Abstract
Three compounds were obtained from Sabia parviflora Wall., and their structures were identified through nuclear magnetic resonance (NMR) spectroscopy, particularly 2-dimensional (2D)-NMR. The molecular masses were determined using quadrupole-time-of-flight-mass spectrometry. Electronic circular dichroism spectra were used to determine the absolute configuration of compound 1. The 3 new compounds were identified as 2,2,7,7-tetramethyl-8S,10-dihydroxy-2,7,8,9-tetrahydro-2H-naphtha[2,3-b]pyran-6-one-10-O-β-D-glucopyranoside, (2Z)-4-(3-carboxy-4-hydroxyphenyl)-2-methylbut-2-enoic acid, and (2Z)-5-(3-carboxy-4-hydroxyphenyl)-5-oxo-2-methylpent-2-enoic acid. The liver protective activities of these compounds were tested by HepG2 and LO2 cell lines, which were induced using free fatty acids.
Introduction
Sabia parviflora Wall., family Sabiaceae, is a plant mainly distributed in southwest China. It is a traditional Chinese herbal medicine that is mainly used for the treatment of “damp heat jaundice, hepatitis and hemostasis.”1,2 Numerous chemical studies have shown that the plant contains triterpenoids,1,3 flavonoids,4,5 alkaloids,1,6 phenolic acids, phenylpropane, and polysaccharides. 7 Previous pharmacological studies have revealed that the plant's compounds possess antiviral, 8 antiinflammatory,5, 9 antiliver injury,5,9 and antioxidant 10 activities.
Based on the results of earlier experimental research, 11 our group has continued to explore the constituents of the plant and their biological activities. In this study, we examined the dried stems of the species and isolated 3 new compounds: 2,2,7,7-tetramethyl-8S,10-dihydroxy-2,7,8,9-tetrahydro-2H-naphtha[2,3-b]pyran-6-one-10-O-β-D-glucopyranoside (1), (2Z)-4-(3-carboxy-4-hydroxyphenyl)-2-methylbut-2-enoic acid (2), and (2Z)-5-(3-carboxy-4-hydroxyphenyl)-5-oxo-2-methylpent-2-enoic acid (3).The structures of the compounds are shown in Figure 1.

The 3 new compounds (1, 2, 3).
Finally, the biological activities of these 3 compounds were determined, and their hepatoprotective effects were investigated using HepG2 and LO2 cell lines.
Results and Discussion
