Abstract
Introduction
Constipation is one of the most common chronic gastrointestinal disorders that affect individuals of both sexes and all ages.1–3 Characteristics of constipation include infrequent or difficulty in defecation, hard stools and incomplete evacuation.4,5 It is estimated that ∼16% adults are suffering from constipation worldwide 6 ; thus, medical care services face high treatment costs annually. 7 Moreover, its prevalence has been reported as >20% in certain populations and regions, including Asia, due to high mental stress levels and environmental factors.2,8,9 In addition, severe constipation is associated with retentive fecal incontinence, mental health issues and reduced quality of life.10,11 Constipation in women is also associated with obesity and hormonal disorders.12,13 At present, numerous therapies have been used for the treatment of various degrees of constipation, such as dietary fiber, 14 laxatives,15,16 biofeedback therapy 17 and surgery. 18 However, the toxic side effects and efficacy of these clinical drugs remain a concern. Thus, the development of effective treatment options with manageable toxic side effects is required for use in the clinic.
Traditional Chinese Medicine (TCM) has been widely used in the treatment of constipation. For example, Zhao et al 19 reviewed the treatment with modified RunChang-Tang in functional constipation using a meta-analysis. Chen et al 20 demonstrated the use of Rhei Radix et Rhizoma (rhubarb) in the treatment of constipation. Chen et al 21 also investigated the tissue distribution, pharmacokinetics and pharmacodynamics of Dahuang-Gancao in mice with constipation. The present study aimed to investigate the use of Zhi Zhu Ma Ren Pill (ZZMRP), which includes dried young fruit of Citrus aurantium L. (Zhishi, ZS), Atractylodes macrocephala Koidz. (Baizhu, BZ), dry ripe fruit of Cannabis sativa L. (Huomaren, HMR), Cynanchum otophyllum Schneid. (Baishao, BS) and Aster tataricus L. f. (Zi-wan, ZW). BZ accounts for the largest dosage in ZZMRP, and this is believed to improve the function of the spleen. ZS is believed to relieve tension in the gastrointestinal tract, acting as a ministerial medicine. Based on the aforementioned factors, it was hypothesized that ZZMRP may exert protective effects on constipation. Furthermore, the targets and potential mechanisms underlying ZZMRP in the treatment of constipation require further analysis.
Network pharmacology analysis is a powerful tool for drug discovery and associated mechanisms. 22 Using a network pharmacology approach, the associations between drug, component, target, pathway and disease can be explored, and the underlying therapeutic mechanisms of drugs in diseases can be predicted. This may lead to the development of a novel theoretical basis for the mechanisms underlying TCM therapy in diseases. 23 In the present study, the underlying targets and molecular mechanisms of ZZMRP in constipation were investigated, using network pharmacology and molecular docking (Figure 1).

Experimental workflow of the present study.
Materials and Methods
Collection of the Primary Active Components and Associated Targets of the Herbs in ZZMRP
The TCM Systems Pharmacology (TCMSP) database and analysis platform database is a systematic pharmacology website that supplies details of the absorption, distribution, metabolism and excretion of herbal medicines, including drug likeness (DL), oral bioavailability (OB), Caco-2 permeability and blood–brain barrier. 24 Thus, the TCMSP database (https://tcmspw.com; accessed on December 4, 2021) 25 was used in the present study to determine the active components and targets involved in ZS, BZ, HMR, BS, and ZW. As OB and DL are the most crucial parameters for drug delivery, 26 OB was set to ≥30% and DL to ≥0.18 to obtain the active components and targets of ZS, BZ, HMR, BS, and ZW. Subsequently, the Chinese names of each herb “Zhi Shi,” “Bai Zhu,” “Huo Ma Ren,” “Bai Shao,” and “Zi Wan” were searched using the search tool. All attained targets were exported into the UniProt database (https://www.uniprot.org/; accessed on December 4, 2021) to obtain the gene symbol.
Target Screening of Constipation
Disease-related targets were acquired from the GeneCards (https://www.genecards.org), Disgenet (https://www.disgenet.org/), Online Mendelian Inheritance in Man (OMIM; https://omim.org), DrugBANK (https://go.drugbank.com/drugs), and Therapeutic Target Database (TTD; http://db.idrblab.net/ttd/) databases using “constipation” as the keyword, in which the relevance score ≥0.85 was set as the screening index in the GeneCards database.27–31 GeneCards is a searchable, integrative database that provides comprehensive, user-friendly information on all annotated and predicted human genes. 32 DisGeNET is a discovery platform containing one of the largest publicly available collections of genes and variants associated with human diseases. 33 OMIM is an online catalog of human genes and genetic disorders. 34 DrugBank is a web-enabled database containing comprehensive molecular information about drugs, and their mechanisms, interactions and targets. 29 TTD is a database providing information about the known and explored therapeutic protein and nucleic acid targets, the targeted disease, pathway information, and the corresponding drugs directed at each of these targets. 35 Microsoft Excel software was used to remove duplicates, to obtain a total of 1501 final disease-related targets.
