Abstract
At present, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has spread all over the world, and the best way to effectively carry out drug diagnosis and treatment presents difficulties for all medical staff. In China, some traditional Chinese medicines (TCMs) have been successfully applied to the treatment of SARS-CoV-2 and have achieved good clinical results, including the Reyanning mixture. In this study, we systematically analyzed the mechanism of the Reyanning mixture and its effects against SARS-CoV-2 based on the method of network pharmacology. Here, we used the TCM Systems Pharmacology database and employed a similarity algorithm to screen and identify the bioactive ingredients and potential targets of the Reyanning mixture. The GeneCards database was used to predict and screen the disease targets and build the active ingredient target network diagram. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses were performed to construct the target signal pathway associations. The STRING tool was used to reconstruct the protein-protein interaction network. As a result, 27 candidate targets, such as tumor necrosis factor, interferon gamma, tumor protein P53, C-reactive protein, and peroxisome proliferator-activated receptor gamma, were identified among the 33 bioactive ingredients of the 4 TCMs in the Reyanning mixture with effects on treating SARS-CoV-2. These targets were significantly enriched in 20 KEGG pathways and associated with 48 diverse GO terms. All of these targets may play a role in inhibiting inflammatory reactions, regulating immune function, and reducing lung injury to achieve the purpose of treating SARS-CoV-2.
Infection with the novel coronavirus 2019 (nCoV), also known as severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), causes coronavirus disease 2019 (COVID-19) and was first observed in Wuhan, Hubei, at the end of 2019. With the spread of the disease, cases in China and other regions have been found. To date, there have been more than 2 million cases of infection worldwide according to statistics from the World Health Organization, and there is a trend of global spread, causing worldwide panic. 1 -3 To control the epidemic situation as soon as possible and protect human health, the Chinese government has formulated a series of “diagnosis and treatment plans for pneumonia caused by SARS-CoV-2 infection.” 4,5 It is mentioned in the fifth version that traditional Chinese medicine (TCM) treatment should be included in the diagnosis and treatment of pneumonia caused by SARS-CoV-2 infection, and TCM has achieved good results in treatment. 6,7
The Reyanning mixture is a kind of Chinese pharmacopeia. It is composed of 4 herbs: Taraxacum officinalis, family Asteraceae (PU GONG YING); Polygonum cuspidatum, family Polygonaceae (HU ZHANG); Patrinia villosa, family Valerianaceae (BAI JIANG CAO); and Scutellaria barbata, family Lamiaceae (BAN ZHI LIAN). It has the function of clearing away heat and detoxification. It is mainly used in the treatment of wind heat, cold, exogenous wind heat, sore throat, dry mouth, cough, and yellow phlegm caused by internal depression and fire melting, purulent tonsillitis, acute pharyngitis, acute bronchitis, and simple pneumonia. The effect of the Reyanning mixture in the treatment of respiratory tract infection, cold, fever, acute pharyngitis, pneumonia, and many other respiratory diseases has been shown to be significant. 8 The Reyanning mixture has been applied in the clinical observation and clinical treatment of SARS-CoV-2 in hospitals in several provinces in China. Clinical statistics show that nCoV pneumonia can be significantly improved by this treatment regimen, and the proportion of severe cases is reduced.
Recently, the pharmacodynamics of the Reyanning mixture have been evaluated in vivo by using a model of mice with lung disease (pneumonia) caused by coronavirus. 9 It was found that the Reyanning mixture can better treat BALB/c mice with coronavirus pneumonia and can improve the lung lesions, enhance the gastrointestinal function of mice, improve the autoimmune function of mice, and reduce the expression of inflammatory factors in vivo. Network pharmacology is a new subject based on the theory of system biology that analyzes the network of biological systems and selects specific signal nodes for multitarget molecular drug design. 10,11 It emphasizes the multichannel regulation of the signaling pathway, improves the treatment effect of the drug, reduces side effects to improve the success rate of the clinical trials of new drugs, and reduce research and development costs. According to the potential value of the Reyanning mixture for treatment of SARS-CoV-2, it is necessary to study the mechanism of the Reyanning mixture in treating nCoV pneumonia by using network pharmacology. This may provide some theoretical support for its wide application in clinical diagnosis and treatment.
