Abstract
COVID-19 mainly causes the collapse of the pulmonary system thereby causing a dearth of oxygen in the human body. Patients infected with this viral disease have been reported to experience various signs and symptoms associated with brain dysfunction, from the feeling of vagueness to loss of smell and taste to severe strokes. These neurological problems have been reported by younger COVID-19 infected patients mainly in their thirties and forties. Various researchers from around the globe have discerned numerous other brain dysfunctions, such as headache, dizziness, numbness, major depressive disorder, anosmia, encephalitis, febrile seizures, and Guillain-Barre syndrome. The involvement of the CNS by this viral infection has been predicted to be for a longer period of time, even if the patient recovers from COVID-19. The neuronal cell damage caused by COVID-19 is a potent factor responsible for cognitive, behavioral, and psychological problems among its sufferers. The hypoxic conditions can also trigger the formation of beta-amyloid plaques and tau-tangles and thus the virus can even induce Alzheimer’s in patients in the near future. The virus affects the brain directly, thereby causing encephalitis. This pandemic has also been shown to have a negative psychological toll on people. This research aims to highlight the brain dysfunction associated with the ACE2 receptor that is known to be a crucial player in the COVID-19 pandemic using genetic networking approaches. Furthermore, we have identified herbal drug candidates that bind to the ACE2 receptor in order to identify potential treatments for the neurological manifestations of COVID-19.
Introduction
COVID-19 is a non-bigotry viral infection that is transmitted in humans of all age groups and is responsible for causing serious respiratory tract infections, in severe cases, even leading to death1, 2. It originated initially in Wuhan city of China in 2019 where it was discerned to be of Beta-BAT-SARS-CoV-2 lineage; however, it has become a source of distress globally since then 3 . Various online social platforms and the global research fraternity have produced humongous data in myriad aspects namely—social, technological, global news, healthcare, political domain, vaccination discovery, and drive, helping to understand the myriad aspects of the ongoing pandemic. Quite recently, many researchers have found that COVID-19 has negative implications on various neurological disorders. Different strains of coronaviruses have been shown to cause neurotropism and neuro-invasive symptoms leading to various brain dysfunctions in COVID-19 affected populations 4 .
SARS-CoV-2 belongs to the Nidovirales order. Common features of the viruses belonging to this category are—(i) conserved and highly organized genomic contents - the replicase gene comes first, followed by the structural and accessory genes; (ii) non-structural gene expression that is caused by ribosomal frameshifting; (iii) enzyme dynamics taking place within the replicase-transcriptase polyprotein; and (iv) 3′-subgenomic mRNA formation resulting in downstream gene expression 3 . Until last year, COVID-19 was thought to be fatal for elderly people or for individuals with co-morbidities. However, in early 2021, various strains were identified that were tagged to be catastrophic for younger people and infants too. As per the CDC report, there are mainly 5 variants of concern (VoC) of COVID-19 that cause severe damage namely - a) B.1.1.7 (initially detected in South Africa, but severely affected the US in early 2021), b) B.1.351 (affected the US in early 2021, but was first rectified in South Africa), c) P.1 (was first observed in travellers from Brazil who tested positive in initial COVID-19 screening in Japan; this variant also caused severe damage in the US population, as reported in January 2021), d) B.1.427 and B.1.429 (these were first identified in California in February 2021 and were classed as a variant of concern (VoC) in March) 5 .
Several researchers have worked hard to identify potential drugs that can treat COVID-19 and related maladies. Today, we have many verified synthetic drug candidates such as riboflavin, lopinavir, oseltamivir, lopinavir/ritonavir 6 , minocycline, tocilizumab, ribavirin, and niclosamide7, 8. Moreover, many natural drug candidates have also been utilized to treat COVID-198–10. However, there are no herbal compounds that can help in curing the neurological manifestations of COVID-19. Several recently published studies have revealed that after getting infected with SARS-CoV-2, various different neurological dysfunctions, such as stroke, brain stem impairments, neuroinflammation, brain fog, and delirium were observed in patients 11 . SARS-CoV-2 has also been revealed to deploy angiotensin-converting enzyme 2 (ACE2) as an entry point to the cells. This simply means that SARS-CoV-2 binds to human ACE2 for entry and attachment inside the host cells 12 . The expression of ACE2 in human neurons suggests the neuro-encroaching strength of SARS-CoV-2, which further reassures that the infected patient's neuronal system could affect the overall respiratory function 13 . Additionally, ACE2 is known to be the receptor for SARS-CoV-1 as well as that for SARS-CoV-2. Reports also discern that ACE2 is expressed in neural cells which in turn allows SARS-CoV-1 to cause brain dysfunctions in humans 14 .