Protein-Protein Interaction Network Construction and Analysis
The intersection targets of the herbs in ZZMRP and constipation were collected using Venny 2.1 (https://bioinfogp.cnb.csic.es/tools/venny/index.html; accessed on December 5, 2021). Subsequently, the STRING database (https://string-db.org/; accessed on December 5, 2021) was utilized to construct the Protein-protein interaction (PPI) network of the intersection target, where confidence scores were set as >0.7, with no changes to the other variables. 36 The STRING database is often used to predict both direct and indirect interactions of proteins. 37 Subsequently, the inputted PPI network in a TSV format was assessed for its topology properties using Cytoscape 3.8.0 software (https://cytoscape.org/). 38 As the most prevalent topological parameter, degree centrality (DC) indicated the central attribute of nodes in the PPI network. The core targets were verified using DC ≥2 × the median.
Functional and Pathway Enrichment Analysis
The Gene Ontology (GO) terms and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways39,40 are used for predicting the roles of underlying targets in gene function and signaling pathways. Thus, GO and KEGG were used in the present study using DAVID 6.8 (https://david.ncifcrf.gov/; accessed March 5, 2021) 41 to explore the mechanisms underlying ZZMRP in the treatment of constipation.
Herb, Component, Target, and Pathway Analysis of ZZMRP
The herb, active components, targets, and associated pathways were imported into Cytoscape 3.8.0, and as 4 categories of nodes to build the herb, component, target and pathway network. Cytoscape is an open-source platform for visualizing complex networks and integrating these with any type of attributed data. 42
Molecular Docking
The binding activity between the active ingredient and core target was predicted using molecular docking. The structural formula of components was retrieved from PubChem (https://pubchem.ncbi.nlm.nih.gov/; accessed March 10, 2021) as an SDF file. The central target conformations were collected from the Protein Data Bank database (https://www.rcsb.org/; accessed March 10, 2021). The screening conditions were as follows: (1) protein structure determined using x-ray crystal diffraction; (2) protein crystal resolution >3; (3) preferential choice of protein structures with unique ligands and (4) Homo sapiens as the organism. The structures of the protein receptors and ligands were collected, and molecular docking was subsequently performed using AutoDockTools 1.5.6. 43 The docking results were visualized using PyMol 2.3.2 (https://pymol.org/2/).44,45
Results
Active Components and Targets of the Herbs in ZZMRP
The active components and targets of ZS, BZ, HMR, BS and ZW were accessed using the TCMSP database. Sixty-one active components were obtained using OB ≥30% and DL ≥0.18, once duplicate compounds were removed. The numbers of active compounds in ZS, BZ, HMR, BS and ZW were 22, 7, 6, 13 and 19, respectively (Table S1). After deleting the active compounds with no corresponding genes in the Uniprot database and targets, 44 active components were still retained with 19, 4, 6, 8, and 13 active compounds in ZS, BZ, HMR, BS and ZW, respectively (Table 1). Moreover, these 44 active components corresponded to 249 associated targets. Subsequently, these active components and associated putative targets were analyzed using Cytoscape. As demonstrated in Figure 2, the component-target network of ZZMRP comprised 298 nodes and 751 edges (Table S2). Each edge signified the reciprocities between herbs and compound molecules, or compound molecules and targets. Thus, the node degree corresponds to the number of edges associated with the node in the network, and the node size is positively associated with the degree value. Results of our network analysis demonstrated that the average degree value of the nodes was 5.0, each compound interacted with an average of 15.9 targets, and the average number of compounds per target was 2.8. Therefore, the results demonstrated that every compound may interact with multiple targets, and different compounds may function on the same targets in ZZMRP. This indicated the pharmacological mechanisms underlying coactions between multiple components and targets of ZZMRP. Moreover, 40.9% of the compounds interacted with ≥10 targets, and 12 compounds interacted with ≥20 targets. Notably, quercetin in ZW exhibited the highest number of associations with 139 target proteins, followed by kaempferol in BS and ZW, and luteolin in ZS, HMR and ZW with 58 and 55 target proteins, respectively. In addition, 7 compounds interacted with ≥30 target proteins; namely, β-sitosterol in BS and ZW, isorhamnetin in ZW, naringenin in ZS, nobiletin in ZS, arachidonic acid in HMR, tetramethoxyluteolin in ZS and stigmasterol in HMR with 37, 35, 35, 33, 33, 31, and 30 target proteins, respectively. Also, 3 target proteins interacted with ≥20 compounds. PTGS2 was the target protein that had the highest degree value with 29 compounds. It was followed by NCOA2 and PTGS1, with 22 and 21 compounds, respectively. Consistently, these data demonstrated the mechanisms of the multiple targets and components of ZZMRP.