Materials and Methods
Screening of Bioactive Ingredients in the Reyanning Mixture
To examine the 4 herbs of the Reyanning mixture, including PU GONG YING, HU ZHANG, BAI JIANG CAO and BAN ZHI LIAN, we used the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP, http://tcmspw.com/tcmsp.php) to obtain the molecular information on each herb, including the molecular name, relative molecular weight (MW), oil-water partition coefficient (AlgP), drug-likeness (DL), oral bioavailability (OB, %) and drug half-life (HL) for all the bioactive ingredients. 12 Then we screened the effective ingredients based on the pharmacokinetic parameters according to the criteria of OB ≥30%, DL ≥0.18, and HL ≥10.
Target Prediction of the Reyanning Mixture in the Treatment of SARS-CoV-2
According to the bioactive ingredients selected above in the TCMSP database, the potential targets were matched one by one and identified with the UniProt database (https://www.uniprot.org/). Here we used a similarity-based method to predict the potential targets of TCM ingredients that was first proposed by Perlman et al to rank potential drug-target interactions based on their similarity to known drug-target interactions. 13,14 Then, the drug targets were scored based on the similarity algorithm. The similarity score calculation formula was as follows.
The optimization of the scoring parameter r was performed in a cross-validation setting (0 ≤ r ≤ 1). A target with a score greater than 20 was considered as an essential target in the bioactive ingredients of TCM.
Next, we used the keyword “novel coronavirus” to search the NCBI and GeneCards databases, and the resulting proteins related to SARS-CoV-2 were retrieved. Furthermore, we determined the overlap of these disease-related targets with the essential targets of the bioactive ingredients in the 4 herbs. The common targets were identified to be potential targets of the Reyanning mixture with effects against SARS-CoV-2.
Gene Set Enrichment Analysis of Common Targets
Gene set enrichment analysis (GSEA) of Gene Ontology (GO), including biological process (BP), cellular component (CC), and molecular function (MF) terms, as well as the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways of the common targets identified above, was performed by using the Database for Annotation, Visualization and Integrated Discovery v6.8. 15,16 The cutoff of the P value was set as 0.01, and the false discovery rate (FDR) was set as 0.01.
Protein-Protein Interaction Network Construction
The functional protein association network of the common targets was constructed using STRING (https://string-db.org) v11.0. 17 We imported multiple proteins by using the UniProt names of our selected targets and chose the species “Homo sapiens.” The minimum required interaction score was set as a medium confidence 0.4. The protein-protein interaction (PPI) network was visualized by Cytoscape version 3.7.2. 18
Molecular Docking of SARS-CoV-2 Proteins and Small Molecules
The crystal structures of the SARS-CoV-2 3 Cl hydrolase (protein data bank [PDB] ID: 6LU7) and SARS-CoV-2 spike glycoprotein (S protein, PDB ID: 6VSB) were obtained from the PDB (https://www.rcsb.org/). The “sdf” file of small molecules was obtained from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/). Molecular docking analysis of SARS-CoV-2 proteins and small molecules was carried out with AutoDock Vina 1.1.2. 19 We first defined the potential binding sites of the 2 proteins (the x, y, and z center coordinates and radius were −13.2, 15.4, 73.1, and 5.9 Å for the SARS-CoV-2 3 Cl hydrolase and 226.4, 229.3, 222.8, and 83.8 Å for the SARS-CoV-2 spike glycoprotein, respectively) in Discovery Studio 2016, and then we ran AutoDock Vina with its default parameters and chose the docking result with the highest affinity and the lowest binding energy.
