Abstract
Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) was initially reported in the Wuhan province of China, spread throughout the world, and caused massive destruction in the form of a global pandemic that started back in 2020. SARS-CoV-2 is an RNA virus with various proteins like main protease, spike protein, NSP15 endoribonuclease, RNA-dependent RNA polymerase, and papain-like protease targeted to screen and find the novel drug candidate that can potentially work against the virus. Previous studies have reported multiple drugs after screening and validation against a single target and reported multiple medications. Nevertheless, many drugs are being used to date but do not have enough potential to work against SARS-CoV-2 and curb the spread and death rate. In this study, with the hypothesis of 1 drug and multiple targets, we have taken 5 main target proteins and screened the Asinex's complete BioDesign library (1,70,269 compounds) and identified N-{2-[(2S)-2-Amino-3-methylbutoxy]-6-propylbenzoyl}-L-phenylalanyl-L-serine (Butoxypheser) as multitarget inhibitor against SARS-CoV-2. Also, Butoxypheser has shown excellent docking scores, hydrogen bonding, and other bonding configurations like van der Waals force and water bridges. The stability and interaction pattern of the compound was validated with structural interaction fingerprints (SIFts) and molecular dynamics (MD) simulation. The Butoxypheser has performed flawlessly throughout the study, and the same results were used to compare the compound's activity against multiple targets. After a thorough theoretical comparative analysis, Butoxypheser can be treated as a multitargeted inhibitor candidate against SARS-CoV-2. Further, this study needs to be validated experimentally before human use.
Introduction
COVID-19 stands for coronavirus disease 2019, needs no introduction in the 21st century, especially in this decade caused by severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2). The first known case was initially reported from Wuhan province of China in December 2019, since it spread worldwide, leading to the ongoing pandemic.1-5 Symptoms of COVID-19 vary but repeatedly include fever, cough, headache, fatigue, breathing difficulties, and loss of smell and taste. Symptoms mainly occur between 1 and 14 days after exposure to the virus. At least a third of infected people do not develop noticeable symptoms and are counted under non-symptomatic, and mostly, they recover without any or slight medications. 5 Those who develop symptoms noticeable enough to be classed as patients develop mild-to-moderate symptoms (up to mild pneumonia).5-8
In contrast, few develop severe symptoms (dyspnea, hypoxia, or more than 50% lung involvement on imaging), and few suffer acute symptoms (respiratory failure, shock, or multiorgan dysfunction). Apart from this, many testing methods have been developed to diagnose the condition. Numerous vaccines have been authorized by WHO and countries at the individual level and distributed through various scenarios in 2 and sometimes in 3 as a booster dose, which has initiated mass vaccination campaigns.9, 10 However, its role in curve spread and the death rate is still questioned. Other protective measures include physical or social distancing, quarantining, ventilation in indoor spaces, masking the face, covering coughs and sneezes, hand washing, and keeping hands away from the face or after washing. Face masks or coverings or face masks and sanitization has been recommended in public settings to minimize the risk of transmissions.4, 10, 11 During all of these in the early months of the COVID-19 pandemic, a very different situation came across the healthcare workers analyzing test results of the patients who had already recovered from COVID-19 would sometimes test positive on a PCR test a week or even months later.8, 11, 12
Nevertheless, it was not the case here that they were catching reinfection because no live virus was isolated from their samples. Even some studies also found false-positive cases while holding patients in quarantine. So, the integration of SARS-CoV-2 genetic material into the human genome might be the reason behind that.13, 14 It might also be responsible for the long-term autoimmune problems that some patients experience. Previous work also mentioned that viral RNA might be integrating into cellular DNA through more than 1 mechanism. Although SARS-CoV-2 is not the only virus that can integrate into the human genome, around 8% of human DNA comprises the archaic viruses’ relics.15, 16 Some viruses like retroviruses depend on our genome to replicate themselves. Also, the frequency of integration of SARS-CoV-2 into the human genome is still unknown. Considering all this, it might be possible that our body records all the RNA viruses that ever infected it. 17 It will eventually open a very new angle to immune memory. A recent study suggested that some viral RNA can be reverse transcribed and inserted into the human genome. 13 It means some people who had already recovered from COVID-19 would sometimes inexplicably test positive on PCR test weeks or even months later, and that also can be classified long-term effect of the virus. These studies also suggested moving toward RNA/DNA-based drug designing instead of just for protein-based or single target-based drug designing. As the cases and new variants evolve daily, fear continues, and there is still no medication on the market.6, 7
Most of the previous studies were from single target and multiple drug backgrounds.18, 19 Nevertheless, in this study, we are reporting N-{2-[(2S)-2-Amino-3-methylbutoxy]-6-propylbenzoyl}-L-phenylalanyl-L-serine (Butoxypheser) 20 as the multitargeted inhibitor that has been identified after comprehensive pipeline-based research. We have performed multilevel screening to reduce the computational costs, ie, High-Throughput Virtual Screening (HTVS), Standard Precision (SP), Extra Precise (XP), and Molecular Mechanics/Generalized Born Surface Area (MMGBSA). Further, the study extended to structural interaction fingerprints (SIFts) and molecular dynamics (MD) simulation. Through the complete study, we have seen the steady performance of the compound without much deviation or fluctuation.
Methods
The methods involve the collection of multiple targets of the SARS CoV-2 and a huge library from the Asinex database and its preparation. Further, analysis involves numerous steps to screen and validate the compounds. We have provided the graphical abstract in Figure 1 to make the modus operand easier to follow.
