Abstract
Keywords
Background
Ulcerative colitis (UC) is not a specific intestinal inflammatory disease. Its lesions involve the mucosa and submucosa of the colon and rectum, and it has been listed as one of the modern refractory diseases. 1 In China, the incidence of UC is increasing yearly, and its symptoms include recurrent or continuous diarrhea, abdominal pain, mucous, pus, bloody stools, and tenesmus. 2 The course of the disease is protracted, and it does not heal, which has an impact on patients’ quality of life. There are many types of drugs for the treatment of UC, including salicylic acid preparations, immunosuppressive agents, hormones, and biological agents. However, their curative effect is not satisfactory, and toxic side effects are prone to occur.3,4 Currently, there is no effective treatment for UC. The principle of treatment is to control the onset of the disease, stabilize the condition, reduce the recurrence rate, and reduce complications. Therefore, it is of great clinical significance to explore the pathogenesis of UC and find effective intervention methods.
Traditional Chinese medicine has attracted increasingly more attention in the treatment of UC, which belongs to the category of “dysentery,” “intestinal dysentery,” and “long dysentery” in Chinese medicine. Pulsatilla decoction (PD) is a classic prescription for the treatment of dysentery. Many studies have confirmed that this prescription has a definite effect in the treatment of UC.5,6 Professor Sun Hongwen, a well-known Chinese medical doctor in Jiangsu Province, supplemented PD with herbs for regulating qi and blood. She concluded that modified Pulsatilla decoction (MPD) can clear heat and dampness, remove blood stasis, protect membranes, cool blood, and stop dysentery. Clinically, MPD is often used to assist in the treatment of UC patients. Previous animal studies find that MPD can promote the colonization of intestinal dominant Bifidobacterium in intestinal epithelial cells and increase Bifidobacterium adhesion and adhesion receptors. MPD can also downregulate the level of interleukin-8 (IL-8), tumor necrosis factor-α (TNF-α), diamine oxidase, D-lactic acid, and deoxycholic acid in serum, and increase the serum level of ursodeoxycholic acid. 7 This may be related to regulating the barrier of the intestinal mucosa and repairing the damage to it.
Traditional Chinese medicine is a multitargeted synergy with a strong system and integrity. Although previous studies have confirmed that MPD has good clinical efficacy in UC, there remains a lack of systematic research. Network pharmacology is a new subject integrating systems biology, multidirectional pharmacology, computational biology, and network analysis. It emphasizes the multichannel regulation of signal pathways to improve the therapeutic effect of drugs and reduce toxic side effects.8-10 Through the construction of a multilevel network, the relationship between drugs and diseases can be explored from a holistic perspective, and the mechanism of the action between drugs and the body can be analyzed.
Therefore, the aim of this study was to explore the target and signal pathways of MPD in the treatment of UC with the help of network pharmacology and molecular docking technology. In this way, the relationship between drugs and related genes can be evaluated, and new ideas and methods can be provided for follow-up research. The flowchart of this study is shown in Figure 1.

Whole framework based on network pharmacology.
Materials and Methods
Screening the Active Ingredients of MPD
By searching the Traditional Chinese Medicine System Pharmacology Database (TCMSP https://tcmspw.com/tcmsp.php, Version 2.3), we gathered the main ingredients of Radix Pulsatilla, Rhizoma Coptidis, Phellodendri chinensis cortex, Fraxini cortex, hairyvein agrimony, Panax notoginseng, Rhizoma Bletillae, and Radix Arnebiae in the MPD. The screening conditions were set as oral BiOAvailability (OB)≥30% and druglikeness (DL)≥0.18 to obtain active ingredients with better druggability. 11
Screening Targets Common to MPD and UC
According to the TCMSP database, the targets involved in the active ingredients in MPD are sorted. We obtained the MPD gene targets using the UniProt database (https://www.uniprot.org/). 12 Using UC as a keyword, we searched for the genes related to UC and screened them through GeneCards (https://www.genecards.org/), OMIM (https://www.omim.org/), PharmGKB (https://www.pharmgkb.org/), DisGeNET (https://www.disgenet.org), and DrugBank databases (https://www.drugbank.ca).13-17 Finally, the genes corresponding to the active ingredients of MPD were mapped with UC-related target genes, and common genes were screened out. We used the Venny 2.1.0 tool to create the Venn diagram.
