Abstract
This study aimed at exploring the active components and mechanisms of Jinhua Qinggan granules (JQG) in the prevention and treatment of coronavirus disease 2019 (COVID-19) using network pharmacology and molecular docking technology. These efforts were accomplished by employing the holistic approach of traditional Chinese medicine (TCM) and considering the virus-host interaction consisting of viral characteristics, the entry pathway into the host, and the resulting immune response. The chemical constituents and molecular targets of the 12 herbs from JQG were obtained using the TCM Systems Pharmacology database and analysis platform. UniProt was used to search for genes corresponding to JQG protein targets and Cytoscape 3.7.2 to construct the component-target (gene) network. Database for Annotation, Visualization and Integrated Discovery was used to perform enrichment analysis of gene ontology functions and the Kyoto Encyclopedia of Genes and Genomes pathways to predict the mechanism of action. The components ranked high in the network, and the major active components of the principal medicines, based on published literature, were docked with the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) 3CL hydrolase, SARS-CoV-2 spike glycoprotein (S protein), angiotensin conversion enzyme II (ACE2), and suppressor of cytokine signaling 1 (SOCS1). Visualization analysis demonstrated that the core active components of JQG had a strong affinity for SARS-CoV-2 3CL hydrolase, SARS-CoV-2 S protein, ACE2, and SOCS1. These data imply that the potential active components of JQG may act on multiple signaling pathways by binding to targets such as SARS-CoV-2 3CL hydrolase, S protein, ACE2, and SOCS1, thereby inhibiting virus replication and targeting cell binding, reducing host inflammation, and activating antiviral immunity to a certain extent.
Keywords
Pneumonia caused by the novel coronavirus disease 2019 (COVID-19), severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), mainly spreads through respiratory droplets and close contact with an infected individual. SARS-CoV-2 is a highly contagious disease that makes the global population generally susceptible, although some infected individuals remain asymptomatic. Currently, the pandemic is rampant around the world, posing a great threat to the lives and health of the global population. The number of infections in many countries has been sharply increasing, and the World Health Organization has raised the global risk level of COVID-19 to the highest level. 1 COVID-19 has grown to be a global public health emergency. The timeline for future vaccines remains unknown, and thus, there exists an urgent demand for a safe and effective treatment for infected patients.
Traditional Chinese medicine (TCM) has been an important means to prevent and control epidemics since ancient times, Namely, TCM has made great contributions to the prevention and treatment of many infectious diseases in the 21st century. 2 For several months of antiepidemic practice in China, TCM was involved in the treatment of COVID-19 in undiagnosed, mild, and severe cases around the country and has shown positive effects for the improvement of clinical symptoms of COVID-19. 3 Studies have shown that it can shorten the duration of illness (corresponding to the length of fever and length of hospital stay), improve the clinical cure rate, and reduce the progression of patients from moderate to severe disease. 4 Presently, the majority of the population is either susceptible or suspected to be infected with the virus. Timely and effective intervention in this population can “intercept and reverse” the duration of the disease and limit it to the early stage. Even if the disease is severe, early intervention may be beneficial to improve and minimize its severity for patients. Seasonal changes are an important consideration since there is also a high incidence of a variety of types of influenza. The treatment and convalescence of suspected COVID-19 patients to prevent concurrent influenza infection remain equally as important, thereby highlighting the necessity to use corresponding prescriptions.
Jinhua Qinggan granules (JQG), which are representative of the effective drugs introduced by the State Council Information Office of China for the prevention and treatment of COVID-19, are a new class 6.1 Chinese medicine developed by multiple departments in Beijing and are composed of Ma Xing Shi Gan and Yin Qiao San Decoctions, which are used to treat influenza A. 5 The formulation consists mainly of 12 TCMs: Lonicerae japonicae Flos, Forsythiae Fructus, Gypsum, Herba Ephedrae, Scutellariae Radix, Armeniacae Semen Amarum, Artemisiae annuae Herba, Fritillariae thunbergii Bulbus, Anemarrhenae Rhizoma, Fructus Arctii, Menthae Haplocalycis Herba, and Glycyrrhizae Radix et Rhizoma. Of these, Lonicerae Japonicae Flos, Herba Ephedrae, and Forsythiae Fructus are the principal medicines that contribute to the major effect of the JQG formula. According to TCM theory, JQG is effective for the prevention and treatment of influenza with wind and fever, and its effect is similar to that of Tamiflu. JQG was recommended as a Chinese patent medicine to treat the clinical symptoms of fatigue and fever during the observation period of COVID-19 in the 4th-7th edition of the COVID-19 diagnosis and also for a treatment program issued by the National Health Commission of China. According to Zhang Boli, 6 research on JQG for the treatment of COVID-19 demonstrated that the formula is effective for mild and moderate patients.