Network of active components and targets. The regular hexagon indicates the active components of 5 herbs (BZ, ZS, HMR, BS, and ZW). The green rhombus represents the targets of active components. A1, B1, C1, D1 and D2 represent the mutual components of the herbs. The edges demonstrate the interaction between the components and targets. Abbreviations: BZ, Baizhu; ZS, Zhishi; HMR, Huomaren; BS, Baishao; ZW, Zi-wan.
Information on the 44 Active Compounds.
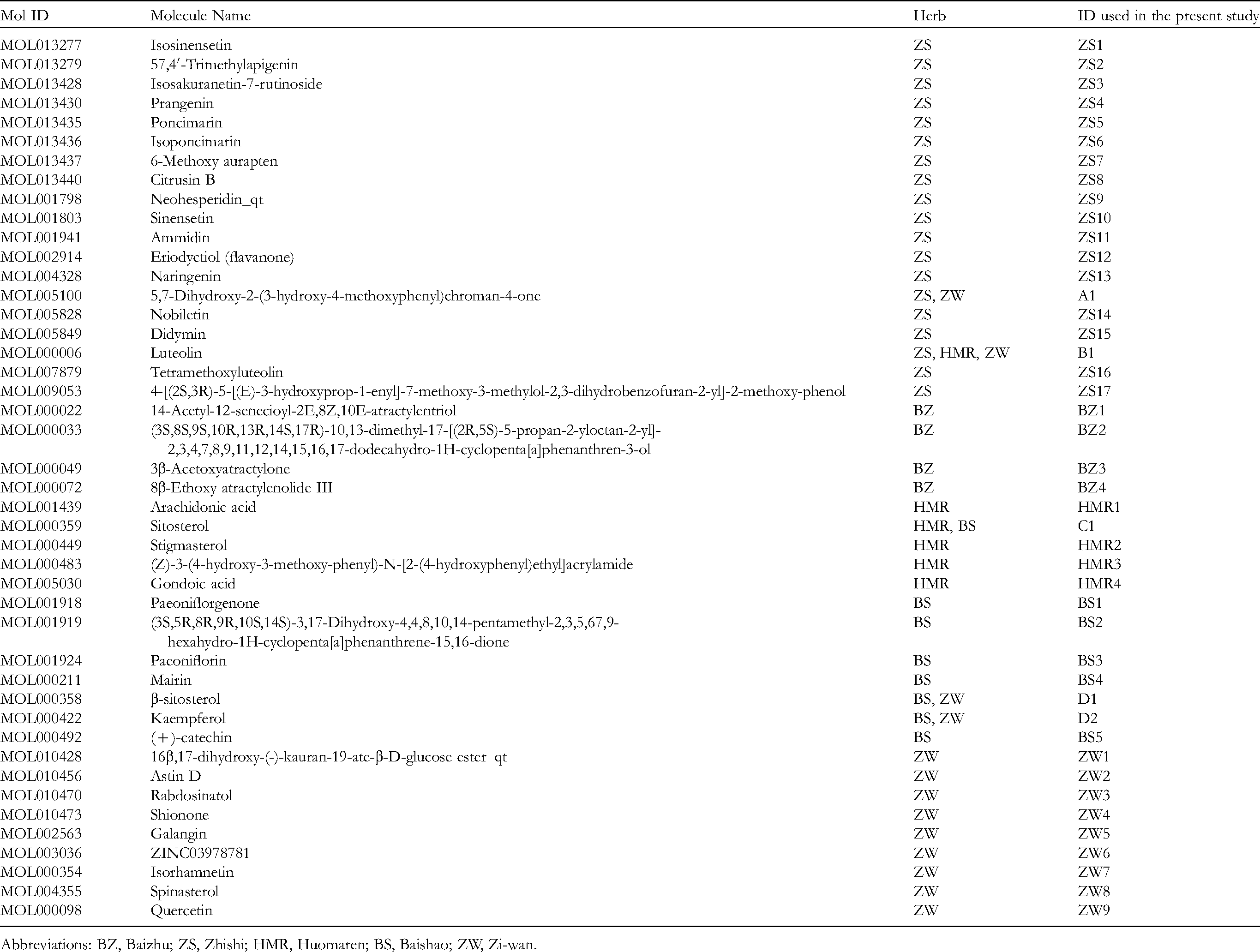
Abbreviations: BZ, Baizhu; ZS, Zhishi; HMR, Huomaren; BS, Baishao; ZW, Zi-wan.