Results and Discussion
Identification of Bioactive Ingredients and Potential Targets in the Reyanning Mixture
Based on the screening of the 4 herbs from the Reyanning mixture in the TCMSP database, a total of 50 bioactive ingredients were identified for which OB ≥30%, DL ≥0.18, and the similarity algorithm score ≥20. There are 8 bioactive ingredients in PU GONG YING, 22 bioactive ingredients in HU ZHANG, 17 bioactive ingredients in BAI JIANG CAO, and 3 bioactive ingredients in BAN ZHI LIAN, as shown in Table 1. After deleting the duplicates, a total of 491 potential targets in the Reyanning mixture were obtained. In addition, we screened 346 SARS-CoV-2 related targets from the National Center for Biotechnology Information and GeneCards databases and defined them by using the UniProt database. By comparing 491 potential targets in the Reyanning mixture with 346 SARS-CoV-2 related targets, 27 common targets were identified that may be recognized as candidate targets in the Reyanning Mixture with effects against SARS-CoV-2. The comparison results are shown in Figure 1, and the information for the 27 candidate targets is shown in Table 2. According to the SARS-CoV-2 disease relevance scores, the 27 candidate targets are tumor necrosis factor (TNF), interferon gamma (IFNG), tumor protein P53 (TP53), C-reactive protein (CRP), peroxisome proliferator-activated receptor gamma (PPARG), mitogen-activated protein kinase 14 (MAPK14), transforming growth factor beta 1 (TGFB1), fibroblast growth factor 2 (FGF2), heat shock protein family A (Hsp70) member 5 (HSPA5), BCL2 associated X, apoptosis regulator (BAX), interleukin 1 beta (IL1B), prostaglandin-endoperoxide synthase 2 (PTGS2), nuclear factor kappa B subunit 1 (NFKB1), apolipoprotein E (APOE), ADAM metallopeptidase domain 17 (ADAM17), albumin (ALB), heme oxygenase 1 (HMOX1), adenosine deaminase (ADA), serpin family E member 1 (SERPINE1), catalase (CAT), promyelocytic leukemia (PML), protein kinase C alpha (PRKCA), interleukin 13 (IL13), prostaglandin-endoperoxide synthase 1 (PTGS1), ATPase Na+/K+ transporting subunit alpha 1 (ATP1A1), protein kinase C beta (PRKCB), and Spi-1 Proto-Oncogene (SPI1). The release of proinflammatory cytokines, including interleukin (IL)-1β and IL-6, and lung inflammation could be induced by SARS-CoV-2 infection; thus, IL-38 is recognized as a potential therapeutic cytokine that inhibits IL-1β and other IL-family members. 20 Elevated CRP and albumin (ALB) were highly correlated with acute lung injury and were reported to be potential diagnostic biomarkers for the treatment of 2019-nCoV infection. 21

Venn diagram of the comparison between the Reyanning mixture targets and the SARS-CoV-2 targets. The Venn diagram showed that there were 27 overlapping genes between the 491 traditional Chinese medicine targets of the Reyanning mixture and the 346 SARS-CoV-2 targets. The gene set is listed below the venn diagram.
The Effective Bioactive Ingredients and Targets of 4 Herbs in Reyanning Mixture.

The Information of 27 Candidate Targets.

These 27 candidate targets were distributed among 35 bioactive ingredients, as shown in Table 3. For PU GONG YING, all 8 of the selected bioactive ingredients have candidate targets. Both taraxasterol and taraxerol target NFKB1, BAX, and PML. Esculetin targets PPARG, PTGS1, and PTGS2. Esculetin was found to inhibit T-cell activation at a site distal to the production of IL-2 and IL-2 receptor expression. 22 Choline targets HMOX1 and CRP. A link between choline and macrophage phospholipid metabolism and inflammation has been established that is mediated by the choline transporter-like protein-1 (CTL1) transporter in polarized cells. 23 Scopoletin targets PRKCA and PRKCB. As one component of Artemisia capillaris, scopoletin was reported to suppress ConA-induced and LPS-induced adaptive immune cell activation. 24 Caffeic acid is the target of IFNG, and the modulatory effects of caffeic acid on human monocytes have been evaluated; caffeic acid is partially involved in propolis action. 25 Both chrysanthemaxanthin and flavoxanthin are targets of ADA.
The Candidate Targets in Each Bioactive Ingredient.