Constructing a Network of Active Drug Ingredients and Disease Targets
The active ingredients and common genes data of MPD were imported into Cytoscape version 3.7.1 software to construct the “active ingredients-target-disease” network. A node in the network represents a target or active ingredients, and an edge represents the interactions between nodes. The degree value of a node indicates the No. of routes connected to other nodes in the network. A node with a high degree value may play a key role in the network and can be used to evaluate the importance of the node to screen out the important ingredients in MPD.
Construction of the PPI Network and Screening of Core Genes
The protein-protein interaction (PPI) network is formed by the interaction between proteins and other protein molecules in the network. This interaction is the basis of cell metabolism and is essential for maintaining the function of life. The systematic analysis of protein interactions in biological systems is highly significant for understanding the changes in biological signals and the functional relationships between proteins under disease conditions. The STRING database (https://string-db.org/) collects many protein interactions and contains a series of data confidence (eg, low confidence: < 0.4, medium confidence: 0.4-0.7, and high confidence: > 0.7), and so can be used to predict protein interactions. We imported the common genes into the STRING database, set the species search to Homo sapiens with a confidence of 0.7, and hid the isolated target to perform PPI network analysis on the common genes. The result was stored in Tab Separated Values format and imported into Cytoscape software. We used the CytoNCA app to evaluate the topological properties of nodes in the interactive network. The core targets were selected according to degree centrality (DC), betweenness centrality (BC), closeness centrality (CC), eigenvector centrality (EC), network centrality (NC), and local average connectivity (LAC). These six parameters measured the importance of nodes in the network and indicated the nature of the nodes in the network. High DC, BC, CC, EC, NC, and LAC values indicate that a node is vitally important to the network. According to the analysis results, a value greater than the median is selected as the condition for screening, and the core target is finally obtained after 2 screenings. 18
GO and KEGG Enrichment Analysis
First, we imported all common genes into the Database for Annotation, Visualization, and Integrated Discovery (DAVID, http://david.nifcrf.gov/) .19,20 Then, the data were analyzed for gene ontology (GO) function enrichment (molecular function, biological process, cellular ingredient) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis. In the programming language, pvalueCutoff = 0.05, qvalueCutoff = 0.05, and the emapplot function were used to support hypergeometric tests and to obtain the results of the target enrichment analysis. The relationships among the overlapping targets based on the enriched GO terms were displayed. Differentially expressed genes that overlap 2 GO terms represent 2 nodes that have overlapping relationships. In the diagram, the overlapping targets are connected with lines, and the top 30 targets in the network are shown. In the KEGG enrichment analysis, the top 20 items, determined by the highest enrichment degrees, were selected and displayed in the form of bar graphs.
Molecular Docking Steps and Results Evaluation
We used Discovery Studio software to perform molecular docking and the Research Collaboration for Structural Bioinformatics Protein Data Bank Database (https://www.rcsb.org/) was used to obtain the core target structures. 21 We obtained the pretreated protein structures using the Prepare Protein tool, and the parameters were set to the default values, including pretreatments such as dehydration, hydrogenation, and ring building. We used the Define and Edit Binding Site tool to define the active center of the target molecule and determined the location and size of the active center based on the positive ligand position for the protein as obtained from the Protein Data Bank and related crystal information in the literature.
In the Chemical Book database (https://www.chemicalbook.com/), the active compound structures were accessed in mol2 format. We generated different conformations of the active compounds using the Prepare Ligands tool to compile a small-molecule ligand library. The LibDock method in the Dock Ligands module was used for molecular docking. The docking preferences were set to high quality, the conformation method was fast, and the parallel processing was true. Other parameters were selected as the default values. The docking results were given by the scoring function LibDock score. The higher the LibDock score, the higher the binding activity of the predicted ingredients with the target.
Animals
A total of 50 male SD rats, SPF grade, weighing 180 to 220 g, were purchased from Suzhou Sino Biotechnology Co., Ltd. The rats were randomly assigned to 6 groups (n = 10): control group, model group, high-MPD (H-MPD) group, middle-MPD (M-MPD) group, and low-MPD (L-MPD) group. All animals were maintained under a 12 h light/dark cycle environment. The room temperature was controlled at 20 to 24 °C, with 45% to 70% humidity, and rats partook freely after disinfection of drinking water and feed. The study protocol was approved by the Ethics Committee of Suzhou Hospital of TCM (ECMUC2019005AO).