Primary Targets of ZZMRP in Constipation Treatment
A total of 1501 associated targets of constipation were obtained from GeneCards, Disgenet, OMIM, DrugBANK and TTD databases, following the removal of duplicates. The top ten targets of constipation were tumor protein 53 (TP53), catenin β 1, epidermal growth factor receptor (EGFR), steroid receptor coactivator, RAC-α serine/threonine-protein kinase (AKT1), signal transducer and activator of transcription 3, mitogen-activated protein kinase 3 (MAPK3), E1A Binding Protein P300, β-actin and insulin, in sequence. Subsequently, 122 intersection targets of ZZMRP and constipation were acquired using the Venn diagram (Figure 3). Following construction using the STRING database, a PPI network was generated with 113 nodes, 807 edges and an average node degree of 14.3, using Cytoscape 3.8.0. The darker the color and the larger the node, the greater the degree. The top ten targets of ZZMRP in constipation treatment were TP53, AKT1, caspase-3 (CASP3), JUN, EGFR, tumor necrosis factor (TNF), MAPK3, heat shock protein 90 kDa α [cytosolic], class A member 1 (HSP90AA1), interleukin 6 (IL6) and MAPK1. The DC of each node was analyzed using the function “network analyzer.” A total of 18 main targets of ZZMRP in constipation treatment were screened using DC ≥24 (2 × 12; Figure 4). The detailed information of these 16 core targets is displayed in Table 2. These results suggested that these targets may play a significant role in constipation treatment.

Intersection targets of Zhi Zhu Ma Ren Pill (ZZMRP) and constipation. The blue circle represents targets for active components of ZZMRP, and the yellow circle represents targets for constipation.

Filter of the key targets. A total of 122 intersection targets of ZZMRP and constipation were input into the STRING database, and 113 targets were obtained with high confidence (confidence score, >0.7). Subsequently, 18 key targets were obtained using DC ≥24 from 113 targets. Abbreviations: ZZMRP, Zhi Zhu Ma Ren Pill; DC, degree centrality.
Detailed Information of the 18 Core Targets.
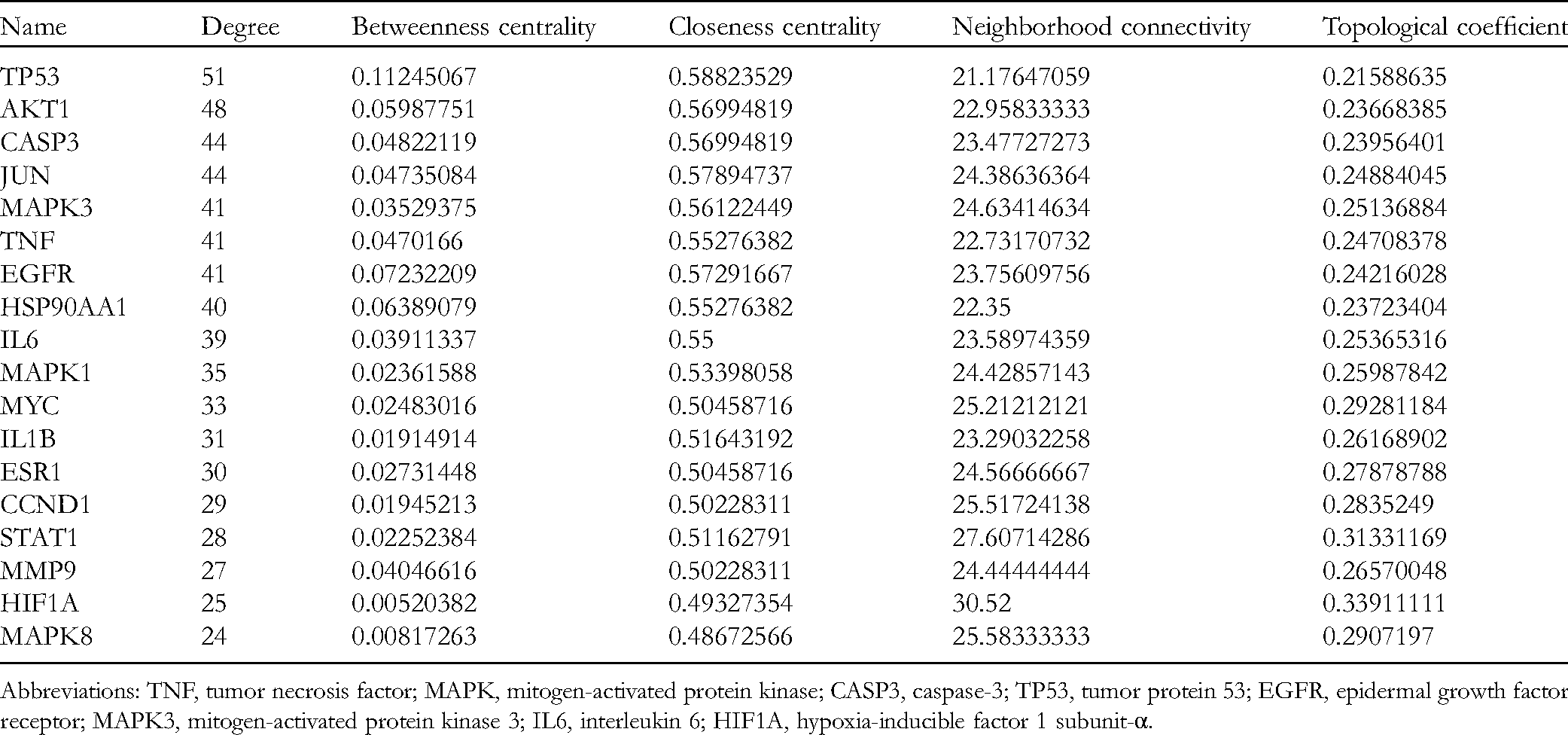
Abbreviations: TNF, tumor necrosis factor; MAPK, mitogen-activated protein kinase; CASP3, caspase-3; TP53, tumor protein 53; EGFR, epidermal growth factor receptor; MAPK3, mitogen-activated protein kinase 3; IL6, interleukin 6; HIF1A, hypoxia-inducible factor 1 subunit-α.
Constructing and Analyzing the PPI Network
The PPI network of these 18 key targets was obtained according to the STRING database (Figure 5a). Subsequently, the network was imported into Cytoscape to analyze the DC value that may denote the importance of a target in the PPI network. The results revealed that targets, including TP53, AKT1, JUN, CAPS3, MAPK3, TNF, EGFR, HSP90AA1, and IL6 demonstrated a high level of significance in the network (Figure 5b).

Analysis of the protein-protein interaction (PPI) network of the 18 key targets. (a) The PPI network of the 18 key targets obtained using the STRING database. Nodes represent core targets, and various edges represent different interactions. (b) The PPI network was constructed using Cytoscape 3.8.0.