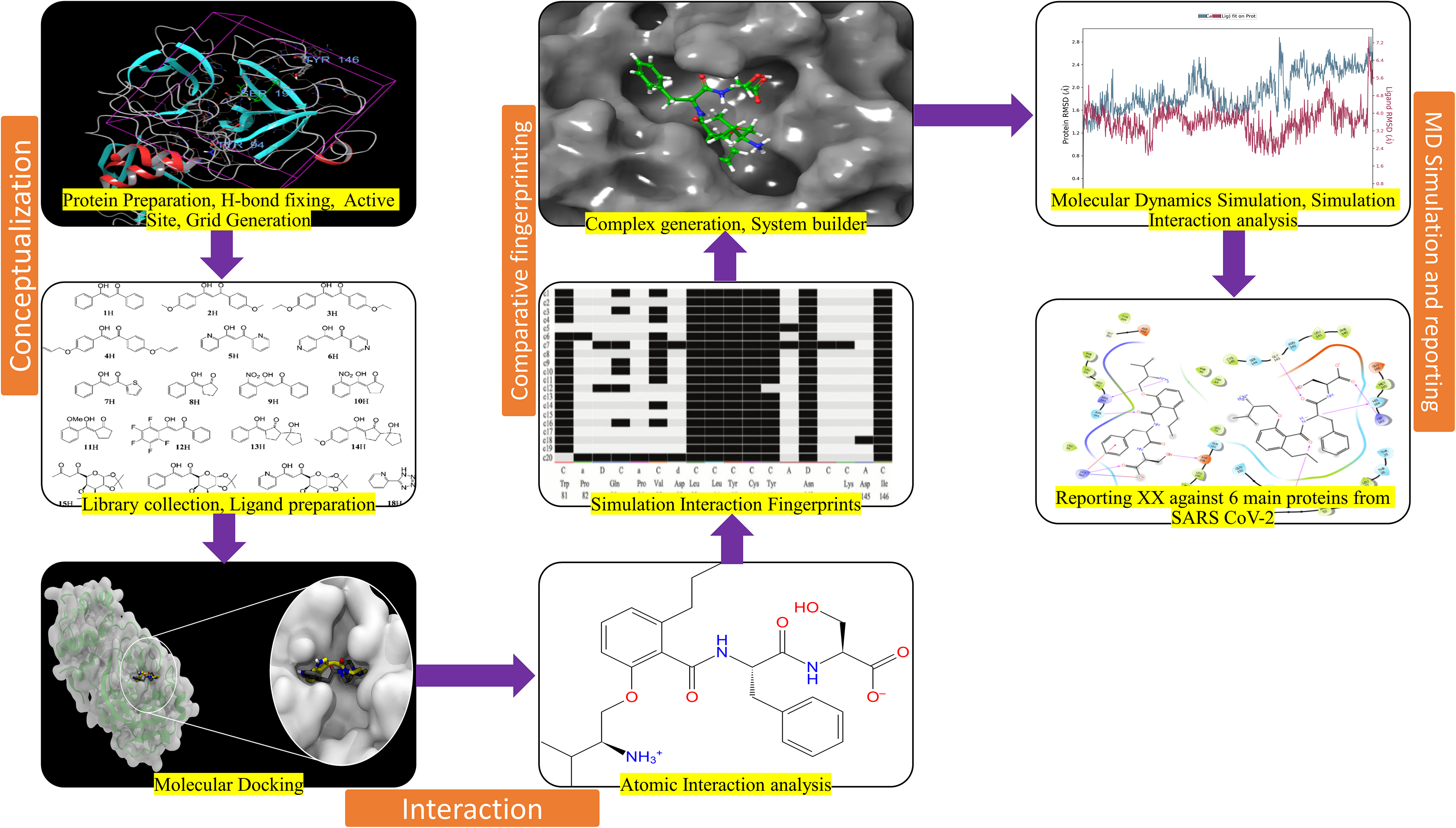
Graphical abstract of the complete study reporting N-{2-[(2S)-2-Amino-3-methylbutoxy]-6-propylbenzoyl}-L-phenylalanyl-L-serine (Butoxypheser) as a multitargeted inhibitor against severe acute respiratory syndrome coronavirus-2 (SARS CoV-2).
Ligand Library Collection and Preparation
The Asinex (https://www.asinex.com/) is a dynamic database that provides the drug library in a well-classified format in one click to download. We downloaded the BioDesign library, which has 1,70,269 natural compounds updated in February 2021. 21 The data was imported to Schrodinger's maestro workspace and prepared using LigPrep.22-24 While preparing the library, the OPLS4 force field was used. The Epik ionizer was selected with the metal-binding state, including the original state. 25 Further, tautomers and stereoisomer (maximum 32 from each) computations were kept producing the ultimate possible state that could show an enhanced binding state.
Data Collection and Protein Preparation
To choose the crucial pathway proteins of the SARS-CoV-2, we went through various literature reviews and identified PDBID: 6LU7, 6VW1, 6M71, 6VWW, and 6W9C26-29 as main drug targets and downloaded them from the RCSB database to import into the Maestro workspace for the preparation using protein preparation wizard. 30 PDBID: 6LU7 has 2 chains, A and C; chain C was very small or misidentified, so we have removed that and kept only chain A for next-level studies. For PDBID: 6VW1, 4 chains, A, B, E, and F, are there, and all chains were deleted except chain A as the complete protein is a dimer and heavy to work on, which could also impose substantial computational costs. Further in PDBID: 6M71, 6VWW, and 6W9C, only chain A was kept and the rest of chains B, C, and D from 6M71, chain B from 6VWW, and chains B and C was deleted from 6W9C, respectively. All proteins were prepared with the same parameters without bias to get a reliable and fixed structure for further studies. In preprocess tab, assigned bond orders against the CCD database, hydrogens were added, zero-order bonds to metals, waters were removed beyond 3 Å, and missing side chains and loops were filled using the prime module. The structures were refined with sample water orientations, and PROPKA at 7.0 pH was used to optimize the H-bonds to fix bondings with proteins. Further, the energy of the protein was minimized with the OPLS4 force field to get the lowest state.24, 30
Molecular Docking
For molecular docking, grid generation is important as the screening work within the defined parameters using sampling algorithms. For PDBIDs: 6LU7, 6VW1, and 6M71, the native ligand site was chosen, while the complete protein was taken into the grid box in the case of PDBID: 6VWW and 6W9C to perform active site-based and blind docking, respectively, to produce the best poses in each condition. Further, the Virtual Screening Workflow (VSW) was used for the docking-based screening. The prepared library was used as the ligand source and filtered with the QikProp tool to expel the compounds not satisfying the parameters of Absorption, Distribution, Metabolism, Excretion, And Toxicity (ADMET). 31 Lipinski's rule was applied as the next filter level, using the ADMET properties generated by the QikProp tool.31, 32 In the docking tab, Epik state penalties were checked and further docked with HTVS, SP, and XP, followed by MMGBSA.33, 34 Only the top 5% of HTVS data was taken for the SP. After this round of screening, only 10% of data was kept for the next screening level with XP to reduce the computational cost drastically. For the MMGBSA, 100% of XP data was taken to calculate the binding free energy and surface. Additionally, we have exported the data into CSV format to calculate and sort the data to identify the compound with the best binding potential among all the compounds against all selected protein targets.
