that SA could decrease the SOD and GSH-px levels of HepG2 cells, while increasing the MDA levels, with statistical significance (P < .05). The results showed that SA could induce oxidative damage and apoptosis of HepG2 cells, indicating these as one of the important ways of its anti-liver cancer activity.
Liver cancer is a highly vascularized malignant disease, and angiogenesis is crucial for the growth, invasion, and metastasis of HegG2 cells. 34 Among the above factors, vascular endothelial growth factor (VEGF) is an important regulatory factor regulating tumor angiogenesis. It has been reported that VEGF is abnormally expressed in liver cancer tissues and cells. 35 Therefore, inhibition of VEGF expression is an important way to alleviate the occurrence and development of tumors. In addition, cytokine TGF-β1 can promote the proliferation, migration, invasion, and other biological behaviors of liver cancer cells by inducing epithelial–mesenchymal transformation-like changes in liver cancer cells and may be a key target for inhibiting liver cancer.36,37 The results of this study show that SA treatment caused the downregulation of VEGF and TGF-β1 mRNA expression in HepG2 cells, suggesting that SA, having good antitumor potential, could reduce tumor angiogenesis and inhibit the growth, invasion, and metastasis of liver cancer cells by reducing the expression of VEGF and TGF-β1.
Network pharmacology and molecular docking provide a new perspective to elucidate the molecular mechanism of the network between the component and target, the target and disease, and the free binding ability of the component and proteins, which is also in line with the characteristics of “multicomponent,” “multitarget,” and “multipathway” of TCM. 38 Network pharmacology suggests that the PI3K/Akt signaling pathway may be the key one for the anti-liver cancer of SA. Molecular docking results show that SA has stable binding states with the target proteins of PI3K and Akt, which may be the prerequisite for the function of this monomer. The PI3K/Akt signaling pathway is a signal transduction mechanism that plays an important role in the process of growth, metabolism, and various life activities of the body, especially in the process of liver cancer. 39 The results of this study indicated that SA could downregulate the expression of PI3K and Akt mRNA genes in HepG2 cells. Meanwhile, Western blotting (WB) results showed that p-PI3K/PI3K and p-Akt/Akt proteins were significantly decreased after the intervention of SA, which verified the results of network pharmacology at both gene and protein levels.
The present study has its limitations; it has only been tested on one type of cell but needs to be tested on other cell lines and further verified in animals. More techniques are needed to verify the correctness of this study. There are other signaling pathways and key targets that have not been fully validated, and these need to be investigated. It is believed that in the near future, SA will be further made into a new drug for the clinical treatment of liver cancers.
Conclusion
Based on the network pharmacology and molecular docking technology, this study preliminarily identifies the possible targets and key pathways of SA in the treatment of liver cancer, and further experimental verification confirms the scientific nature of the predicted targets and pathways, providing a new reference for the development and pharmacological research of relevant targets and drugs in the treatment of complex diseases with TCM in the future.
Supplemental Material
sj-tif-1-npx-10.1177_1934578X231176916 - Supplemental material for Effect and Mechanism of Schizandrin A in the Treatment of Liver Cancer Using Network Pharmacology, Molecular Docking, and Target Validation
Supplemental material, sj-tif-1-npx-10.1177_1934578X231176916 for Effect and Mechanism of Schizandrin A in the Treatment of Liver Cancer Using Network Pharmacology, Molecular Docking, and Target Validation by Xiaohui Wang, Lin Zhou, Tao Zhang, Hui Chen, Xinhao Song and Feng Wang in Natural Product Communications
Footnotes
Data Availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by funds from the Key Scientific Research Project of Colleges and Universities in Henan Province (23A320050; 23A360018), Medical Science and Technology Project of Henan Province (LHGJ20210283; LHGJ20210348), and Science and Technology Development Program of Henan Province (No. 212102311107).
Ethical Approval
Ethical approval is not applicable to this article.
Statement of Human and Animal Rights
There are no human and animal subjects in this article, and the relevant rights are not applicable.
Informed Consent
Informed consent is not applicable.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
