Abstract
Glioblastoma (GBM) is the most common type of primary malignant tumor that develops in the brain, with 0.21 million new cases per year globally and a median survival period of less than 2 years after diagnosis. Traditional Korean medicines have been increasingly suggested as effective and safe therapeutic strategies for GBM. However, their pharmacological effects and mechanistic characteristics remain to be studied. In this study, we employed a computational network pharmacological approach to determine the effects and mechanisms of the traditional Korean medicinal formula FDY003 on GBM. We found that FDY003 treatment decreased the viability of human GBM cells and increased their response to chemotherapeutics. We identified 10 potential active pharmacological compounds of FDY003 and 67 potential GBM-related target genes and proteins. The GBM-related targets of FDY003 were signaling components of various crucial GBM-associated pathways, such as PI3K-Akt, focal adhesion, MAPK, HIF-1, FoxO, Ras, and TNF. These pathways are functional regulators for the determination of cell growth and proliferation, survival and death, and cell division cycle of GBM cells. Together, the overall analyses contribute to the pharmacological basis for the anti-GBM roles of FDY003 and its systematic mechanisms.
Introduction
Glioblastoma (GBM) is the most common type of primary malignant tumor that develops in the brain, with 0.21 million new cases per year worldwide and a median survival period of less than 2 years after diagnosis. 1 The current standard therapies for GBM include surgery, chemotherapy, radiotherapy, and immune checkpoint inhibition. 2 Unfortunately, these therapies have limited efficacy in improving the survival of patients with GBM. 2 Herbal drugs have been increasingly suggested as effective and safe therapeutic strategies for GBM since they can effectively suppress the malignant activities of GBM cells and can elevate the treatment efficacies of standard anti-GBM therapies. 3
FDY003 is a traditional Korean medicinal formula composed of
Numerous previous studies have proven that computational network pharmacology is an efficient approach for understanding drug mechanisms of action.8-10 The pharmacological effects of herbal drugs are determined by multiple interactions between their active pharmacological compounds and therapeutic targets.8-10 Hence, network pharmacology elucidates the mechanisms of herbal drugs by analyzing the characteristics of their relevant networks, which are the integration of comprehensive compound–target interactions.8-10 In this study, we employed a computational network pharmacology approach to determine the mechanisms underlying the role of FDY003 as a potential anti-GBM therapy.
Results and Discussion
Examination of FDY003 Activity in GBM
To examine FDY003 activity in GBM, we treated U-87 MG human GBM cells with either FDY003, temozolomide (a chemotherapy drug for GBM treatment 2 ), or their combination. FDY003 significantly decreased the viability of GBM cells with an IC50 value of 74.25 μg/mL (Figure 1A) and further enhanced the pharmacological effects along with temozolomide (Figure 1B), suggesting that FDY003 may possess anticancer properties in GBM.

Effects of FDY003 treatment on the viability of human glioblastoma cells. (A) U-87 MG human glioblastoma cells were treated with indicated doses of FDY003 and their viability was measured. (B) U-87 MG cells were treated with indicated doses of FDY003 and/or temozolomide (TMZ) for 72 h, and their viability was measured. Data represent the mean ± SEM of five replicates.
**
Active Pharmacological Compounds and Targets of FDY003
To identify active pharmacological compounds, we screened the FDY003 compounds with Caco-2 permeability ≥ −0.4, drug-likeness ≥ 0.18, and oral bioavailability ≥ 30% to determine their activity (Supplementary Tables S1 and S2).10,11 In addition, some compounds known to possess potent anticancer effects were selected as active compounds even if their pharmacokinetic parameters did not meet the inclusion criteria (Supplementary Tables S2 and S3). Then, we identified the therapeutic targets of the active pharmacological compounds using databases and programs used for the investigation of the molecular binding between chemicals and genes/proteins, as well as tools containing large-scale information regarding cancer biology and clinical oncology (Supplementary Table S3). In the identified FDY003 targets, those that are closely involved in the pro-tumorigenic processes of GBM were determined for the further network pharmacological analyses (Figure 2 and Supplementary Table S3).
