Abstract
Objective
The molecular mechanism of Salviae Miltiorrhizae Radix et Rhizoma (SMRR) in the treatment of knee osteoarthritis (KOA) was analyzed based on network pharmacology.
Methods
Active components and potential targets of SMRR were obtained from the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform. KOA targets were obtained from the OMIM, DisGeNET, DrugBank, PharmGKB, and GeneCards Databases. The potential targets of SMRR in the treatment of KOA were identified by the Venn diagram. A protein-protein interaction network was generated with the STRING database. Visualization of the interactions in a potential pharmacodynamic component-target network was accomplished with Cytoscape software. The Database for Annotation, Visualization, and Integrated Discovery database and R software were used for Gene Ontology enrichment and Kyoto Encyclopedia of Genes and Genomes pathway annotation analyses of common targets. Molecular docking of the potential leading components, as determined by efficacy with the core target molecules, was performed with Discovery Studio.
Results
Fifty-seven potential pharmacodynamic components and 58 potential targets of SMRR in the treatment of KOA were found. Bioinformatics analyses showed that the interleukin (IL)-17, hypoxia-inducible factor-1 (HIF-1), and tumor necrosis factor (TNF) signaling pathways, as well as the advanced glycation end product-receptor for advanced glycation end product signaling pathway in cases of diabetic complications, are related to the molecular mechanism of SMRR in the treatment of KOA. Molecular docking results showed that luteolin, tanshinone IIA, cryptotanshinone, and other components of SMRR had a strong affinity for MYC, signal transducer and activator of transcription 3, caspase-3 (CASP3), JUN, cyclin D1, prostaglandin endoperoxide synthase 2 (PTGS2), epidermal growth factor receptor (EGFR), mitogen-activated protein kinase 1 (MAPK1), protein kinase B, vascular endothelial growth factor A, and other targets.
Conclusion
SMRR indirectly regulates IL-17, HIF-1, TNF, and other signal transduction pathways by regulating the expression of proteins, including PTGS2, MAPK1, EGFR, and CASP3, thus playing a role in promoting chondrocyte proliferation, improving microcirculation, eliminating free radicals, and inhibiting inflammatory factors.
Keywords
Knee osteoarthritis (KOA) is a common disease in middle-aged and elderly people. The main clinical manifestations are knee joint pain, stiffness, and limited joint mobility. In severe cases, it may be disabling. 1 As the rate of China’s aging population accelerates, KOA has gradually become a major health issue affecting people’s physical and mental health. 2 More than one-half of old people aged 60‐75 years have undergone imaging to verify KOA, and furthermore, more than 4/5 of those older than 75 years have been diagnosed with KOA. 1,3
Salviae Miltiorrhizae Radix et Rhizoma (SMRR) is a Chinese herbal medicine (CHM) that has been recorded in the Compendium of Materia Medica. Medicinal salvia is derived from SMRR consisting of the dry roots and rhizomes of
The main components of SMRR are water-soluble phenolic acids and fat-soluble tanshinone compounds. In 1934, Japanese scholars extracted tanshinone I, tanshinone II, and tanshinone III from SMRR. 6 For the next 30 years, the fat-soluble components of SMRR became a hot research topic as a drug. It was not until 1979 that Junyan Yao obtained the water-soluble components in his research on SMRR. 7
Tanshinone I is an active component of SMRR that reduces cartilage degradation and synovitis inflammation, which may be associated with the inhibition of inflammatory cytokine expression, reduced levels of nuclear factor kappa-light-chain-enhancer of activated B cells (NF-κB), and attenuated degradation of the extracellular matrix (ECM). 8 Cryptotanshinone can prevent interleukin (IL)-1-induced inflammation and improve osteoarthritis (OA) progression. 9 It has been reported that cryptotanshinone is a cyclooxygenase-2 (COX-2) inhibitor that can reduce the synthesis of prostaglandin E2 (PGE2) and the generation of reactive oxygen species (ROS) catalyzed by COX-2. 10 Salvianolate has an anti-inflammatory effect, which may be related to a decrease in inflammatory cell infiltration and the content of IL-6 and tumor necrosis factor (TNF)-α. Salvianolic acid B is a hydrophilic SMRR component that has been reported to have a regulatory effect on osteoblastic activity. 11,12
Oxidative stress (OS) can activate a variety of intracellular signaling pathways, leading to apoptosis or cell overgrowth and thus plays important roles in a variety of physiological and pathological processes. 13 ROS has 2 effects on OS. At low endogenous and dose levels, ROS are active components of cell signaling pathways, but at high doses, they are cytotoxic agents. 14 SMRR can inhibit the secretion of lipopolysaccharide (LPS)-induced TNF-α expression in peritoneal mouse macrophages, a response that may be related to the inhibition of ROS production and the activation of the NF-κB pathway. 15,16 Studies have shown that, in abnormal circumstances, SMRR can interfere with OS. 17,18 It has been claimed that SMRR can prevent articular cartilage degeneration in OA rabbits by inhibiting OS. 19 In addition, SMRR can alleviate OA by inhibiting NF-κB signal transduction and regulating the Janus kinase (JAK2)/signal transducer and activator of transcription 3 (STAT-3), and protein kinase B (AKT) signal transduction pathways. 20 However, information on how the active components of SMRR participate in the regulation of KOA is still lacking. The therapeutic effect of SMRR on KOA and its underlying mechanism need to be further clarified.