For HU ZHANG, the candidate targets were distributed among 13 bioactive ingredients. 3,5-Dimethyl-4-methoxybenzoic acid (DMMBA) has the most targets, and there were 10 candidate targets, including PTGS1, PTGS2, TP53, TNF, NFKB1, HSPA5, PPARG, SPI1, MAPK14, and FGF2. Rhein has 6 targets, including PPARG, PTGS1, PTGS2, SERPINE1, IL-1B, and IL-13. It was reported that rhein can induce oxidative stress and apoptosis in mouse blastocysts and has immunotoxic effects during embryonic development. 26 Trans-resveratrol has 5 targets, including PTGS1, PTGS2, ATP1A1, TGFB1, and ADAM17. Resveratrol has 3 targets, including ATP1A1, TGFB1, and ADAM17. Resveratrol may play potential roles in the immune response by suppressing Toll-like receptor (TLR) and proinflammatory gene expression. 27 Gallic acid also has 5 targets, including PRKCA, PRKCB, PTGS1, PTGS2, and PPARG. As one of the natural phenolic compounds, gallic acid has antioxidant and antitumor effects, which attenuate thymic involution via stimulation of FoxN1 expression. 28 Quillaic acid, polygalacic acid, and anthraquinone have 2 targets, PTGS1 and PTGS2. For mixed Th1/Th2 immune responses, compound 5 of Quillaja saponaria (QS)-7 can serve as a structurally defined synthetic vaccine adjuvant. 29 Polygalacic acid has been demonstrated to have immunomodulatory and antitumoral effects. 30 The subchronic toxicity and immunoregulatory and antibreast tumor effects of anthraquinone have also been evaluated in vitro and in vivo. 31 (+)-Catechin, (−)-3-hydroxy-4-methoxy-8-9-methylenedioxy pterocarpan (HMMP), and 7-hydroxy-4-methoxy-5-methylcoumarin (HMM) have 2 targets, PRKCA and PRKCB. Both emodin and emodin anthrone have the same target as TGFB1. As an anthraquinone derivative, emodin has been investigated to be potential in anti-inflammatory and antiproliferative effects in cells and mouse models. 32
For BAI JIANG CAO, there were 13 bioactive ingredients associated with candidate targets. The targets of CAT, APOE, and IL1B were distributed among α-bergamotene, patrinene, limonene, α-pinene, isopatrinene, β-pinene, α-muurolene, and copaene. Esculetin has 3 targets of PPARG, PTGS1, and PTGS2. β-Sitosterol has 3 targets of NFKB1, BAX, and PML. Scopoletin has 2 targets of PRKCA and PRKCB. Both villosolside and villoside have the same 2 targets of ALB and ATP1A1. For BAN ZHI LIAN, there was only 1 bioactive ingredient, p-hydroxybenzyl acetone, containing 3 targets of PTGS1, PTGS2, and TGFB1.
Functional Enrichment of Candidate Targets in the Reyanning Mixture
Based on GSEA of the above candidate targets in the Reyanning mixture, 20 significantly enriched KEGG pathways were identified to be associated with the effects of the Reyanning mixture in treating nCoV pneumonia. The details are shown in Table 4 and Figure 2. According to the ranking of the P value and the FDR, the associated terms were hsa05140: Leishmaniasis, hsa04380: Osteoclast differentiation, hsa05200: pathways in cancer, hsa05142: Chagas disease (American trypanosomiasis), hsa05146: Amoebiasis, hsa04010: MAPK signaling pathway, hsa05164: influenza A, hsa05321: inflammatory bowel disease (IBD), hsa04071: sphingolipid signaling pathway, hsa05143: African trypanosomiasis, hsa05161: Hepatitis B, hsa04066: hypoxia-inducible factor-1 signaling pathway, hsa05014: amyotrophic lateral sclerosis (ALS), hsa05152: tuberculosis, hsa05205: proteoglycans in cancer, hsa04664: Fc epsilon RI signaling pathway, hsa05168: herpes simplex infection, hsa04668: TNF signaling pathway, hsa05145: toxoplasmosis and hsa05144: malaria.

The summary of 20 significantly enriched KEGG pathways The barplot showed the significance of 20 KEGG pathways, which were significantly enriched in functions of candidate targets in the Reyanning mixture against severe acute respiratory syndrome coronavirus 2. The X-axis represents the -log10(FDR) value of each pathway. The Y-axis represents the KEGG name of each pathway, the numbers in brackets indicate the number of candidate genes in each pathway. FDR, false discovery rate; KEGG, Kyoto Encyclopedia of Genes and Genomes.
The Significantly Enriched KEGG Pathways.