Drugs and Reagents
The composition of the MPD was 30 g of Fraxini cortex, 10 g of Phellodendri chinensis cortex, 30g of Radix Pulsatillae, 10g of Rhizoma Coptidis, 30g of hairyvein agrimony, 6 g of Panax notoginseng, 30 g of Radix Arnebiae, and 20 g of Rhizoma Bletillae. All the traditional Chinese medicine decoction samples were provided by the pharmacy department of Suzhou Traditional Chinese Medicine Hospital and were identified as qualified traditional Chinese medicine decoction samples. We took one dose of MPD, soaked it for 30 min, decocted it twice according to the conventional decocting method, and filtered and merged the extracts, which were concentrated in a rotary evaporator under vacuum (temperature 80 °C) to the equivalent of 2 g/mL of crude drug. Finally, we stored the liquid medicine in a 4 °C refrigerator for subsequent use. Dextran sulfate sodium (DSS) was obtained from MP Biomedicals (Santa Ana, CA, USA), rabbit anti-human antibodies against p-p38MAPK, p-MLC, β-actin, and secondary antibodies from Abcam, and pentobarbital sodium from Sigma (Sigma-Aldrich).
Treatment Protocol for DSS-Induced Colitic Rats
After 1 week of adaptive feeding, the drinking water of the remaining 40 rats (except the rats in the blank group) was replaced with DSS solution for 6 days. 22 When the model rats displayed effects such as weight loss, mental lethargy, loose stools, and mucus purulent bloody stools, we deemed the modeling successful. 23 The rats were dosed on the third day after establishing a model according to the conversion of body surface area between people and rats. The H-MPD, M-MPD, and L-MPD groups were given 17.28 g/kg, 8.64 g/kg, and 4.32 g/kg MPD, respectively, and the control group and model group were given the same volume of saline enema. The dosage volume of each group was 3 mL once a day for 14 consecutive days. Rats in each group fasted for 24 h after the last administration. After weighing, the rats were anesthetized with pentobarbital sodium (50 mg/kg) and then euthanized. Then, 4 to 8 cm of colon tissue was removed from the anal margin upward and a cut was made along the longitudinal axis of the mesentery. The tissue was rinsed with distilled water, and then the intestinal mucosal tissue morphology was observed before the tissue was refrigerated at −80 °C for later use.
Histomorphology of Colon Mucosa
The colon tissues were fixed, embedded, sectioned, and stained with uranium acetate and lead citrate before the ultrastructure of the colon under an electron microscope was observed. The degree of colonic mucosal injury was observed macroscopically, and the colonic mucosal damage index (CMDI) was evaluated. The scoring standard was as follows 24 : no damage to the colonic mucosa was scored zero; local congestion and edema of the mucosa without thickening of the intestinal wall or ulcer was scored one; mucosal hyperemia, edema, and thickening of the intestinal wall without ulcer formation was scored 2; a single ulcer formation was scored 3; ulcers or local inflammation in multiple locations scored 4; and an ulcer along the long axis of the colon >1cm was scored 5.
Western Blotting Analysis
Radioimmunoprecipitation lysis buffer with the protease inhibitor cocktail (Beyotime Biotechnology, China) was used to extract the total protein from the colon tissue. The sample protein content was determined according to the BCA method and the loading amount was calculated. The separation gel and concentrated gel were configured according to the molecular weight of the target protein, the sample was loaded according to the calculated loading amount, and electrophoresis began. When the dye reached the bottom of the gel, the electrophoresis was stopped. To the transferred membrane was added the primary antibody and the secondary antibody, and Tanon 5200 gel image processing system was used to analyze the expression of the target band after being exposed to film.
Statistical Processing
SPSS 23.0 statistical software (IBM Corp.) was used to analyze the data. GraphPad Prism 7.0 software (GraphPad Software Inc.) was used for the calculations. All results are expressed as means ± SD. Student's t-test was used to evaluate the paired data, and ANOVA was used to make multiple comparisons. A P value < .05 was considered statistically significant.