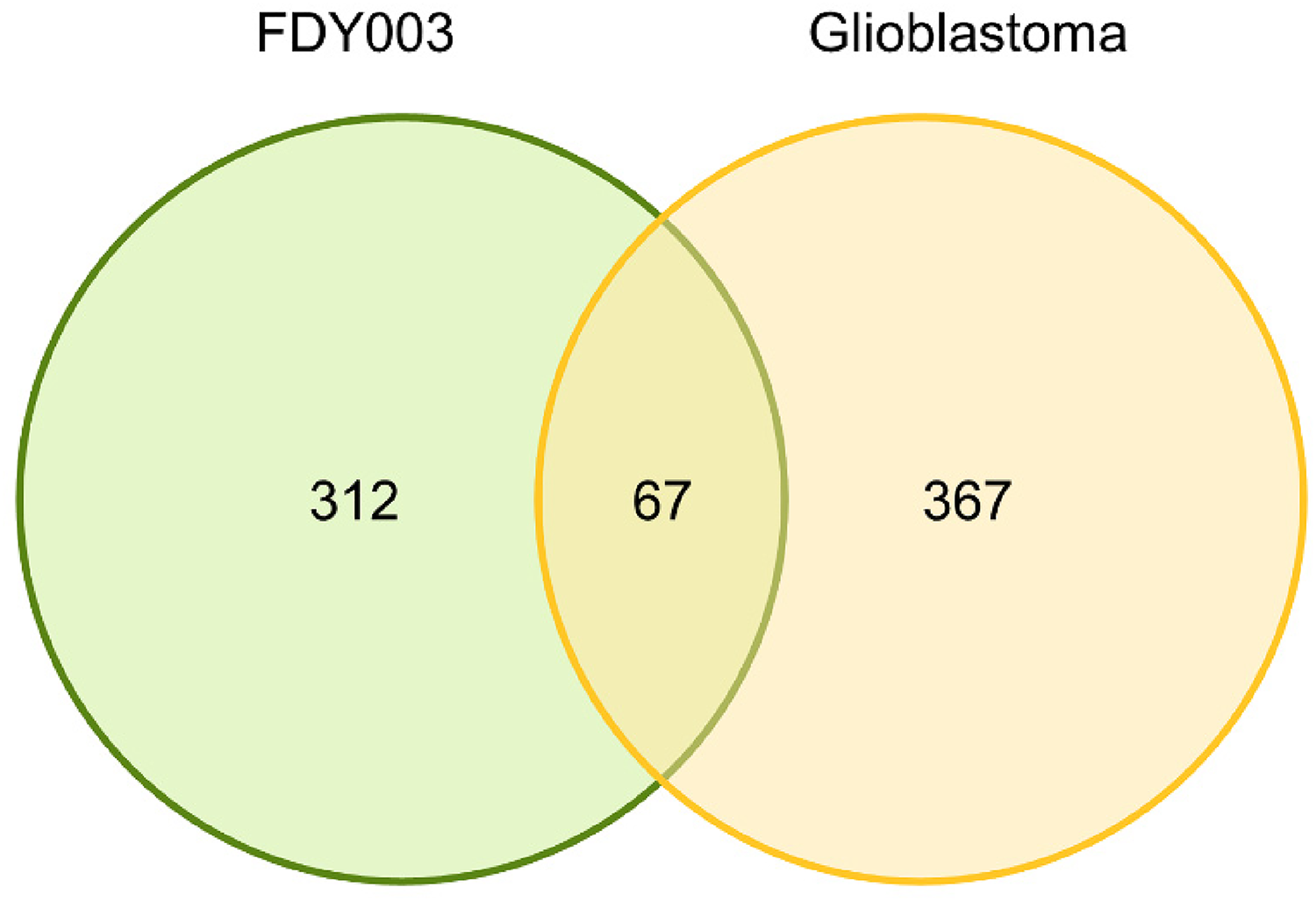
A venn diagram of FDY003 targets (green circle) and GBM-associated genes (yellow circle).