However, the active substances and mechanism of action of this formula in the treatment of COVID-19 are unclear. It is well known that viruses interact with the host as the basis of infection. In infectious disease prevention and control, there are many international drug studies that only focus on agents which target the host, using the law of interaction between the pathogen and host cells, especially targeting signaling proteins in order to explore the mechanism of disease prevention and treatment. 7 Current studies have shown that SARS-CoV-2 can trigger host immune regulation for the duration of COVID-19 progression. 8 Therefore, the prevention and mechanism of action of drugs targeting COVID-19 can be explored from 3 different aspects: active substances against the virus itself, the entry pathway, and the binding of targeted proteins in the host immune response.
Network pharmacology is widely used in pharmacodynamic investigations to elucidate molecular mechanisms of TCM formulas because of its systematic approach that can describe the interactions between complex biological systems and drugs. This resonates with the holistic view of TCM for syndrome differentiation and treatment to understand the nature of the disease. 9,10 The “disease-targets-drug” multilevel network directly illustrates the relevance of TCM formulas and diseases. Molecular docking is used to predict ligand-protein binding patterns and affinity by simulating ligand binding with receptor proteins. 11 The purpose of this study was to screen the targets of JQG through network pharmacology and to predict the core active compounds of JQG. These findings were used to conduct metabolic pathway analysis, followed by molecular docking of the active compounds with their chemical target. The results from metabolic pathway analysis and molecular docking provide insight to explore the molecular mechanism of JQG for the prevention and treatment of COVID-19 and serve as a reference for future clinical research. Figure 1 depicts a flowchart detailing the technical strategy and research idea of this study.
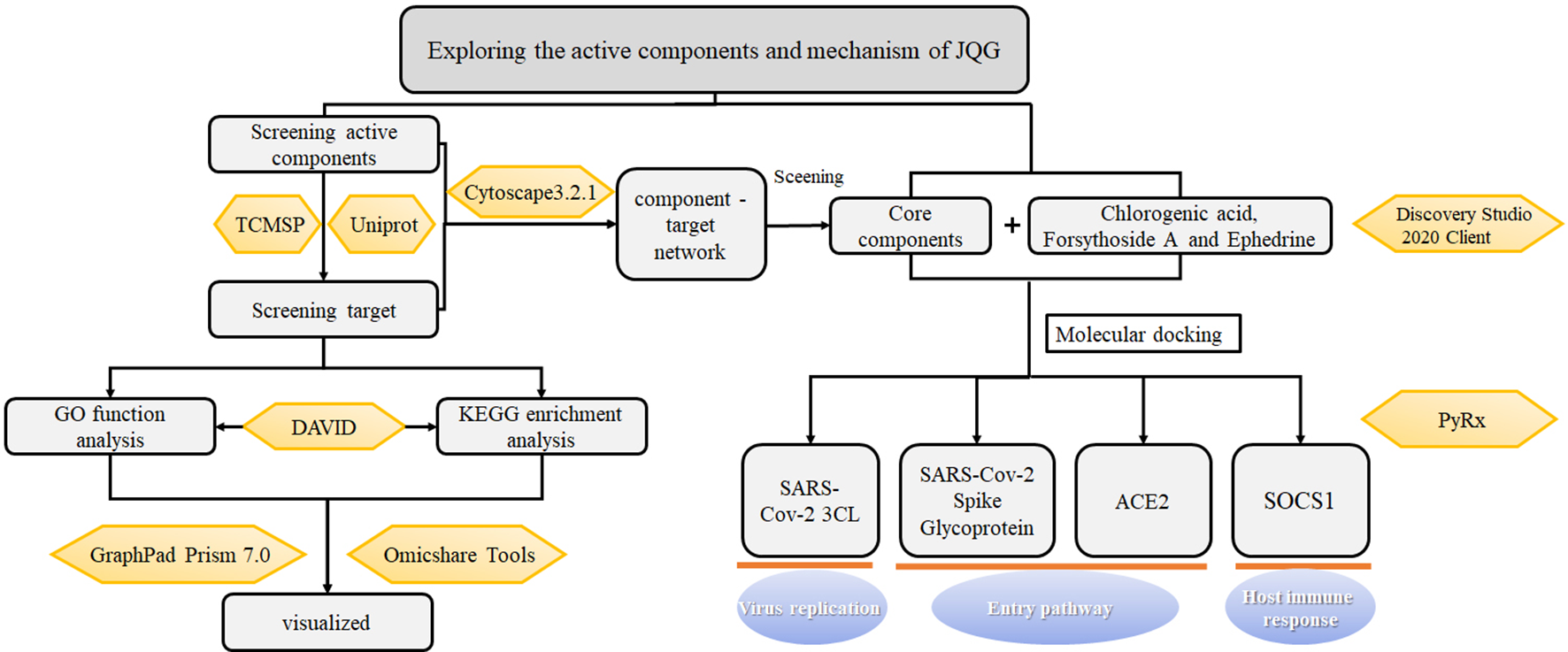
Flowchart detailing the technical strategy and research idea. ACE2, angiotensin conversion enzyme II; DAVID, Database for Annotation, Visualization and Integrated Discovery; GO, gene ontology; JQG, Jinhua Qinggan granules; KEGG, Kyoto Encyclopedia of Genes and Genomes; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; SOCS1, suppressor of cytokine signaling 1; TCMSP, traditional Chinese medicine Systems Pharmacology.