This research work aims to highlight the neurological dysfunctions that are associated with the ACE2 receptor active in COVID-19. Also, using epinformatics-based network analysis, we have identified the direct network associates of ACE2, and thus have established why ACE2 is responsible for COVID-19 related neurological manifestations in individuals. By deploying a gene-drug mapping approach, we have highlighted a few synthetic and alternative herbal drug candidates that can be seen as possible treatment strategies for treating the brain dysfunctions associated with COVID-19. Additionally, we executed pharmacological screening, molecular docking, and essential electrostatics analysis on the best suitable phytochemicals to validate their usefulness as a therapeutic and stable affinity with the ACE2 receptor.
Methods & Methodology
Retrieval of Sequence
We used the Angiotensin-converting enzyme 2 (ACE2) which has been discerned to be the entry receptor for SARS-CoV-2 from the National Center of Biotechnology Information (NCBI) (https://www.ncbi.nlm.nih.gov/), (accession number NM_001371415.1 and Gene ID: 59272). The protein sequence of ACE2 was also retrieved (accession ID NP_001358344.1) from Protein Data Bank (PDB Id: 7RPV). This is the crystal structure of the affinity-enhancing and catalytically inactive ACE2 in complex with SARS-CoV-2 RBD, respectively. The ACE2 receptor was separated from the complex of spike protein (S1), 2-acetamido-2-deoxy-beta-D-glucopyranose, and attached zinc ion using Pymol.
Motif Identification, Enrichment & Annotation Analysis
This was executed using CTCFBSDB software15, 16; while for enrichment analysis, we deployed the MEME suite's CentriMo software 17 and comparison TomTom software, available in MEME suite 18 . For genomic annotation purposes, we used Rapid Annotation using Subsystem Technology (RAST) software 19 and curated a pipeline that can screen for seed genes and essential genomic annotations for ACE2 transcript files (NCBI Gene ID: 59272). We used the updated annotation scheme named RASTk with call features repeat region SEED minimum identity as 95 and a minimum length of 100; CDS-glimmer3 was used for a minimum training length of 2000. Along with this, we used the kmer dataset release70 for annotation purposes.
Genetic Network Analysis
The ACE2 protein sequence (NP_001358344.1) was deployed to the WebMGA 20 web server to identify the cluster of orthologs to annotate its function. The network interactors were identified using STRING software 21 by tuning the network parameter settings for displaying only the physical network which has a good confidence score taken from experimentally identified sources, literature, and text mining approaches. The network was classified using the k-means clustering algorithm.
Gene-Drug Mapping
We used DGIdb 22 , DrugBank 23 , and PubChem 24 to explore the potential drug candidates that can serve the purpose of treating COVID-19. The initial criteria to search for the drugs were (a) having a strong interaction with the target seed genes, (b) interaction score should be ≥ 5.0, and (c) the identified drugs must be reported in the literature. We executed it in a keyword-based search strategy method, where identified seed genes were separately searched in the databases.
Identification of Alternative Herbal Compounds as Potential Drug Candidates
An extensive literature search was executed using PubMed (https://www.ncbi.nlm.nih.gov/pmc/) to identify alternative herbal compounds as potential drug candidates.
Pharmacological Screening Using ADMET
The identified phytochemicals were retrieved from ChEMBL 25 and stored in SMILES format (Simplified molecular-input line-entry system). SwissAMDE webserver 26 was used to assess the phytochemicals for their physicochemical, pharmacokinetic (PK), pharmacodynamic (PD), medicinal chemistry, solubility, and lipophilicity properties.