Compound 1, a yellow powder, had a relative molecular mass of C23H30O9 based on its molecular ion peak at m/z 451.1943 [M + H]+(calcd. for C23H30O9, 450.1890) in its quadrupole time-of-flight tandem mass spectrum (Q-TOF-MS). The 1H nuclear magnetic resonance (NMR) spectrum of compound 1 exhibited signals for an aromatic proton of a penta-substituted benzene (δH 7.47 s, 1H), 2 olefinic protons of a 1,2-disubstituted alkene (δH 5.77, d, J = 10.0 Hz, 1H and δH 6.41, d, J = 10.0 H z, 1H), 1 methylene group (δH 3.21 m, 1H and 3.33 m, 1H), 1 methine proton (δH 3.91, dd, J = 3.5, 6.2 Hz, 1H), 4 methyl protons (δH 1.48, 1.49, 1.16, 1.17 s, each 3H), and 1 anomeric proton (δH 4.96, d, J = 7.7 H z, 1H). The 13C-NMR spectrum of compound 1 showed 23 signals: a penta-substituted benzene (δC 122.4, 122.7, 125.6, 137.9, 143.1, and 151.1), a carbonyl carbon (δC 203.6), 2 double-bond carbons (δC 132.4 and 122.6), 1 methylene carbon (δC 30.4), 1 methine carbon (δC 75.6), 2 quaternary carbons (δC 79.7 and 48.5), 4 methyl carbons (δC 28.8, 28.5, 24.1, and 23.1), and 6 glucosyl carbons (δC 105.3, 75.9, 78.2, 71.8, 78.4, and 62.7). Furthermore, C-10 was found to be connected to the oxygen atom, which was determined by the chemical shift of δC 143.1 (C-10). The 1H–13C-NMR spectra of compound 1 revealed structural similarity between compound 1 and sabianin C, 11 except for subtle differences in the 7 position of the methyl or hydroxymethyl substituents. The heteronuclear multiple bond correlation (HMBC) cross peak (δ 4.96/143.1) indicated that the glucose moiety was connected to C-10. Accordingly, acid hydrolysis of compound 1 gave D-glucose as the sugar residue, which was determined through gas chromatographic (GC) analysis of its corresponding trimethylsilylated L-cysteine adduct. The sugar in compound 1 was a β-glucose moiety according to the proton signal at δH 4.96 (d, J = 7.7 Hz). The nuclear overhauser effect spectroscopy spectrum showed 2 configurations for 1:2,2, 7,7-tetramethyl-8S,10-dihydroxy-2,7,8,9-tetrahydro-2H-naphtha[2,3-b] pyran-6-one-10-O-β-D-glucopyranoside and 2,2,7,7-tetramethyl-8R,10-dihydroxy-2,7,8,9-tetrahydro-2H-naphtha[2,3-b]pyran-6-one-10-O-β-D-glucopyranoside. The experimental circular dichroism (CD) spectrum and calculated CD(8S-C) exhibited the same trend at the 200-280 nm wavelength. The result established the absolute configuration as 2,2,7,7-tetramethyl-8S,10-dihydroxy-2,7,8,9-tetrahydro-2H-naphtha[2,3-b]pyran-6-one-10-O-β-D-glucopyranoside. The 1H and 13C NMR spectroscopic data of compound 1 are listed in Table 1, the HMBC spectrum in Figure 2, and the electronic circular dichroism (ECD) spectrum in Figure 3.

Important heteronuclear multiple bond correlation (HMBC) information (h→c) of the 3 new compounds 1, 2 and 3.
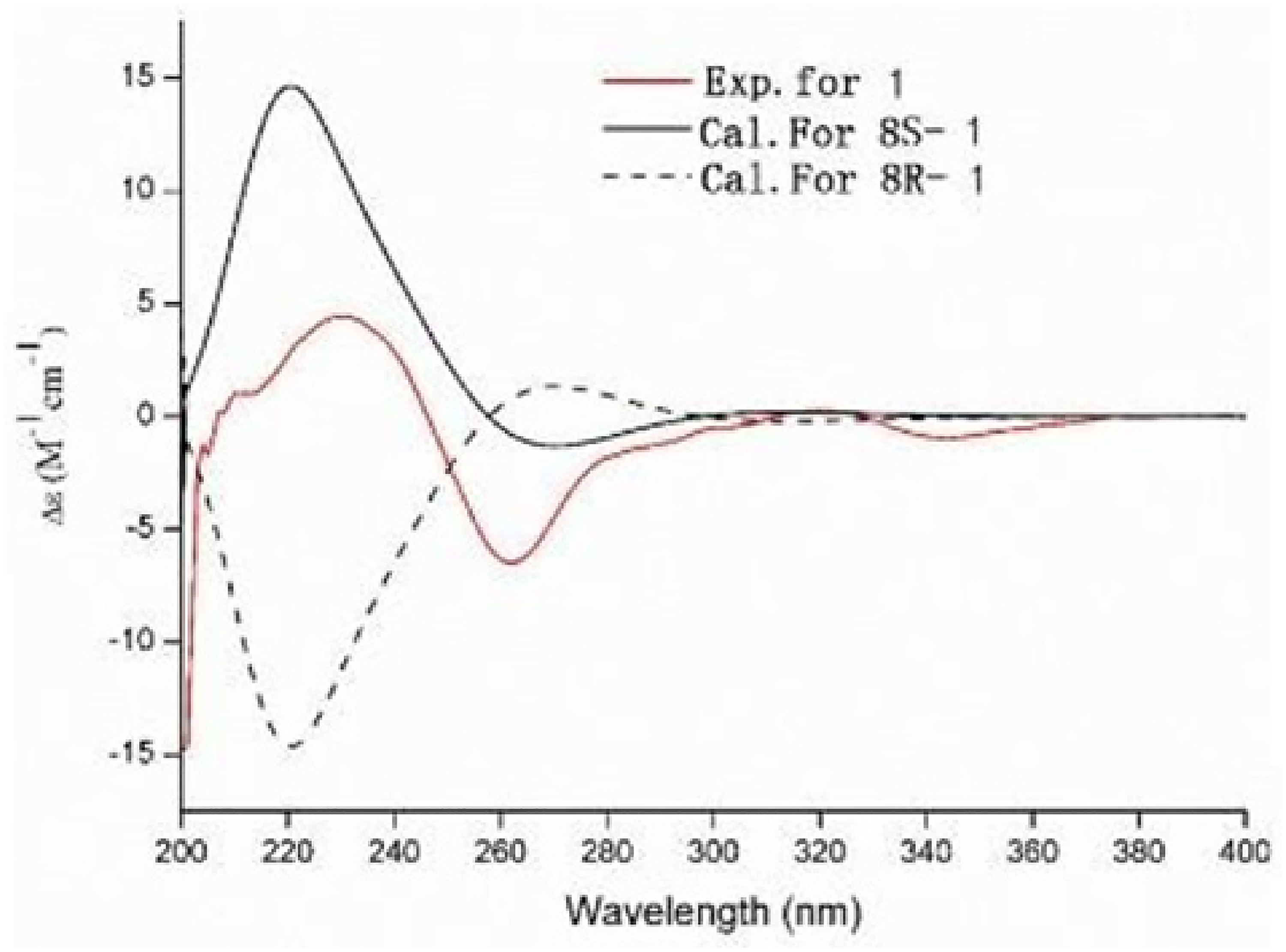
Electronic circular dichroism (ECD) spectra of compound 1.