It remains difficult to study CHM systematically. Currently, most studies on traditional Chinese medicine focus on single components, single targets, and single diseases.
Moreover, CHM lacks a reasonable, scientific and efficient drug efficacy, and safety evaluation system, which makes it difficult to conduct comprehensive and systematic research. With the development of genomics, proteomics, and metabolomics, the advancements to systems biology and bioinformatics have accelerated, with network pharmacology based on systems biology and computer technology quickly following. The concept of network pharmacology was proposed by Hopkins, a British pharmacologist, 21 and network pharmacology is currently an emerging discipline that is based on the theory of systems biology and network analyses of biological systems to explain the effects of drugs and their mechanisms. Network pharmacology emphasizes the multicomponent, multitarget, and multipath synergistic relationships between drugs, targets, and diseases. These analyses’ characteristics align with the systemic and holistic treatment concepts of traditional Chinese medicine.
To reveal further the mechanism of action of SMRR in the treatment of KOA, the intervention effect of the effective components of SMRR on KOA is discerned. This research uses network pharmacology to study the targets and related pathways of SMRR in the treatment of KOA, providing new ideas for the development of drugs for treating KOA. The flow chart of the research is shown in Figure 1.

Whole framework based on network pharmacology. GO, gene ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; KOA, knee osteoarthritis; PPI, protein-protein interaction; SMRR, Salviae Miltiorrhizae Radix et Rhizoma.
Materials and Methods
Screening the Active Components and Targets of SMRR
The active components of SMRR were searched through Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP; https://tcmspw.com/tcmsp.php, Version 2.3).
22
According to the pharmacokinetic parameters (absorption, distribution, metabolism, and excretion) in the TCMSP database, the active components of
Screening Targets Common to KOA and SMRR
Through the OMIM database 24 (https://www.omim.org/), DisGeNET database 25 (https://www.disgenet.org), GeneCards database 26 (https://www.genecards.org/), DrugBank database 27 (https://www.drugbank.ca/), and PharmGKB database 28 (https://www.pharmgkb.org/), targets related to KOA were obtained.
The targets were uploaded from these 5 databases to the UniProt database (http://www.UniProt.org/), and the search was limited to “Human” species to obtain the corresponding gene information and verify the accuracy of the targets. Then, the targets corresponding to active components and the KOA targets were mapped, the intersection targets were screened, and the common targets of SMRR and KOA were obtained. These common targets may be a collection of potential targets for SMRR to treat KOA. A Venn diagram was generated online through the Venny 2.1.0 tool (https://bioinfogp.cnb.csic.es/tools/venny/index.html).
Constructing a Network of Active SMRR Components and KOA Targets
Cytoscape is an open-source software platform used to visualize complex networks and integrate them with various types of attribute data. 29 The screening results of the active component targets and the KOA targets of SMRR were imported into Cytoscape software to construct a drug-disease target network. A node in the network represents a target or active component, and an edge represents the interactions between nodes. The core architecture of the Cytoscape software is a network in which each node is a gene, protein, or molecule, and the connection between nodes represents the interactions between these biological molecules. The degree value of a node represents the number of nodes connected to that node in the network. The greater the degree value, the more likely the node represents a key target of the compound.
Construction of the PIN and Screening of Core Genes
Protein-protein interaction (PPI) refers to the correlation between protein molecules. For life-sustaining functions, this correlation represents the basis of cell metabolism. A protein interaction network (PIN) is formed by the interaction of each protein with the other proteins in the network. The systematic analysis of the interactions of a large number of proteins in biological systems is of great significance for understanding the reaction mechanisms of biological signals and energy metabolism under special physiological conditions, such as diseases, and for understanding the functional connections between proteins. The STRING database (https://string-db.org/) is a compendium of a large number of protein interaction relationships and thus can be used to predict protein interaction relationships. On this basis, we imported common genes into the STRING database for analysis, limited the species search to
GO Function Enrichment and KEGG Pathway Enrichment Analyses
The Database for Annotation, Visualization, and Integrated Discovery (DAVID, http://david.nifcrf.gov/) is a large-scale gene and protein tool that can be used to find the most significantly enriched biological annotations.