Molecular Docking
Binding Site Identification
PrankWeb 27 was used to identify binding pockets within the target ACE2 receptor.
Preparation of Target and Ligand Files
PyRx software 28 was used for virtual screening and docking purposes. Using the software, polar charges were added to the target receptor ACE2 and were checked for any additional co-factors/metals/ions bound with the receptor. If any, they were removed and then converted from .pdb to. pdbqt. Similarly, the ligands were first energy minimized by conjugant gradient optimization algorithm with the universal forcefield in the background. After minimization, the ligands were saved from the. mol/.sdf to. pdbqt files. Multi-drug docking took place with the grid box set at 60 Å × 60 Å × 60 Å and a spacing of 0.375 Å. The total number of steps was set as 200 with stopping criteria of energy difference less than 0.1.
Electrostatics Computation
Molecular mechanics generalized Born surface area (MM-GBSA) was used to compute the binding free energy (delta G) using the adaptive Poisson-Boltzmann solver (APBS) plugin available in PyMOL software (https://pymolwiki.org/index.php/APBS_Electrostatics_Plugin). Moreover, other electrostatic features were also calculated using Bluues software, 29 and macromolecular energy landscape was computed using the Frustratometer webserver 30 .
Results
Motif Identification & Enrichment
Myriad motifs have been already identified in the past year in the Wuhan-based SARS-CoV-2 genome. However, we aimed at identifying the motifs that were present in the main entry receptor ACE2 that has been identified to play a direct role in neurological dysfunction caused by SARS-CoV-2. Therefore, we used CTCFBSDB software that works on the CCCTC-binding factor (CTCF), a highly conserved transcription regulator that is found in all organisms ranging from Drosophila melanogaster to Homo sapiens and is said to associate itself with DNA sequences using the 11-zinc fingers. These binding fingers have been said to play a pivotal role in the adaptive divergence of DNA sequences. Researchers have reported these factors to be associated with myriad epigenetic mechanisms such as genomic imprinting and X-chromosome inactivation15, 16.
We deployed the ACE2 receptor mRNA sequence (accession Id—NM_001371415.1) to CTCFBSDB and identified six major motifs. Table 1 displays the detailed results of the motif identification using the CTCFBSDB software. Only two motifs were found to be significant, namely motifs M2 and M5, based on their high scores. Motif M2 (AGTACTGCC, score = 10.6676) is present on the negative strand and has a length of 9 nucleotides, while motif M5 (AGTACTGTAGATGGTGCTCA, score = 11.8863) is also located on the negative strand and has a length of 20 nucleotides. Since these two motifs had higher scores, we proceeded with our motif enrichment analysis using these only. After enrichment analysis, we discerned that the two motifs- M2 and M5 are highly conserved in the mouse genome and there are about 579 non-redundant motifs that are present matching our identified motifs in the JASPAR database 31 , while 386 similar matches were observed in the Uniprobe mouse database 32 . Also, A-T nucleotide content was found to be 0.2908 while C-G was 0.2092.
Six major motifs present in the ACE2 mRNA sequence.

For comparing the two motifs M2 and M5, we subjected them to MEME suite's TomTom server that predicted two different matches for each of the two motifs. Table 2 represents the detailed results of the motif comparisons.
Motif enrichment of the best-identified ACE2 mRNA sequence (NM_001371415.1).

It was observed that these compared motifs were matched to zinc fingers, DNA-binding domains, heart and neural crest derivatives, tRNA-methyltransferases, and paired like homeoboxes present in the human, mouse, and yeast genome.
The genomic annotation revealed some essential characteristics and information such as the closest neighbours, GC content that directly affects the genome functioning, and species ecology of the ACE2 gene. In total, 280 characteristic features were identified in the ACE2 gene. The GC content was observed to be 38.5% displaying the quadratic relationship with the genomic size of the ACE2 gene. The analogy is simple - the larger the genomic size, the lower the GC content viz., because of the higher biochemical expenditure of GC base formation. There were 15 out of the total 280 characteristic features that were present in the ACE2 gene present mainly in the coding sequence regions. Table 3 encapsulates the detailed report of the 15 best features retrieved, as identified by our genomic annotation.