GO and KEGG Enrichment Analysis
To explore the mechanisms underlying ZZMRP in the treatment of constipation, the 18 core targets were introduced into DAVID 6.8 for GO and KEGG enrichment analysis. The results revealed a total of 248 items, of which 196 were associated with biological processes, 16 with cell composition and 36 with molecular function. Subsequently, the top ten items based on the gene ratio were selected for imaging; these are displayed in Figure 6a. The biological processes mainly included positive regulation of transcription from RNA polymerase II promoter, positive regulation of transcription, DNA-templates, negative regulation of apoptotic process, transcription, signal transduction, response to drugs, positive regulation of smooth muscle cell proliferation, positive regulation of protein phosphorylation, positive regulation of nitric oxide biosynthetic processes and positive regulation of gene expression. The cell composition primarily contained the nucleus, nucleoplasm, cytosol, cytoplasm, protein complex, mitochondrion, extracellular space, extracellular region, nuclear chromatin and microtubule cytoskeleton. The molecular functions involved protein binding, identical protein binding, enzyme binding, transcription factor binding, adenosine triphosphate binding, transcription factor activity, DNA binding, sequence-specific DNA binding, nitric-oxide synthase regulator activity, and double-stranded DNA binding. In addition, 99 pathways were found during the KEGG enrichment pathways analysis, of which the top 20 items were selected for further analysis. These included the MAPK, TNF, Toll-like receptor, phosphoinositide 3-kinase (PI3K)-protein kinase B (Akt), estrogen and thyroid hormone signaling pathways, and osteoclast differentiation (Figure 6b).

Go and KEGG analysis of the 18 core targets. Abbreviations: GO, gene ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes.
Herb, Compound, Target and Pathway Network
To determine the associations between herbs, active compounds, targets and pathways, the herb, compound, target and pathway network was built using a total of 5 herbs, all 44 active compounds, 18 core targets of ZZMRP in constipation treatment, and 7 underlying pathways. The network included 75 nodes and 190 edges (Figure 7). The results demonstrated that the degree value of the MAPK signaling pathway was the highest, and hypoxia-inducible factor 1 subunit-α demonstrated the highest degree value among the targets. Table 3 displays the targets associated with the pathways.

The herb, compound, target and pathway network of ZZMRP in the treatment of constipation. The regular hexagon indicates the associated signaling pathway, the purple circle represents the 18 core targets, the green rhombus represents the active components, the orange square represents the 5 herbs, and the red triangle represents Zhi Zhu Ma Ren Pill (ZZMRP).
