Abstract
Colorectal cancer (CRC) originates from the uncontrolled growth of epithelial cells in the colon or rectum. Annually, 1.9 million new CRC cases are being reported, causing 0.9 million deaths worldwide. The suppressive effects of the herbal prescription FDY003, a mixture of
Introduction
Colorectal cancer (CRC) is the second leading cause of cancer-related deaths worldwide. 1 Approximately 1.9 million new CRC cases and 0.9 million CRC-related deaths are reported every year. 2 The primary pharmacological strategies for CRC treatment are cytotoxic chemotherapy and targeted therapy. 1 Despite advances and breakthroughs that have improved the safety and effectiveness of treatment regimens, the subpopulation of patients who benefits from CRC therapeutics is still limited. 1 Development and investigation of herbal medicines and natural products are part of efforts to overcome these limitations.3,4 Herbal medicines and natural products have been known for tumor-suppressing activities in diverse types of cancers and malignant tumors, including CRC, as well as chemo- and radiosensitivity to anticancer therapies, while alleviating toxic effects.3,4 They enhance survival, tumor response, immune function, and quality of life and repress the emergence of adverse reactions and relevant symptoms in patients with CRC who received standard anticancer therapies.5,6
FDY003 is an herbal formula containing three medicinal herbs,
Multiple-compound-multiple-target pharmacological mechanisms are the most distinct features of herbal medicines compared with other types of therapeutics. 14 They generally contain a wide array of pharmaceutical compounds that may regulate the expression levels and activities of many genes and proteins closely related to the pathology of a disease. 14 The pharmacological effects of herbal medicines are then determined through the functional influences given to the genetic and protein–protein interaction (PPI) networks that are made up of complex interactions among the corresponding targets of the pharmaceutical compounds present in the herbal medicines. 14 The network pharmacology (NP) approach has appeared to be the most powerful tool for investigating multiple-compound-multiple-target pharmacological mechanisms of action of herbal medicines. 14 NP analyzes the physicochemical and functional connection between the pharmaceutical compounds of herbal medicines and their targeted genes and proteins, and the disease-related pathways that consist of the targets in a form of “herbal drug-pharmaceutical compound-targeted genes and proteins-signaling pathway” network, and investigates the polypharmacological mechanisms of herbal medicines using by integrating diverse network analysis methodologies. 14 Thus, by applying the NP technique, we studied the multiple-compound-multiple-target activity of FDY003 in CRC treatment from a network perspective.
Materials and Methods
Cell Culture
HT-29 human CRC cell line (Korean Cell Line Bank, Seoul, Korea) was grown using Dulbecco's modified Eagle's medium (Gibco BRL, Grand Island, NY, USA) that contained 10% fetal bovine serum (Gibco BRL, Grand Island, NY, USA), 100 µg/mL streptomycin (Gibco BRL, Grand Island, NY, USA), and 100 U/mL penicillin (Gibco BRL, Grand Island, NY, USA) at 37 °C-5% CO2.
Preparation of FDY003 Herbal Formula
AcT (150.0 g), Cm (100.0 g), and LjT (150.0 g), which were purchased from Hanpure Pharmaceuticals (Pocheon, Korea), were dried, mixed, and milled together. The powdered herbs were reflux-extracted with 70% ethanol (500 mL) for 3 h at 80 °C. The extract was filtered and successively purified with 80% and 90% ethanol. This solution was then lyophilized at − 80 °C, yielding 50.4 g of dried extract, which was stored at − 20 °C and dissolved in distilled water prior to the experiment.