Best 15 characteristic contigs present in the ACE2 gene (Gene ID: 59272).

Network Analysis
We employed the protein sequence of ACE2 (NP_001358344.1) for network analysis. Our main objective was to identify the cluster of orthologs and accordingly interpret the functional annotation of the ACE2 receptor. We identified 25 clusters of orthologs (COGs) for our ACE2 protein that play important biological roles in various cellular processes such as - cell cycle regulation; translation; RNA processing and modification; chromatin structure and dynamics; post-translational modifications (PTMs); intracellular trafficking, secretions, and vesicular control. However, only one COG was found to have a good coverage score of 0.85 with a good abundance score of 0.99 named - Oligoendopeptidase F (COG1164).
The ACE2 protein was further subjected to the network analysis in order to identify its associated interacting partners. There were 10 significant interacting partners of our query protein ACE2, namely AQPEP, ST3GAL4, DENR, NPEPPS, AGTRAP, ITPKB, SOCS2, REN, GHRHR, and SLC15A1. Fig. 1 shows the interacting network of ACE2 protein based on the k-means clustering algorithm. There are 21 nodes, with 32 edges; the average node degree is 3.05 with an average local clustering coefficient of 0.756. Table 4 showcases the detailed results of the interaction analysis.

Network interactors of ACE2 protein receptor.
Significant 10 interactors found in ACE2 physical network.

Amino peptides have been reported to play a crucial role in the formation and metabolism of neuropeptides essential for neurological health 33 . Various studies suggest that amino peptides are sensitive to puromycin and hold myriad central nervous system (CNS) functions such as memory 34 , amnesia 35 , apoptosis 36 , and schizophrenia 37 . On the other hand, ST3GAL4 has been said to promote Parkinson's disease (PD) because of its decreased expression 38 . The density-regulated protein (DENR) has been shown to promote various diseases in humans namely cancer, neurological disorders, and viral infections 39 . Moreover, overexpression of NPEPPS is responsible for Alzheimer's and other neurodegenerative dementias collectively called tauopathies 40 . Type-1 angiotensin II receptor-associated proteins have been studied widely for elucidating medical conditions such as blood pressure, electrolyte balance, and renal and neuronal, as well as endocrine functions that are associated with cardiovascular control 41 . Another 2014 study reports that Inositol trisphosphate 3-kinase B is increased in the human cerebral cortex of Alzheimer's patients 42 . Studies have shown that SOCS2 plays an essential role in the central nervous system (CNS), metabolism, immune response, mammary gland development, oncology, and cytokine-dependent signaling pathways 43 . Renin or brain rennin-angiotensin system (RAS) has been proven to have implications in a much broader range of neural effects. Also, the brain RAS is composed of the ACE2, Ang17, and prorenin and Mas receptors responsible for many neural effects 44 . It is not a new concept that the growth hormone (GH) and gonadotropin-releasing hormone (GnRH) in the brain are implicated in disorders in the cerebral cortex, hypothalamus, hippocampus, cerebellum, spinal cord, and neural retina, and are also responsible for brain tumors. Both GH and GnRH have shown the potential to cause neurotrophic, neuroprotective, and neuro-regenerative action in the human brain 45 . On the other hand, solute carrier transporters (SLC) play a pivotal role in promoting neuro-degenerative disorders such as Alzheimer's disease, amyotrophic lateral sclerosis, Huntington's disease, Parkinson's disease, depression, post-traumatic stress disorder, dementia, schizophrenia, and epilepsy 46 . Our interaction analysis discerns the evidence: ACE2 and its 10 most significant interactors are related and play a role in promoting neuronal disorders in humans, while ACE2 is also the main entry receptor of SARS-CoV-2. Therefore, it is important to refer here that ACE2 is directly responsible for neurological disorders such as Alzheimer's disease, Parkinson's disease, and epilepsy that have been lately observed to rise in COVID-19 patients.
