Abstract
Keywords
Introduction
Globally, liver disorders is a major cause of illness and death in humans. The liver disorders include hepatitis, nonalcoholic liver disease, drug-induced liver injury, liver cirrhosis, and hepatocellular carcinoma. Recently, there has been a significant increase in the incidence and prevalence of liver diseases. Although drug therapy is still the main choice of treatment for liver diseases, most drugs are associated with unwanted side effect. 1 Traditional Chinese medicine (TCM) has accumulated abundant clinical experience in the prevention and treatment of liver diseases. However, for most hepatoprotective drugs containing the Chinese medicine ingredients to be oral preparation, the slow absorption and efficacy are susceptible to gastrointestinal function and content. 2 Therefore, there is inevitable to explore new and more effective drugs for liver protection.
Shuganning injection (SGNI) is one of the Chinese patent medicines approved by the State Food and Drug Administration of China, which is developed based on “Yinchenhao Decoction,” and composed of different Chinese medicinal herbs: Artemisia capillaries (YC), Gardeniae Fructus (ZZ), Baicalin (HQG), Isatidis Radix (BLG), and Ganoderma (LZ). The effects of SGNI for clearing heat, detoxification, and hepatoprotective function are used for the treatment of acute and chronic hepatitis, as well as jaundice in clinical. SGNI was approved as a traditional Chinese patent medicine by the China Food and Drug Administration in 2002, with national medicine approval number Z0025660 and implemented in accordance with the China National Food and Drug Administration National Drug Standard (WS-10467(ZD-0467)-2002). Since SGNI has become one of the most excellent drugs in protecting liver, 3 it is applied for the treatment of liver function damage, fatty liver and cholangitis, clinical hepatitis, and high bilirubin hematic disease. In addition, it is used as an adjuvant treatment for cancer, especially for hepatic carcinomas. 4 Currently, the effective action of components such as Artemisia capillaries from SGNI have been reported to inhibit the increase of lipid peroxide, scavenge oxygen free radicals, and increase activity in levels of superoxide dismutase, glutathione peroxidase, and catalase for liver protection.5,6 The achievement of hepatoprotective effect of baicalin by increasing the level of peroxisome proliferator-activated receptor gamma,7,8 and the significantly reduced liver messenger RNA as well as protein expression of transforming growth factor β1, collagen type I, and α-smooth muscle actin using Gardeniae Fructus to attenuate hepatocellular injury and fibrosis. 9 However, the synergistic mechanism of multiple compounds in the SGNI for liver protection is not completely understood.
Network pharmacology is a novel discipline based on systems biology theory, which reveals the pathophysiology and treatment strategies of many disorders by integrating related fields like systems biology and pharmacology.10,11 Network pharmacology allows us to grasp how multiple ingredients of an herbal formula interact and what effect they can have on multiple targets of a disease through constructing the network of drugs, proteins, and pathways. 12 The application of network pharmacology to explore the mechanism of TCM has achieved significant results in elucidating some potential bioactive compounds and mechanisms of several traditional medicines in treatment of complex diseases in the last few years.13,14
Therefore, in this investigation, a comprehensive network pharmacology strategy and molecular docking approach were used in combination with in vitro experimental validation to examine the mechanism of SGNI in protecting liver (study flow diagram is shown in Figure 1).

Systems pharmacology approach workflow.
Results
Screening of Active Compounds and Targets of SGNI
In the present study, a total of 312 active compounds of SGNI were obtained. Furthermore, it was found that there were 34, 50, 1, 81, and 146 active compounds in Artemisia capillaries, Gardeniae Fructus, Baicalin, Isatidis Radix, and Ganoderma, respectively (see Table S1 for details). In addition, it was noted that different herbs contained similar compounds. For instance, β-sitosterol was found in Artemisia capillaries, Gardeniae Fructus, Isatidis Radix, and Ganoderma. Hirsutrin, rutin, and chlorogenic acid were revealed in Artemisia capillaries and Gardeniae Fructus. Stigmasterol was derived from Gardeniae Fructus, Isatidis Radix, and Ganoderma. Tetracosane and adenosine were found in both Isatidis Radix and Ganoderma.
Prediction of Hepatoprotective Targets of SGNI
A total of 408 targets of SGNI were obtained using Venny2.0, and they were used to construct the “compound-target” network which consisted of 860 nodes and 5990 edges. Among the active compounds in the SGNI, the top 5 active compounds were highly connected to the targets, including quercetin (degree = 152), piceol (degree = 102), hexacosane (degree = 101), nonacosane (degree = 99), and hentriacontan (degree = 99), as shown in Figure 2A.