CRC Cell Viability Assay
To test cell viability, 1.0 × 104 cells/well were seeded in a 96-well plate at 37 °C-5% CO2 and then incubated for 72 h after treatment with FDY003 and/or 5-fluorouracil (5-FU; Sigma-Aldrich, St. Louis, MO, USA). The cells were then treated with 10 µL of water-soluble tetrazolium salt solution (WST-1; Daeil Lab Service Co. Ltd, Seoul, Korea) and incubated at 37 °C-5% CO2 for 2 h. Cell viability was assessed by reading the 450-nm absorbance of the microplates using an xMark microplate absorbance spectrophotometer (Bio-Rad, Hercules, CA, USA). Statistical significance was determined using Student's
Pharmaceutical Compounds of FDY003 and Their Interacting Genes
The pharmacokinetic characteristics of the chemical compounds present in FDY003 were retrieved from the Anticancer Herbs Database of Systems Pharmacology, 15 Traditional Chinese Medicine Systems Pharmacology, 16 and Bioinformatics Analysis Tool for Molecular Mechanism of Traditional Chinese Medicine. 17 The compounds of FDY003 capable of pharmacological action were identified based on Caco-2 permeability (CP), drug-likeness (DL), and oral bioavailability (OB), where CP ≥ −0.4 indicates strong in vivo permeability of a compound, DL score ≥ 0.18 indicates potential in a compound to be a drug, and OB value ≥ 30% indicates effective absorption and distribution.7,16,18
For target screening with respect to the pharmaceutical compounds of FDY003, we downloaded the simplified molecular-input line-entry system (SMILES) notation of the pharmaceutical compounds from PubChem 19 database, and imported them into the Similarity Ensemble Approach, 20 Search Tool for Interactions of Chemicals 5, 21 PharmMapper, 22 and SwissTargetPrediction 23 platforms. After obtaining the targets of the pharmaceutical compounds from the aforementioned databases and integrating them, we eliminated the duplicates and standardized the unoverlapped targets using the UniProt database. 24
To screen genes and proteins that potentially contribute to CRC pathology, we used the keywords “colorectal cancer” in the following databases: Human Genome Epidemiology Navigator, 25 GeneCards, 26 Therapeutic Target Database, 27 Comparative Toxicogenomics Database, 28 DrugBank, 29 Pharmacogenomics Knowledgebase, 30 Online Mendelian Inheritance in Man, 31 and DisGeNET. 32 After integrating the CRC-related target genes obtained from these databases, we eliminated duplicates and standardized the non-overlapped target genes using the UniProt database. 24
Networks for the Investigation of Anti-CRC Mechanisms of FDY003
A network generally consists of nodes that refer to individual components and factors of network systems and edges that refer to their interactions and associations. 33 “Degree” refers to the quantity of edges that a node carries, and generally nodes having relatively more degrees show a tendency to play more important roles in a network. 33
In the case of the PPI network, the nodes are CRC-related genes targeted by the pharmaceutical compounds of FDY003, and the edges are interaction data between the targets obtained from the STRING database
34
(confidence score ≥ 0.9) for
The building and analysis of D-P-T, D-P-T-S, and PPI networks were made using the Cytoscape software. 35
Determination of the Gene Ontologies and Pathways Enriched Target Genes of FDY003
The gene ontology (GO) terms and pathways, significantly (
Molecular Docking Study for Core Targets and Pharmaceutical Compounds of FDY003
The chemical structures of the compounds of FDY003 (in a structure data file format) were acquired from PubChem. 19 The 3D protein structures (in a protein data bank format) of the core targets of FDY003 were acquired from the RCSB Protein Data Bank. 38 Elimination of water molecules and ligands, hydrogenation, and charge redistribution of the structures were conducted using PyMol. 39 Subsequently, the binding energies between the compounds and their targets were obtained using AutoDock Vina. 40 Compound-target pairs exhibiting binding energies of ≤ −5.0 kcal/mol were considered to express strong binding affinities.7–13
Results
Treatment Effects of FDY003 on CRC Cell Viability
The viability of HT-29 human CRC cells decreased significantly (
Pharmaceutical Compounds of FDY003 and Their Target Genes
Among the various chemical compounds that FDY003 contains, we identified those that play major roles in exhibiting the pharmacological effects of the herbal prescription using the pharmacokinetic characteristics of the individual compounds (see Materials and Methods) (Supplementary Tables S1 and S2). Some compounds whose pharmacokinetic parameters were outside the range of the selection criteria were nevertheless listed as pharmaceutical compounds due to their antitumor activities (Supplementary Tables S1 and S2). Subsequently, the catalog of genes interacting with and targeted by the pharmaceutical compounds of FDY003 was made based on the protein–chemical interaction data (Supplementary Table S3), which revealed 289 FDY003 targets, of which 90 were CRC-related (Figure 1 and Supplementary Table S3).