Structural Interaction Fingerprints
The SIFts were calculated using the Interaction Fingerprints tool in Maestro for the receptor–ligand complex for all 5 cases together after aligning sequences to get a common interaction diagram and matrix to understand the interaction pattern among the residues of the protein and ligand's atoms. Further, any contact, backbone interaction, side–chain interaction, polar residues, hydrophobic residues, hydrogen bond acceptor, hydrogen bond donor, aromatic residue, and charged residue were selected to generate the graph for the fingerprinting parameters. 35 The similarity pattern was calculated in the similarity tab, and in the cluster tab, the clustering of residues was calculated. Further, the data were plotted with the help of the Matplotlib library of the Python and provided the color to the main plot by residue number, sorted the data with the docking score to arrange it correctly in the figure and make it understandable, and checked for only interacting residues.
MD Simulation
The MD simulation is a stand-alone set of algorithms that provides a considerable advantage in the stability and flexibility of the atom in biological as well as in material sciences.36, 37 For the MD simulation studies, we have used the Desmond package available from the Schrodinger's Maestro developed by the DE Shaw Research (https://www.deshawresearch.com/), which is considered the fastest and most accurate calculations among all the MD algorithmic calculations. 38 System builder tool used for preparing the system using the SPC water model with orthorhombic boundary condition of 10 × 10 × 10Å of distance in the buffer. Further, after excluding the ion and salt placement within 20Å and adding ion placement to neutralize the system and minimize to build the complete system for the MD simulation. In 6LU7; 3Na+, 6VW1; 25Na+, 6M71; 9Na+, in 6VWW; 12Na+, and 6W9C; 3Cl− were added to neutralize the system to biasedness data. Further, 100ns of simulation production run for each complex were kept at 300K temperature and pressure (bar) 1.01325, with a recording interval (ps) trajectory of 100 to generate the 1000 frames at NPT ensemble class. Additionally, the produced trajectory was analyzed using the simulation interaction diagram (SID) tool. 24
Results
Data Collection and Preparation
The raw compound library initially consisted of 1,70,269 compounds prepared using the LigPrep tool that produced 6lacs + compounds, used for the screening with all 3 HTVS, SP, and XP algorithms. The proteins of the SARS CoV-2 were prepared and kept only clean files for the next level of the studies to generate the grids. After molecular docking studies, the data were exported to the CSV format to extract the important data to calculate the compounds’ activity. The same CSV file was used to find the common drug against multiple SARS CoV-2 and identified Butoxypheser.
Screening and Drug Interaction Analysis
The strict parameters in molecular docking produced from 111 to 117 potential candidates individually, used for the analysis and to sort against docking score, led to identifying the Butoxypheser as a possible drug candidate. The ligand interaction diagram analyses each protein's bonding configuration individually with the ligand Butoxypheser. The Butoxypheser with main protease (6LU7) has shown a docking score of −7.327 and MMGBSA score of −62.94 (Table 1) while interacting with hydrogen bond among GLY143, HIP163, GLN189 with O atoms, and HIS164 is interacting with the NH atom of the ligand (Figure 2A). Interaction with the receptor-binding domain (6VW1) has shown a docking score of −7.676 and a MMGBSA score of −30.45 (Table 1) while interacting with the hydrogen bonds among NH3+atoms with ARG393; 2 O atoms are interacting with the ASN394, LYS562 individually and the OH atoms is interacting with the ASP206. LYS562 also forms 1 pi-cation interaction with the benzene ring and 1 salt bridge with the O− atom (Figure 2B). The RNA-dependent RNA polymerase (6M71) has produced a docking score of −9.584 and MMGBSA score of −28.07 (Table 1) with 4 hydrogen bonds among O atom and ASP623, ARG553 while OH atom is interacting with THR556, NH3+ is interacting with ASP623 (Figure 2C). The same NH3+ with ASP623 is also forming 1 salt bridge. O− atom is interacting with the LYS545, ARG553, and ARG555 with a salt bridge, and while summing up in this instance, there are 4 hydrogen bonds, and 4 salt bridges are incorporated to make the structure stable.

Showing the surface view and ligand interaction diagram of N-{2-[(2S)-2-Amino-3-methylbutoxy]-6-propylbenzoyl}-L-phenylalanyl-L-serine (Butoxypheser) (ligand) with severe acute respiratory syndrome coronavirus-2 (SARS CoV-2). (A) Main protease (6LU7), (B) receptor-binding domain (6VW1), (C) RNA-dependent RNA polymerase (6M71), (D) NSP15 endoribonuclease (6VWW), and (E) papain-like protease (6W9C).
Tabular Representation of the Docking Score of N-{2-[(2S)-2-Amino-3-Methylbutoxy]-6-Propylbenzoyl}-L-Phenylalanyl-L-Serine (Butoxypheser) Against Each Protein (PDB) With Arranged Molecular Mechanics/Generalized Born Surface Area (MMGBSA) Score and its Bonding Configuration and Types.

The NSP15 endoribonuclease (6VWW) has produced a docking score of −6.72 and an MMGBSA score of −30.8 (Table 1) while forming the hydrogen bonding among TYR279, LYS90with O atom. NH3+ interacts with ASP268 via a hydrogen bond and 1 salt bridge. O−creates 2 salt bridges with LYS71 and LYS90 (Figure 2D). The papain-like protease (6W9C) has produced a docking score of −4.935 and the MMGBSA score of −30.61 (Table 1) while interacting with 5 hydrogen bonds and 2 salt bridges. NH3+ and O− interact with the salt bridges with GLU214 and LYS217, and the same LYS217 forms 1 hydrogen bond with the O atom. GLU214 forms 4 hydrogen bonds with NH3+, OH, and 2 NH atoms (Figure 2E). Further, the docking score, along with the other interaction types, is provided in Table 1, while the direction of the arrows shows the electron exchanging mechanisms and oversight, and all the bonding types are in different colors. Further, the ADMET values are provided in Table 2, along with the standard values of the QikProp tool that can be used for comparative understanding. All the values satisfy the standard criteria and give a taste that the drug compound Butoxypheser can be used against the SARS-CoV-2, and it will not give any side effects at a major level; anyhow, the experimental validation is a must before proceeding with the human use.