Compound 2, isolated as a colorless oil, had a molecular formula of C12H12O5 based on its molecular ion peak at m/z 235.0608 [M-H]− (calcd. for C12H12O5, 236.0685) in its Q-TOF-MS. The 1H-NMR spectrum of compound 2 showed signals for 3 aromatic protons of a trisubstituted benzene (δH 7.70, dd, J = 8.2, 2.2 Hz, 7.71, d, J = 8.2 Hz, 6.90 d, J = 8.2 Hz, each 1H), 1 olefinic proton (δH 6.01, dd, J = 7.68 Hz, 1H), 1 methine (δH 3.70, d, J = 7.6 Hz, 2H), and 1 methyl (δH 1.85 s, 3H). The 13C-NMR spectrum of compound 2 showed 12 signals: 1 benzene ring (δC 131.7, 126.7, 129.8, 121.7, 160.0, and 115.2), 2 carboxyl carbons (δC 167.8 and 169.8), double-bond carbons (δC 138.3 and 129.0), 1 methine carbon (δC 30.0), and 1 methyl carbon (δC 21.1). The HMBC cross peak (δ 3.70/126.7) suggested that the methene was connected to C-1 (δH 6.01/δC 30.0), and the double bond to C-4′ (δH 6.01/δC 169.9 and δH 1.85/δC 129.0). Furthermore, the HMBC cross peak also indicated that the carboxyl carbon and methyl carbon were connected to C-2′. The HMBC cross peaks (δH 3.70/δC 131.8, 129.9) and (δH 7.70/δC 121.7), and the ABX coupling system suggested that carboxyl and hydroxyl carbons were connected to C-3 and C-4, respectively. In the ROESY experiment, the cross peak of δH 6.01, H-3′/δH1.85, and H-1′a supports the Z-configuration of C-2′-C-3′. Compound 2 was thus established as (2Z)-4-(3-carboxy-4-hydroxyphenyl)-2-methylbut-2-enoic acid.
Compound 3, isolated as a yellow oil, had a molecular formula of C13H12O6, based on the molecular ion peak at m/z 263.0570 [M-H]−(calcd. For C13H12O6, 264.0634) in its Q-TOF-MS. The 1H-NMR spectrum of compound 3 showed signals for 3 aromatic protons of a trisubstituted benzene (δH 7.69 m, 6.93, d, J = 8.2 Hz, 7.67 d, J = 8.2 Hz, each 1H), 1 olefinic proton (δH 6.00, t, J = 6.0 Hz, 1H), 1 methene (δH 3.7, d, J = 6.0 Hz, 2H), and 1 methyl (δH 1.88 s, 3H). The 13C-NMR spectrum of compound 3 showed 13 signals: 1 benzene ring (δC 130.7, 127.3, 138.5, 115.4, 161.3, and 125.1), 2 carboxyl carbons (δC 169.5 and 169.6), 2 double-bond carbons (δC 131.5 and 128.9), 1 methene carbon (δC 30.1), and 1 methyl carbon (δC 21.1). The 1H- and 13C-NMR spectra of compound 3 were similar to those of compound 2, except for subtle differences at C-5′. The signal at δC 192.1 was used to define the carbonyl group. The HMBC cross peak (δ 7.67/192.1) indicated that the carbonyl carbon was connected to C-1. Furthermore, in the ROESY experiment, the cross peak at δH 6.00, H-3′/δH 1.88, H-2′a supports the Z-configuration of the C-2′-C-3′ double bond. Therefore, compound 3 was established as (2Z)-5-(3-carboxy-4-hydroxyphenyl)-5-oxo-2-methylpent-2-enoic acid. The NMR data of compounds 2 and 3 are listed in Table 2, and the HMBC spectrum is shown in Figure 2.