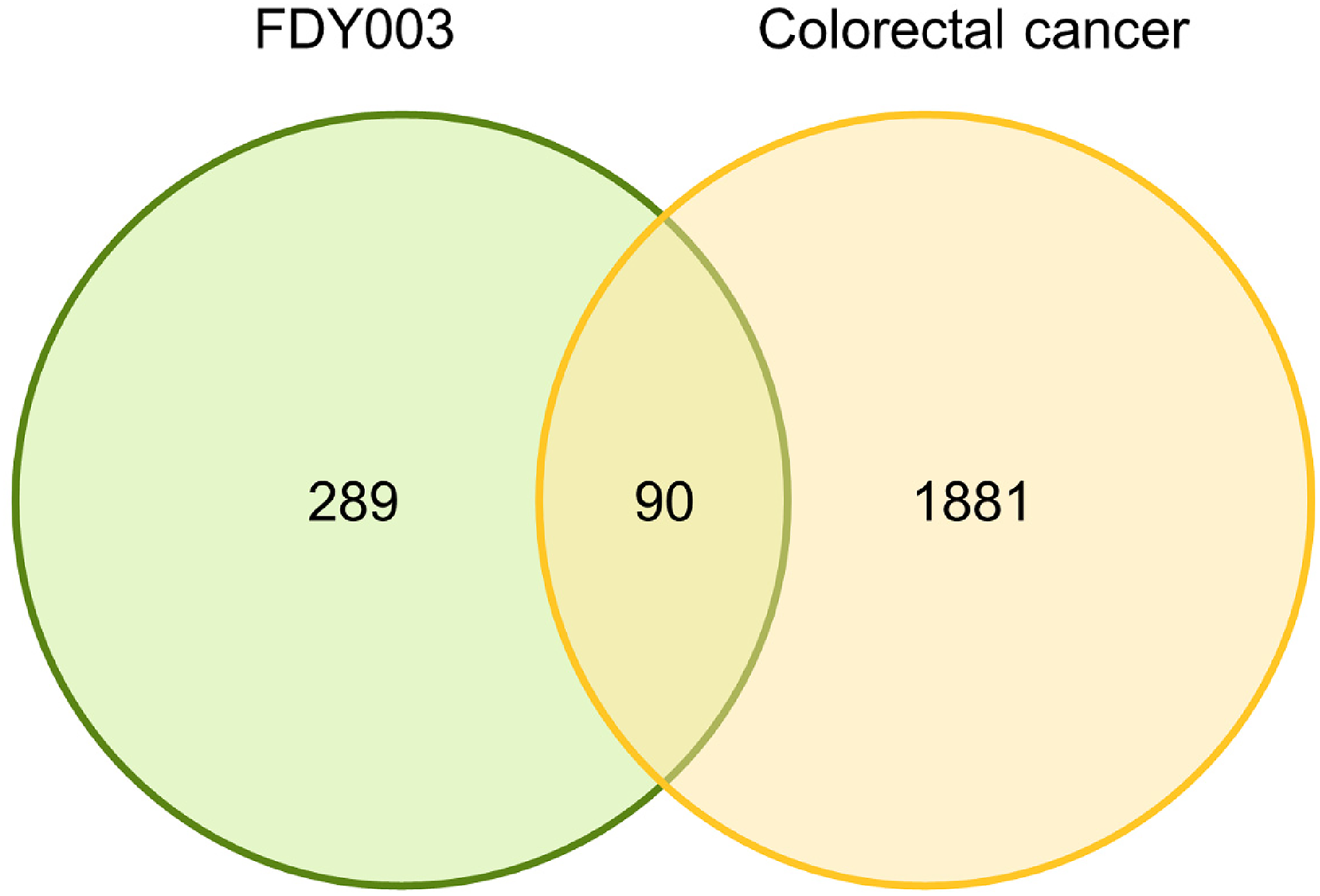
Venn diagram of FDY003 targets (green circle) and CRC-related genes (yellow circle).
Drug-Pharmaceutical Compound-Target Gene Network of FDY003 Describing its Anti-CRC Mechanism
The D-P-T network of FDY003, which was generated based on the relationships and interactions between its herbal components, pharmaceutical compounds, and CRC-related targeted genes, describes the multiple-compound-multiple-target pharmacological features of the herbal prescription for CRC treatment (Figure 2 and Supplementary Table S3). There were 17 pharmaceutical compounds of FDY003 that targeted 90 CRC-related genes, and all compounds interacted with ≥ 2 of the target genes (Figure 2 and Supplementary Table S3), demonstrating its polypharmacological aspect.