31
All common targets were imported into the DAVID database, and then, gene ontology (GO) function enrichment and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were performed. The background database and gene list were defined by the “
Molecular Docking
Discovery Studio is a new-generation tool for molecular modeling and simulation environments for life science research and is based on the Windows/Linux system and available for use on personal computers. 32 It serves experimental biologists, pharmaceutical chemists, structural biologists, computational biologists, and computational chemists working in life science fields and is applied to protein structure and function research and drug discovery.
We used this software to perform molecular docking. The RCSB Protein Data Bank Database (https://www.rcsb.org/) was used to obtain the core target structures. With the Prepare Protein tool, the parameters set to the default values, and pretreatments such as dehydration, hydrogenation, and ring building included, we obtained the pretreated protein structures. The Define and Edit Binding Site tool was used to define the active center of the target molecule. The location and size of the active center were determined based on the positive ligand position for the protein as obtained from the PDB and related crystal information in the literature.
In the Chemical Book database (https://www.chemicalbook.com/), the active compound structures were accessed in mol2 format. Using the Prepare Ligands tool, different conformations of the active compounds were generated to compile a small-molecule ligand library. The LibDock method in the Dock Ligands module was used for molecular docking. The preconditioned target receptor was docked to each small-molecular ligand in the library. Then, the output LibDock scores were obtained. The higher the score, the stronger the binding affinity of the small molecule ligand for the protein target. This score was used to evaluate the binding avidity of the active components of SMRR and the target proteins. After obtaining all the LibDock scores, a heat map was generated. After obtaining all the LibDock scores, all of them were normalized. This process was realized by TBtools 33 v1.046 software. The first step was to open the TBtools software and select HeatMap, and then import the obtained LibDock scores. Next, the parameters were set, including Row Scale, Cluster Rows, and Cluster Cols, and the default values were taken for other parameters.
Results
Screening of the Active SMRR Components
The TCMSP database was used for screening OB ≥30% and DL ≥0.18. A total of 65 active SMRR components were found. Then, the relevant targets were retrieved manually, no targets corresponding with KOA were deleted, and 57 active components were ultimately included. See Supplemental Table 1 for details.
Identifying Effective Active Components and the Construction of the KOA Target Library of SMRR Components
A total of 932 component targets were retrieved from the TCMSP database. The UniProt database was used to obtain the gene names of the targets, and invalid and repeated targets were deleted. A final 108 targets of the active components were obtained. Through OMIM, DisGeNET, GeneCards, DrugBank, and PharmGKB database searches and results integration, 1952 KOA gene targets were obtained, as shown in Supplemental Table 2. One hundred and eight active drug targets were mapped with 1952 KOA target genes to obtain 58 common targets. The Venn diagram is shown in Figure 2, and the common target information is shown in Supplemental Table 3.

Intersection of action targets of SMRR and KOA targets. KOA, knee osteoarthritis; SMRR, Salviae Miltiorrhizae Radix et Rhizoma.
A Network of “Drug Components and Disease Targets” was Generated
Fifty-seven active drug components were matched with 58 disease targets, and the active components without corresponding targets and duplicate targets were deleted. The data were imported into Cytoscape software to construct a drug target-disease intersection network. The detailed information is shown in Supplemental Table 4. This network includes 115 nodes and 284 edges, as shown in Figure 3.

SMRR treatment of KOA active ingredient—common target network. The blue quadrilateral represents the common target and the green quadrilateral the active component of SMRR. KOA, knee osteoarthritis; SMRR, Salviae Miltiorrhizae Radix et Rhizoma.
Target PIN Analysis and Core Target Screening
The common genes were imported into the STRING database to determine their interaction relationships, and the data were saved in the TSV file format. The files were imported into Cytoscape software for visualization. Then, CytoNCA was used to analyze the PPIs and obtain the DC, BC, and CC values. The topological feature analysis of the aforementioned PPI afterward was based on the 3 major parameters, DC, BC, and CC, with selected targets above the median values as key targets. The screening process was carried out twice. The following criteria were used for the first screening: DC ≥ 11, BC ≥ 15.461, CC ≥0.514, and a total of 55 nodes and 367 edges. The following criteria were used for further screening of 22 targets: DC ≥ 22, BC ≥ 76.787, CC ≥0.600, and a total of 22 nodes and 164 edges. The second screening resulted in 10 nodes and 40 edges. The following core targets were obtained: MYC, STAT3, caspase-3 (CASP3), JUN, cyclin D1 (CCND1), prostaglandin endoperoxide synthase 2 (PTGS2), epidermal growth factor receptor (EGFR), vascular endothelial growth factor A (VEGFA), mitogen-activated protein kinase (MAPK1), and AKT1. The core target screening process is shown in Figure 4, and the topology parameters of the PPI network are shown in Supplemental Table 5.