Network Pharmacological Dissection of Anti-GBM Mechanisms of FDY003
For the network pharmacological dissection of the anti-GBM mechanism of FDY003, its H-A-T network was generated by integrating comprehensive data (Figure 3 and Supplementary Table S3). In the network, 76.12% (51 of 67 targets) of targets belonged to quercetin (Figure 3 and Supplementary Table S3), which suggests that it may have the most important pharmacological role. Additionally, 89.55% (60 of 67 targets) belonged to ≥ 2 pharmacological compounds (Figure 3), which indicates that FDY003 shows multiple compound-multiple target characteristics.

The herbal medicine–active pharmacological compound–target network for FDY003. Green node, herbal medicine; red node, active pharmacological compound; blue node, glioblastoma-related target.
Analyzing the interaction mechanisms of the druggable targets may facilitate the investigation of the pharmacological effects and regulatory mechanisms of therapeutic agents.12,13 Hence, we integrated the interaction data between the GBM-related targets of FDY003 into a protein–protein interaction (PPI) network and investigated its topological characteristics to search for hub nodes within the network (Figure 4). Of note, hub nodes are those whose degrees greatly exceed the average node degree and are effective for mediating the pharmacological effects of drugs in cells and tissues.14,15 In the analysis, nodes were defined as hubs if their degree was ≥2x average node degree.4-7 The hub targets identified were AKT1, PIK3CA, PIK3R1, SRC, STAT3, TP53, and VEGFA (Figure 4), suggesting their function as key mediators for conferring FDY003 activity for GBM treatment.

The protein–protein interaction network for glioblastoma-related targets of FDY003. Purple node, hub; blue node, non-hub.
We examined the functional enrichment of the targets to explore the potential regulatory mechanisms of FDY003 for the anti-GBM activity of the identified targets. The FDY003 targets were modulators for the cell cycle process, proliferation, and survival of GBM cells (Supplementary Figure S1), which could potentially explain its therapeutic activity for GBM treatment. In addition, FDY003 pharmacologically interacted with various pathways that are implicated in the carcinogenesis and progression of GBM (Figure 5 and Supplementary Figure S1), suggesting the involvement of multiple pathways in the action mechanism of FDY003. Moreover, we found that FDY003 may affect the gene expression levels of the crucial components of those targeted pathways in human GBM cells (Supplementary Figure S2).

The herbal medicine–active pharmacological compound–target pathway network for FDY003. Green node, herbal medicine; red node, active pharmacological compound; blue node, glioblastoma-related target; orange node, glioblastoma-related pathway.
Examination of the Molecular Binding Interaction Between the Pharmacological Compounds and the Targets of FDY003
To examine the molecular binding interaction between the pharmacological compounds and the identified targets of FDY003, we investigated their binding affinities by calculating the molecular docking scores. The binding energies of the pharmacological compounds and their targets were ≤ −5.0 kcal/mol (Figure 6 and Supplementary Figure S3), which suggests their significant binding capacity.

Evaluation of the binding activities between the active pharmacological compounds of FDY003 and their targets. (A) Isorhamnetin-AKT1 (binding energy = −6.4 kcal/mol). (B) Kaempferol-AKT1 (binding energy = −6.7 kcal/mol). (C) Kaempferol-Src (binding energy = −8.5 kcal/mol). (D) Kaempferol-STAT3 (binding energy = −9.0 kcal/mol). (E) Kaempferol-TP53 (binding energy = −8.8 kcal/mol). (F) Luteolin-AKT1 (binding energy = −6.6 kcal/mol). (G) Luteolin-STAT3 (binding energy = −7.7 kcal/mol). (H) Luteolin-TP53 (binding energy = −9.1 kcal/mol). (I) Luteolin-VEGFA (binding energy = −8.4 kcal/mol). (J) Quercetin-AKT (binding energy = −6.9 kcal/mol). (K) Quercetin-PIK3CA (binding energy = −8.2 kcal/mol). (L) Quercetin-PIK3R1 (binding energy = −9.7 kcal/mol).