Networks of SGNI in liver protection. (A) Compound-target network of SGNI in liver protection. Triangles represent the compounds, and red diamonds represent the targets. The light purple triangles represent the Artemisia capillaries, green triangles represent the Gardeniae Fructus, dark purple triangles represent the Baicalin, blue triangles represent the Isatidis Radix, and yellow triangles represent the Ganoderma. (B) Protein-protein interaction network of SGNI in protecting liver. A node represents a target. The color of the node represents the value of the degree. The larger the value, the darker the color. (C) Core compound-target network of SGNI. The blue circles represent the compounds, and the red rectangles represent the targets.
Analysis of the Protein-Protein Interaction Network
Construction of the Protein-Protein Interaction (PPI) network was carried out in the present study using 408 targets of SGNI (Figure 2B). It was found that there were 341 nodes and 1888 edges in the network. Furthermore, a total of 60 targets with degree and betweenness centrality above average were screened out. Among them, the top 5 targets were amyloid-beta precursor protein (APP, degree = 56), tumor protein p53 (p53, TP53, degree = 49), proto-oncogene tyrosine-protein kinase Src (SRC, degree = 46), transcription factor Jun (JUN, degree = 43), and nuclear factor NF-kappa B p65 subunit (RELA, degree = 40). In addition, a total of 82 compounds and 97 targets with degree and betweenness centrality above average were obtained after merging the PPI network and “compound-target” network. Construction of a “core compound-target” network was then conducted after adding the main active compounds and targets reported in the literature. Therefore, a total of 60 active compounds and 131 targets were obtained (Figure 2C). The analysis of the parameters using a network analyzer, and the top 10 compounds and targets were as presented in Table 1. The structural formulas of the top 10 of the core compounds are shown in Figure 3.

The structural formulas of the top 10 core compounds and the core active compounds of SGNI. A, B, C, D, E, F, G, H, I, and J were the top 10 core compounds, and K, L, M, N, and O were the active compounds of SGNI.
Top 10 of the Core Compounds and Targets of SGNI in Protecting Liver.

Each number represents a compound of herbs in Supplementary Table S1 with the corresponding number. The degree value represents the number of nodes connected by a node, and the higher the degree, the more important the point is in the network.
Abbreviations: AR, androgen receptor; BLG, Isatidis Radix; CHRM1, muscarinic acetylcholine receptor M1; ESR1, estrogen receptor; JUN, transcription factor Jun; LACC1, laccase domain-containing protein 1; LZ, Ganoderma; NOS2, nitric oxide synthase, inducible; PRKACA, cAMP-dependent protein kinase catalytic subunit alpha; PTGS1, prostaglandin G/H synthase 1; PTGS2, prostaglandin G/H synthase 2; RELA, nuclear factor NF-kappa B p65 subunit; SGNI, Shuganning injection; YC, Artemisia capillaries; ZZ, Gardeniae Fructus.
Gene Ontology Analysis and Kyoto Encyclopedia of Genes and Genomes Pathway Enrichment Analysis of SGNI
Gene Ontology (GO) analysis identified 1532 biological processes (BPs), 44 molecular functions (MFs), and 29 cellular compounds (CC). In BP, the targets mainly involved the positive regulation of transcription from RNA polymerase II promoter, the positive regulation of nitric oxide biosynthetic process, negative regulation of apoptosis process, and inflammatory response. In CC, the targets mainly involved the plasma membrane, cytosol, cell surface, and synapse. In MF, the targets mainly involved enzyme binding, transcription factor binding, and nitric-oxide synthase regulator activity.
Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis in the current study revealed that the targets were closely related to the pathway in cancer, nitrogen metabolism, hepatitis B signaling pathway, sphingolipid signaling pathway, hepatitis C signaling pathway, NF-kappa B(NF-kB) signaling pathway, tumor necrosis factor (TNF) signaling pathway, nonalcoholic fatty liver disease (NAFLD) signaling pathway, and Toll-like receptor (TLR) signaling pathway.