KEGG Pathway Enrichment Analysis of the key Pathways Involved in the Herb, Compound, Target and Pathway Network.
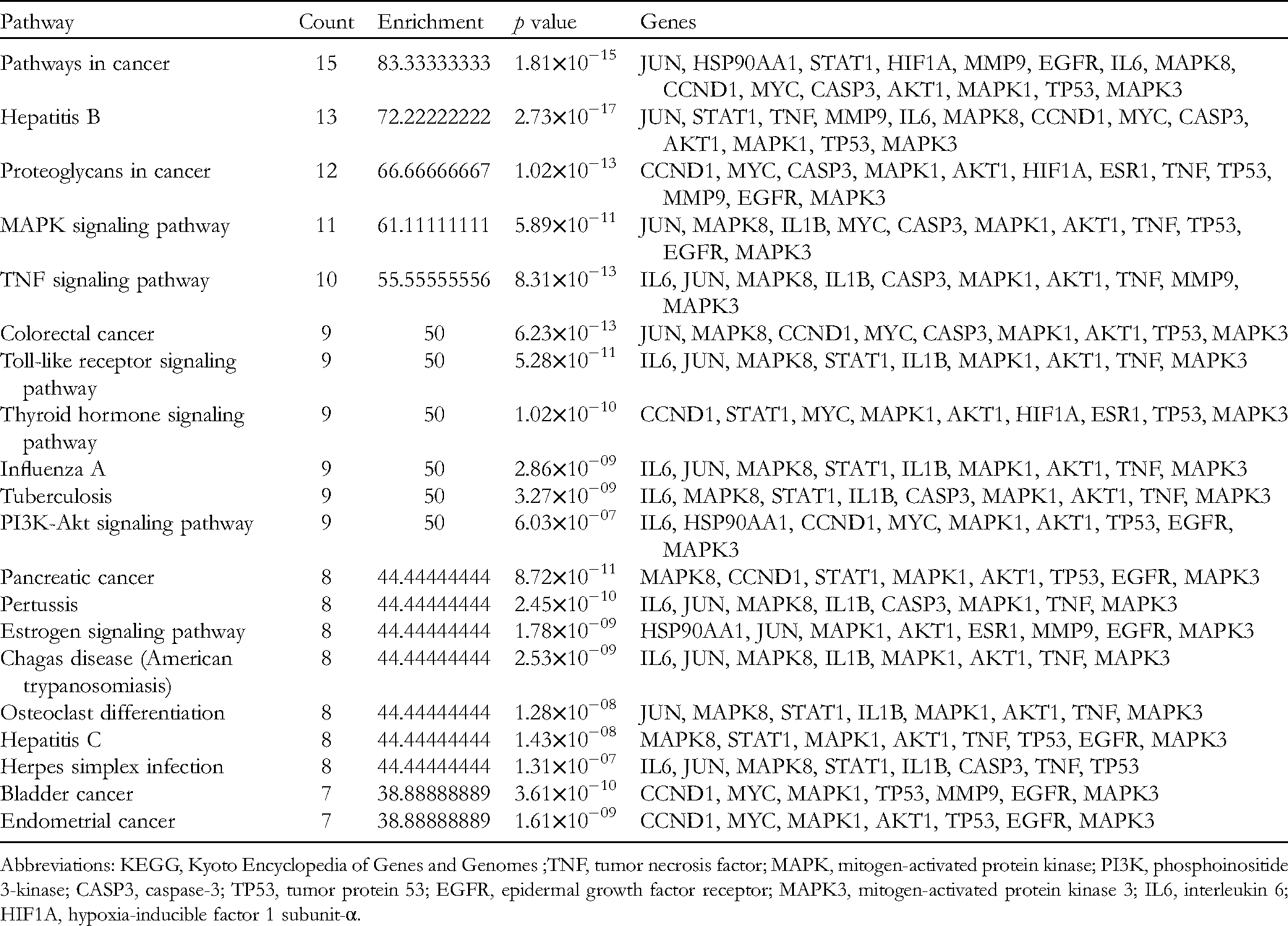
Abbreviations: KEGG, Kyoto Encyclopedia of Genes and Genomes ;TNF, tumor necrosis factor; MAPK, mitogen-activated protein kinase; PI3K, phosphoinositide 3-kinase; CASP3, caspase-3; TP53, tumor protein 53; EGFR, epidermal growth factor receptor; MAPK3, mitogen-activated protein kinase 3; IL6, interleukin 6; HIF1A, hypoxia-inducible factor 1 subunit-α.