Showing the Absorption, Distribution, Metabolism, Excretion, and Toxicity (ADMET) Properties of the N-{2-[(2S)-2-Amino-3-Methylbutoxy]-6-Propylbenzoyl}-L-Phenylalanyl-L-Serine (Butoxypheser) Against the Standard Values of the QikProp Tool.

Filtering and Stepping into SIFts
The merged protein–ligand complexes were aligned together to generate the interaction fingerprints to understand the residue-wise comparative coordination that can potentially deliver the inhibition potency in the viral system. In Figure 3, the main plot is colored and sorted with the different color codes against residue numbers showing the ligand interaction counts. The interaction matrix shows ligand interaction counts. The RNA-dependent RNA polymerase (6M71) has shown the maximum interaction, and the same protein has also depicted the lowest value in the docking score table. At the same time, the papain-like protease (6W9C) has shown the lowest interaction counts. The comparative fingerprinting also depicts the functional coverage among the residues or similarity matrix among protein residues and its position while interacting with the ligand Butoxypheser. The fingerprinting also suggested better interaction coordination, and the compounds may perform better while simulating and performing the apoptosis and experimental analysis. So, the complete complex simulated the SPC water model to find its stability, coordination, and possible positioning with the ligand atoms with protein residues.
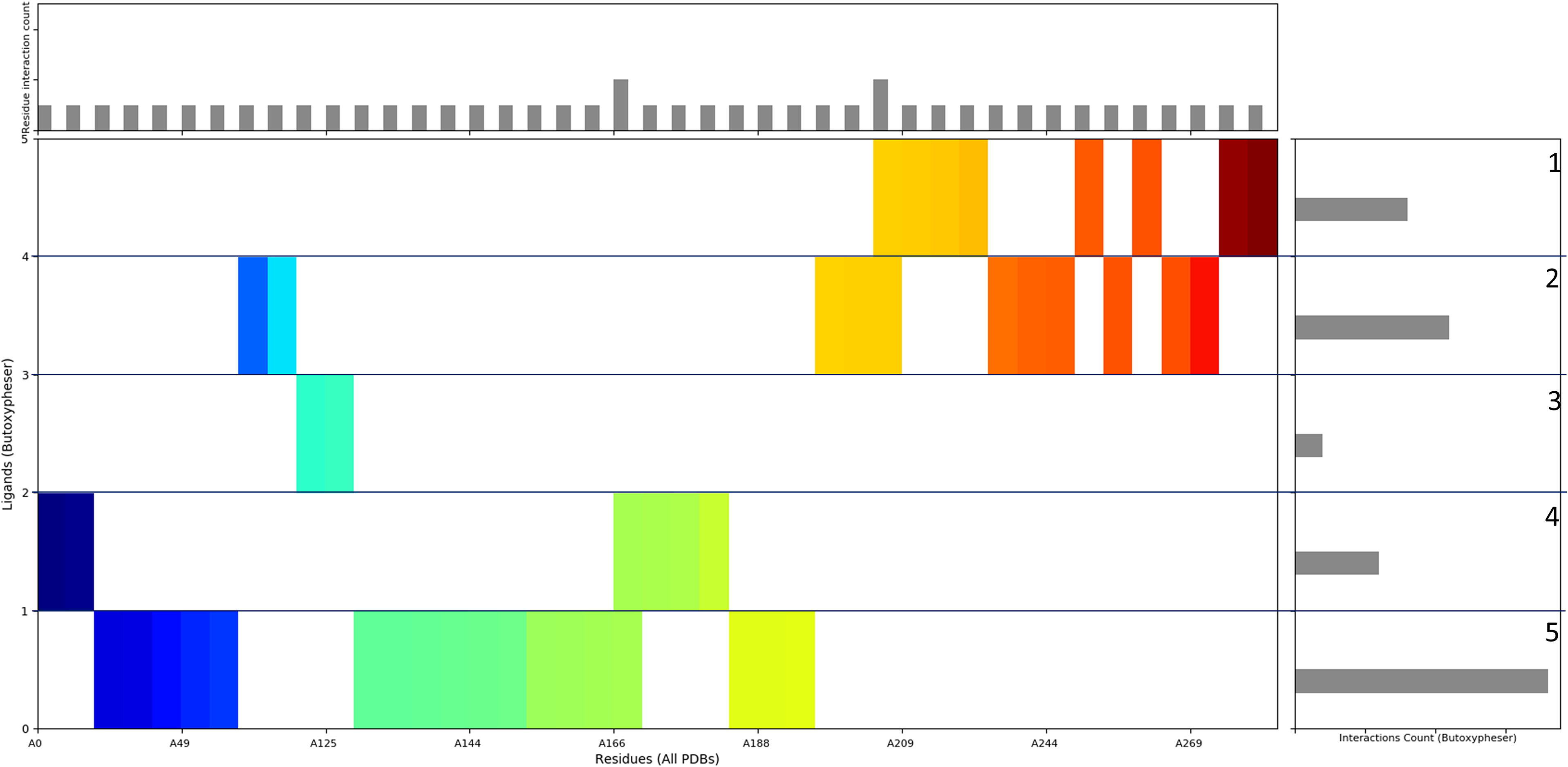
Structure interaction fingerprints, the right plot is for the N-{2-[(2S)-2-Amino-3-methylbutoxy]-6-propylbenzoyl}-L-phenylalanyl-L-serine (Butoxypheser) interaction count with respective proteins. The colored (main plot) plot shows the Butoxypheser interaction against the residue number in the upper plot (gray).