Among these pathways, it was noted that hepatitis B signaling pathway was involved in 17 targets, including retinoblastoma-associated protein (RB1), JUN, chemokine (C-X-C motif) ligand 8 (CXCL8), signal transducer and activator of transcription 1 (STAT1), SRC, protein kinase C alpha type (PRKCA), fos proto-oncogene (FOS), TNF, matrix metalloproteinase-9 (MMP9), phosphoinositide-3-kinase catalytic gamma polypeptide (PIK3CG), RELA, nuclear factor NF-kappa-B p105 subunit (NFKB1), cyclin-A2 (CCNA2), Caspase 8 (CASP8), G1/S-specific cyclin-D1 (CCND1), Caspase 3 (CASP3), and TP53. TNF signaling pathway was involved in 13 targets, including JUN, endothelin-1 (EDN1), vascular cell adhesion protein 1 (VCAM1), FOS, prostaglandin G/H synthase 2 (PTGS2), mitogen-activated protein kinase 14 (MAPK14), TNF, MMP9, PIK3CG, RELA, NFKB1, CASP8, and CASP3. Sphingolipid signaling pathway was involved in 13 targets, including delta-type opioid receptor (OPRD1), nitric oxide synthase, endothelial (NOS3), PRKCA, MAPK14, TNF, PIK3CG, RELA, NFKB1, adenosine receptor A3 (ADORA3), adenosine receptor A1 (ADORA1), tyrosine-protein kinase Fyn (FYN), sphingosine 1-phosphate receptor 2 (S1PR2), and TP53. TLR signaling pathway was involved in 11 targets, including JUN, CXCL8, CASP8, STAT1, TLR9, FOS, MAPK14, TNF, RELA, NFKB1, and PIK3CG. The NF-kB signaling pathway was involved in 8 targets, including VCAM1, CXCL8, Bcl-2-related protein A1 (BCL2A1), urokinase-type plasminogen activator (PLAU), PTGS2, TNF, RELA, and NFKB1. It was also found that visualization of 20 critical items is associated with the hepatoprotective effect of SGNI, as shown in Figure 4. Furthermore, the top 3 pathways were manually integrated shown in Figure 5.
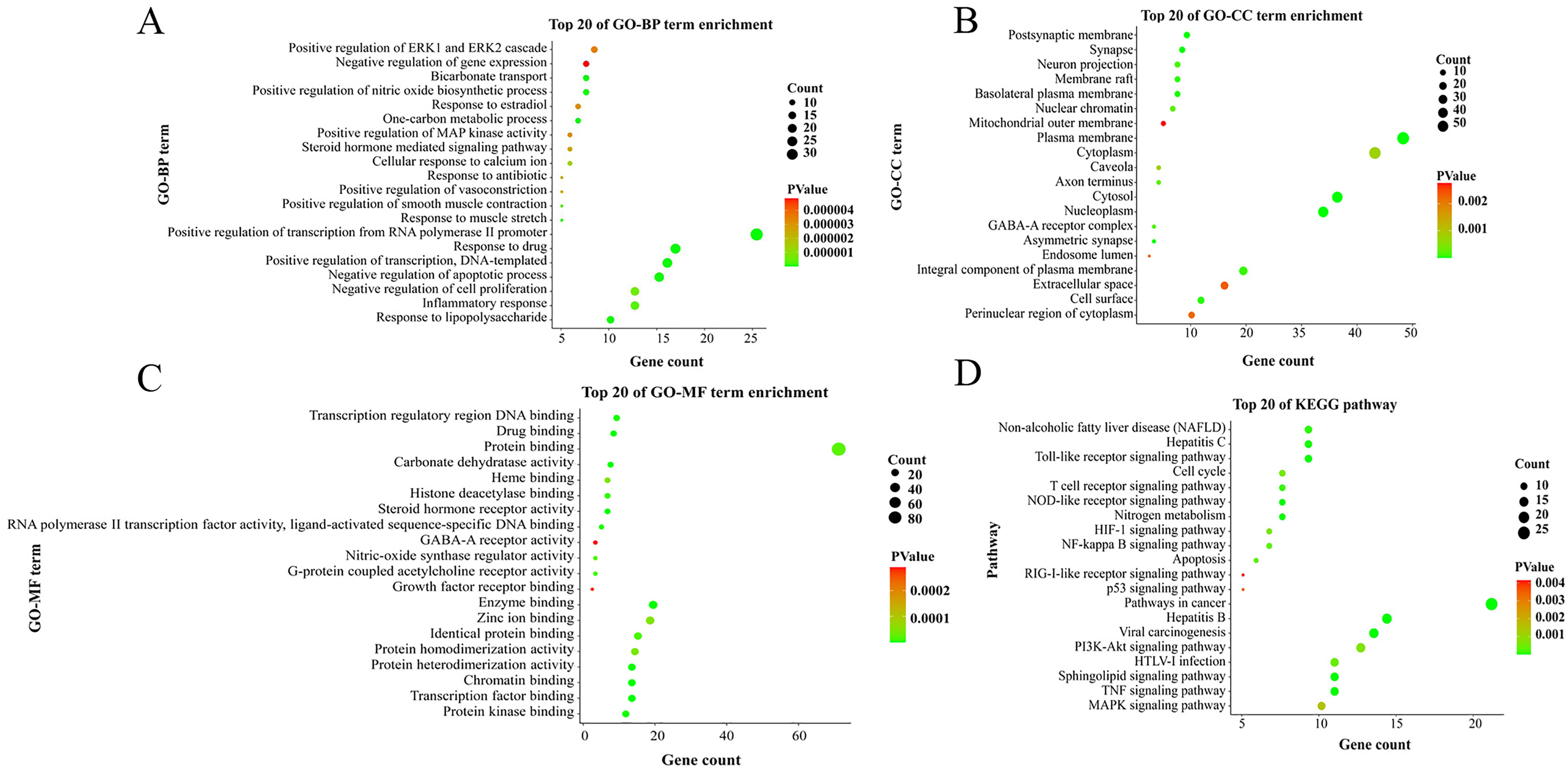
Bubble diagram of GO enrichment analysis and KEGG pathway analysis. (A) Gene ontology biological processes (GO-BP) analysis of SGNI. (B) Gene ontology cellular compound (GO-CC) analysis of SGNI. (C) Gene ontology molecular function (GO-MF) analysis of SGNI. (D) Bubble diagram of the KEGG pathway enrichment analysis of SGNI.