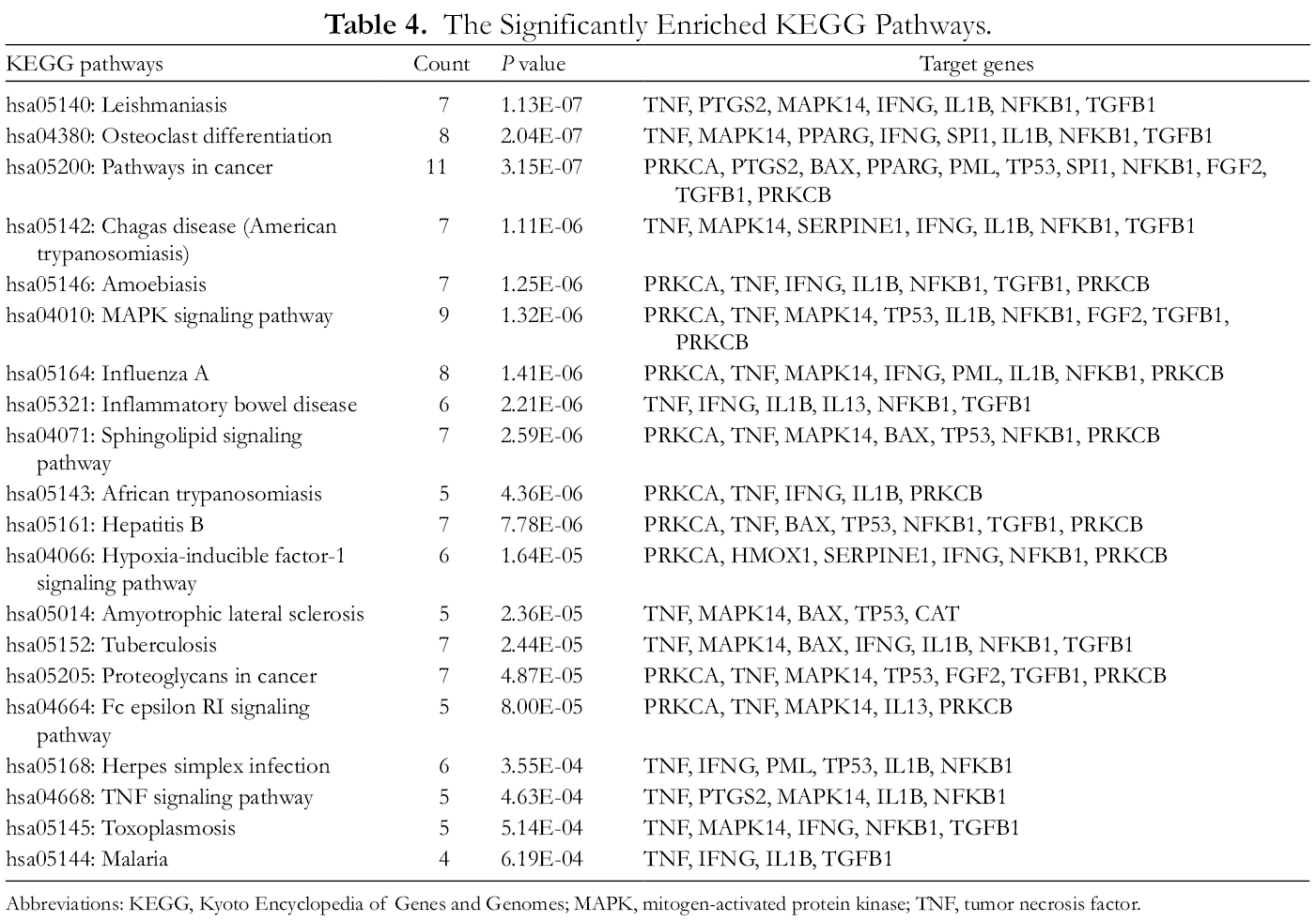
Abbreviations: KEGG, Kyoto Encyclopedia of Genes and Genomes; MAPK, mitogen-activated protein kinase; TNF, tumor necrosis factor.