MD Simulation
The MD simulation studies consume huge computational costs due to their algorithmic and looping complexity. So, it provides enough evidence that can be trusted to perform the in vitro and other experimental research. In this study, we kept a 100ns production run to study the protein–ligand deviation and fluctuation along with the ligand interaction while being simulated in the water. Further, the stepwise in-depth analysis is as follows:
Root-Mean-Square Deviation
The root-mean-square Deviation (RMSD) values provide the deviation in the protein and ligand at the Å level with time durations at the nanosecond level, which means the RMSD provides evidence of steady performance and compound's (complex's) stability as a whole. Main protease (6LU7) in complex with Butoxypheser has initially fluctuated of 1.10Å in protein's case while in the ligand case, it shows 4.22Å at 1.10 ns and after that, during the entire simulative period, it has shown the steady performance. At 100ns, the ligand shows a deviation of 5.98Å, and the protein has demonstrated a deviation of 2.62Å. After ignoring the first 1 ns, the main protease's RMSD can be trusted at 1.52Å, and for the Butoxypheser, it will be 1.76Å (Figure 4A). The receptor-binding domain (6VW1) complex with Butoxypheser has initially fluctuated and then stabilized. The initial fluctuation was noted till 28.50ns at 3.08Å. For ligand, it shows a deviation of 3.95Å, and after analyzing the complete 1000s trajectories of the production run, at 100ns, the protein deviated at 3.57Å, and for the ligand, it went up to 4.24Å. After ignoring the initial phase of deviation, the noted deviation in the protein's case will be 0.49Å, and for the ligand, it will be at 0.29Å (Figure 4B). RNA-dependent RNA polymerase (6M71) in complex with Butoxypheser, the initial fluctuation was higher than in the previous 2 simulations. The RMSD at 0 ns was 0 Å, and at 13ns, the protein RMSD reached 3.66Å, while the ligand RMSD was noted at 3.58Å and after stabilizing the complex. At 100ns, the protein has produced a deviation at 4.78Å while the ligand has produced a deviation at 4.26Å, meaning the protein RMSD can be understood at 1.12Å while for the ligand 0.68Å after ignoring the initial deviations (Figure 4C).
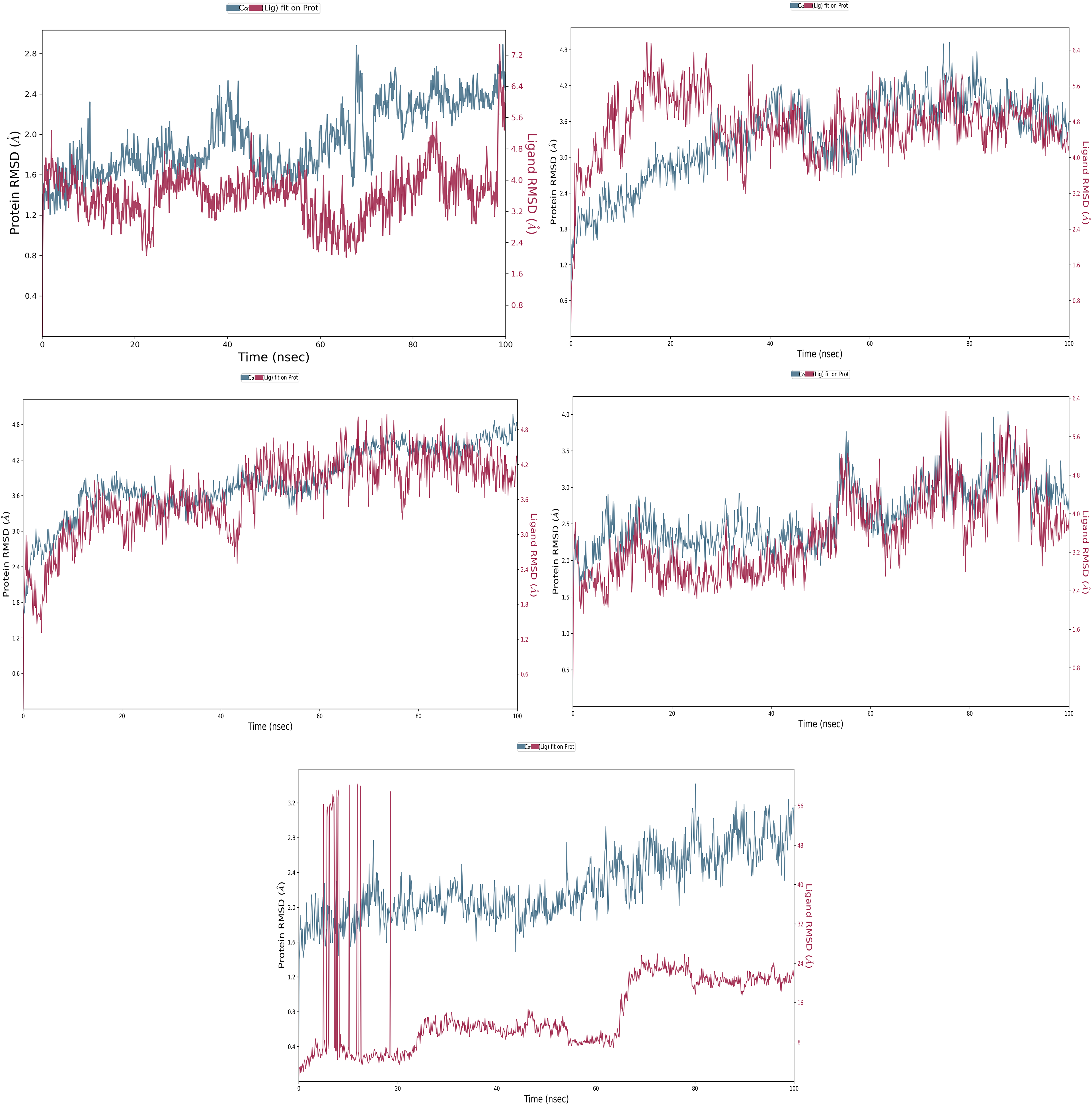
The root-mean-square deviation (RMSD) of N-{2-[(2S)-2-Amino-3-methylbutoxy]-6-propylbenzoyl}-L-phenylalanyl-L-serine (Butoxypheser) (ligand) with severe acute respiratory syndrome coronavirus-2 (SARS CoV-2) proteins. (A) Main protease (6LU7), (B) receptor-binding domain (6VW1), (C) RNA-dependent RNA polymerase (6M71), (D) NSP15 endoribonuclease (6VWW), and (E) papain-like protease (6W9C).