In this study, we employed a network pharmacological approach to elucidate the mechanisms underlying the anti-GBM characteristics of the traditional Korean medicinal formula FDY003. The overall network pharmacological analyses determined active pharmacological compounds and targets of FDY003 for its anti-GBM activity and further investigated its molecular and signaling mechanisms. The results may contribute to the understanding of the pharmacological basis of the anti-GBM properties of FDY003, along with its systematic mechanisms.
The key GBM-associated targets of FDY003 may participate in the pathological processes of GBM, and their targeting has been shown to exhibit substantial anticancer effects.
The pathways involving the FDY003 targets are important signaling mechanisms for the pathological processes underlying GBM. The calcium signaling pathway modulates the growth, mobility, tumor microenvironment organization, cancer stemness, and cancer-promoting inflammation of GBM cells. 36 The chemokine pathway plays a key role in tumor microenvironment remodeling to induce glioblastomagenesis, metastasis, tumor progression, neovascularization, angiogenesis, microglia and macrophage polarization, invasion, and migration of GBM cells and tumors. 37 This pathway serves as a potential target site for anti-angiogenic and immunotherapeutic strategies for GBM treatment. 37 The erythroblastic leukemia viral oncogene homolog (ErbB), mitogen-activated protein kinase (MAPK), nuclear factor kappa-light-chain-enhancer of activated B cells (NF-κB), phosphoinositide 3-kinase (PI3K)-Akt, and Ras pathways are involved in tumorigenicity, migration, proliferation, stemness, survival, cell cycle control, therapy sensitivity, angiogenesis, invasiveness, and inflammation of GBM cells and tumors.38-46 The expression and activity of the signaling components of these pathways may predict the prognosis of patients with GBM and may function as relevant action sites for anti-GBM therapeutic interventions.38-46 Targeting the focal adhesion and fork head box O (FOXO) pathways can suppress proliferation, migration, stemness, survival, angiogenesis, motility, and invasion of GBM cells and tumors while elevating the treatment outcomes.47-53 Their expression profiles have been related to the clinical prognosis of patients with GBM.47-53 Various cancerous and tumorigenic mechanisms, including cell immortalization, invasion, metastasis, angiogenesis, migration, metabolic reorganization, proliferation, and stemness, are linked to the hypoxia-inducible factor (HIF)-1 pathway in the pathological processes of GBM.54,55 The janus kinase (JAK)-signal transducer and activator of transcription (STAT) pathway participates in the carcinogenesis and progression of GBM by inducing migration, invasion, EMT, cancer stem-like abilities, inflammation, immunosuppression, angiogenesis, proliferation, and therapeutic resistance. 56 This activity of the pathway correlates with the survival and prognosis of patients with GBM and is a potential target for cytotoxic and immune checkpoint therapies. 56 Dysregulation of the p53 pathway, the most frequent event in GBM pathology, causes metastasis, growth, self-renewal potential, and anti-apoptosis of GBM cells and tumors, and its activity status is a predictor of treatment response and prognosis of patients with GBM. 30 The programmed death-ligand 1 (PD-1)/programmed cell death-ligand 1 (PD-L1) pathway is an immunoregulatory cascade whose deregulation can cause the escape from the immune surveillance of GBM cells and tumors and targeting the pathway can restore and enhance immunity.57-59 The tumor necrosis factor (TNF) pathway controls not only the cancerous inflammation but also the radiosensitivity, chemosensitivity, migration, invasiveness, and growth of GBM cells and tumors.60-64 The TNF pathway also functions as a predictor for prognosis of survival rate and disease progression in patients with GBM.60-64 The vascular endothelial growth factor (VEGF) pathway is a primary angiogenic driver of GBM cells and tumors, and it coordinates their immune responses, migration, proliferation, and invasion.65-67 This pathway may also predict the survival, prognosis, metastasis, and recurrence of GBM patients, and it is the main druggable site for anti-angiogenic treatment strategies against GBM.65-67