Distribution of target proteins of SGNI on the compressed “hepatoprotective effects.” Three pathways from the compressed hepatoprotective pathway. Arrows represent activation effect, T-arrows represent inhibition effect, and segments show activation effect or inhibition effect.
Molecular Docking Analysis
Autodock is an open-source molecular simulation software, which is mainly used to perform ligand-protein molecular docking. In the present study, 1 active compound of each herb was selected to be docked with 7 targets using AutoDock1.5.6 software. The binding energy was less than 0, which indicated that these active compounds could be combined with the core targets. Among them, chlorogenic acid bound well to JUN, TP53, and CASP3. Furthermore, baicalin bound well to TP53, NFKB1, and CASP3, whereas indirubin bound well to RELA, JUN, TP53, TNF, CASP3, CASP8, and NFKB1. In addition, ganoderma acid A bound well to the target proteins of JUN, TP53, CASP3, CASP8, and NFKB1. The docking results are shown in Figure 6.

The result of molecular docking of SGNI in liver protection. (A) Molecular docking pattern of potential compounds and targets of SGNI. (B) The heatmap of molecular docking results of SGNI act on liver protection. Color plot indicates scale of correlation strength.
Experimental Verification
Effects of SGNI on the Proliferation of Acetaminophen-Injured Cells
The acetaminophen (APAP)-induced liver injury in HepaRG cells in a dose-dependent manner, as shown in Figure 7. The 50% inhibitory concentration of APAP was 23.44 mM, whereby 25 mM of APAP was used for subsequent experiments.
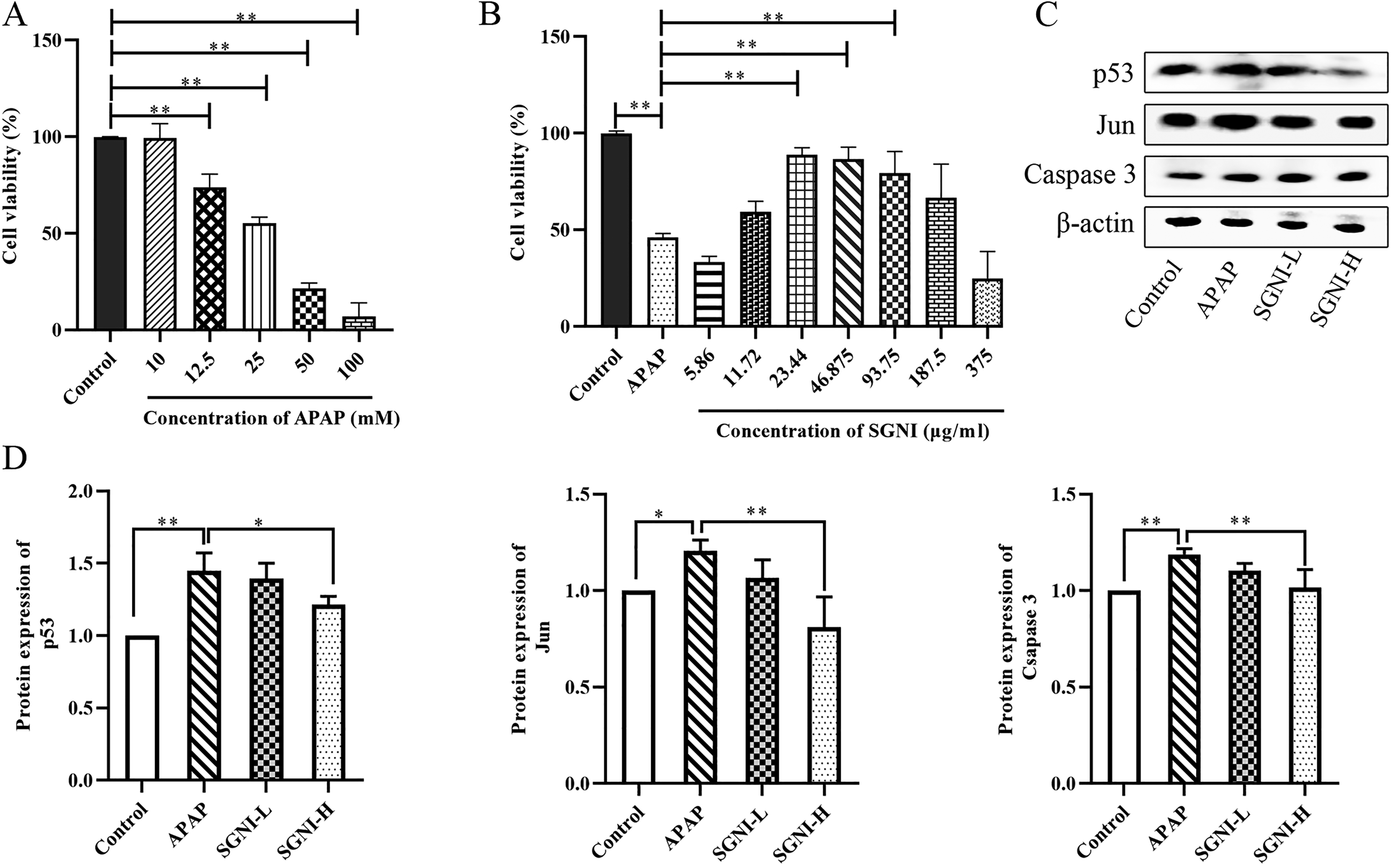
The result of experimental verification of SGNI in liver protection. (A) Viability of HepaRG cells following exposure to APAP (the y-axis represents percentage of inhibition, and the x-axis indicates inhibitor concentration [mM]). The values are expressed as the mean ± SD (n = 3 *P < 0.05, **P < 0.01). (B) Protecting effect of SGNI in the APAP-induced liver injury model (the y-axis represents percentage of inhibition, and the x-axis indicates inhibitor concentration [μg/ml]). The values are expressed as the mean ± SD (n = 3 *P < 0.05, **P < 0.01). (C) Western blot image showing p53, Jun, and Caspase 3 expression in HepaRG cells. (D) Expression of p53, Jun, and Caspase 3 by western blot assay. The values are expressed as the mean ± SD (n = 3, *P < 0.05, **P < 0.01).
The APAP group showed lower cell viability compared with the control group, whereas the cell viability was increased in the groups treated with 11.72, 23.44, 46.875, 93.75, and 187.5 μg/ml of SGNI than in the APAP group. Based on this, it was found that the concentration of SGNI was 11.72 μg/ml in the low-dose group (SGNI-L) and 23.44 μg/ml in the high-dose group (SGNI-H), as shown in Figure 7