Molecular Docking
To investigate the underlying drug-target interactions, molecular docking was performed to forecast the binding affinities between the active components and core targets in the herb, compound, target, and pathway network. The first 3 active components based on their degree values were quercetin, kaempferol and luteolin, which were selected as ligands. The 8 core targets of ZZMRP in constipation treatment with DC ≥40, which were TP53, AKT1, CASP3, JUN, MAPK3, TNF, EGFR and HSP90AA1, were selected as receptors. A lower affinity indicated improved binding activity between the compound and target. A docking score of ≤−5 kcal/mol among the 24 pairs of target-compound combinations revealed their precise binding activity (Table 4). Moreover, the corresponding images were visualized using Pymol software (Figure 8).

Results of molecular docking were visualized using Pymol software. Cayan indicate the ligands (quercetin, kaempferol and luteolin), and green represents the amino acids that interacted with the ligands.
Results of Molecular Docking.

Abbreviations: TNF, tumor necrosis factor; MAPK, mitogen-activated protein kinase; CASP3, caspase-3; TP53, tumor protein 53; EGFR, epidermal growth factor receptor; MAPK3, mitogen-activated protein kinase 3.
Discussion
Constipation is one of the most prevalent chronic gastrointestinal diseases, that greatly impacts the physical and mental health of patients. Previous studies have demonstrated that Chinese herbal compounds may improve constipation. For example, MaZiRenWan increased colonic motility, thus improving the symptoms of constipation by acting on multiple targets and pathways. 46 Yiqi Kaimi Recipe modulated the levels of substance P proteins and NOS1 in the enteric plexus, which enhanced the contraction amplitude and frequency of the smooth muscle. This facilitated colon motility and alleviated colonic slow transit constipation. 47 Moreover, Ji et al 48 recently reviewed the positive efficacy and safety of Chinese herbal compounds in the treatment of functional constipation. In the present study, a Chinese herbal compound, named ZZMRP, was used. This includes ZS, BZ, HMR, BS and ZW, based on previous studies that mitigate constipation. Treatments composed of BZ and ZS, which were first recorded in the Synopsis of the Golden Chamber 1700 years ago, have the ability to regulate gastrointestinal activity and promote gastrointestinal motility. 49 HMR and BS are common drugs used for treating constipation. 50 In addition, patients with constipation often exhibit mental and psychological symptoms. It is believed that BS functions by softening the liver and nourishing the liver yin, which help to smooth the liver qi. It is also believed that the lungs and the large intestine are on the outside, and the normal suppression of lung qi is essential for the normal functioning of the large intestine. Moreover, ZW is believed to nourish the lungs and lower qi. Notably, multiple previous studies have elucidated the positive effects of these herbal medicines on intestinal diseases, including constipation. For example, a network pharmacology analysis revealed the use of ZS volatile oil in the treatment of slow transit constipation. 51 In addition, a metabolic study demonstrated the therapeutic effects of ZS and BZ on slow transit constipation. 52 Gao et al 53 demonstrated that intestinal diseases with major symptoms of constipation, dyschesia, and abdominal fullness, may be treated with numerous herbal medicines, and this is believed to be through moistening the intestine and invigorating qi. Among them, BZ was believed to clear Fu-organs, and HMR was believed to moisten the intestine. In addition, ethanol extracts of HMR demonstrated laxative properties that may affect Cl− and Na+ movement in the intestinal epithelia cells in rats. 54 Notably, total glucosides of paeony isolated from BS ameliorated Sjögren's syndrome-elicited intestinal inflammation and constipation in mice. 55 Thus, the present study aimed to analyze the underlying targets and pharmacological mechanisms of ZZMRP on constipation, using network pharmacology and molecular docking.
Forty-four active components were obtained using TCMSP, with 249 associated targets. Among them, quercetin, kaempferol, luteolin, β-sitosterol, isorhamnetin, naringenin, nobiletin, arachidonic acid, tetramethoxyluteolin and stigmasterol were determined to be active components. Previous pharmacological studies revealed that quercetin modulates the mAChRs downstream signal to enhance mucin secretion and gastrointestinal motility, which ameliorates chronic constipation. 56 The results of a previous study also reported that quercetin protects rats from loperamide-induced constipation. 57 Ali et al 58 demonstrated that kaempferol, a flavonoid constituent of Capparis decidua, generates Ca2+ antagonist-like activity, therefore promoting the plant's utilization in the treatment of constipation. Naringenin exhibited anti-constipation roles through enhancing the expression of c-Kit, SCF, and AQP3. 59 In addition, naringenin promoted Cl− secretion in the colonic epithelium via the PKA and cAMP signaling pathways to improve constipation. 60 Moreover, Chen et al 20 Discovered that arachidonic acid metabolism was associated with constipation in rats. HPLC analysis demonstrated that stigmasterol, an active component of Carissa carandas, exerted notable effects in constipation. 61 Additionally, numerous proteins, such as PTGS2, NCOA2, PTGS1, HSP90AA1, and PRKACA are major putative targets for treating constipation. PTGS2 is a pivotal enzyme for the formation of PGE2, that plays a role in prostanoid biosynthesis, and is associated with mitogenesis and inflammation. It is also involved in the activation of the NF-κB signaling pathway. 62 Therefore, PTGS2 may act as a potential target for nonsteroidal anti-inflammatory drugs. A previous network pharmacology study used molecular docking analysis to demonstrate that PTGS2 was a target protein for the treatment of slow transit constipation. 51 Moreover, both the transcriptional and translational levels of PTGS2 were markedly upregulated, while those of PTGS1 were significantly downregulated in both patients with slow transit constipation and cellular models.63,64 A further network pharmacology analysis also demonstrated that HSP90AA1 was one of main targets for the treatment of constipation. 65 Collectively, these results indicated that ZZMRP may exert anti-constipation effects through multiple components and targets.
