Abstract
Introduction
Parkinson’s disease (PD) is a neurodegenerative disorder that poses a serious threat to physical and mental health of middle-aged and elderly people.1,2 The disease gradually attacks patient's motor cortex and leads to long-lasting impairment in motor functions, such as movement and coordination. 3 The main treatment for PD is symptomatic. Chinese herbal medicine was reported effective in alleviating clinical symptoms, such as tremor and paralysis, thereby improving the quality of life of PD patients.4,5 Ginseng is one of the most widely used traditional Chinese medicines (TCMs) for treating PD.6,8 Although its efficacy is well documented, the specific mechanism underlying its action in PD treatment is still unclear. Rapid advancement in computer science, molecular biology and bioinformatics, as well as network pharmacology has led to the development of novel types of models for disease-drug prediction. In recent years, tremendous progress has been made to elucidate the mechanisms of TCM using network pharmacology,9,10 making it a powerful tool for exploration of the specific targets and pathways for the treatment of diseases. 11 The purpose of this study was to evaluate the potential effective components and underlying mechanism of ginseng in the treatment of PD using network pharmacology and molecular docking.
Materials and Methods
Acquisition and Formulation of Ginseng
The composition of ginseng was screened from the traditional Chinese medicine systems pharmacology database and analysis platform (TCMSP). The criteria for effective compound screening were oral bioavailability (OB) ≥30% and drug-like properties (DL) ≥0.18, which allowed identification of all the effective ingredients of ginseng. Potential drug targets of the effective ingredients were collected from TCMSP, while the genes matching these targets were identified from the UniProt database (https://www.uniprot.org/uniprot/).
Screening for PD-Related Genes
PD-related gene targets were searched using “Parkinson's disease” as the keyword in the GeneCards (https://www.genecards.org/), OMIM (https://www.omim.org/), PharmGkb (https://www.pharmgkb.org/), Therapeutic Targets (http://bidd.nus.edu.sg/BIDD-) Databases/TTD/TTD.asp), and DrugBank (https://www.drugbank.ca) databases. The final targets were filtered by combining all the results from each database.
PD Targets Prediction and PPI Network Construction
VENNY software (http://bioinfogp.cnb.csic.es/tools/venny/index.html) was employed to map the identified effective ginseng ingredients obtained in section 1.1 and PD-related targets are identified in section 1.2. Then a Venn diagram was drawn to assess the interaction of the ingredients and targets. Nodes in the network represent the genes, proteins or molecules that constitute either the active ingredients or PD-related targets, whereas connections between nodes denote interactions between these ingredients.
The STRING database (https://string-db.org) contains a large No. of protein–protein interaction (PPI) relationships. These interactions are classified according to the parameters into low, medium, and high confidence levels, with confidence levels of < 0.4, 0.4–0.7, and >0.7, respectively. Potential targets of PD were uploaded to the STRING database and then a PPI analysis was performed to generate a network. The network's minimum linkage score between targets was set to 0.9, while isolated nodes in the network were hidden. Topological analysis was performed to calculate the interacting parameters between target proteins.
GO and KEGG Gene Pathway Enrichment Analysis
Gene Ontology (GO) function and Kyoto Gene and Gene Encyclopedia (KEGG) enrichment analysis of the potential targets was performed using R software. Three indicators of GO function were calculated: Biological process (BP), cellular component (CC), and molecular function (MF). Twenty pathways (
Molecular Docking
Molecular structures for the effective ingredients and protein crystal structures of PD-related targets in ginseng were downloaded from Chemical Book (https://www.chemicalbook.com) and RCSBPDB (https://www.rcsb.org) databases. Then, we employed Discovery Studio 2018 for molecular docking of the core targets with the ginseng effective ingredients. Finally, we calculated LibDock scores to quantify the binding activity between ginseng effective ingredients and core targets.
Results
Identification of Effective ginseng Ingredients and Target Genes
After screening the TCMSP database, 97 and 2025 potentially effective ginseng ingredients and targets were obtained (Supplementary Table 1). Moreover, 212 target genes that corresponded with the effective ingredient targets were finally obtained from the Uniprot database after manual deletion of the invalid and duplicated targets.
Identification of PD-Related Genes
After screening the GeneCards, OMIM, PharmGkb, therapeutic targets, as well as the DrugBank databases, 7722 PD-associated disease targets were identified after duplicated merging (Supplementary Table 2).
Prediction of Effective ginseng Targets in PD
VENNY2.1 software was employed to map 212 effective drug targets to the 7722 PD-related target genes and finally 168 intersection genes were obtained (Figure 1, Supplementary Table 3). In addition, a compound-target action network with 261 nodes and 1251 edges was obtained (Figure 2). It was found that one active component of the drug could correspond to one or more targets of disease, whereas one target of PD could also correspond to multiple effective drug ingredients. Therefore, this suggested that ginseng has a cross action in the treatment of PD.