Process of topological screening for PPI network. BC, betweenness centrality; CC, closeness centrality; DC, degree centrality; PPI, protein-protein interaction.
GO and KEGG Pathway Enrichment Analyses
The common targets enriched in MF, BP, and CC GO categories were analyzed, and the results showed that the target of SMRR for KOA was enriched in 85 GO categories. The targets were enriched in the following biological functions and processes: cytokine receptor binding, phosphatase binding, ubiquitin-like protein ligase binding, peptide binding, nuclear receptor activity, growth factor receptor binding, RNA polymerase II transcription factor binding, cytokine activity, steroid hormone receptor, activated transcription factor binding, steroid binding, NF-κB binding, cyclin-dependent protein serine/threonine kinase regulation activity, and cysteine-type endopeptidase activity, which is involved in the process of apoptosis. With

Gene ontology biofunctional enrichment analysis of component-disease targets. The larger the node and the smaller the

KEGG enrichment analysis performed on the common targets. The smaller the p.adjust, the greater the correlation. HIF-1, hypoxia-inducible factor-1; IL, interleukin; KEGG, Kyoto Encyclopedia of Genes and Genomes; TNF, tumor necrosis factor.
Molecular Docking
The LibDock tool was used to first dock the processed active components with the core targets and then calculate the LibDock score for each docking. The docking results revealed the 2-dimensional (2D) and 3D docking poses of luteolin with each core target and presented the LibDock scores in a heat map. Successful docking indicates that the active component may act on the relevant target and is a potential effective component with certain pharmacodynamics. However, the specific pharmacodynamic effects need to be verified by further in vivo and in vitro experiments. Despite some limitations, this molecular docking strategy revealed that the effective active components of SMRR were good matches with the core target. The docking process is shown in Figure 7, the heat map in Figure 8, and the results of the ligand-receptor protein molecular docking in Supplemental Table 6.

Molecular docking of compounds with core targets. The left side is the 3-dimensional docking diagram, and the right side is the 2D docking diagram. Docking process of luteolin with (A) AKT1; (B) CASP3; (C) CCND1; (D) EGFR; (E) JUN; (F) MAPK1; (G) MYC; (H) PTGS2; (I) STAT3; and (J) VEGFA.
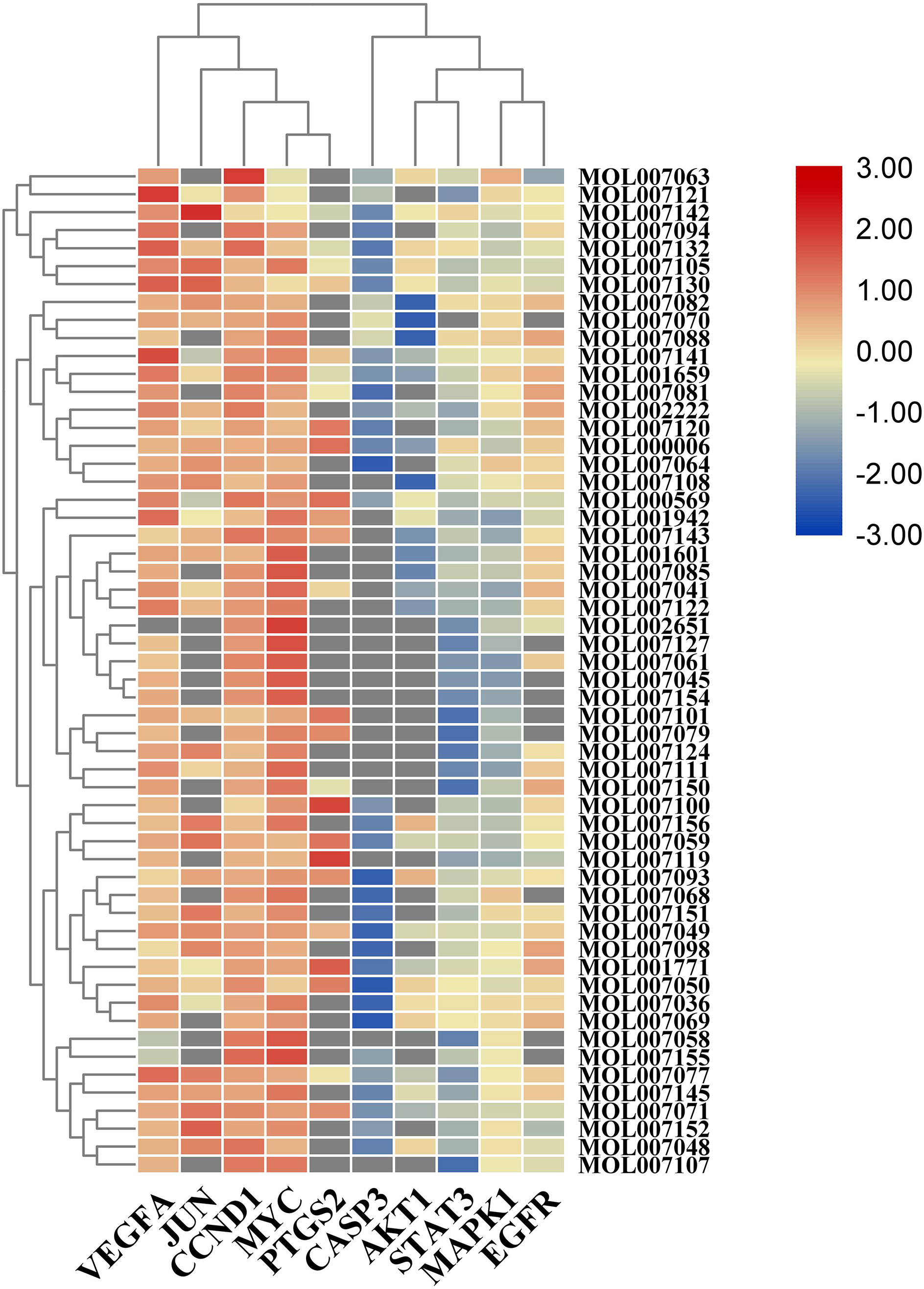
Heat map of the Libdock Score for molecular docking. The horizontal axis represents the core target and the vertical axis represents the active components of SMRR, Salviae Miltiorrhizae Radix et Rhizoma.
Discussion