Effects of SGNI on Key Proteins
As shown in Figure 7, it was found that there were significantly increased protein expressions of p53, Jun, and Caspase 3 in the APAP group as compared with the control group (P < 0.05 or P < 0.01). On the other hand, it was noted that there was a significant decrease in protein expressions of p53, Jun and Caspase 3 in the SGNI group (P < 0.05 or P < 0.01) after treatment with different doses of SGNI.
Discussion
Although hepatoprotective effect of SGNI for acute and chronic viral hepatitis, liver cirrhosis, and liver failure has been previously reported, the mechanism of SGNI is not understood. In the present study, network pharmacology and molecular docking were performed to screen the active compounds and targets of SGNI. Furthermore, the mechanism of action of SGNI was validated using in vitro experiments in this study.
Analysis of the Core Active Compounds of SGNI
Quercetin, chlorogenic acid, indirubin, baicalin, ursolic acid, geniposide, and ganoderic acid A were the core active compounds of SGNI involved in liver protection through the network analysis, and the structural formulas of core active compounds of SGNI are shown in Figure 3. Of these compounds, it was evident that quercetin and chlorogenic acid exerted their hepatoprotective effect through their antioxidant, and anti-inflammatory properties, 15 and chlorogenic acid inhibited expression of NF-kB and activator protein-1 (AP-1) alleviating acute liver injury caused by CCL4. 16 It was found that chlorogenic acid combined with geniposide improved fatty liver, which confirmed the good curative efficacy of SGNI protecting the liver by the synergy of a variety of TCM. 17 It was evident baicalin was the only specific monomer compound of SGNI with a definite hepatoprotective effect. In cooperation with the active compounds of the monarch and minister drugs, baicalin played a role in the anti-inflammatory action. Previous studies have shown that baicalin had hepatoprotective effects against several types of liver diseases, including viral hepatitis, xenobiotic induced liver injury, fatty liver disease, cholestatic liver injury, and hepatocellular carcinoma. 18 Similarly, baicalin could inhibit the message RNA levels of inflammatory cytokines TNF-α, interleukin-1β, PTGS2, Caspase 3, and Caspase 11. Furthermore, it has been reported that baicalin inhibits cell apoptosis of Zearalenone-induced liver injury in chicks. 19 In addition, other compounds of SGNI such as indirubin, ganoderma acid A, and ursolic acid exerted hepatoprotective effect mainly related to the synergistic anti-apoptotic and anti-inflammatory effects.20–24 Finally, the hepatoprotective effect of the studied compounds, such as chlorogenic acid and indirubin were validated through molecular docking.