Results
Screening of Active Ingredients in MPD
According to the conditions of OB ≥ 30% and DL ≥ 0.18, 82 active ingredients were found in MPD. After deleting 31 that were not associated with UC, 51 active ingredients were ultimately included. See Supplementary Table 1 for details.
Target Acquisition for Active Ingredients and Disease-Related Effects
The targets involved in the active ingredients of MPD were retrieved from the TCMSP database. The UniProt database was used to obtain the gene name of the target, and 204 active ingredient targets were obtained after correction and deduplication. As shown in Supplementary Table 2, 4622 UC-related gene targets were obtained after searching in multiple databases with “ulcerative colitis” as the search term. We mapped 204 active ingredient targets with 4622 UC-related targets and obtained 141 common genes. See Supplementary Table 3 for details. The Venn diagram is shown in Figure 2.

Common genes of modified Pulsatilla decoction (MPD) and ulcerative colitis (UC).
“Active Ingredients-Target-Disease” Network of MPD
We matched 141 common genes with 51 active ingredients and imported them into Cytoscape software to construct an active ingredients-target-disease network. The detailed information is shown in Supplementary Table 4. The network has 192 nodes (51 active ingredients, 141 common genes) and 884 edges, as shown in Figure 3. The quadrilateral represents the active ingredients and the ellipse represents the common genes. Figure 3 shows that one target can correspond to one or more active ingredients, and multiple targets can correspond to the same active ingredient, which indicates that MPD has the characteristics of a multi-ingredient and multi-target treatment of UC.

The “active ingredients-common gene-disease” network. The blue quadrilateral represents the common genes and the green quadrilateral represents the active component of modified Pulsatilla decoction (MPD).
PPI Network Construction and Core Gene Screening
We uploaded common genes to the STRING database to construct a protein interaction network. After that, we imported the file into Cytoscape for visualization. We used CytoNCA to calculate the topological parameters of the network nodes to get the DC, BC, and CC parameters. The first screening threshold was DC ≥ 16, EC ≥ 0.065, LAC ≥ 7.124, BC ≥ 173.773, CC ≥ 0.443, and NC ≥ 10.113. The results show that there are 26 nodes and 230 edges in total. The second screening threshold was DC ≥ 37.767, EC ≥ 0.158, LAC ≥ 14.110, BC ≥ 590.397, CC ≥ 0.546, and NC ≥ 24.624. The second screening results were 6 nodes and 12 edges, including mitogen-activated protein kinase 1 (MAPK1), mitogen-activated protein kinase 8 (MAPK8), RAC-alpha serine (AKT1), vascular endothelial growth factor-A (VEGFA), transcription factor AP-1 (JUN), and interleukin-6(IL-6). The core target screening process is shown in Figure 4.
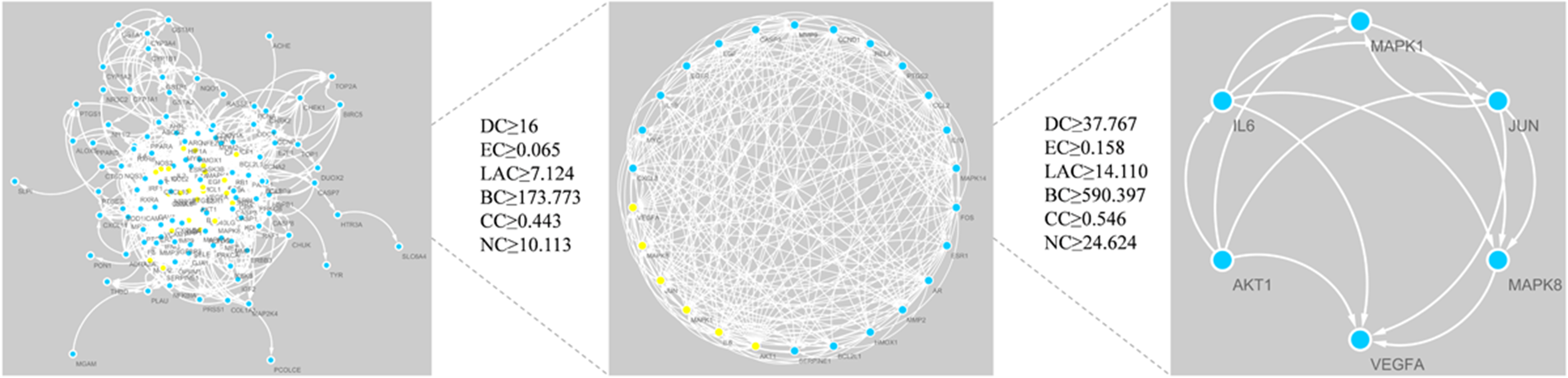
Process of topological screening for PPI network.