The active pharmacological compounds identified in FDY003 act as therapeutic components for GBM treatment. Cm induces apoptosis and autophagy in GBM cells and arrests their growth and cell cycle process by targeting caspase, B-cell lymphoma 2 (Bcl-2), autophagy-related (Atg) 5, microtubule-associated protein 1A/1B-light chain 3 (LC3), p53, p21, extracellular signal-regulated kinase (ERK), and PI3K/Akt cascades. 68 Cordycepin represses migration, proliferation, and survival of GBM cells through the pharmacological modulation of focal adhesion kinase (FAK), paxillin, integrin, Akt, p38, caspase, Bcl-2, Fas, Fas ligand (FasL), Bcl2-antagonist/killer (Bak), and poly (adenosine diphosphate-ribose) polymerase (PARP) signaling.69,70 Eriodyctiol intervenes in the activities of PI3K/Akt, glycogen synthase kinase (GSK)-3β, NF-kappa B, cadherin, p38, and zinc finger E-box binding homeobox 1 (ZEB1) pathways in GBM cells, which leads to the inhibition of their migration, invasion, EMT, metastasis, survival, and proliferation. 71 Kaempferol promotes oxidative stress, autophagy, pyroptosis, and death of GBM cells while repressing their drug resistance, growth, and migration through the regulation of reactive oxygen species (ROS), multidrug resistance-associated protein 1 (MRP1), superoxide dismutase 1 (SOD-1), thioredoxin, Bcl-2, caspase, PARP, interleukins (ILs), monocyte chemoattractant protein 1 (MCP-1), and regulated on activation, normal T cell expressed and secreted (RANTES) cascades.72, 73 Luteolin decreased the ability of GBM cells to induce cell viability, cell adhesion, invasion, migration, angiogenesis, cancer stemness, colony formation, chemoresistance and radioresistance, cell cycle progression, EMT, and inflammation. Therapeutic modulation of signaling cascades, including the musashi RNA binding protein 1 (MSI1), epidermal growth factor receptor (EGFR), insulin-like growth factor 1 receptor (IGF1R), MAPK, PI3K/Akt, caspase, PARP, matrix metalloproteinases (MMPs), tissue inhibitor of metalloproteinases (TIMPs), ILs, cyclooxygenase-2 (COX-2), NF-κB, survivin, Bcl-2, induced myeloid leukemia cell differentiation protein (Mcl-1), cell division control protein 42 homolog (CDC42), protein kinase C (PKC) α, X-linked inhibitor of apoptosis protein (XIAP), inducible nitric oxide synthase (iNOS), and cyclin pathways are the key signaling mechanisms underlying the anti-GBM activities of luteolin.74-80 Quercetin acts on various signaling pathways, such as p53, mouse double minute 2 homolog (MDM2), Bcl-2, Bcl-2-associated X protein (Bax), PARP, caspase, cyclin, ERK, p38, c-Jun N-terminal kinase (JNK), PI3K/Akt, VEGF-A, MMPs, fibronectin, Axl, NF-κB, phospholipase D1 (PLD1), IL-6/STAT3, COX-2, Beclin-1, LC3, Atg7, heat shock proteins (HSPs), endoplasmic reticulum (ER) stress, and ROS pathways, to demonstrate its anti-growth, anti-survival, anti-migratory, anti-invasive, anti-angiogenic, chemosensitizing, and radiosensitizing effects in GBM cells. 81
In summary, we utilized a network pharmacological approach to evaluate the effects and mechanisms of FDY003 on GBM. Further extensive studies investigating the mechanisms of FDY003 targeting various pathological behaviors of cancer cells are important. Understanding the cancer responses of various traditional Korean medicinal formula also can help expand the pharmacological role and usage of traditional Korean medicines as anti-tumor therapies.
Materials and Methods
A list of abbreviations used in the study are provided in Supplementary Table S4.
Cell Culture
The U-87 MG human GBM cell line was purchased from the Korean Cell Line Bank (Seoul, Korea) and was incubated in Dulbecco's Modified Eagle Medium (WELGENE Inc., Gyeongsang, Korea) with 10% fetal bovine serum (WELGENE Inc.) and penicillin-streptomycin (Thermo Fisher Scientific, Inc., Waltham, MA, USA) at 37 °C-5% CO2.