NSP15 endoribonuclease (6VWW) in complex with Butoxypheser has shown a steady performance after the initial protein's deviation of 2.39Å. The ligand noted 3.82Å at first 0.50 ns of the simulative period, and after performing stably at 100ns, it has shown a deviation of 2.74Å in protein's case and for the ligand's case went at 3.74Å. After ignoring the initial 0.50ns, the overall fluctuation noted the protein deviation of 0.35Å. At the same time, for the ligand, it can be assumed −0.8 Å, meaning the complex is very much stable. Still, there was residual deviation (Figure 4D) after noticing it correctly, but that will not affect the structure's performance on the higher side. The papain-like protease has shown the least docking and interaction fingerprints with Butoxypheser and has not performed well as other complexes. At initially 0.30 ns, the protein deviation was 1.91Å. It ended at 3.15, meaning it has produced an overall deviation of 1.24Å, meaning it is stable and the structure does not have any mismatched bonding configurations. However, Butoxypheser has shown an incredible range of fluctuation that went up to 60Å at a few periods. Still, after 18ns, it also started stabilizing, and at 100 ns, it produced a deviation of 21.59Å (Figure 4E). After ignoring the initial deviations, overall performance shows that the ligand Butoxypheser has performed better in each case while remaining under the deviation of less than 2 Å. The initial deviation was because the system directly imposes a new system. Due to initial heating, the atoms and residues deviate, and the solvation methods predict the biased results due to their level of solubility. The comparative understanding of the deviation shows that the Butoxypheser has performed better in main protease and NSP-15 endoribonuclease against the other 3 protein–ligand complexes.
Root Mean Square Fluctuation
The root mean square fluctuation (RMSF) shows the protein and ligand's fluctuation at the atomic and residual levels. In this analysis, we have only analyzed the protein fluctuation at residue level concerning residue number and interaction with the Butoxypheser. The residues of main protease (6LU7) have performed so steadily that only a few residues have gone beyond 2 Å, and those residues are ACE0, SER1, GLY2, LEU50, ASN51, SER301, GLY302, VAL303, THR304, PHE305, GLN306, and while simulating the ligand molecule interacted with the protein 24 times at THR26, LEY27, HIS41, SER46, MET49, SER149, PHE140, LEU141, GLY143, SER144, CYS145, HIS163, HIS164, MET165, GLU166, LEU167, PRO168, HIS172, GLN189, THR190, GLN192, ARG222, THR224, and ASP263 (Figure 5A). The receptor-binding domain (6VW1) also performed in the stable during the simulation period, and the ligand atoms interacted with the protein 30 times involving LEU39, LYS68, LEU73, VAL93, LYS94, LEU97, GLN98, ALA99, LEU100, GLN101, GLN102, ASN103, ALA193, TYR196, TYR202, GLY205, ASP206, GLU208, ASN210, ASP350, ASP382, ALA384, PHE390, LEU391, LEU392, ARG393, ASN394, ASN394, GLU398, ILE513, and GLY561 residues (Figure 5B). The RNA-dependent RNA polymerase (6M71) residue fluctuates slightly more than the above 2 complexes. ACE30, ARG55, PHE56 to LYS73, PHE104 to VAL111, LEU895 to ASP910, and HIS928 to LEU931 are the residues showing the fluctuation of more than2Å; the remaining amino acids are showing less fluctuation, meaning they are stable. These regions of residues are only interacting with the ligand, meaning the interaction is not hampering even after fluctuations (Figure 5C). The Butoxypheser is interacting 25 times with SER451, ASP454, LEU544, ASN552, ARG553, ALA554, ARG555, THR556, VAL557, ALA558, ASP618, PRO620, LYS621, CYS622, ASP623, ARG624, ALA625, GLY778, GLY679, THR680, SER681, GLY683, ASP684, ALA688, and SER758 residues of the protein during 100ns of simulation. NSP15 endoribonuclease (6VWW) residues have stable performance except for ASN0, MET1, GLY30, GLY170, GLU171, andALA172, VAL173, LYS174, THR175, and GLN176. The ligand interacted with the protein 26 times with GLU69, LYS71, LYS90, GLN160, THR196, SER198, ARG199, ASN200, LEU201, GLN202, LEU252, LEU255, LEU266, GLU267, ASP268, ILE270, PRO271, MET272, ASP273, SER274, THR275, LYS277, TYR279, VAL295, ASP297, and ASP324 residues of the protein (Figure 5D).

The root mean square fluctuation (RMSF) of N-{2-[(2S)-2-Amino-3-methylbutoxy]-6-propylbenzoyl}-L-phenylalanyl-L-serine (Butoxypheser) (ligand) with severe acute respiratory syndrome coronavirus-2 (SARS CoV-2) proteins. (A) Main protease (6LU7), (B) receptor-binding domain (6VW1), (C) RNA-dependent RNA polymerase (6M71), (D) NSP15 endoribonuclease (6VWW), and (E) papain-like protease (6W9C).
The papain-like protease (6W9C) has the best stability among all proteins. Anyhow a few fluctuated residues are CYS224 to GLN229, and THR313 to LYS315. In this condition, the interacting periods were noticed to be relatively higher from the beginning to the end, meaning the interaction distribution was quite flexible. The interacting residues are GLN29, ASP40, THR42, LYS43, ASN186, VAL187, VAL188, CYS189, LYS190, THR191, GLN194, SER212, TYR213, GLU214, GLN215, LYS217, LYS218, GLN221, THR231, TYR233, TYR251, GLU252, LEU253, HIS255, GLY256, THR257, PHE258, SER278, LYS279, GLU295, TYR305, LYS306, GLU307, ASN308, SER309, TYR310, ILE314, and LYS315, in sum, have incorporated a total of 38 times during the simulative period (Figure 5E). Some residues showed higher fluctuations in the complete protein-RMSF analysis but did not affect the secondary structure. The interactions were not noted specific to the range of fluctuated amino acids, and the width of the line (green color) shows the interaction duration. While summarizing the complete RMSF, the protein was stable. It showed many interactions with the ligand Butoxypheser, believing that the ligand can get and fit into the pocket to block the protein's activity and ultimately seize the viral system.