GO and KEGG Pathway Enrichment Analysis
GO enrichment analysis was performed on 141 common genes. The results showed that the key targets of MPD in the treatment of UC were highly enriched in 132 GO terms. The top 30 GO analysis results were screened with P < .05 as the threshold. The biological functions and processes included BH domain binding, oxidoreductase activity, ubiquitin-protein ligase binding, DNA-binding transcription activator activity, RNA polymerase II-specific, phosphatase binding, peroxidase activity, kinase regulator activity, cysteine-type endopeptidase activity involved in the apoptotic process, RNA polymerase II transcription factor binding, antioxidant activity, receptor–ligand activity, tetrapyrrole binding, heme binding, cytokine activity, cytokine receptor binding, steroid hormone receptor, transcription factor activity, and nuclear receptor activity. The results are shown in Figure 5. A cluster profiler was used for KEGG pathway enrichment analysis, and 158 pathways were enriched. The first 20 pathways with more enriched genes were screened with P < .05 as the threshold. They include interleukin-17 (IL-17) signaling pathway, tumor necrosis factor (TNF) signaling pathway, advanced glycation end products-receptor for advanced glycation end products (AGE-RAGE) signaling pathway in diabetic complications, C-type lectin receptor signaling pathway, viral infection-related signaling pathways, and some cancer pathways. See Figure 6 for details.

GO enrichment analysis of common genes. The larger the node and the smaller the P value, the higher the correlation.

KEGG pathway enrichment analysis of common genes. The smaller the P value, the greater the correlation.
Target Path Analysis
We used a KEGG mapper tool to obtain the pathway map of MPD in the treatment of UC, and the AGE-RAGE signaling pathway is shown in Figure 7. The target of the pathway is marked in white, and the marker of MPD in the treatment of UC in red. As shown in the figure, the effect of MPD in the treatment of UC includes the MAPK signaling pathway, the phosphatidylinositol 3-kinase (PI3K)-AKT signaling pathway, the NF-κB signaling pathway, the JNK/c-Jun signaling pathway, the Jak-STAT signaling pathway, the TGF-βsignaling pathway, and the ERK signaling pathway, involving 31 effective targets of MPD in the treatment of UC. It was suggested that MPD may play a role in the treatment of UC by regulating several aspects, and its target may be located in these pathways.

The AGE-RAGE signaling pathway of MPD in the treatment of UC.
Molecular Docking Analysis
The results of molecular docking indicate that the active ingredients of MPD may act on the target of UC and are potentially effective ingredients with certain pharmacodynamics. The results of the ligand–receptor protein molecular docking are shown in Supplementary Table 5. Figure 8 shows the effect of quercetin docking with JUN (Figure 8a), AKT1 (Figure 8b), MAPK8 (Figure 8c), IL-6 (Figure 8d), and MAPK1 (Figure 8e). The results show that the results of molecular docking are consistent with the screening results of network pharmacology, and the reliability of network pharmacology is verified by molecular docking.