Preparation of Drugs
First, we obtained all the dried raw medicines that comprise FDY003 (ie, Act, Cm, and LjT) from Hanpure Pharmaceuticals (Pocheon, Korea). Subsequently, we mixed and ground AcT (6.25 g), LjT (4.16 g), and Cm (6.25 g), placed them in 70% ethanol (500 mL), and performed an 80 °C reflux extraction process for 3 h. The extract was filtered using a 1-μm pore size-filter paper (Hyundai Micro, Seoul, South Korea). After concentration to dryness by rotary evaporation, 80% ethanol (500 mL) was added to the dried residue and stirred for 1 h. After filtration using a 1-μm pore size-filter paper (Hyundai Micro, Seoul, South Korea) and concentration by rotary evaporation, 90% ethanol (500 mL) was added to the dried material and further stirred for 1 h. This mixture was subsequently concentrated by rotary evaporation. After adding distilled water (500 mL) to the dried material, the solvent was sequentially filtered through 1-μm and 0.6-μm pore size-filter papers (Hyundai Micro, Seoul, South Korea), and cryodesiccated at − 80 °C. The extract was stored at − 20 °C and dissolved in distilled water immediately before use.
Determination of the Cell Viability of Drug-Treated Cells
The effects of drug treatment on cell viability were investigated using a water-soluble tetrazolium salt (WST-1) assay. In brief, we seeded 1.0 × 104 cells in the 96-well plates and treated them with either FDY003, temozolomide (Sigma-Aldrich, St. Louis, MO, USA), or both for 72 h. Subsequently, we incubated the drug-treated cells for 2 h with WST-1 solution (Daeil Lab Service Co. Ltd, Seoul, Korea) at 37 °C-5% CO2. We determined cell viability by examining the absorbance at 450 nm using an xMark microplate absorbance spectrophotometer (Bio-Rad, Hercules, CA, USA).
Quantitative Real-Time PCR
RNA isolation from the harvested cells was performed using an easy-BlueTM RNA Extraction Kit (iNtRON Biotech, Seongnam, Korea), according to the manufacturer's protocols. Then, the complementary DNA (cDNA) synthesis of the samples (total RNA 2.5 μg) was performed using a cDNA Synthesis Kit RT (iNtRON Biotech, Seongnam, Korea), according to the manufacturer's protocols. Quantitative real-time polymerase chain reaction (PCR) was performed with primers (Supplementary Table S5) through a StepOnePlus PCR system instrument (Applied Biosystems, Waltham, MA, USA) by using SYBR Green Master Mix (Applied Biosystems, Waltham, MA, USA). The transcription levels of the genes of interest were normalized to those of
Screening of the Active Pharmacological Compounds and Targets of FDY003
To screen the active pharmacological compounds of FDY003, we searched the following databases: the Bioinformatics Analysis Tool for Molecular Mechanism of Traditional Chinese Medicine, 82 Traditional Chinese Medicine Systems Pharmacology, 11 and Anticancer Herbs Database of Systems Pharmacology. 83 We obtained a list of the pharmacological compounds in FDY003 and retrieved detailed information on their pharmacokinetic properties. Among the investigated compounds, we screened the compounds that are active on the basis of their drug-likeness, oral bioavailability, and Caco-2 permeability, which are the most useful pharmacokinetic parameters for active compound screening.10,11 Drug-likeness examines the potential of chemicals to be designed and developed as drugs based on their biophysical characteristics. 11 Chemicals whose drug-likeness is ≥ 0.18 possess a strong potential to act as drugs. 11 Oral bioavailability examines the effectiveness of chemical delivery to the drug action sites after oral administration. 11 Chemicals whose oral bioavailability is ≥ 30% are generally effectively delivered to the targeted tissues and organs. 11 Caco-2 permeability examines the ability of chemicals to permeate and be absorbed in the intestines. 11 Chemicals whose Caco-2 permeability is ≥ −0.4 can permeate effectively and become absorbed in intestines. 11 Based on these reports, compounds with drug-likeness ≥ 0.18, oral bioavailability ≥ 30%, and Caco-2 permeability ≥ −0.4 were included in the list of active pharmacological compounds of FDY003.10, 11