Drug-pharmaceutical compounds-target gene network of FDY003 for CRC treatment. Green nodes, herbal components; red nodes, pharmaceutical compounds; blue nodes, CRC-related targeted genes.
The pharmacological activity of a specific drug is conferred through the genetic and PPI network consisting of the genes and proteins that are the therapeutic target molecules of the compounds contained in the drug.42,43 Meanwhile, identifying the core therapeutic targets of a specific drug among the diverse cellular genes, metabolites, and proteins has been proven to be useful in dissecting its pharmacological mechanisms.42,43 For the exploration of core targets from the 90 CRC-related targets of FDY003, we constructed a PPI network (66 nodes and 561 edges) for the targets.42,43 Subsequently, degree centrality (DC), which is one of the most efficient measures used to define nodes with core functions in a network,42,43 was assessed for the FDY003 targets in the PPI network. DC evaluates the connections of a node,42,43 and hub nodes with higher degrees are considered favorable drug targets because targeting them can induce strong pharmacological influence in the cellular networks after drug administration.42,43 We defined hubs to be the nodes with degrees possessing a number of connections ≥ 2 × average degree value per node (6.17).7–13 DC analysis indicated that the following nodes, AKT1 (DC = 17), CYP1A1 (DC = 13), CYP3A4 (DC = 14), EGFR (DC = 13), ESR1 (DC = 15), JUN (DC = 15), PIK3R1 (DC = 17), PIK3R1 (DC = 17), SRC (DC = 20), TP53 (DC = 21), and VEGFA (DC = 14), satisfied the inclusion criteria for being considered a hub (Figure 3), suggesting the core targets for the anti-CRC activity of FDY003.

PPI network of CRC-related targeted genes of FDY003. Claret nodes, hub nodes; blue nodes, non-hub nodes.
For the understanding of the mechanisms of action of FDY003 in CRC treatment by targeting CRC-related genes, target-enriched GO terms were surveyed. The results showed that pharmaceutical compounds in FDY003 may target CRC-related genes that are determinants of a number of cellular phenotypes, including survival, death, proliferation, stress response, and chemical response of CRC cells (Figure 4). Moreover, the KEGG pathways significantly (

Gene ontology and pathway enriched with the colorectal cancer-related genes targeted by FDY003. (A) The bubble chart of the results of the gene ontology enrichment analysis. (B) The bubble chart of the results of the KEGG pathway enrichment analysis.
Molecular Docking Study for Determining the Binding Between the Pharmaceutical Compounds of FDY003 and the Proteins Encoded by Their Interacting Target Genes
We performed a molecular docking study to evaluate the binding between the pharmaceutical compounds of FDY003 and the proteins encoded by their interacting hub target genes (eg, AKT1, CYP1A1, CYP3A4, EGFR, ESR1, JUN, PIK3R1, SRC, TP53, and VEGFA) to assess their interaction affinities and binding sites. Notably, the binding energy of ≤ −5.0 is commonly used to prove that a given compound and a target shows effective binding affinity.7–13 The analysis results suggested the effective binding of the pharmaceutical compounds of FDY003 to the proteins encoded by their hub target genes (binding energies ≤ −5.0 kcal/mol) (Figure 5A to C and Supplementary Figure S2A to K). AKT1 has residues Thr82, Tyr83, Glu17, Glu83, Gly16, Arg86, Ile84, Thr87, Rrg15, and Lys20 that may interact with quercetin (Figure 5A). CYP1A1 has residues Asp320, Phe224, Thr321, Phe123, Ser122, Ser120, Asp313, Leu312, Leu314, Ser116, Ala317, Gly316, Phe319, Phe258, and Leu254 that interact with