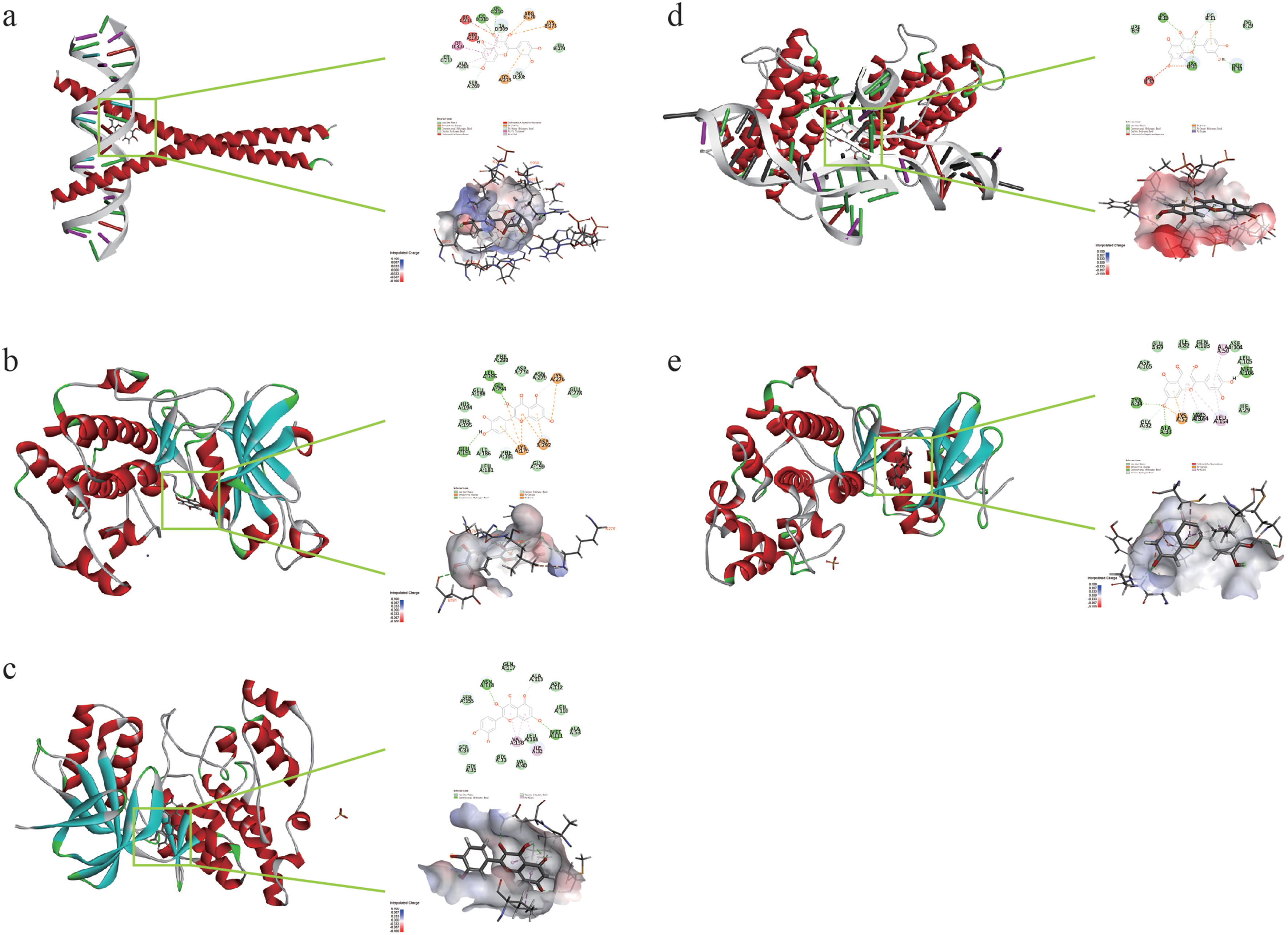
Molecular docking of compounds with core targets. The left side is the 3D docking diagram and the right side is the 2D docking diagram. (a) Docking process of quercetin with JUN. (b) Docking process of quercetin with AKT1. (c) Docking process of quercetin with MAPK8. (d). Docking process of quercetin with IL-6. (e) Docking process of quercetin with MAPK1.
Morphology of Rat Colonic Mucosa
The sliced specimens were observed under the electron microscope. In the control group, the microvilli were arranged neatly and tightly. No inflammatory cells were seen. In the model group, the microvilli were uneven, the gap was widened, and a large No. of inflammatory cells were infiltrated. Compared with the model group, the morphology of rat colonic mucosa in the MPD groups was improved, accompanied by epithelial repair and glandular hyperplasia (Figure 9A). Compared with the control group, the CMDI score of the model group was significantly higher (P < .01). Compared with the model group, the CMDI scores of the H-MPD group and M-MPD group were significantly decreased (P < .01), as shown in Figure 9B.

Morphology of rat colonic mucosa. (A) Observation of rats’ colon tissues in each group ( × 20,000 magnification). (B) The colonic mucosal injury index of rats.