Moreover, there were 37 BP terms, 4 CC terms, and 7 MF terms identified to be significantly enriched in the functional annotation of candidate targets involved in the effects of the Reyanning mixture on SARS-CoV-2, which are shown in Table 5. For the functional category of biological process, the top 5 terms were GO:0071260—cellular response to mechanical stimulus, GO:0045944—positive regulation of transcription from RNA polymerase II promoter, GO:0001666—response to hypoxia, GO:0042493—response to drug, and GO:0045080—positive regulation of chemokine biosynthetic process. The significant terms associated with cellular components were GO:0005615—extracellular space, GO:0005783—endoplasmic reticulum, GO:0070062— extracellular exosome, and GO:0005576—extracellular region. The significant terms associated with molecular function were GO:0019899—enzyme binding, GO:0005125—cytokine activity, GO:0051087—chaperone binding, GO:0042803—protein homodimerization activity, GO:0016209—antioxidant activity, GO:0005515—protein binding, and GO:0042802—identical protein binding.
The Significantly Enriched GO Terms.

Abbreviations: BP, biological process; CC, cellular component; FDR, false discovery rate; GO, gene ontology; MAPK, mitogen activated protein kinase; MF, molecular function.
Protein-Protein Interactions of Candidate Targets in the Reconstructed Network
The functional association network was reconstructed to represent 168 protein-protein interactions of 27 candidate targets by the STRING tool and is shown in Figure 3. In the network, the known interactions were obtained from curated databases or were experimentally determined, and the predicted interactions were related to the gene neighborhood, fusions, co-occurrence, and other interactions based on text mining, coexpression, and protein homology. According to the combined scores calculated on the basis of this information, the top 5 interactions were between TNF and IL13, TP53 and MAPK14, TNF and IL1B, BAX and TP53, and TNF and MAPK14.
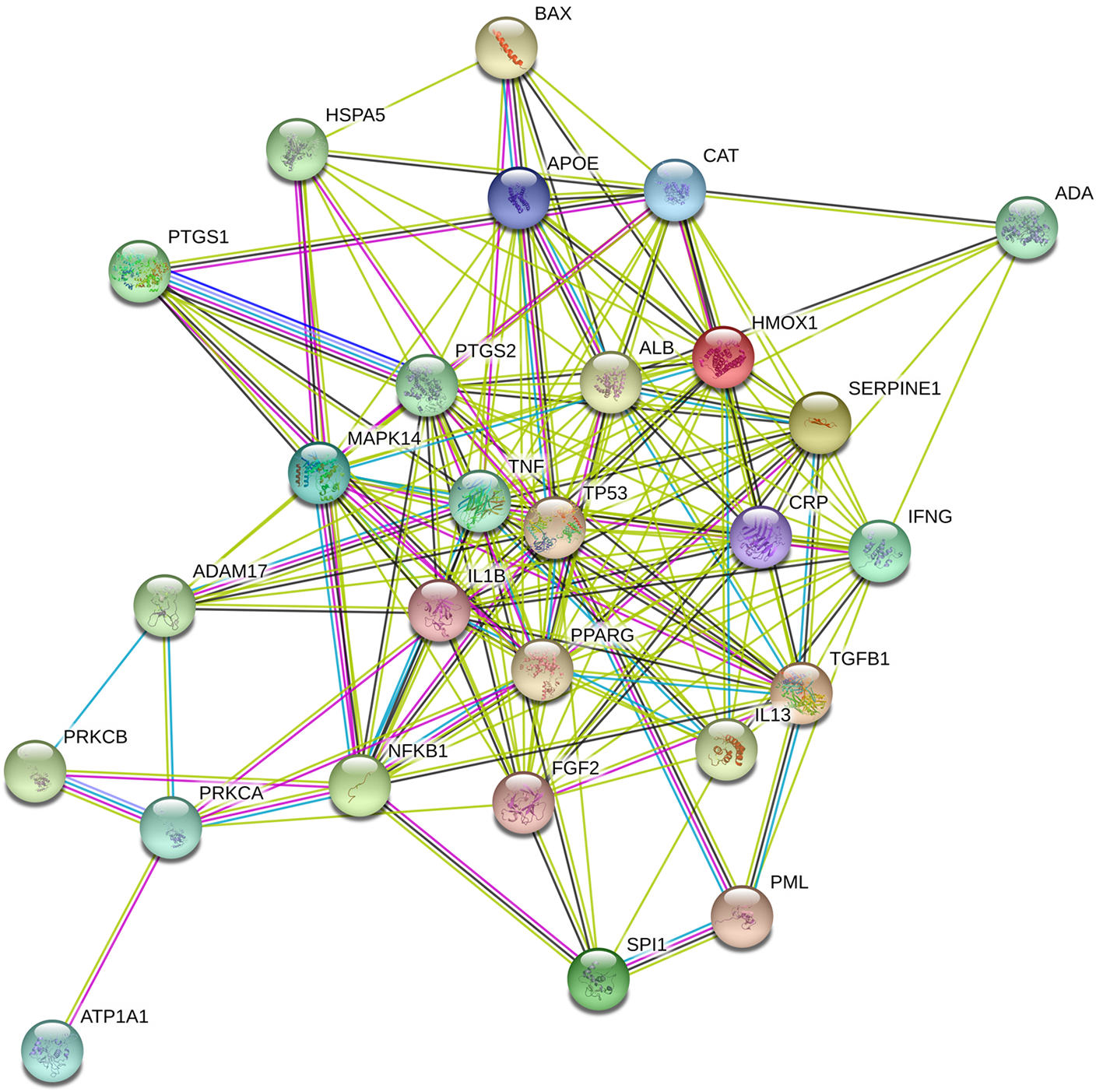
The protein interaction network of the 27 candidate targets. The protein-protein interaction network of the 27 candidate targets was constructed by using the STRING tool.