Detailed 1H- and 13C-NMR Spectroscopic Data for Compound 1 and Sabianin C (δ in ppm, in CD3OD, 600 and 150 MHz).
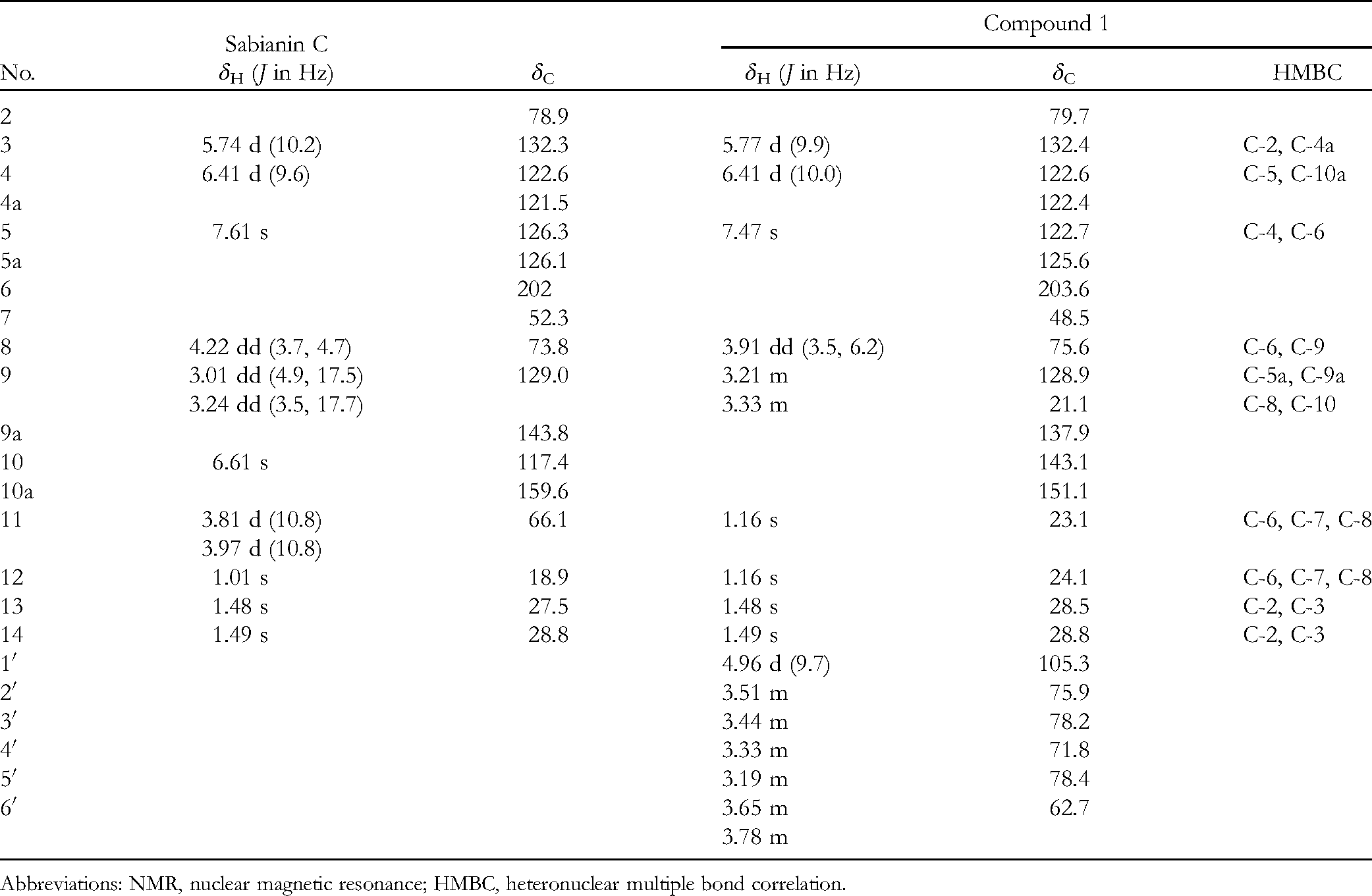
Abbreviations: NMR, nuclear magnetic resonance; HMBC, heteronuclear multiple bond correlation.