Protein Expression of p-p38MAPK and p-MLC in Colon Tissue of Rats
The protein expression of p-p38MAPK and p-MLC in the model group was significantly higher than that in the control group (P < .01). Compared with the model group, the protein expressions of the H-MPD and M-MPD groups were significantly decreased (P < .01), as shown in Figure 10A and B.

The effect of MPD on the protein expression of p-p38MAPK and p-MLC in colon tissue of DSS-induced UC rats. (A) Protein electrophoresis band. (B) Relative protein expression of p-p38MAPK and p-MLC.
Discussion
Experts in Chinese medicine believe that the pathogenesis of UC is the accumulation of damp and heat. The evil of dampness and heat invades the colon, blood stasis blocks the colon, and then flesh rots into pus. The evil persists for a long time, and the intestinal conduction is abnormal. In “Treatise on Febrile and Miscellaneous Diseases” Zhang Zhongjing wrote that patients with febrile diarrhea (dysentery) accompanied by tenesmus should be treated with PD. 25 Professor Sun Hongwen, a well-known Chinese medicine practitioner in Jiangsu Province, is the inheritor of Wumen Chinese Medicine. She believes that repeated attacks and the inability to cure UC are consistent with the “long illness enters the collaterals” theory, which was created by Ye Tianshi (a representative of Wumen Chinese Medicine). Regarding PD, Professor Sun added traditional Chinese medicine for regulating qi and blood and recommended MPD to treat UC patients. The decoction is composed of Radix Pulsatilla, Rhizoma Coptidis, Phellodendri chinensis cortex, Fraxini cortex, hairyvein agrimony, Panax notoginseng, Rhizoma Bletillae, and Radix Arnebiae. It has the effects of clearing heat and dampness, removing blood stasis, protecting membranes, cooling blood, and stopping dysentery. This study scientifically investigated the pharmacological mechanism of MPD in the treatment of UC through network pharmacology, docking analysis, Western blotting, and in vivo animal study. Network pharmacology was used to screen and predict the active ingredients, core targets, and signal pathways of MPD in the treatment of UC. Then we used molecular docking analysis to simulate the binding of the active ingredients and the core target. The results of network pharmacology were verified by Western blot technology.
Through the ingredient-target-disease network, we can draw several conclusions. First, the treatment of UC by MPD is achieved using multiple ingredients. Second, there are many target genes involved in this process, and the relationship between genes is complicated. This shows that the pathogenesis of UC is complex, and MPD has a multitarget effect on UC which reflects the characteristics and advantages of traditional Chinese medicine. The compounds reported to be most active were quercetin, β-sitosterol, luteolin, kaempferol, and stigmasterol. Quercetin, a bioflavonoid, has anti-inflammatory effects in various diseases. Quercetin, luteolin, and kaempferol are compounds with anti-inflammatory activities and are considered important candidate drugs for IBD treatment. 26 Quercetin can inhibit the intestinal inflammatory response in UC mice, improve the cell structure of colon tissue and maintain the integrity of the intestinal epithelial barrier.27,28 It can also block MAPK and NF-κB signaling pathways to alleviate inflammatory response. 29 Luteolin is a flavonoid that exists in many plants and has anti-inflammatory and anti-allergenic activities. Studies have shown that luteolin can reduce the inflammatory phenotype and oxidative stress of tissue cells by inhibiting the NF-κB pathway. 30 It has also been reported that luteolin's anti-inflammatory mechanism may be related to changing the diversity and composition of intestinal microbiota in UC rats. 31 Kaempferol, a natural flavonoid, is believed to have anti-inflammatory activity and a potential immunomodulatory effect. 32 It is confirmed that kaempferol plays a protective role in colitis mice by regulating intestinal microbiota and toll-like receptor-related signaling pathways. 33 Stigmasterol has been shown to be an immunomodulator with great therapeutic potential. 34 Stigmasterol plays an important anti-inflammatory role and can reduce oxidative stress in a variety of diseases.35,36 Stigmasterol can also reduce the score of colitis, the expression of Cyclooxygenase-2(COX-2), and colony-stimulating factor-1, which indicates that stigmasterol can improve colitis. 37