Venn diagram of Parkinson’s disease (PD) target and ginseng therapeutic target.

The correlation network between effective ingredient and potential target.
Profiles of the PPI Network and Core Targets
Potential treatment targets were uploaded to STRING database and a PPI network was generated. It showed that the No. of nodes and edges were 144 and 579, respectively. Notably, 14 core targets and 61 edges were finally screened out in the network after target filtering (Figure 3). An edge represents the association between the targets, whereas the nodes represent the targets. The final core targets screened included STAT3, JUN, AKT1, MAPK1, MAPK3, RELA, MAPK14, IL6, EGFR, FOS, MAPK8, CXCL8, IL4, and ESR1.

Process of topological screening for the protein–protein interaction (PPI) net.
GO and KEGG Pathway Enrichment
GO enrichment analysis revealed that the effect of ginseng for treatment of PD was mainly enriched in 165 GO terms. A bubble diagram was plotted using the top 20 of these (Figure 4). Results from the KEGG pathway enrichment analysis revealed that the treatment was enriched in 181 signaling pathways. A summary of the top 20 KEGG enrichment pathways is shown in Figure 5.
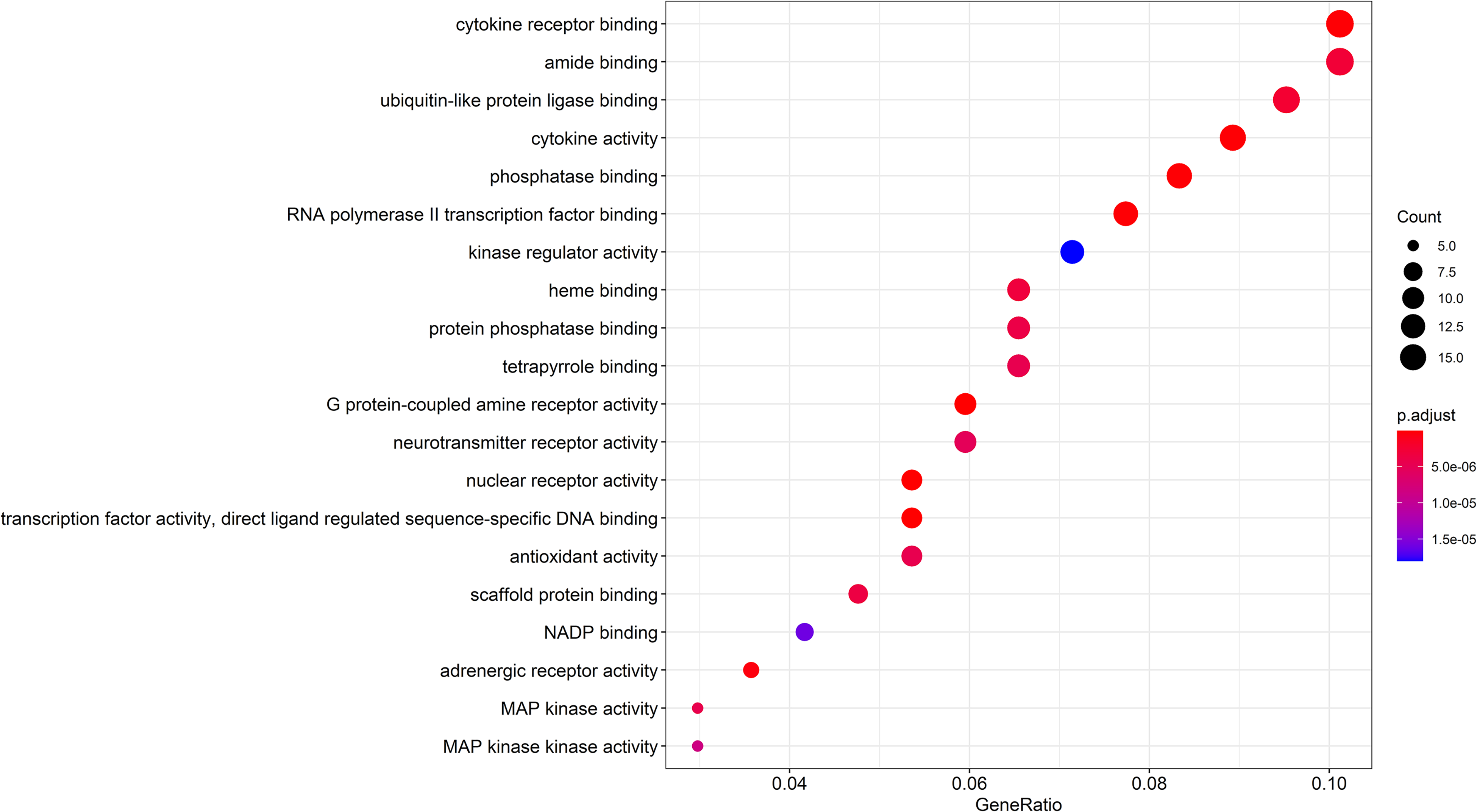
Gene Ontology (GO) enrichment of active ingredients of ginseng in the treatment of common targets of Parkinson’s disease (PD).