Existing studies have shown that SMRR not only can prevent OA cartilage degeneration through antioxidative stress but can also reduce OA cartilage damage and thus be used to treat OA. 19,34 The effective active SMRR component-target gene relationship network showed that treatment of OA by SMRR is achieved through multiple components, multiple pathways, and multiple targets. Furthermore, the results showing that the target genes involved with OA indicated that there are multiple targets for OA treatment. Therefore, it is particularly important to clarify the target genes and pathways of the active components of SMRR in OA through network pharmacology. Although some studies on the mechanism of action of SMRR in the treatment of KOA have been carried out, most studies are limited to a focus on the pharmacological effects of drug monomers, and there is still much remaining for exploration in this field. In this study, the network pharmacology strategy using various related databases was adopted to screen 57 active components of SMRR and 58 potential targets for preventing and treating KOA. The chemical components of SMRR were divided into fat-soluble diterpene quinones and water-soluble phenolic acids. These compounds include flavonoids, phenolic acids, diterpenoids, sugars, proteins, alkaloids, and volatile oils. Among these compounds, luteolin, tanshinone IIA cryptotanshinone, dihydrotanshinlactone, isotanshinone II, and dan-shexinkum d play important roles in the prevention and treatment of KOA.
Luteolin is a natural flavonoid that exists in a variety of plants and has various pharmacological activities, such as anti-inflammatory, antiallergic, and uric acid activity. Studies have shown that luteolin can reduce inflammation, protect chondrocytes, and delay cartilage degradation, which may be achieved by inhibiting the proliferation of OA chondrocytes and downregulating the expression of JNK and p38 MAPK in OA chondrocytes. 35 Similarly, luteolin can reduce IL-1β-induced inflammatory factor production and inhibit IL-1β-induced NF-κB phosphorylation, as shown by reducing cartilage destruction in OA rats. 36 In addition, experimental studies in rats showed that luteolin could improve the effect of superoxide dismutase and catalase induced by selenite, enhance the activity of glutathione peroxidase and glutathione reductase, and enhance the activity of the antioxidant enzyme system in organisms. 37 It has also been reported that luteolin can regulate the expression, secretion, and activity of matrix metalloproteinase (MMP)-3 by directly acting on articular chondrocytes. 38
Two fat-soluble terpenoids from SMRR were identified. Tanshinone IIA protects ATDC5 cells from LPS-induced inflammatory injury by downregulating microRNA-203a and inhibiting the JAK/STAT and JNK pathways. 38 Tanshinone IIA can also inhibit the proliferation of chondrocytes, and the degree of inhibition is positively correlated with the concentration of tanshinone IIA. As the concentration of tanshinone IIA increases, the ratio of chondrocytes in G0/G1 phases increases, while the ratio in the S phase decreases. 39 Cryptotanshinone is another important component of SMRR. It can inhibit the activation of the NF-κB and MAPK signaling pathways induced by LPS in macrophages, prevent IL-1β-induced human osteoarthritis chondrocyte inflammation, and attenuate the progression of mouse osteoarthritis. 9,40
According to the topological analysis of the PPI network, there are 10 core targets: MYC, STAT3, CASP3, JUN, CCND1, PTGS2, EGFR, MAPK1, AKT1, and VEGFA. These may be potential targets of SMRR for the treatment of OA.