After the active pharmacological compound screening, we entered the simplified molecular-input line-entry system notation (investigated from PubChem 84 ) of the active pharmacological compounds of FDY003 to the following databases: the Search Tool for Interactions of Chemicals 5, 85 PharmMapper, 86 SwissTargetPrediction, 87 and Similarity Ensemble Approach 88 to obtain their targets. The GBM-associated genes and proteins that participate in their pathological mechanisms were obtained from the following databases: Comparative Toxicogenomics Database, 89 DrugBank, 90 Pharmacogenomics Knowledgebase, 91 Therapeutic Target Database, 92 Online Mendelian Inheritance in Man, 93 GeneCards, 94 Human Genome Epidemiology Navigator, 95 and DisGeNET. 96
FDY003-Related Network Construction
Biological networks, including FDY003-related networks, contain nodes and links describing their molecular and functional interactions. 97 In the herbal medicine–active pharmacological compound–target (H-A-T) network for FDY003, its medicinal materials, pharmacological compounds, and their targets act as nodes, and the interaction relationships among these nodes act as links. The H-A-T-Pathway (H-A-T-P) network of FDY003 was built by adding the connections between the FDY003 targets and their GBM-associated signals. In the PPI network for FDY003, its targets act as nodes, and their interactions (with a confidence score ≥ 0.9 investigated using STRING 98 ) act as links. The number of links that a node has is called a degree. 97 Network production, drawing, and analysis were performed using Cytoscape. 99
Functional Analysis of FDY003 Targets for Their Enrichments in Gene Ontology and Pathways
We used Database for Annotation, Visualization and Integrated Discovery (DAVID)
100
and Kyoto Encyclopedia of Genes and Genomes (KEGG)
101
for the functional analysis of FDY003 targets for their enrichments in gene ontology and pathways, respectively. The results with
Evaluating the Molecular Interaction Affinities of Pharmacological Compounds and Targets of FDY003
To examine the binding interaction between the pharmacological compounds and the targets of FDY003, we investigated the molecular docking scores of compound–target pairs. In brief, we entered the structural data of FDY003 compounds (obtained from RCSB Protein Data Bank 102 ) and their targets (obtained from PubChem 84 ) to Autodock Vina 103 to calculate their scores for molecular docking. Following previous studies,4-7 binding energy ≤ −5.0 kcal/mol was indicative of strong molecular interaction affinities for the compounds and their targets.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X221126311 - Supplemental material for Unveiling the Mechanism of the Traditional Korean Medicinal Formula FDY003 on Glioblastoma Through a Computational Network Pharmacology Approach
Supplemental material, sj-docx-1-npx-10.1177_1934578X221126311 for Unveiling the Mechanism of the Traditional Korean Medicinal Formula FDY003 on Glioblastoma Through a Computational Network Pharmacology Approach by Ho-Sung Lee, In-Hee Lee, Sang-In Park, Minho Jung, Seung Gu Yang, Tae-Wook Kwon and Dae-Yeon Lee in Natural Product Communications
Footnotes
Author Contributions
Conceptualization: Ho-Sung Lee, In-Hee Lee, Dae-Yeon Lee. Methodology: Ho-Sung Lee, In-Hee Lee, Dae-Yeon Lee. Data collection: Ho-Sung Lee, In-Hee Lee, Sang-In Park, Minho Jung, Seung Gu Yang, Tae-Wook Kwon. Data analysis and investigation: Ho-Sung Lee, In-Hee Lee, Dae-Yeon Lee. Writing: Ho-Sung Lee, In-Hee Lee, Dae-Yeon Lee. All authors read and approved the final manuscript.
Data Statement
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Ethical Approval is not applicable for this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (No. 2021R1F1A1049472).
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