1H- and 13C-NMR Spectroscopic Data for Compounds 2 and 3 (δ in ppm, in DMSO-d6, 600 and 150 MHz).

NMR, nuclear magnetic resonance; DMSO-d6, dimethylsulifde-d6; HMBC, heteronuclear multiple bond correlation.
Compounds 1-3 were evaluated for their hepatoprotective activities. The results demonstrated that the 3 compounds had no hepatoprotective activity with no significant difference being observed between the cells in the presence or absence of these 3 compounds.
Experimental
Materials
Analytical methanol and glacial acetic acid were purchased from Sinopharm Chemical Reagent Co., Ltd. Dimethylsulfide-d6 (DMSO-d6) was used as a solvent to dissolve monomers. All chemicals used in this study were of analytical grade. Deionized ultrapure water was obtained using a MillQ Millipore Ultrapure Water Purification System (USA).
Methanol and acetonitrile for separation and purification were from Shanghai McLean Company, silica gel (300-400 mesh) for chromatographic separation from Qingdao Haiyang Chemical Group, and DIAION HP-20 macroporous resin from Mitsubishi. The medium-pressure liquid chromatography system was equipped with a Lisui EZ Purify III system, which was from Shanghai Lisui Chemical Engineering. The RP-C18 (25 µm) chromatographic column was purchased from Merck Company. The Ryong C18 column (10 µm, 30 × 250 mm) was equipped with a preparative high performance liquid chromatography (HPLC) system. We also used a Ryong C18 column (5 µm, 20 × 25 0 mm) for further purification. A Cosmosil C18 column (5 µm, 4.6 × 50 mm) was used in the component purity test. A Shimadzu LC-30A HPLC was connected to a TOF™5600 + mass spectrometer. The important NMR data were obtained using a Bruker Avance spectrometer. Palmitic acid (33.3%) and oleic acid (66.6%) were purchased from Sigma.
The 2 cell lines used in this study, HepG2 and LO2, were purchased from American Tissue Culture Collection.
Plant Material
The stems of S parviflora were collected in Ziyun City and identified by Professor Guo-Yue Zhong, Jiangxi from University of Traditional Chinese Medicine. A voucher specimen is deposited in the corresponding author's laboratory, the Center of National Medicine Resource, Jiangxi University of Traditional Chinese Medicine, which is located on Yangming Road, Donghu District, Nanchang City, Jiangxi Province, China
Extraction and Isolation
The dried stems of S parviflora (dry weight: 17 kg) were extracted with 70% ethanol (40 L × 3) through hot reflux. After removal of the solvent, the dry extract (380 g) was separated using macroporous resin HP20 (20 L) and eluted with an ethanol/water (30%→95%) gradient, thereby obtaining 4 fractions (frs 1-4). Fr 1 (253 g) was placed on a silica gel column and eluted with different proportions of CH2Cl2/CH3OH (20:1→1:1 v/v), and at the end of the process, 20 fractions were obtained (frs 1.1-1.20). Fr 1.1 was eluted with ethanol/water (10:90→0:100 v/v, 30 mL min−1), and 11 subfractions were obtained (Frs 1.1.1→1.1.11). Subfraction fr 1.1.3 (2.4 g) was separated using silica gel column chromatography where different proportions of CH2Cl2/CH3OH (20:1→1:1 v/v) were used. Finally, compound 2 (5.1 mg, TR 21.5 min) was eluted with 22% acetonitrile (5 mL min−1) through a reversed-phase HPLC-C18 chromatography column. Fr 2 (94 g) was chromatographed on a silica gel column with CH2Cl2/CH3OH (20:1→1:1 v/v), and 16 subcomponents (frs 2.1-2.16) were obtained. Using medium- and low-pressure chromatographic columns (10 cm × 60 cm), 10 subcomponents (frs 2.1.1-2.1.10) were obtained with ethanol/water elution (20:80→0:100 v/v, 30 mL min−1). Then, compounds 1 (2.0 mg, TR 42.5 min) and 3 (8.9 mg, TR 23.6 min) were purified from fr 2.1.8 (1.2 g) by semipreparative HPLC-C1 column chromatography (15% acetonitrile 5 mL min−1).