Materials and Methods
Active Component Screening, Target Prediction, and Component-Prediction Target Network Construction
Chemical constituents of each TCM component of the JQG formula were obtained using TCM Systems Pharmacology (TCMSP) database (http://tcmspw.com/). 12 We screened for the chemical components and corresponding targets at the same time with reference to the oral availability (OB) and drug-likeness (DL). The genes corresponding to the target proteins were queried using the UniProt database (https://www.uniprot.org/). 13 The screened components and corresponding targets were imported into Cytoscape 3.7.2 (http://www.cytoscape.org/), 14 and the component-target network was built by analyzing the component and target network.
Target Path Analysis
To explore further the function of the target and the signaling pathways involved, the results of the screening were imported into the Database for Annotation, Visualization and Integrated Discovery (DAVID) (http://david.ncifcrf.gov/home.jsp), 15 and anthropogenic, gene ontology (GO) enrichment of biological process analysis, and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis were performed. GraphPad Prism 7.0 and Omicshare Tools (http://omicshare.com/ Tools /index.php) 16 were used to visualize the results.
Component-Target Molecular Docking
The 2-dimensional (2D) structure SDF file format was obtained from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/). 17 The 3D structures (Figure 2) of SARS-CoV-2 3 Cl hydrolase (protein data bank [PDB] ID: 6LU7), SARS-CoV-2 spike glycoprotein (S protein; PDB ID: 6VSB), angiotensin conversion enzyme II (ACE2; PDB ID: 1R42), and suppressor of cytokine signaling 1 (SOCS1; PDB ID: 6C7Y) 18 were obtained from the Research Collaboratory for Structural Bioinformatics PDB database (https://www.rcsb.org/). Data were preprocessed using the Discovery Studio 2020 Client software. The components were processed using PyRx software to minimize their energy, and then the AutoDock Vina software was run for docking analysis. The ligand is assumed to bind spontaneously to the receptor if the binding energy is less than 0 kJ/mol.