Simulative Interaction Analysis
The SID shows the ligand contacts with the protein during the simulative periods and how the electron transfer occurs, providing a better ligand binding pose to understand the positioning from the dynamic angle. The main protease (6LU7) in complex with Butoxypheser has shown a tremendous bonding combination involving 1 pi-pi stacking among the benzene ring of the ligand and HIS41, and the same residue is forming 1 direct hydrogen bonding with O atoms and 1 water bridge. GLN189 is involved with 1 water bridge and 1 direct hydrogen bond with the O atom, HIS163 is forming 1 hydrogen bond, and GLU166 is forming 1 hydrogen bond with the O− and 2 water bridge contacts with the same O− atoms and 1 water bridge with the O atom. CYS145 involves 1 water bridge O atom, and GLY143 involves 2 hydrogen bonds, O and OH atoms. SER144 creates 2 hydrogen bonds with O and OH atoms and 1 water bridge with the O atom of the ligand. There are 7 water molecules involving balancing the interaction (Figure 6A). The receptor-binding domain (6VW1) in complex with Butoxypheser involves 10 water molecules. NH3+ forms 1 pi–cation interaction with PHE390, ARG393, and 1 hydrogen bond with ASN394, and the same amino acid interacts with a hydrogen bond with O atoms and 1 water bridge with the same atom. There is 1 intramolecular interaction between the NH atom and O atom. LEU391 and LEU95 interact with NH atom and OH, respectively. GLN98 forms 1 direct interaction and 1 water bridge with the same OH atom. LYS562, GLU208, ASP206, and GLY205 form multiple direct hydrogen interactions, water bridges with O and O− atoms in a web form to stabilize the structure, and ARG514 forms 1 salt bridge with O− atom (Figure 6B).
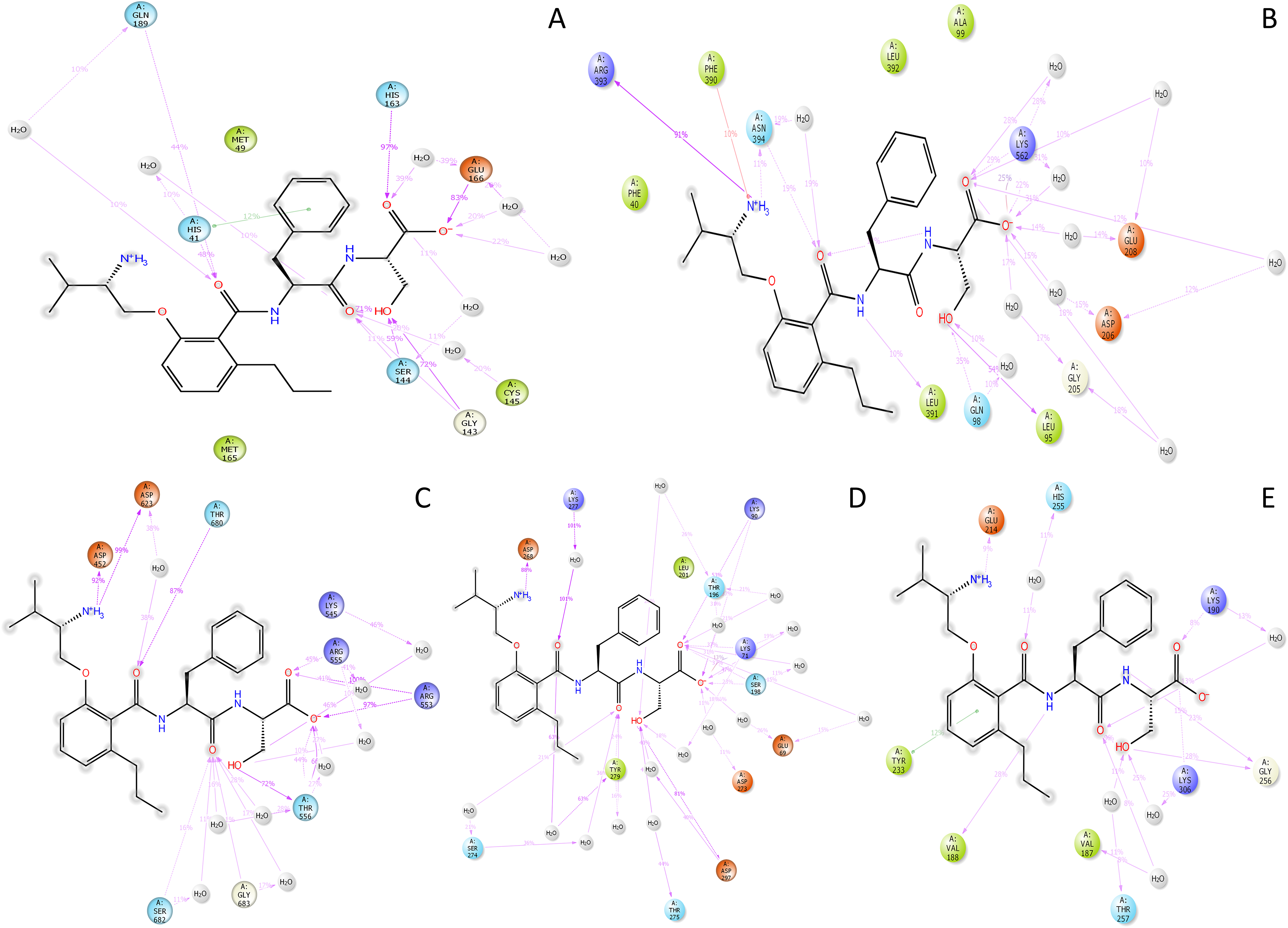
The interaction diagram of N-{2-[(2S)-2-Amino-3-methylbutoxy]-6-propylbenzoyl}-L-phenylalanyl-L-serine (Butoxypheser) (ligand) with severe acute respiratory syndrome coronavirus-2 (SARS CoV-2) proteins. (A) Main protease (6LU7), (B) receptor-binding domain (6VW1), (C) RNA-dependent RNA polymerase (6M71), (D) NSP15 endoribonuclease (6VWW), and (E) papain-like protease (6W9C).