Cytokines, including proinflammatory factors, chemokines, and other signal molecules, can activate signal transduction and regulate various biological functions by binding with cell surface receptors. 41 Proinflammatory factors are widely present in joints. For example, IL-1β, IL-6, TNF-α, and transforming growth factor-β can promote the development of inflammation, indirectly or directly affecting sensory neurons, and sensitizing nociceptors and pain. 42,43 Chemokines and their receptors are closely related to peripheral hyperalgesia. In particular, peripheral sensory neurons can be activated directly by chemokines. 44
There is a consensus suggesting that the characteristics of KOA, the degeneration of articular cartilage, and the changes in synovium and subchondral bone lead to joint pain and joint dysfunction. Aging, metabolic dysfunction, and mechanical injury can lead to further degeneration of articular cartilage. Interestingly, it seems that the difference in age and OA stage drives EGFR to play a dual role in articular cartilage. 45 On the one hand, EGFR stimulates the proliferation of chondrocytes through anabolism and then plays a role in maintaining cartilage. In the early stages of OA, EGFR promotes the production of joint lubricating molecules. On the other hand, EGFR can inhibit the expression of Sox9 and reduce the synthesis of cartilage matrix protein. 46
MAPK1 is a serine/threonine protein kinase that is mainly involved in the regulation of cell proliferation, differentiation, growth, and apoptosis. The activation of the MAPK signaling pathway can affect the activity of a variety of transcription factors, thereby regulating the expression of cytokine genes such as TNF, IL-1, and IL-6. 47 The KEGG enrichment analysis showed that MAPK1 directly or indirectly participates in the TNF, MAPK, phosphoinositide-3-kinase (PI3K)-Akt, and many other signaling pathways, forming complex interaction relationships within the “SMRR active component-signaling pathway-KOA” network.
With respect to the pathological progression of OA, VEGF is increased in articular cartilage, synovial membranes, and subchondral bones. Moreover, VEGF in articular fluid is related to the severity and pain degree of OA in KOA patients; therefore, VEGF may be an OA biomarker. 48,49 A genome-wide association study and meta-analysis also showed that VEGF is significantly related to OA. 50
STAT3 is a DNA-binding molecule that can regulate the levels of many cytokines. A report showed that the activation of STAT3 in chondrocytes of patients with OA was significantly higher than that in the normal controls. 51 Studies have shown that injection of STAT3 inhibitors can reduce the expression of inflammatory mediators and chemokines in cartilage areas. Moreover, in MIa-induced OA rats, pain level and TRPV1 production in the dorsal root ganglion were decreased with the injection of a STAT3 inhibitor. 52 An analysis of 16 OA dysfunction modules and the aggregation of 3239 module genes showed that STAT3 may be the core transcription factor of OA and may promote the development of OA through NF-κB signaling. 53 These reports suggest that STAT3 inhibitors may have therapeutic potential for inflammatory diseases.