3-Dimensional structure diagram of target protein. ACE2, angiotensin conversion enzyme II; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; SOCS1, suppressor of cytokine signaling 1.
Results
Active Component Screening
The components of JQG were searched in the TCMSP database and screened based on OB ≥30% 19 and DL ≥0.18. 20 After the components lacking target prediction data were removed, a total of 280 active ingredients remained. Of these, 24 were from Lonicerae japonicae Flos, 23 from Forsythiae Fructus, 23 from Ephedrae Herba, 36 from Scutellariae Radix, 22 from Artemisiae annuae Herba, 10 from Menthae Haplocalycis Herba, 20 from Armeniacae Semen amarum, 92 from Glycyrrhizae Radix et Rhizoma, 15 from Anemarrhenae Rhizoma, 8 from Fructus Arctii, and 7 from Fritillariae Thunbergii Bulbus. The active ingredient of gypsum could not be found in the database. The basic information of some of the active components found in JQG is shown in Table 1.
Basic Information of Some of the Active Components of JQG.

Abbreviations: TCM, traditional Chinese medicine; OB, oral availability; DL, drug likeness; JQG, Jinhua Qinggan granules.
The screening results show that there is some crossover of the active components contained in the 11 herbs of the JQG formula. This cross-distribution is shown in Figure 3.

Cross-distribution of components in JQG (A. Menthae Haplocalycis Herba; B. Glycyrrhizae Radix et Rhizoma; C. Lonicerae Japonicae Flos; D. Forsythiae Fructus; E. Ephedrae Herba; F. Armeniacae Semen Amarun; G. Scutellariae Radix; H. Anemarrhenae Rhizoma; I. Fritillariae Thunbergii Bulbus; J. Artemisiae Annuae Herba; K. Fructus Arctii). JQG, Jinhua Qinggan granules.
Component-Target Interaction Network
There are 195 unique active components after taking cross-distribution into account, and 295 target proteins of these components were identified in humans. The JQG component-target network has a total of 490 nodes, including 295 target nodes and 195 component nodes. There are 2674 edges between the component nodes and target nodes, as shown in Figure 4. In the network, the mean degree value (the number of associated targets) of the components is 13.7, and the mean component per target is 9.1, reflecting the mechanism of multicomponent and multitarget interactions of TCM. In the network, the degree value of a node represents the number of nodes that are connected to it. The higher the degree value, the more nodes that are related to that particular node. Analyzing the component-target network data of JQG with a set degree value, it was found that 14 components have ≥30 targets, including quercetin, luteolin, kaempferol, baicalein, 7-methyl-2 methyl isoflavones, naringenin, β-sitosterol, formononetin, dehydrated icariin, medicarpin, isorhamnetin, licochalcone A, and beans sterol, and their degree values are 143, 57, 57, 42, 42, 39, 35, 35, 34, 33, 33, 31, 31, 30, and 30, respectively.

The component-target network in JQG. (Light blue: all target nodes; rose purple: components contained in Glycyrrhizae Radix et Rhizoma; gray circles: components contained in Lonicerae Japonicae Flos; pink square: components contained in Forsythiae Fructus; orange square: components contained in Ephedrae Herba; brown green circles: components contained in Armeniacae Semen Amarun; light purple square: components contained in Scutellariae Radix; red square: components contained in Anemarrhenae Rhizoma; fluorescent green square: components contained in Fritillariae Thunbergii Bulbus; yellow circles: components contained in Artemisiae Annuae Herba; blue square: components contained in Menthae Haplocalycis Herba; dark green: components contained in Fructus Arctii). JQG, Jinhua Qinggan granules.
GO Function and KEGG Enrichment Analysis of Target Proteins
DAVID was used for GO and KEGG enrichment analysis of the target proteins. The data were screened based on

GO enrichment analysis of the protein targets of JQG. ACE2, angiotensin conversion enzyme II; BP, biological process; CC, cell composition; DAVID, Database for Annotation, Visualization and Integrated Discovery; GO, gene ontology; JQG, Jinhua Qinggan granules; KEGG, Kyoto Encyclopedia of Genes and Genomes; MF, molecular function; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; SOCS1, suppressor of cytokine signaling 1; TCMSP, traditional Chinese medicine Systems Pharmacology.