According to a study on the regulation of acute lung inflammatory injury by endogenous IL-13, the bronchoalveolar lavage levels of TNF-α were reported to be significantly increased caused by anti-IL-13. 33 In MCF-7 cells, p38 kinase plays an important role in the phosphorylation of TP53, which regulates ltraviolet-induced apoptosis mediated by p53. 34 The production of both TNF-α and IL-1β is considered to be important, as both are target molecules of immune interventions in Crohn’s disease. 35 Mitochondrial membrane permeabilization and apoptosis have been proven to be mediated through the direct activation of Bax by p53, which binds to BclXL via its deoxyribonucleic acid-binding domain. 36,37 The migration of vascular smooth muscle cells induced by TNF-α is reported to be dependent on MAPK. 38
Pharmacological Network of the Targets of the Reyanning Mixture With Effects Against SARS-CoV-2
Based on the integration of the above-analyzed results, we mapped the pharmacological network of bioactive ingredients and their targets in the Reyanning mixture with effects against SARS-CoV-2 by using Cytoscape. The detailed information is shown in Figure 4. In total, there were 27 candidate targets, such as TNF, IFNG, TP53, CRP, and PPARG of 33 bioactive ingredients in the 4 TCMs in the Reyanning mixture involved in treating SARS-CoV-2. All of these factors may play a role in inhibiting inflammatory reactions, regulating immune function, and reducing lung injury to achieve the purpose of treating SARS-CoV-2.

Pharmacological network of bioactive ingredients and their targets in the Reyanning mixture with effects against severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). The pharmacological network showed the bioactive ingredients of 4 Chinese herbs in the Reyanning mixture with effects against SARS-CoV-2 and their target information.
Molecular Docking of the Active Components of the Reyanning Mixture With SARS-CoV-2 Proteins
Based on the molecular docking analysis, the binding energy of each interaction between the main active components of the Reyanning mixture and the SARS-CoV-2 3 Cl hydrolase or SARS-CoV-2 S protein is given in Table 6. The docking results showed that the core components of the Reyanning mixture all have a good affinity with the SARS-CoV-2 S protein but have different affinities for the SARS-CoV-2 3 Cl hydrolase. The top 5 components with the lowest binding energies are α-pinene, β-pinene, choline, DMMBA, and gallic acid.
The Binding Energy Values of Some Components of Reyanning Mixture With Severe Acute Respiratory Syndrome Coronavirus 2 S Proteins.

Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article. We acknowledge financial supports by the National Natural Science Foundation of China (#81872832), Natural Science Foundation Project of Anhui Province (#1 908 085MC87) and the Scientific Research Foundation and Academic & Technology Leaders Introduction Project, and 211 Project of Anhui University (#10117700023).