Moreover, 18 main targets of ZZMRP in the treatment of constipation were obtained in the present study, and these were primarily involved in cell proliferation, apoptosis and inflammation. Interstitial cells of Cajal (ICCs) may serve as pacemakers of gastrointestinal muscles, exerting crucial roles in modulating gut motility. 66 The healthy phenotypes and functions of ICCs rely on the activation of tyrosine kinase receptor c-Kit protein expressed on the cell surface. 67 Therefore, c-Kit acts as a marker for ICCs. 68 He et al 69 previously demonstrated that astragaloside IV improved slow transit constipation in mice, and this was involved in the proliferation of ICCs. Similarly, germinated barley exerted effects on colonic epithelial proliferation, to impact loperamide-induced constipation in rats. 70 The results of a further study demonstrated that microRNA-222 modulated the growth, apoptosis and autophagy of ICCs isolated from rats with slow transit constipation, by targeting c-kit. 71 The reduced apoptosis of colonic smooth muscle cells demonstrated the protective role of bacterial cellulose in rats with diphenoxylate-induced constipation. 72 Moreover, numerous previous studies demonstrated that the protective effects of various drugs or herbs on constipation were associated with the downregulation of inflammation.73–75 Thus, the aforementioned studies revealed that these may act as targets in the treatment of constipation.
The GO and KEGG enrichment analyses of the 18 main targets of ZZMRP in the treatment of constipation demonstrated that numerous signaling pathways, such as MAPK, TNF, and PI3K-Akt, played vital roles in treating constipation using ZZMRP. The MAPK signaling pathway is a central inflammation-related signaling pathway that is activated by TNF-α. 76 The expression of cytokines associated with the MAPK pathway was increased in rats with loperamide-induced constipation, and upregulation of the MAPK and PI3K/AKT signaling pathways markedly improved constipation in rats. 77 Choi et al 78 demonstrated that polystyrene microplastics reduced chronic constipation by regulation of the MAPK signaling pathway. TNF is also an important mediator of inflammation, and the results of a previous study demonstrated that this is markedly upregulated in constipation. 77 A similar analysis also indicated that the TNF signaling pathway played a critical role in the use of Shanzha in treating constipation. 66 Moreover, molecular docking indicated that quercetin, kaempferol, and luteolin most likely affected certain pathways by combining with TP53, AKT1, and CASP3, ultimately reducing the levels of constipation.
In conclusion, the results of the present study demonstrated that numerous active ingredients, such as quercetin, kaempferol, and luteolin, were significantly active components of ZZMRP in the treatment of constipation, with a series of relevant targets, including PTGS2, NCOA2, PTGS1, HSP90AA1, and PRKACA. Moreover, the MAPK, TNF, and PI3K-Akt signaling pathways played vital roles in the treatment of constipation using ZZMRP. The results of the present study also suggested that ZZMRP may treat constipation using multiple components, targets, and pathways. Although the results of the present study revealed the potential therapeutic effects of ZZMRP on constipation, further experiments are required for verification. Thus, the impact of dose and composition of ZZMRP must be investigated in subsequent in vivo assays. Collectively, the results of the present study provide a novel theoretical basis for the use of ZZMRP or associated herbs in the treatment of constipation.
Footnotes
Author Contributions
Yong Wen, Yu Zhan, and Xuegui Tang designed the experiments, and analyzed and interpreted the data. Shiyu Tang, Jian Kang, and Rong Wu collected the data. Yong Wen, Yu Zhan, and Xuegui Tang wrote and revised the manuscript. All authors have approved the final manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The present study was supported by the National Natural Science Foundation of China (grant nos. 82074429 and 82004173).
Data Availability Statement
The datasets used and/or analyzed during the present study are available from the corresponding author on reasonable request.