Kyoto Gene and Gene Encyclopedia (KEGG) enrichment of active ingredients of ginseng in the treatment of targets of Parkinson’s disease (PD).
Furthermore, the KEGG Mapper tool was used to generate a pathway map of ginseng for the treatment of PD. Notably, the mitogen-activated protein kinase (MAPK) pathway was identified (Figure 6), comprised of the p53 and Wnt signaling pathways.

Mitogen-activated protein kinase (MAPK) pathway of ginseng in the treatment of Parkinson’s disease (PD).
Molecular Docking Analysis
The docking results were analyzed as a screening criterion in LibDock score. The heatmap showed that these components bind well with the core genes MAPK3, MAPK14, and EGFR (Figures 7 and 8). The obtained results for molecular docking in the present study were in agreement with those obtained in network pharmacological analysis.

Molecular docking process of active ingredients of ginseng and Parkinson’s disease (PD)-related targets.

Heatmap of the molecular docking results.
Discussion
In the present study, we used network pharmacology to identify 97 active ginseng compounds and predicted 168 possible targets in PD, including dopamine (DA) receptors, DA transporters, 5-hydroxytryptamine receptors, cholinesterase, and hypersensitive response proteins, among others. We found a strong interaction between these targets, mainly involving MAPK3, MAPK14, EGFR, and IL17. Our results revealed that ginseng contains a large No. of active ingredients, including ginsenosides, ginseng polypeptides, ginseng polysaccharides, and vitamins, among others. Specifically, ginsenosides are considered the main active ingredients of this drug, with recent studies demonstrating that these compounds exert biological activity, regulating immunity, and having anti-cancer and antioxidant effects, not only in the nervous system, but also on the cardiovascular and immune systems.8,12 Based on our findings, saponins can inhibit cell death by regulating many signal pathways in vivo, such as inhibiting the aggregation and activation of glial cells, thus enhancing the viability of nerve cells. This may be the main therapeutic mechanisms of ginseng in the treatment of PD.
Through analysis of the topological network and PPI, we found that ginseng saponins, polypeptides, and polysaccharides occupy the core position in the network diagram. The potential targets of EGFR, IL-17, MAPK 1, and MAPK 14 have high degree scores. The active components may exert neuroprotective effects by attenuating the microglia-mediated neuroinflammatory response through these potential targets. The main pathological change in PD is degeneration and death of DA neurons in the substantia nigra of the midbrain, which results in a significant decrease in striatal DA content and causes PD.12,13 Activation of MAPK regulates the axonal structure and maintains normal brain function. 12 EGFR has been shown to play a key role in PD progression. 13 Therefore, both EGFR and MAPK are the main targets for the treatment of PD. In addition, EGFR mainly upregulates eNOS in endothelial cells through the advanced glycation end products (AGE)-receptors of advanced glycation end products (RAGE) signaling pathway, thereby causing vascular endothelial dysfunction, and vasodilatation imbalance, as well as vascular remodeling and proliferation.14,16 Consequently, this phenomenon causes an interaction with metabolic function and further damages vessels, leading to ischemic and hypoxic changes in intracranial neurons. Interleukin-17 (IL-17) is an important pro-inflammatory cytokine that is secreted by helper T 17 cells (Th17) and innate immune cells.17,18 It plays a key role in a variety of inflammatory responses in autoimmune diseases. 19 Functionally, IL-17 receptor (IL-17R) activates downstream NF-kB, JNK, and other signaling pathways through IL-17R-Act1-TRAF6.20,21 Previous studies have shown that TNF and IL-17 inhibitors can significantly elevate striatal DA levels and reduce neuronal necrosis. They generate a protective effect in PD models and are considered to be potential therapeutic targets for PD.14,22,24 All the aforementioned targets are mainly related to neuronal function, vascular proliferation, and the inflammation process, indicating that these active ingredients are core components in treating PD by modulating EGFR, IL-17, MAPK 1, MAPK 14, and other potential targets.