Gene-Drug Mapping
Gene drug mapping is an essential step toward understanding how drug compounds function when used to treat various diseases. Every drug candidate has its own potential to bind to a gene/protein target. We deployed the three best gene-drug mapping resources – DGIdb, DrugBank, and PubChem to identify the best drug candidates that have been validated to bind to the seed genes. Most of the synthetic drugs that were identified were mainly inhibiting the seed genes or were antagonists. Table 5 summarises the gene-drug mapping.
Gene-drug mapping of each seed gene.

Identification of Alternative Herbal Compounds
The search for herbal medicines has become necessary to prevent adverse drug reactions (ADRs) and various other side effects that are encapsulated by synthetic medications. Alternative medicine is an umbrella term for a number of therapies that involve the use of dietary supplements, hormones, vitamins, proteins, and herbs. Alternative herbal medications are gaining ground due to serious risks posed by highly effective synthetic drugs. We have identified the alternative medications to synthetic drugs via extensive literature mining studies. The identification of alternative herbal compounds as a potential treatment for the neurological manifestations of COVID-19 can be a safe and effective therapy for the said disease manifestations. The herbal source, along with its bioactive phytochemical components, are mentioned in Table 6.
Alternative herbal compounds.

Alkaloids, flavonoids, saponins, terpenoids, and tannins are major phytochemical classes that can be used to treat the neurological dysfunctions that are associated with COVID-19 in patients. Phytochemicals have already been established to provide protection against numerous diseases such as cancer, diabetes, cardiac problems, and even neurological disorders9, 10, 47, 48, 51–53. Alkaloids are known to provide protection against neurological diseases such as febrile epilepsies, cerebral ischemia, oblivion, major depressive disorders (MDD), anxiety, and autism 55 . Flavonoids on the other hand associate with different pathways such as ERK and PI3-kinase/Akt and regulate the dynamics, in turn producing neuroprotective effects. They have a tendency to disturb and inhibit neuronal apoptosis that is triggered by neurotoxic substances in neurodegenerative diseases. Because of the inhibitory actions of flavonoids, these are prescribed mainly to treat dementia or oblivion or age-related neurological disorders (Alzheimer's disease) in people 50 .
Tannins, also known as polyphenols, encapsulate myriad health benefits. They are omnipresent in vegetables and fruits 56 . They are known to possess anti-oxidant, anti-aging, anti-inflammatory, and anti-cancer properties, but can also be deployed to treat neurological problems such as –Alzheimer's disease (AD) and Parkinson's disease (PD) 57 . Similarly, another important group of bioactive phytochemicals, saponins, are simply either glycosides of triterpenoid or steroidal aglycones. They have been widely deployed in traditional Chinese medicines (TCMs). Evidence suggests that saponins are beneficial to treat neurodegenerative and psychological diseases such as stroke, Alzheimer's disease, Parkinson's disease, and Huntington's disease 54 . Terpenoids have been suggested to be anti-convulsants that can be deployed to treat schizophrenia, insomnia, anxiety, pain, and cognitive problems 49 .
Pharmacological Screening of Phytochemicals
The identified phytochemicals were then subjected to ADMET screening. For the pharmacological assessment, the main criteria for the selection of the phytochemicals were i) Druglikeness, ie, based on the Lipinsky's Rule of 5 (RO5), where the compounds must have a) logP < = 5 (milog), b) molecular weight < = 500 (MW), c) number of hydrogen bond acceptors < = 10 (noN), d) number of hydrogen bond donors < = 5 (nOHNH) and e) the number of molecules violating more than 1 of these rules should be 0; ii) Medicinal chemistry that refers to the lead-likeness of a compound, bioavailability score and synthetic accessibility. Out of 35 identified phytochemicals, only 6 passed the ADMET evaluation. These were andrographolide, 14-deoxy-11,12-didehydroandrographolide, terphenolide, safranal, zingerone and p-cineole. Table 7 depicts the pharmacological screening results for the six qualified phytochemicals. The detailed result of the evaluation has been provided in Supplementary Table S1.