2,2,7,7-Tetramethyl-8S,10-dihydroxy-2,7,8,9-tetrahydro-2H-naphtho(2,3-b) pyran-6-one-10-O-β-D-glucopyranoside (yellow powder); Table 1 presents the 1H- and 13C-NMR data. Q-TOF-MS: m/z 451.1943 [M + H]+ (calcd. for C23H30O9, 450.1890). (2Z)-4-(3-carboxy-4-hydroxyphenyl)-2-methylbut-2-enoic acid (colorless oil); Table 2 presents the 1H- and 13C-NMR data. Q-TOF-MS: m/z 235.0608 [M−H]− (calcd. for C12H12O5, 236.0685). (2Z)-5-(3-carboxy-4-hydroxyphenyl)-5-oxo-2-methylpent-2-enoic acid (yellow oil); Table 2 presents the 1H- and 13C-NMR data. Q-TOF-MS: m/z 263.0570 [M − H]− (calcd. for C13H12O6, 264.0634).
Cell Viability Assay
HepG2 and LO2 cells were preserved in Roswell Park Memorial Institute 1640 (RPMI 1640) medium under a humidified atmosphere of 5% CO2 at 37°C. The medium contained 5% fetal bovine serum, penicillin (100 μm L−1), and streptomycin sulfate (100 μm L−1). HepG2 and LO2 cells were seeded and inoculated on 96-well plates at a density of 1 × 104 cells/well for 24 h. Then, 100 μL of medium containing different concentrations of the 3 compounds was added to each well, except for the blank and model wells. After 4 h, free fatty acids (1 mmol L−1) were added to all groups except the blank. After 24 h, the supernatant was discarded and a 10% CCK8 reagent (100 μL/well) was added and incubated for 2 h. In our experiment, a microplate reader was used to measure the absorbance of the sample at 450 nm. Cell viability is presented as a relative percentage of the untreated control.12,13
Hydrolysis of Compound 1 and GC Analysis
As reported in the previous literature, 14 compound 1(3 mg) was hydrolyzed with 2 mol L−1 HCl–dioxane (1:1, v/v, 5 mL) at 85°C for 8 h. The reaction mixture was extracted with CHCl3 (2 mL × 4). The aqueous layer was neutralized with 2 mol L−1 NaOH and dried to obtain the monosaccharides. Then, in the presence of pyridine (2 mL), the dry powder was added to L-cysteine methyl ester hydrochloride (1.5 mg) and kept at 60°C for 1 h. Next, trimethylsilylimidazole (1.5 mL) was added to the reaction mixture and kept at 60°C for 30 min. The supernatant (4 μL) was directly subjected to GC and compared with a glucose standard. GC was performed using the following experimental conditions: Agilent-7890b ECD detector, Chromatographic column: HP-5 (30 m × 0.25 nm, 0.25 μm), flow: 2 mL min−1; split ratio 30:1. temperature: 250°C.
Conclusion
Three new compounds were isolated from S parviflora. These 3 compounds showed no significant activity regarding liver protection. Further research is needed in order to understand other possible biological activities of these compounds.
Footnotes
Acknowledgements
Author Contributions
Wrote or contributed to revising the manuscript: Huan He, WeiDong Du. Conceived and designed the study: Huan He, WeiDong Du, ZhiFeng Li, Qiang Zhou. Do experiment: Huan He, WeiDong Du, Qiang Zhou.Experimental technical support: Yuanying Fang, Qi Wang, YuLin Feng.Data processing and analysis: Huan He, WeiDong Du, ZhiFeng Li.All authors have read and approved this version of the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (No. 81960701), the funding programs for academic and technical leaders of major disciplines in Jiangxi Province (20194BCJ22020), Jiangxi Province 5511 R&D projects (No. 20165BCB19009), Jiangxi Province talent Project (No. [2016]332), and Nanchang innovative talent team (No. [2018]274), and Grant for Nanchang innovative talent training project (Grant No. [2020]137). This work was also supported by the General project of Chongqing Natural Science Foundation (cstc2021jxyj-msxmx0665) titled “Studies on the active components and anti-hepatitis B virus activity of Sabia parviflora from Chongqing.”
Ethical Approval
Not applicable, because this article does not contain any studies with human or animal subjects.
Statement of Human and Animal Rights
Not applicable, because this article does not contain any studies with human or animal subjects.
Informed Consent
Not applicable, because this article does not contain any studies with human or animal subjects.