PTGS, also known as COX, is a key enzyme in prostaglandin biosynthesis. PTGS has 2 isoenzymes, PTGS1 (COX-1) and PTGS2 (COX-2). PTGS2 expression is induced by cytokines and growth factors and upregulated during inflammation. 54 PTGS2 is activated to produce PGE2, which acts on a variety of cell signaling pathways, such as cell proliferation, apoptosis, and immunosuppression. 55 PTGS2 plays an important role in the destruction of bone tissue, and COX-2-dependent PGE2 synthesis is considered to be an important mediator of tissue destruction in inflammatory bone diseases. 56,57
The PI3K/Akt signaling pathway can maintain cell survival in a variety of ways. Akt not only can inhibit the activation of proapoptotic molecules but can also act as the substrate of CASP3. 58 CASP3 is an important link in the process of apoptosis. Studies have shown that the pathological progression of OA is related to apoptosis and the endoplasmic reticulum stress response, and CASP3 activation is related to cartilage degeneration and chondrocyte apoptosis. 59 Akt can activate cell survival signals and maintain cell survival. 60,61 The increase in PI3K activity can also upregulate the activity of the Akt signaling pathway and promote the activity of Akt. 62
Aging is one of the risk factors in the pathological progression of OA. AGEs are a series of highly active end products formed in the context of nonenzymatic glycosylation reactions between the amino groups of proteins, fatty acids and nucleic acids, and the aldehyde group of reducing sugars. In the aging process, especially in the elderly, AGEs accumulate in the articular cartilage. 63,64 AGE may also increase the levels of PGE2 and nitric oxide through induction of the MAPK pathway to enhance the inflammatory response of OA chondrocytes. 65,66 Other studies have shown that the activation of RAGE stimulates inflammation-related signaling pathways. 67 In OA patients, RAGE activation can stimulate chondrocytes and synovial cells, leading to an increase in MMP-1. 68
IL-17 is a proinflammatory cytokine, particularly IL-17A. IL-17A can act through NF-κB, MAPK, and other signaling pathways. 69 In the pathogenesis of KOA, IL-17 may be involved with a variety of cytokines and pathways, such as IL-1, IL-8, IL-6, and macrophage inflammatory proteins. 70 Studies have found that IL-17 can activate NF-κB and promote cell apoptosis through this pathway. 71,72 IL-17 can also promote bone resorption and collagen degradation; enhance the expression of chondrocytes MMP-9, MMP-13, and other metal matrix proteases; strengthen cartilage plate destruction and cartilage tissue decomposition; and induce joint destruction. 73,74 Studies have shown that IL-17 can enhance the formation of blood vessels in arthritic cartilage while activating and increasing vascular endothelial growth factor, aggravating inflammation, and causing bone fibrosis and hyperplasia. 75 Moreover, clinical studies have shown that the expression level of IL-17 in OA synovium and synovial fluid is increased, and the increase in IL-17 level in OA synovial fluid is related to the severity of OA. 76 -78
As an important regulator of hypoxia, HIF-1α is expressed in articular chondrocytes and plays an important role in regulating the metabolism of articular chondrocytes, maintaining their survival, and expressing specific phenotypes. 79 VEGF is an important target gene of HIF-1α. In the process of cartilage development, VEGF can promote the formation of new blood vessels, maintain the survival of chondrocytes, and accelerate bone turnover. 80,81 Studies have shown that the expression of HIF-1α increases with the degeneration of articular cartilage. 82 In addition, the activated HIF-1 pathway may also interfere with the combination of osteoblasts and osteoclasts, thereby inhibiting the production of osteoclasts. 83
The relationship between TNF-α and inflammation has been established. TNF-α is a multifunctional inflammatory cytokine that can stimulate the production of PGE2 in synovial cells and can stimulate human chondrocytes to secrete plasmin, thereby aggravating arthritis damage and leading to the destruction of bone and cartilage. 84 In addition, TNF-α is an important mediator of cartilage matrix degradation and can play an important role in the process of synovitis and cartilage tissue degradation. TNF-α participates in inflammation by binding to the receptors TNFR1 and TNFR2. 85 It combines with TNFRI to induce COX-II and stimulate synovial cells to produce PGE and chondrocytes to produce peroxidation, leading to the destruction of bone and cartilage. 86 Furthermore, TNF-α activates the NF-κB signaling pathway by binding to receptors TNFR1 and TNFR2, leading to IκBα phosphorylation. The subsequent nuclear translocation of P65 leads to the activation of the NF-κB signaling pathway, leading to further enhanced proinflammatory function. 87
In conclusion, this study used network pharmacology to predict preliminarily that SMRR may regulate MYC, STAT3, CASP3, JUN, CCND1, PTGS2, EGFR, MAPK1, AKT1, VEGFA, and other related targets through active components such as luteolin, tanshinone IIA, and cryptotanshinone. Through the IL-17, HIF-1, and TNF signaling pathways, as well as the AGE-RAGE signaling pathway in the context of diabetic complications signaling pathway, the proliferation of chondrocytes, improved microcirculation, elimination of free radicals, and inhibition of inflammatory factors are promoted. However, this study still has limitations. Due to the limitations of the development of computer science, systems biology, network science, and mathematics, there is still room for exploration in the methods of this research. In addition, in order to confirm further the results of this research and determine the future application scenarios and scientific value of SMRR, further experimental research is needed. However, this study predicts some of the potential targets, biological function modules, and signaling pathways affected by SMRR in the treatment of KOA, which provide a theoretical basis and direction for further research.