Best phytochemicals obtained after the pharmacological screening.

Binding Cavity Prediction and Molecular Docking
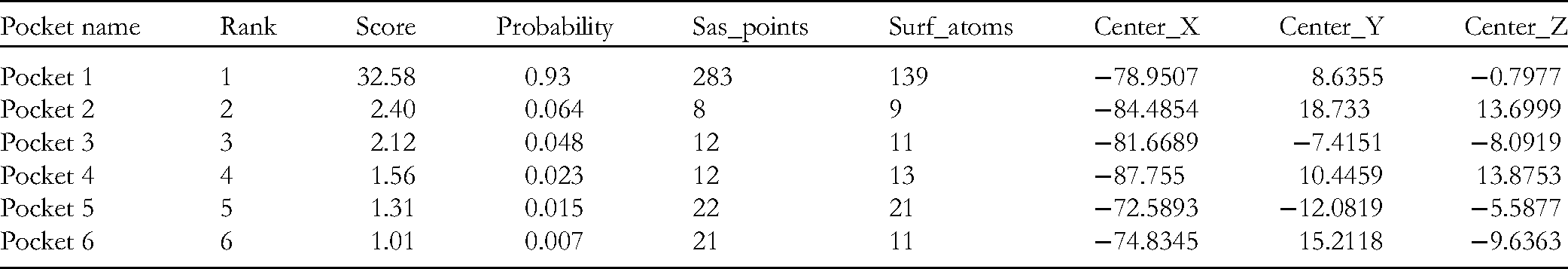
There were mainly six pockets that were identified in receptor ACE2. The highest scoring pocket was pocket 1, which had 139 surface atoms. This pocket had a good probability of fitting a ligand (P = 0.93). It spanned a larger surface area of the ACE2 protein and suggested a central ligand docking. That simply indicates that any ligand/molecule irrespective of its size can easily fit inside this cavity. Furthermore, pocket 1 includes 139 residues. Out of these, position 273 is known to be a binding site for substrate 58 ; position 374 is a metal binding site 58 ; position 375 is discerned to be an active site58, 59; and position 515 is the substrate binding site region 58 . The rest of the binding cavities identified are mostly present on the surface. Table 8 displays the details of each binding cavity. Supplementary file S2 shows a video that displays the two main pockets – 1 (in blue) and pocket 2 (in red) (Fig. 2).

Six best phytochemicals docked to receptor ACE2.
Predicted binding cavities present in ACE2 receptor.
After docking, we observed that only andrographolide (score = −7.8) and 14-deoxy-11,12-didehydroandrographolide (score = −8.0) had a greater binding affinity towards receptor ACE2 (Table 9). Additionally, it is noteworthy that both these phytochemicals are from Andrographis paniculata, and occupied a cavity space of 926, indicating their bigger ligand size. The bigger the ligand size, the greater the protein structural conformations are observed.
Docking scores of the six phytochemicals with ACE2 receptor.

Electrostatic Computation
After molecular docking, it was evident that only andrographolide and 14-deoxy-11,12-didehydroandrographolide have a greater binding affinity toward the ACE2 receptor. Therefore, only these two complexes were further selected for an electrostatic analysis to check the overall energy of the systems along with the macromolecular energy landscape. Energy assessment plays an important role after molecular docking and simulation. It is so because when a ligand is docked to a receptor, especially a protein structure, it tends to alter the conformation of the protein. This alteration in the protein pocket eventually dictates the effectiveness or ineffectiveness of a ligand. If the docked ligand comfortably fits inside the pocket of the protein receptor, it indicates that the complex will not have enormous unstable energy. It will also tend to have little to no steric hindrance. Table 10 summarizes the electrostatics computed for the two selected complexes.
Electrostatics of the two complexes.