The complex of the Butoxypheser and RNA-dependent RNA polymerase (6M71) involves 9 water molecules for water bridges. THR680 and ASP623 form 1 hydrogen bond and 1 water bridge with O atom, respectively, whileASP623 and ASP452 interact with NH3+atom. OH, atoms of the ligand interact through a hydrogen bond with THR556 and LYS545, and THR556 through water bridges. The O atom interacts with ARG553 and THR556 through hydrogen bonds and water bridges. Further, 2 O atoms create a web-like interaction structure involving multiple water molecules and GLY683, SER682, THR556, ARG553, and ARG555 residues of the protein (Figure 6C). NSP15 endoribonuclease (6VWW) in complex with Butoxypheser has used 16 water molecules to satisfy the criteria of the bonding configurations. NH3 + interacts with ASP268 through a hydrogen bond. O atom interacts with LYS277 and TYR279 through the water bridge, and with another O atom SER274, TYR279 interacts with 2 water bridges individually. O, OH, and O− interacts with LYS90, THR196, LYS71, SER198, GLU69, ASP273, ASP297, and THR275 through direct and water bridges, creating a web of interactions (Figure 6D). The complex of papain-like protease (6W9C) and Butoxypheser has shown relatively lesser interaction than other complexes. GLU214 forms a hydrogen bond with NH3+, and HIS255 interacts with the O atom through the water bridge. The unique interaction noted in this scenario is pi-pi stacking among the benzene ring and TYR233, while VAL188 interacts with the NH atom of the ligand. LYS190 interacts with the O atom through a hydrogen bond and 1 water bridge to another O atom, and with the same O atom, VAL187 interacts with a water bridge. GLY256 interacts directly with OH and NH atoms, while THR257 and LYS306 interact with OH atoms through water bridges (Figure 6E).
The overall simulative interaction analysis reveals that the O, O−, and OH atoms form multiple interactions together. That lets the Butoxypheser fit appropriately in the pocket of the proteins and destabilizes its core functionalities to inhibit viral growth. Further, the ligand interaction counts are provided below in the bar graph (Figure 7) to understand better the bonding statistics and the compound's efficacy with different proteins individually.

The bar graph for the interaction counts of N-{2-[(2S)-2-Amino-3-methylbutoxy]-6-propylbenzoyl}-L-phenylalanyl-L-serine (Butoxypheser) (ligand) with severe acute respiratory syndrome coronavirus-2 (SARS CoV-2) proteins. (A) Main protease (6LU7), (B) receptor-binding domain (6VW1), (C) RNA-dependent RNA polymerase (6M71), (D) NSP15 endoribonuclease (6VWW), and (E) papain-like protease (6W9C).
Discussion
After so many deaths and enormous economic losses, nobody is there who is not aware of the SARS CoV-2 and its complications. Still, due to improper cooperation and not working on multitargeted inhibitors, no such drug is available that can potentially work against SARS-CoV-2. In this study, we have followed a comprehensive path and screened the BioDesign library of the Asinex database and further data analysis while calculating the average performance the Butoxypheser has found to be suitable that potentially can work against multiple targets (Figure 1). Further, the study extended to structural fingerprinting and MD simulation. After seeing the steady and better performance, it is predicted that the Btoxypheser can be a potential drug candidate against SARS-CoV-2. In Figure 2, we have shown how the ligand Butoxypheser fitted into the pockets of the different proteins. It might reduce the bonding potential of the proteins and lead to the inactivation of the virus. The docking score of the ligand with all proteins individually varies from −9.676 to −4.935, considered a good docking score, and produced an excellent binding free energy in the form of the MMGBSA scores. Further other bonding configurations were noted, giving an immense view of how the drug interacts and the potency of its bonds and types. Overall, RMSD and RMSF were under 2 Å, and up to this level of flexibility is provided in the biological systems as the complex systems are virtually heated with multiple parameters, which causes the minor deviation. The simulative interaction diagram has shown a tremendous result that confirms the ligand's atoms are interacting widely in the simulative period with respective proteins, and this will work in the natural system; the simulative system gets neutralized after adding sufficient negative and positive atoms in the system. Butoxypheser is the candidate that has not been studied much and not reported earlier in the case of SARS CoV-2 and is available from Aurora Fine chemicals (https://aurorafinechemicals.com/) via catalogue number 191.797.122. Further, the experimental validation of the compound is needed as targeting 5 major proteins together can seize the complete viral system, and this compound can end the search for the potential candidate.
Conclusion
The SARS-CoV-2 is an RNA virus that produces multiple proteins treated as the potential target for binding drug molecules. The main target proteins are main protease, spike protein, receptor-binding domain, RNA-dependent RNA polymerase, NSP15 endoribonuclease, and papain-like protease. The previous studies have reported many compounds targeting 1 or 2 targets that can work against SARS-CoV-2. Due to mutational adaptations, a single target-based strategy may fail at some stages. In this study, we have identified a Butoxypheser as a multitargeted inhibitor against SARS-CoV-2. The complete in silico study has shown a good performance of the candidate against all 5 proteins, which suggests that the candidate can be used to treat the patients infected with the SARS-CoV-2. However, this study must be validated in vitro and in vivo, and it must pass through clinical trials before direct use as a medicine.
Footnotes
Acknowledgements
The authors would like to thank SRM University for providing the software solutions to carry out the study. SQ is supported by the DST-INSPIRE fellowship provided by the Department of Science & Technology, Govt. of India.
Author’s Contribution
SA: Conceptualization, data collection/curation, analysis, writing, and reviewing the first draft; NB: Analysis, writing the first draft; SQ: Reviewing, editing; MKY: Software solutions, reviewing and editing, and supervision; KR: Supervision, computational resources, reviewing and editing.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical Approval
This study does not directly involve humans or other organisms as it is a complete in silico study.
Consent for Publication
All authors consent to submit the manuscript to the Natural Product Communications.
Availability of Data and Material
All the data and supplemental material can be made public after publication.