A total of 132 signaling pathways were identified through KEGG pathway enrichment analysis, including the tumor necrosis factor (TNF), hypoxia-inducible factor-1 (HIF-1), T cell receptor, and Toll-like receptor signaling pathways, as well as other signaling pathways related to inflammation
21
(

KEGG pathway enrichment analysis of the protein targets of JQG. JQG, Jinhua Qinggan granules; KEGG, Kyoto Encyclopedia of Genes and Genomes.
Molecular Docking of the Active Components of JQG With SARS-CoV-2 3CL Hydrolase
The 5 components with the highest target degree value identified by network pharmacology and the 3 main active components of clinical antiviral treatments described in the literature 22,23 (chlorogenic acid, forsythoside A, and ephedrine) were docked with SARS-CoV-2 3 Cl hydrolase. The binding energy of each interaction is given in Table 2. It is generally believed that it is the lower the energy of the ligand conformation that yields stable binding to the receptor and the higher the possibility of interaction. The docking results showed that the core components of JQG and the main active components of the clinical treatments all have a good affinity with SARS-CoV-2 3 Cl hydrolase, and the binding energies of the JQG components are either similar to or even lower than those of Western medicine, as shown in Table 2.
Binding Energy Values of the Core Components of JQG and Clinical Treatments with SARS-CoV-2 3 Cl Hydrolase.

Abbreviation: SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; JQG, Jinhua Qinggan granules.
Molecular Docking of the Active Components of JQG With SARS-CoV-2 S Protein and ACE2
Studies have shown that SARS-CoV-2 may invade the host through binding of the SARS-CoV-2 S protein with ACE2. 24 Therefore, the active components of JQG which bind to the 3 Cl hydrolase, associated with ribonucleic acid replication of SARS-CoV-2, were analyzed for affinity with the S protein and ACE2. The results show that most of the components, such as quercetin, luteolin, kaempferol, chlorogenic acid, and forsythiaside A, have low binding energy and high affinity with S protein and ACE2. The corresponding results are shown in Table 3.
Binding Energy Values of Some Components of JQG with SARS-CoV-2 S Protein, ACE2, and SOCS1.

Abbreviation: SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; SOCS1, suppressor of cytokine signaling 1; ACE2, angiotensin conversion enzyme II.
Molecular Docking of the Active Components of JQG With Antiviral Immune-Related Targets
Based on the theory in TCM of dispelling pathogens and strengthening the positive, the host’s antiviral immune response to the virus must be considered. The immune-activating protein STAT1, related to antiviral immunity, and the negative regulatory factor SOCS1 25 were chosen for analysis by molecular docking. The results show that quercetin, luteolin, chlorogenic acid, and forsythiaside A have a high affinity for SOCS1, suggesting that they are conducive to immune activation of STAT1, as shown in Table 3 and Figure 7.

Molecular docking patterns of some components with SARS-CoV-2 3 Cl hydrolase, SARS-CoV-2 S protein ACE2, and SOCS1. ACE2, angiotensin conversion enzyme II; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; SOCS1, suppressor of cytokine signaling 1.
Discussion
Several reports have demonstrated that after the novel coronavirus enters the host, it stimulates the immune system to produce a large number of inflammatory factors that lead to a deleterious cytokine storm. 26 This is followed by severe symptoms that may lead to death, such as ventilation dysfunction, acute respiratory distress syndrome, and acute myocardial injury. Therefore, preventing or attenuating cytokine storms may be the key to the successful treatment of infected patients. The results of network pharmacological analysis suggest that the components of JQG can potentially act on multiple targets in the human body. The KEGG analysis indicates that these targets are involved in multiple pathways in the human body, including Toll-like receptors, retinoic acid-inducible gene (RIG)-I-like receptors, TNF, and HIF-1 signaling pathways, as well as other inflammation-related pathways. This indicates that JQG may treat viral pneumonia by regulating inflammatory-related pathways in the host. In addition, it is worth noting that multiple pathways are involved in antiviral innate immunity, including influenza A, Janus kinase/signal transducer and activator of transcription 1 (STAT1), and other pathways. STAT1 plays an important role in antivirus immunity and immune activation. 27 The results of molecular docking indicate that the core active components of the JQG formula show high binding energy with SOCS1, a negative regulator of STAT1.