The results from GO enrichment analysis revealed that most of the targets are involved in the regulation of DA secretion, the related metabolic processes, chemical synaptic transmission, negative regulation of synaptic transmission, and other biological processes. Moreover, regulation of DA secretion and the related metabolic processes are closely associated with mechanisms of PD. 12 Additionally, the results from KEGG analysis revealed that the toll-like receptor (TLR) was the most prominent biological function during treatment of PD, since it promotes the synthesis and release of cytokines and triggers inflammatory responses. 25 Previous studies have shown that TLRs were expressed on the membrane surface of monocytes and dendritic cells to initiate the synthesis of intranuclear-related genes. 26 Consequently, these molecular factors synthesize cytokines such as IL1, IL 6, IL8, IL12, and TNF-α, leading to increased capillary permeability, and lymphocyte infiltration.27,33 These findings suggest that TLRs are involved in the therapeutic process of PD through anti-inflammatory and neuroprotective effects. In the present study, the enrichment analysis results revealed that the drug may treat PD by targeting the MAPK, IL-17, and AGE-RAGE signaling pathways.
This study has a few limitations. First, we only employed network pharmacology for mining of information from the relevant databases, and hence our results may be limited by the differences across databases and mining algorithms. Second, we did not validate the underlying mechanisms of these targets using animal experiments. Therefore, future explorations are expected to perform
Conclusion
In summary, we employed network pharmacology and a molecular docking approach to identify the main compounds, targets, and signaling pathways of ginseng for the treatment of PD. Based on the results, we speculated that ginsenosides, polypeptides, and polysaccharides are the active components in ginseng. They may regulate MAPK3 EGFR, IL4, and other targets through the MAPK, IL-17, and AGE-RAGE signaling pathways. Ginseng plays an important role as an anti-inflammatory, with anti-immune and anti-vascular atrophy activity. Therefore, this study provides a theoretical basis for further clinical trials in the treatment of PD with ginseng.
Supplemental Material
sj-xlsx-1-npx-10.1177_1934578X221102029 - Supplemental material for Network Pharmacology and Molecular Docking-Based Prediction of the Molecular Targets and Signaling Pathways of Ginseng in the Treatment of Parkinson's Disease
Supplemental material, sj-xlsx-1-npx-10.1177_1934578X221102029 for Network Pharmacology and Molecular Docking-Based Prediction of the Molecular Targets and Signaling Pathways of Ginseng in the Treatment of Parkinson's Disease by Wei Zhang, Jingya Chen and Hongquan Liu in Natural Product Communications
Supplemental Material
sj-xlsx-2-npx-10.1177_1934578X221102029 - Supplemental material for Network Pharmacology and Molecular Docking-Based Prediction of the Molecular Targets and Signaling Pathways of Ginseng in the Treatment of Parkinson's Disease
Supplemental material, sj-xlsx-2-npx-10.1177_1934578X221102029 for Network Pharmacology and Molecular Docking-Based Prediction of the Molecular Targets and Signaling Pathways of Ginseng in the Treatment of Parkinson's Disease by Wei Zhang, Jingya Chen and Hongquan Liu in Natural Product Communications
Supplemental Material
sj-xlsx-3-npx-10.1177_1934578X221102029 - Supplemental material for Network Pharmacology and Molecular Docking-Based Prediction of the Molecular Targets and Signaling Pathways of Ginseng in the Treatment of Parkinson's Disease
Supplemental material, sj-xlsx-3-npx-10.1177_1934578X221102029 for Network Pharmacology and Molecular Docking-Based Prediction of the Molecular Targets and Signaling Pathways of Ginseng in the Treatment of Parkinson's Disease by Wei Zhang, Jingya Chen and Hongquan Liu in Natural Product Communications
Supplemental Material
sj-xlsx-4-npx-10.1177_1934578X221102029 - Supplemental material for Network Pharmacology and Molecular Docking-Based Prediction of the Molecular Targets and Signaling Pathways of Ginseng in the Treatment of Parkinson's Disease
Supplemental material, sj-xlsx-4-npx-10.1177_1934578X221102029 for Network Pharmacology and Molecular Docking-Based Prediction of the Molecular Targets and Signaling Pathways of Ginseng in the Treatment of Parkinson's Disease by Wei Zhang, Jingya Chen and Hongquan Liu in Natural Product Communications
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical Approval
Not applicable, because this article does not contain any studies with human or animal subjects.
Informed Consent
Not applicable, because this article does not contain any studies with human or animal subjects.
Trial Registration
Not applicable, because this article does not contain any clinical trials.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