Analysis of Molecular Mechanism of SGNI
KEGG pathway analysis in the current study revealed the targets of SGNI correlated with hepatic pathways, such as hepatitis B signaling pathway, TLR signaling pathway, NF-kB signaling pathway, hepatitis C, and NAFLD signaling pathway. In addition, it was evident that seven targets were involved in most of the 20 critical pathways, including RELA, JUN, TP53, TNF, CASP3, CASP8, NFKB1, and three targets were highly related to active compounds in molecular docking results, including TP53, JUN, and CASP3, which were verified through in vitro experiments.
The p53, as a tumor suppressor and apoptotic factor, is widely expressed in various liver injuries through the regulation of DNA damage and oncogenes. 25 Expression of p53 increases significantly with the generation of oxidative stress in the liver. 26 For APAP-induced liver injury, p53 could activate apoptosis signal pathways, regulate apoptosis regulator BAX, cyclin-dependent kinase inhibitor 1, apoptosis regulator Bcl-2, Caspase 3 and other related proteins, and induce apoptosis in hepatocytes.27–29 In the present study, it was evident that the protein expression of p53 was decreased following SGNI treatment compared with the APAP groups, indicating that SGNI can protect the liver through down-regulating the p53 protein expression and further reducing the apoptotic progress. Caspase 3 acts as an apoptosis executive protein in the caspase family. It is associated with cell death and pro-inflammatory signal transduction, 30 and its main pathway is apoptosis induced by mitochondria pathway. 31 Typically, it has been reported the activation of Caspase 3 is associated with liver injury. In the present study, the protein expressions of Caspase 3 were significantly decreased following SGNI treatment compared with the APAP groups. This indicated that SGNI can protect liver injury through the downregulation of the Caspase 3 protein expression and thus further reduction of the apoptotic progress. Jun, which is also known as c-Jun, belongs to the bZIP family and Jun subfamily. It is the most active transcription factor of the AP-1 transcription factors. Jun is involved in regulating inflammation, apoptosis, and proliferation, as well as plays an important regulatory role in a variety of liver diseases. 32 A decrease in expression of Jun leads to damage to liver and caused the depletion of mitochondrial glutathione as well as amplifies the oxidative stress and mitochondrial dysfunction.33,34 The results of the present study showed that the protein expressions of Jun were significantly decreased after the treatment of SGNI as compared with the APAP groups, indicating that SGNI could protect liver injury through downregulation of the Jun protein expression further inhibit jun-mediated oxidative stress.
The results of pathway enrichment revealed that the molecular signaling pathways of SGNI in protecting liver were closely related to inflammation-based pathways, such as TNF signaling pathway, TLR signaling pathway, and NF-kB signaling pathway. It was evident that TNF signaling pathway is the key among the pathways in liver injury, and TNF is a major mediator of immunity, apoptosis and inflammation. 35 It has been reported that the activation of TNF signaling pathway is involved in NF-kB transduction mechanisms. 36 Similarly, NF-kB signaling pathway is crucial for liver diseases, involving multiple important BPs, such as inflammatory response, and apoptosis.37,38 All chronic liver diseases lead to the activation of factors such as NF-kB or AP-1, and mediated activation of NF-kB signaling pathway.39,40 In the current study, it was evident that SGNI may protect liver by attenuating inflammation through inhibition of TNF and NF-kB signaling pathways. In addition, TLRs are important regulators of immune response, such as TLR2, TLR4, and TLR7, which are involved in the immune response to hepatitis B virus infection. TLRs could hence wound healing and regeneration. Furthermore, TLRs are also involved in the pathogenesis and progression of various inflammatory liver diseases. And most TLRs stimulated the expression of NF-kB and AP-1 by mediating the MyD88-dependent pathway, ultimately leading to increased production of various pro-inflammatory cytokines.41,42 As described earlier, SNGI may inhibit the release of inflammatory factors through the TNF, TLR, and NF-kB signaling pathway, and protect liver. These results of the present study are preliminary findings showing that the hepatoprotective effects of SGNI were related to anti-inflammatory, antioxidant, and apoptosis.