isorhamnetin, and also residues Ile115, Asp313, Leu314, Phe123, Gly316, Ala317, Phe319, Asp320, Phe224, and Asn222 that interact with quercetin (Figure 5B and C). The residues of CYP3A4 interacting with eriodictyol (flavanone) as shown are Phe215, Asp76, Phe220, Thr224, Phe108, Phe106, Glu374, Glu373, Arg372, Ala370, and Met371, and those interacting with quercetin are Phe108, Phe220, Phe215, Arg372, Ala370, Leu373, and Glu374 (Supplementary Figure S2A and B). EGFR has residues Gly721, Ser720, Val726, Gly794, Ala743, Lys745, Met793, and Arg841 that may interact with kaempferol, and the residues Gly724, Phe723, Lys875, Glu872, Glu758, Arg748, and Leu747 that may interact with quercetin (Supplementary Figure S2C and D). The residues of ESR1 that interact with kaempferol include Trp393, Arg394, Leu327, Glu323, Pro327, Glu353, and Leu387 (Supplementary Figure S2E). JUN may interact with luteolin through its residues Ala366, Arg270, Lys273, and Lys271, and further with quercetin through residues Lys295, Val281, Leu273, Ser345, Val340, Met341, Ala293, Thr338, Ser342, and Val323 (Supplementary Figure S2F and G). Moreover, the residues of PIK3R1 that quercetin interacts with are Gln760, Asp258, Glu259, Ala759, Asn756, Pro757, and Leu725 (Supplementary Figure S2H). SRC interacts with quercetin through the residues Tyr340, Met341, Ser342, Gly344, Ser345, Leu273, Lys295, Thr338, Val323, Leu393, Ala463, Val402, Asp404, and Ile336 (Supplementary Figure S2I). Quercetin may interact with TP53 through Asn239, Leu137, Cys275, Gln136, Lys139, Ala276, Cys277, Ser121, Gln165, Gln167, and His168 residues (Supplementary Figure S2J). The residues of VEGFA that interact with quercetin were found to be Leu137, Asn239, His168, Ser121, Cys277, Ala276, Gln136, Lys139, and Cys275 (Supplementary Figure S2K). These results may help verify the interaction relationships between them obtained from the NP analysis.

Molecular docking diagrams of pharmaceutical compounds of FDY003 and the proteins encoded by their hub target genes. (A) AKT1-Quercetin (binding energy = −6.6 kcal/mol). (B) CYP1A1-Isorhamnetin (binding energy = −10.2 kcal/mol). (C) CYP1A1-Quercetin (binding energy = −10.4 kcal/mol). (D) CYP3A4-Eriodyctiol (flavanone) (binding energy = −8.8 kcal/mol). (E) CYP3A4-Quercetin (binding energy = −8.9 kcal/mol). (F) EGFR-Kaempferol (binding energy = −8.0 kcal/mol). (G) EGFR-Quercetin (binding energy = −8.2 kcal/mol). (H) ESR1-Kaempferol (binding energy = −8.6 kcal/mol). (I) JUN-Luteolin (binding energy = −8.8 kcal/mol). (J) JUN-Quercetin (binding energy = −9.2 kcal/mol). (K) PIK3R1-Quercetin (binding energy = −9.7 kcal/mol). (L) VEGFA-Quercetin (binding energy = −8.3 kcal/mol).
Discussion
The aim of this study was to determine the potential effects and mechanisms of action of FDY003 against CRC using the NP technique. FDY003 induced the repression of the viability of CRC cells and also strengthened their chemosensitivity. The NP approach identified 17 potential pharmaceutical compounds and 90 CRC-related genes that the compounds may target. The GO terms enriched with the targets were those involved in the control of a variety of CRC cell phenotypes, such as survival, death, proliferation, stress response, and chemical response of CRC cells. Moreover, the targeted genes of FDY003 were enriched in various KEGG pathways, such as the ErbB, focal adhesion, HIF-1, IL-17, MAPK, PD-L1/PD-1, PI3K-Akt, Ras, TNF, and VEGF, which coordinate the crucial pathological processes of CRC. The overall analysis results obtained from the NP methodology suggest a multiple-compound-multiple-target-multiple-pathway mechanism of FDY003 against CRC.