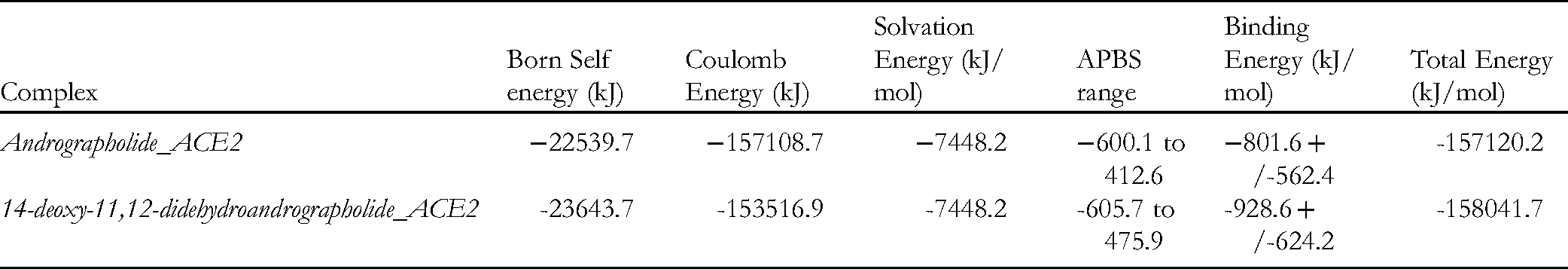
Although, both andrographolide and 14-deoxy-11,12-didehydroandrographolide showcased a better affinity, the latter overshadowed the former with respect to the overall energy of the docked complex. 14-Deoxy-11,12-didehydroandrographolide bound to the ACE2 receptor depicted good stable energy (−158041.7 kJ/mol) with minimal energy fluctuations. Minimal residual fluctuations represent minimal clashes and steric hindrances that occur usually after molecular docking of the ligand with the receptor (refer to Fig. 3). The complex showcased good Coulomb energy (−153516.9 kJ) with stable solvation energy (−7448.2 kJ/mol). The minimal residual fluctuations have been represented in green links, while maximum residual fluctuations have been depicted in red. The MM-GBSA has been represented in the form of an APBS map. The APBS range for the complex was recorded to lie between −605.7 to 475.9. This score further confirms the stability of the phytochemical inside the main pocket (pocket 1) of the ACE2 receptor.

MMGBSA and macromolecular energy landscape of 14-deoxy-11,12-didehydroandrographolide_ACE2.
Discussion
Since the COVID-19 pandemic broke out in early 2020, life has been turned upside-down for one and all. Because of drastic life changes, people have not only been infected by the SARS-CoV-2 virus, but have also experienced neurological and psychological problems, such as depression, anxiety, insomnia, brain fog, and epilepsy. Angiotensin-converting enzyme 2 (ACE2) has been discerned to be the entry receptor of SARS-CoV-2. Therefore, we tried to understand how the ACE2 receptor is involved in the COVID-19 disease by exploring genomic and epinformatics domains. By motif enrichment, we identified only two motifs, motif M2 and M5. Motif M2 (AGTACTGCC, score = 10.6676) is present on the negative strand and has a length of 9 nucleotides, while motif M5 (AGTACTGTAGATGGTGCTCA, score = 11.8863) is also located on the negative strand and has a length of 20 nucleotides. These two motifs are said to be crucial, as they were a perfect match for zinc fingers, DNA-binding domains, heart and neural crest derivatives, tRNA-methyltransferases, and paired like homeoboxes present in the human, mouse, and yeast genome. Furthermore, by analyzing the genomic annotation, we found that the GC content directly affects the genome functioning and species ecology of the ACE2 gene, and it was observed to be 38.5% showing the quadratic relationship with the genomic size of the ACE2 gene. The analogy is simple - the larger the genomic size, the lower the GC content viz., because of the higher biochemical expenditure of GC base formation.