In summary, it is evident that quercetin, chlorogenic acid, baicalin, indirubin, ganoderic acid A, as well as RELA, JUN, TP53, and CASP3 were the core active compounds and targets, respectively. Moreover, related active compounds and targets may play a role through the Hepatitis B signaling pathway, NF-kB signaling pathway, TLR signaling pathway, and TNF signaling pathway. The inhibition of the expression of p53, Jun, and Caspase 3 proteins by SGNI, occurs through anti-inflammatory, anti-apoptotic, and anti-oxidant actions alleviating APAP-induced liver injury in HepaRG cells. Therefore, this verified the feasibility and reliability of the prediction of network pharmacology. However, there is still a need for further studies on the specific mechanism of SGNI.
Conclusion
In conclusion, SGNI has the characteristics of the synergism of multi-compounds, multi-targets, and multi-pathways, producing a superior efficacy, which precisely reflects the holistic view of TCM and the principle of syndrome differentiation and treatment. In this paper, the core active compounds and targets of SGNI were explored through network pharmacology, the binding effect of the core active compounds and targets through molecular docking. Finally, the expressions of p53, Jun, and Caspase 3 proteins were detected to explore active compounds and mechanism of SGNI against APAP-induced liver injury in HepaRG cells, providing a reference for the clinical medication value of SGNI.
Materials and Methods
Materials
SGNI was obtained from Guizhou Ruihe Pharmaceutical Co., Ltd (Guizhou, China); APAP (Aladdin, China); HepaRG cells (GuanDao Biologic, China); Fetal bovine serum (Giboco, USA); RPMI-1640 medium (Giboco, USA); Cell Counting Kit-8 (CCK-8) reagent (Dojindo, Japan); antibody: anti-β-actin (Abbkine, China); anti-p53 (Proteintech, China); anti-Jun (Jun) (Proteintech, China); anti-Caspase 3 (Proteintech, China); Goat anti-Mouse IgG-HRP antibody Goat (HuaBio, China).
Methods
Screening of Active Compounds and Targets of SGNI
Screening of active compounds of Artemisia capillaries, Gardeniae Fructus, Baicalin, Isatidis Radix, and Ganoderma from literature and Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP, https://tcmspw.com/index.phph), and met the standard of drug-likeness (DL) ≥ 0.18%.17,43–47 However, the value of oral bioavailability was not considered due to injection. Targets were then predicted from two sources: (1) TCMSP database; (2) Targetnet (http://targetnet.scbdd.com/), with prediction accuracy ≥ 0.7, predicted probability > 0.9. Finally, all the targets were converted to gene ID by Uniprot database, and the species was set to Homo sapiens (https://www.uniprot.org/).
Prediction of Hepatoprotective Targets of SGNI
All the targets related to liver injury were collected from the Genecards disease database (https://www.genecards.org/), Online Mendelian Inheritance in Man disease database (https://omim.org/), and Comparative Toxicogenomics Database (http://ctdbase.org/). The keywords used were “Liver Injury,” and “Hepatitis.” Subsequently, the SGNI and disease-related targets were intersected using Venny2.0, and construction of the “compound-target” network using Cytoscape 3.7.2 software.
Construction of the PPI Network
After obtaining the retrieved intersection targets, and construction of the PPI network using the STRING11.0 (http://string-db.org), with Homo species as a requisite, the interaction score > 0.9, then visualized and analyzed using Cytoscape 3.7.2 software. Subsequently, construction of the core PPI network was by filtering out targets with the degree and betweenness centrality above the PPI network average.
Construction of “Core Compound-Target” Network and Enrichment Analysis
The “compound-target” and “core PPI” networks were merged in Cytoscape 3.7.2 software. Then, the top 40 targets and active compounds with degree and betweenness centrality above average combined with the corresponding targets of SGNI were reported in the literature to constitute a “core compound-target” network.
To characterize the functional role of the core targets in relevant pathways, GO analysis and KEGG pathway enrichment analysis were performed using the Database for Annotation, Visualization, and Integrated Discovery bioinformatics resource database (https://david.ncifcrf.gov/). Data were presented as Homo sapiens for the selected species and at P < 0.01. Visualization of the data is done using imageGP (http://www.ehbio.com/ImageGP/) software.