A recent study reported that SARS-CoV-2 3 Cl hydrolase is a key enzyme that regulates the replication of COVID-19. 28 Therefore, blocking SARS-CoV-2 virus replication by inhibiting the activity of this enzyme is a promising avenue of research. In this study, 5 core JQG components were selected for molecular docking studies based on the degree value with targets that were identified using network pharmacology. The top 3 components showing the strongest association to target proteins were quercetin, luteolin, and kaempferol. These 3 components are found in the principal medicines (TCM theory considers that these kinds of herbs are the most effective medicine in a formula), Lonicerae japonicae Flos, Ephedrae Herba, and Forsythiae Fructus, and show a higher affinity for SARS-CoV-2 3 Cl hydrolase than synthetic medicines. Moreover, these compounds have been reported as potential active substances in other formulas used to treat COVID-19 and thus warrant further investigation. 29 However, due to the limitations of screening components by network pharmacology, these 3 chemicals are abundant in plants that are not representative components of the TCM found in the JQG formula. Therefore, we also selected active components with known antiviral properties which are representative of the principal medicines of JQG. Consequently, the results suggest that chlorogenic acid (Lonicerae japonicae Flos), forsythiaside A (Forsythiae Fructus), and ephedrine (Ephedrae Herba) have a high affinity for SARS-CoV-2 3 Cl hydrolase and could be potential active ingredients of JQG.
Currently, ACE2 is considered an important factor for the entry of the SARS-CoV-2 virus to the human body. 21 COVID-19 is similar to SARS since both viruses invade host cells through the binding of S protein on the viral surface to ACE2 on the surface of lung cells, 30 but the 2 causative viruses are structurally different. Compared with SARS, the SARS-CoV-2 S protein of COVID-19 has a stronger binding affinity for ACE2, resulting in higher infectivity of COVID-19. In addition to the lung, ACE2 is widely expressed in human tissues, including the heart, liver, kidney, and digestive organs. 31 For this reason, blocking the binding of S protein to ACE2 could prevent the disease from progressing its severity or delaying the development of the disease to provide more time for adequate clinical response to treatment. Based on this assumption, we performed preliminary studies on the binding of core JQG components to S protein and ACE2. The results show that quercetin, luteolin, kaempferol, chlorogenic acid, and forsythiaside A all have a strong affinity to both S protein and ACE2 with respective binding energies much lower than −5 kJ/ mol. This indicates that these potential active components may prevent and treat COVID-19 by destroying the binding ability of S protein to ACE2, thereby reducing the ability of the virus to invade host cells.
Conclusions
The components of JQG may be beneficial for the treatment of COVID-19 in 3 main ways: attenuating cytokine storms, enhancing antiviral immunity, and inhibiting the activity of SARS-CoV-2, including its replication and spread. This study provides evidence for the study of the material basis and mechanism of action of JQG. Moreover, combined with the theory of “homotherapy for heteropathy” in TCM, it draws attention to the possibility of JQG for use not only in treating COVID-19 but also influenza and infections of other respiratory pathogens. Future in-depth studies of the molecular mechanisms of TCM will be helpful to explain the scientific connotations of this medicine and to broaden the application scope in order to potentially benefit the health of the global population.
Footnotes
Acknowledgments
The authors are grateful to Antonio Del Rio Flores for helpful discussions.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Construction Project of the Inheritance Studio of National Famous Chinese Medicine Experts of the State Administration of Traditional Chinese Medicine (Memo on teaching and education of traditional Chinese medicine [2019] No. 41); Key Research and Development Program 2018SZ0078 of Science and Technology Department of Sichuan Province; The Fourth National TCM Doctor (Clinical and Basic) Excellent Talents Research and Study Program (CMM No. 24 [2017]).