The key CRC-related target genes of FDY003 with hub-like characteristics (eg, AKT1, CYP1A1, CYP3A4, EGFR, ESR1, JUN, PIK3R1, SRC, TP53, and VEGFA) are the major inducers and contributors to the pathological procedures of CRC. The oncogenes
Targeted signaling of FDY003 in CRC is a pivotal mechanism for the pathology of CRC. Uncontrolled activities of the ErbB, focal adhesion, HIF-1, MAPK, PI3K-Akt, and Ras pathways are the major factors driving the initiation and progression of CRC because these pathways function as the most crucial modulators of the diverse malignant phenotypes of CRC cells and tumors.44–47,69,70 These oncogenic signaling pathways also contribute to the development of therapeutic resistance against various anti-CRC strategies, which may be circumvented by targeting their misregulated activities.44–47,69,70 IL-17 and TNF signaling are pro-inflammatory cascades associated with CRC progression by triggering epithelial-to-mesenchymal transition (EMT), invasion, metastasis, survival, immune suppression of microenvironment, proliferation, migration, angiogenesis, and tumorigenic potential of CRC cells and tumors.71,72 These pro-inflammatory signaling pathways can also be used to predict the prognosis of CRC patients.71,72 The PD-L1/PD-1 pathway negatively modulates immune cell activation and promotes an immunosuppressive tumor microenvironment, and blocking this immune checkpoint axis can inhibit the immune evasion of CRC cells and tumors.73,74 The VEGF pathway enhances the diverse metastatic and angiogenic properties of CRC cells and tumors, and inhibiting this pathway can block their growth and prolong the survival of patients with CRC.67,68
The medicinal constituents of FDY003 have been shown to exert pharmacological effects on CRC cells and tumors. Cm inhibits the division, growth, and survival of CRC cells and tumors by suppressing Ras/extracellular signal-regulated kinase (ERK) but upregulating caspase, p53, poly (ADP-ribose) polymerase (PARP), B-cell lymphoma 2 (Bcl-2)-associated agonist of cell death (BAD), Bcl-2 homologous antagonist/killer (Bak), and Bim pathways.75,76 Cirsimaritin upregulates PARP and caspase signaling in CRC cells to reduce their proliferation, survival, clonogenicity, and viability. 77 Cordycepin is an anti-CRC compound that decreases the survival, proliferation, invasion, viability, and migration of CRC cells by targeting Bcl-2 associated X-protein (Bax), death receptor (DR)-3, caspase, PARP, p53, c-Jun N-terminal kinase (JNK), p38, BH3 interacting-domain death agonist (Bid), Bim, p53 upregulated modulator of apoptosis (PUMA), prostaglandin E2 (PGE2), EP4, cyclic adenosine monophosphate (cAMP)-response element binding protein (CREB), AMP-activated protein kinase (AMPK), matrix metalloproteinases (MMP), and cyclooxygenase 2 (Cox-2) cascades.78–80 Eupalitin has anti-CRC properties that can activate the caspase pathway to promote apoptosis in CRC cells. 81 Genkwanin exerts antitumor, anti-inflammatory, and immunomodulatory activities in CRC cells and tumors. 82 The anti-growth and death-promoting functions of isorhamnetin are mediated through the heat shock protein (Hsp)-70, caspase, Src, β-catenin, and PI3K-Akt-mammalian target of rapamycin (mTOR) pathways.83–85 Kaempferol induces growth-arrest, death, and chemosensitization of CRC cells and tumors by targeting reactive oxygen species (ROS), p38, p53, PARP, p21, Bax, Bcl-2, thymidylate synthase (TS), phosphatase and tensin homolog (PTEN), PI3K-Akt, polypyrimidine-tract binding protein 1 (PTBP1)- pyruvate kinase M2 (PKM2), TNF-related apoptosis-inducing ligand (TRAIL)/DR-5, FAS, cyclin-dependent kinases (CDKs), cyclins, MAPK, Janus kinase (JAK)-signal transducers and activators of transcription (STAT), nuclear factor kappa-light-chain-enhancer of activated B cells (NF-κB), PUMA, ataxia telangiectasia mutated (ATM), H2A histone family member X (H2AX), insulin-like growth factor 2 (IGF-2), histidine-rich glycoprotein (HRG), insulin-like growth factor 1-receptor (IGF-1R), ErbB3, MMP, VEGF, nitric oxide (NO), and PGE-2 signaling.86–95 Luteolin modulates various cascades, [eg, MAPK, heme oxygenase-1 (HO-1), AMPK, Peroxisome Proliferator-Activated Receptor (PPAR)-γ/organic anion/cation transporter (OCTN)-2, calpain, ubiquitin like with PHD