Molecular Docking
One active compound of each herb from SGNI was selected for molecular docking with seven targets after integrative analysis of the literature, the “core compound-target” network, and the KEGG pathways. The crystal structures of protein were obtained from the Protein Data Bank (http://www.rcsb.org/). In addition, PyMol software was used to process the protein, including removing the water molecules and other ligands of the protein structure. The Mol2 molecular structures of compounds were acquired from the TCMSP database and ChemBio 3D Uitra14.0 software for energy minimization. The AutoDock1.5.6 software was used to modify the compound structures, including hydrogenating the proteins, calculating the charges, adding the protein types, and detecting paths. Finally, molecular docking was performed with AutoDock1.5.6 software and the docked models were analyzed using PyMol. Binding energy < 0 indicated that the ligand and the protein could bind spontaneously; the lower the binding energy, the better the docking was affected. The binding energy ≤−5.0 kcal/mol indicated that the compound binds well to the target.
APAP-Induced Live Injury in HepaRG Cells
The HepaRG cells were cultured at 37 °C in 5% CO2 and saturation humidity in RPMI-1640 medium containing 10% fetal bovine serum.
The HepaRG cells in the logarithmic growth stage were inoculated into 96-well plates at a density of 1.5 × 105 cells/ml. The cells were treated with different concentrations of APAP, 0, 10, 12.5, 25, 50, and 100 mM. After the cells were adhered onto the wall for 12 h, 100 μl of CCK-8 reagent was added to each well and incubated in the incubator for 2 h. Absorbance was measured with a microplate reader at 450 nm, and the relative cell viability was calculated. The 50% inhibitory concentration corresponding to the APAP concentration was used for subsequent experiments.
Experimental Groups and Intervention of SGNI
The HepaRG cells were randomly divided into control group, APAP group (APAP 25 mM), and SGNI treatment groups (5.86, 11.72, 23.44, 46.875, 93.75, 187.5, and 375 μg/ml). The cell in the control group was treated with culture medium, and the APAP and SGNI treatment groups were treated with 25 mM APAP for 12 h, respectively. After the cells adhered to the wall, 100 μl of CCK-8 reagent was added to each well and incubated for 2 h. Absorbance at 450 nm was measured with a microplate reader, and the relative cell viability was calculated. The concentration of SGNI without overt toxicity was chosen for further experiments in the present study.
Western Blotting Analysis
Protein expression of p53, Jun and Caspase 3 in HepaRG cells after SGNI treatment were determined through western blotting analysis to evaluate the accuracy of the network predicted and explore the mechanism of SGNI in protecting liver based on APAP-induced live injury in HepaRG cells.
Cells were placed on ice and washed with pre-cooling phosphate-buffered saline 2 times, and 500 μl cell lysate was added (RIPA lysate:PMSF protease inhibitor:phosphatase inhibitor = 100:1:1). Then, the cells were scraped while shaking for 30 min; collected into Eppendorf tubes; and centrifuged at 1200g at 4 °C for 20 min. The supernatant was recovered, and protein was quantified using the BCA method. In addition, the protein was denatured at 100 °C for 15 min. Samples were run on 10% acrylamide gels and transferred to 0.45 μm PVDF membrane (the concentration gel was 80 V, whereas the separation gel was 120 V). Afterwards, the membrane was blocked in 5% milk in TBST for 1 h, which was then followed by an overnight incubation with primary antibody (β-actin (1:10 000), p53 (1:2000), Jun (1:1000), Caspase 3 (1:1000)) in 4 °C and incubation with secondary antibody for 1 h at room temperature. Furthermore, liquid A and B mixture of luminescence was evenly dropped onto the membrane, which was placed in a Bio-Rad gel imaging system for imaging. Finally, the ImageJ software was used to perform grayscale analysis of protein bands.
Data Processing and Statistical Methods
Data obtained from the current study were expressed as the means ± standard deviation. Graphpad prism 8.0 software and SPSS 18.0 software were used to process the experimental data. The univariate analysis of variance was used to compare the differences in the means among the groups. The statistical significant difference was set at P < 0.05 or P < 0.01.17,43–47
Supplemental Material
sj-docx-1-npx-10.1177_1934578X221124756 - Supplemental material for Active Compounds Screening and Hepatoprotective Mechanism of Shuganning Injection Based on Network Pharmacology and Experimental Validation
Supplemental material, sj-docx-1-npx-10.1177_1934578X221124756 for Active Compounds Screening and Hepatoprotective Mechanism of Shuganning Injection Based on Network Pharmacology and Experimental Validation by Qiyi Wang, Xiaotong Duan, Shan Li, Huaqing Lai, Weina Cheng, Jingwen Ao, Jianyong Zhang and Cancan Duan in Natural Product Communications
Footnotes
Authors’ Contributions
QYW, XTD, SL, CCD, and HQL conceived and designed the experiment. QYW and XTD performed the experiments. WNC, CCD, and JWA analyzed the data. QYW wrote the paper. All of the authors reviewed and approved the submitted version of the paper.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Qian Jiao He (grant number: YJSKYJJ [2021]183).
Data Availability
The data used to support the findings of this study are included within the article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