Supplemental Material
Figure S1 - Supplemental material for Mechanism of Salviae Miltiorrhizae Radix et Rhizoma in the Treatment of Knee Osteoarthritis Based on Network Pharmacology
Supplemental material, Figure S1, for Mechanism of Salviae Miltiorrhizae Radix et Rhizoma in the Treatment of Knee Osteoarthritis Based on Network Pharmacology by Xiaoqing Shi, Haosheng Zhang, Yue Hu, Xiaochen Li, Songjiang Yin, Runlin Xing, Nongshan Zhang, Jun Mao and Peimin Wang in Natural Product Communications
Supplemental Material
Figure S2 - Supplemental material for Mechanism of Salviae Miltiorrhizae Radix et Rhizoma in the Treatment of Knee Osteoarthritis Based on Network Pharmacology
Supplemental material, Figure S2, for Mechanism of Salviae Miltiorrhizae Radix et Rhizoma in the Treatment of Knee Osteoarthritis Based on Network Pharmacology by Xiaoqing Shi, Haosheng Zhang, Yue Hu, Xiaochen Li, Songjiang Yin, Runlin Xing, Nongshan Zhang, Jun Mao and Peimin Wang in Natural Product Communications
Supplemental Material
Figure S3 - Supplemental material for Mechanism of Salviae Miltiorrhizae Radix et Rhizoma in the Treatment of Knee Osteoarthritis Based on Network Pharmacology
Supplemental material, Figure S3, for Mechanism of Salviae Miltiorrhizae Radix et Rhizoma in the Treatment of Knee Osteoarthritis Based on Network Pharmacology by Xiaoqing Shi, Haosheng Zhang, Yue Hu, Xiaochen Li, Songjiang Yin, Runlin Xing, Nongshan Zhang, Jun Mao and Peimin Wang in Natural Product Communications
Supplemental Material
Figure S4 - Supplemental material for Mechanism of Salviae Miltiorrhizae Radix et Rhizoma in the Treatment of Knee Osteoarthritis Based on Network Pharmacology
Supplemental material, Figure S4, for Mechanism of Salviae Miltiorrhizae Radix et Rhizoma in the Treatment of Knee Osteoarthritis Based on Network Pharmacology by Xiaoqing Shi, Haosheng Zhang, Yue Hu, Xiaochen Li, Songjiang Yin, Runlin Xing, Nongshan Zhang, Jun Mao and Peimin Wang in Natural Product Communications
Supplemental Material
Figure S5 - Supplemental material for Mechanism of Salviae Miltiorrhizae Radix et Rhizoma in the Treatment of Knee Osteoarthritis Based on Network Pharmacology
Supplemental material, Figure S5, for Mechanism of Salviae Miltiorrhizae Radix et Rhizoma in the Treatment of Knee Osteoarthritis Based on Network Pharmacology by Xiaoqing Shi, Haosheng Zhang, Yue Hu, Xiaochen Li, Songjiang Yin, Runlin Xing, Nongshan Zhang, Jun Mao and Peimin Wang in Natural Product Communications
Supplemental Material
Figure S6 - Supplemental material for Mechanism of Salviae Miltiorrhizae Radix et Rhizoma in the Treatment of Knee Osteoarthritis Based on Network Pharmacology
Supplemental material, Figure S6, for Mechanism of Salviae Miltiorrhizae Radix et Rhizoma in the Treatment of Knee Osteoarthritis Based on Network Pharmacology by Xiaoqing Shi, Haosheng Zhang, Yue Hu, Xiaochen Li, Songjiang Yin, Runlin Xing, Nongshan Zhang, Jun Mao and Peimin Wang in Natural Product Communications
Footnotes
Acknowledgments
We gratefully acknowledge the contributions of Yingying Yang (Nanjing University of Traditional Chinese Medicine) to study design and data analysis.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: SXQ and ZHS contributed equally to this paper, and they designed the study. Experimental work and data collection were conducted by SXQ and XRL. SXQ and ZHS analyzed and interpreted the data. SXQ and HY drafted the manuscript. WPM and XRL provided critical comments and revised the manuscript. ZNS, YSJ, MJ and WPM helped perform the analysis with constructive discussions. All authors read and approved the final version of the manuscript.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The present study was supported by the National Natural Science Foundation of China (81804123), and the Leading Talents of Traditional Chinese Medicine in Jiangsu Province project (SLJ0207).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