and ring finger domains 1 (UHRF1), DNA methyltransferase 1 (DNMT1), CREB, miR-384/pleiotrophin, cyclin, Wnt, β-catenin, glycogen synthase kinase (GSK)-3β, glutathione (GSH), nuclear factor erythroid 2-related factor 2 (Nrf2), HIF-1, IGF-2, IGF-1R, PI3K-Akt-mTOR, ERK, cell division cycle 25C (CDC25c), forkhead box transcription factor (FoxO), Beclin-1, PARP, Bcl-2, Bax, caspase, autophagy related 5 (Atg5), microtubule-associated protein 1 light chain 3B (LC3B)-I/II, focal adhesion kinase (Fak), Bim, BAD, N-cadherin, sphingosine kinase (Sphk)-1/Sphk-2, Sry-related HMG box (Sox), transient receptor potential vanilloids (TRPVs), VEGF, CDKs, p21, PARP, survivin, p21, myeloid cell leukemia (Mcl)-1, B-cell lymphoma-extra-large (Bcl-xL), mouse double minute 2 homolog (MDM2), Cox-2, MMPs, and tissue inhibitor metalloproteinase-2 (TIMP-2) cascades] to inhibit EMT, growth, carcinogenesis, migration, chemoresistance, invasion, survival, and metastasis of CRC cells and tumors.96–98 Quercetin acts on diverse cancer-associated pathways in CRC cells and tumors, affecting their various cancerous and tumorigenic behaviors, including proliferation, migration, survival, metastasis, invasion, cell division, cell cycle process, angiogenesis, stemness, inflammation, and therapeutic sensitivity.99,100 β-Sitosterol targets p53, p21, β-catenin, caspase, Cox-2, PGE2, TNF-α, NF-κB, Bax, Bcl-2, MDM2, cellular inhibitor of apoptosis 1 (cIAP-1), PARP, and breast cancer resistance protein (BCRP) pathways in CRC cells and tumors, leading to the inhibition of carcinogenesis, pro-tumorigenic inflammation, drug resistance, proliferation, and survival.101–104 Dietary intake of phytochemical compounds, such as quercetin and β-sitosterol, is inversely correlated with the risk of CRC incidence.99,100,105
The present study has several limitations and requires further experimental verification: (1) determination of the quantity of key pharmaceutical compounds contained in FDY003 and their individual pharmacological roles; (2) exploration of a variety of pharmacological roles, such as anti-self-renewal potential, anti-invasion, anti-EMT, anti-metastasis, anti-migration, and anti-angiogenesis effects of FDY003 in cancer cells; and (3) determination of the effectiveness and safety of FDY003 using extensive preclinical and clinical studies.
Conclusion
In this study, we used a NP approach to investigate the molecular mechanisms underlying the anti-CRC activity of FDY003. Seventeen pharmaceutical compounds of FDY003 exhibited anticancer effects by targeting 90 CRC-related genes. FDY003 targets were associated with the modulation of the stress response, chemical response, cell proliferation, growth, survival, and death of CRC cells, as well as signaling pathways associated with tumorigenesis and progression of CRC. The findings of this study make a significant contribution to cancer research and support the use of herbal medicines in CRC treatment.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X221126964 - Supplemental material for A Study on the Mechanism of Herbal Drug FDY003 for Colorectal Cancer Treatment by Employing Network Pharmacology
Supplemental material, sj-docx-1-npx-10.1177_1934578X221126964 for A Study on the Mechanism of Herbal Drug FDY003 for Colorectal Cancer Treatment by Employing Network Pharmacology by Ho-Sung Lee, In-Hee Lee, Sang-In Park, Minho Jung, Seung Gu Yang, Tae-Wook Kwon and Dae-Yeon Lee in Natural Product Communications
Footnotes
Ethical Approval
Ethical approval is not applicable for this article.
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Author Contributions
Conceptualization: Ho-Sung Lee, In-Hee Lee, Dae-Yeon Lee; Methodology: Ho-Sung Lee, In-Hee Lee, Dae-Yeon Lee; Data collection: Ho-Sung Lee, In-Hee Lee, Sang-In Park, Minho Jung, Seung Gu Yang, Tae-Wook Kwon; Data analysis and investigation: Ho-Sung Lee, In-Hee Lee, Dae-Yeon Lee; Writing: Ho-Sung Lee, In-Hee Lee, Dae-Yeon Lee. All authors read and approved the final manuscript.
Data Statement
All data generated or analyzed during this study are included in this published article and its supplemental materials file.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Research Foundation (Grant Number: 2021R1F1A1049472).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