We identified clusters of orthologs (COGs) for the ACE2 protein sequence. The identified COGs play crucial roles in biological processes such as cell cycle regulation; translation; RNA processing and modification; chromatin structure and dynamics; post-translational modifications (PTMs); intracellular trafficking, secretions, and vesicular control. However, only COG Oligoendopeptidase F (COG1164) was found to be more significant for the ACE2 protein receptor. The ACE2 protein was further subjected to network analysis to identify its associated interacting partners. There were 10 significant interacting partners of our query protein ACE2, namely AQPEP, ST3GAL4, DENR, NPEPPS, AGTRAP, ITPKB, SOCS2, REN, GHRHR, and SLC15A1. These network associators provide insights into the possible neurological dysfunctions that could be occurring in the patients who were infected with COVID-19. Therefore, we executed a gene-drug mapping to screen for highly potent synthetic drug candidates for each of these seed genes. Once the synthetic medications were identified, their alternative herbal phytochemicals were screened by literature mining. It was found that alkaloids, flavonoids, saponins, terpenoids, and tannins can be used to treat the neurological dysfunctions that are associated with COVID-19 in patients. On subjecting the identified phytochemicals to pharmacological, pharmacokinetic (PK), and pharmacodynamic (PD) analysis, only 6 out of a total of 35 passed the ADMET evaluation. These were andrographolide, 14-deoxy-11,12-didehydroandrographolide, terphenolide, safranal, zingerone and p-cineole. These six phytochemicals were then submitted for molecular docking using PyRx software. The highest binding affinity with the ACE2 receptor was noted for andrographolide and 14-deoxy-11,12-didehydroandrographolide. Although, both displayed a good binding affinity, 14-deoxy-11,12-didehydroandrographolide bound to the ACE2 receptor with much more stable energy (−158041.728091 kJ/mol), with minimal energy fluctuations when compared to andrographolide with ACE2. Therefore, we discern andrographolide and 14-deoxy-11,12-didehydroandrographolide, both major constituents of Andrographis paniculata, must be further clinically validated for treating neurological dysfunctions associated with the ACE2 receptor in COVID-19 patients.
Herbal compounds are known to be important sources for developing therapeutics for diseases. Often with fewer to no adverse reactions and low molecular weight, they often showcase a good binding towards protein receptors. In order to prevent a disease from occurring or reducing its effect, it becomes important to block the enzyme's catalytic activity. Thus, herbal compounds such as the diterpenoids andrographolide and 14-deoxy-11,12-didehydroandrographolide can be used as natural inhibitors60, 61.
A. paniculata is known for its anti-inflammation, anti-bacterial, anti-oxidant, and anti-cancer properties, and forms one of the important Ayurvedic formulas in Indian Ayurvedic medicine. The plant is native to South Asia and is found in India and Sri Lanka 62 . The constituents of the plant are well-recognized for viral and bacterial infections. However, there is also a need to pay heed to their efficacy towards neurological dysfunctions 63 .
Conclusion
Angiotensin-converting enzyme 2 (ACE2) is known to be the entry receptor of the SARS-CoV-2 virus, and is responsible for various neurological dysfunctions that are associated with COVID-19 in patients. To treat the neurological manifestations associated with the ACE2 receptor, instead of synthetic medications, we discern that the phytochemicals andrographolide and 14-deoxy-11,12-didehydroandrographolide, both major constituents of the plant Andrographis paniculata, must be further clinically validated for treating neurological dysfunctions associated with the ACE2 receptor in COVID-19 patients.
Supplemental Material
sj-xlsx-1-npx-10.1177_1934578X221118549 - Supplemental material for In silico Analysis of ACE2 Receptor to Find Potential Herbal Drugs in COVID-19 Associated Neurological Dysfunctions
Supplemental material, sj-xlsx-1-npx-10.1177_1934578X221118549 for In silico Analysis of ACE2 Receptor to Find Potential Herbal Drugs in COVID-19 Associated Neurological Dysfunctions by Juan Hou, Adil Manzoor Bhat, Shaban Ahmad, Khalid Raza and Sahar Qazi in Natural Product Communications
Footnotes
Acknowledgements
SQ is supported by a DST-INSPIRE fellowship provided by the Department of Science & Technology (DST), Govt. of India.
Conflict of Interest
None
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Author JH acknowledges funding from “Study on the effect of low oxygen inhalation in patients humidification”, and “Study on nursing effect of psychological intervention guidance in emergency patients with multiple injuries”.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