Through PPI network analysis, 6 core targets (eg, MAPK1, MAPK8, AKT1, VEGFA, JUN, and IL-6) were obtained. Inflammatory factors play an important role in the development of UC. The abnormal activation of the PI3K/AKT signaling pathway in UC has been demonstrated to enhance the expression and secretion of proinflammatory cytokines such as TNF-α, IL-1β, and IL-6. 38 TNF plays a vital role in the typical immune response through the regulation of a No. of pathways. 39 IL-6 is significantly activated in the body's response to injury and plays important role in inflammation.40,41 IL-6 can activate the NF-κB and IL-6/STAT3 pathways to induce intestinal epithelial cell barrier injury and regulate barrier function. 42 It is one of the pathogeneses of intestinal inflammation such as UC and bacillary dysentery. 43 MAPK1 and MAPK8 exist in most cells. They belong to the MAPK signaling pathway and are closely related to the proliferation and apoptosis of intestinal epithelial cells. After inflammatory factors activate the MAPK signaling pathway, the transcription and expression of inflammatory genes related to inflammatory bowel disease (IBD) are changed, which intensifies the development of inflammation. 44 In IBD model mice, the expression of MAPK1/3 increased significantly. However, its expression decreased after drug treatment. 45 AKT1 is a core factor in the PI3K/AKT signaling pathway. AKT1 can regulate cell function by phosphorylation of downstream target proteins. It plays an important role in cell survival and apoptosis. 46 The PI3K/Akt pathway, the MAPK/ERK pathway, and the JNK/c-Jun pathways are responsible for regulating a variety of cellular processes including cell growth, migration, invasion, and apoptosis. The 3 pathways are essential to the progression of chronic inflammation and tumors.47-49 Multiple studies have shown that inflammation is closely related to the occurrence and development of UC.50,51 Colorectal cancer is a recognized complication of UC. Inflammation predisposes one to the development of cancer and promotes all stages of tumorigenesis.52,53 In inflamed tissue, the blocked VEGF signaling pathway exacerbates inflammation in a variety of disease models, including IBD. 54 In addition, a retrospective cohort study showed that the expression of VEGF in the intestinal mucosa of UC patients is increased and is related to the development and recurrence of the patient's disease. Therefore, angiogenesis is not only a passive process driven by inflammation but also a hypothesis of active participants in mucosal lesions of UC. 55 These core targets are related to inflammation, angiogenesis, cell proliferation, apoptosis, and carcinogenesis, and may be potential targets for the treatment of UC.
GO function enrichment analysis shows that the molecular functions involved in these genes are complex and diverse. They are involved in multiple processes such as receptor binding, transcription regulation, and post-translational modification, which indicate that MPD can intervene in the occurrence and development of UC in multiple ways from different levels. Through KEGG enrichment analysis, we found that the signaling pathway of MPD in the treatment of UC is mainly related to cell proliferation, apoptosis, and carcinogenesis. The involved pathways include the IL-17 signaling pathway, the TNF signaling pathway, the AGE-RAGE signaling pathway in diabetic complications, viral infection-related signaling pathways, and some cancer pathways. AGEs are formed as a result of nonenzymatic reaction between the free reducing sugars and proteins, lipids, or nucleic acids. AGEs are predominantly synthesized during chronic hyperglycemic conditions or aging. AGEs interact with their receptor RAGE to induce inflammation and immunosuppression by producing proinflammatory cytokines, reactive oxygen species, and reactive nitrogen intermediates. The accumulation of AGEs and the upregulated expression of RAGE are related to various pathological processes. They lead to the injury of vascular endothelial cells, increase the stiffness and fragility of the tissue, and reduce the ability of cells to rebuild the extracellular matrix so that tissues are more vulnerable to mechanical damage. AGEs may also enhance the inflammatory response of cells by inducing the MAPK pathway. RAGE and s-RAGE may be a useful biomarkers of ligand-RAGE pathway activation and cancer.56-58 KEGG enrichment analysis showed that the MAPK pathway directly or indirectly participates in the AGE-RAGE signaling pathway in diabetic complications. Therefore, this pathway may be an important target for MPD to treat UC. The relationship between the TNF-α pathway and inflammation has been confirmed. TNF-α can activate the TNF signaling pathway by binding to the receptors TNFR1 and TNFR2 and ultimately lead to the occurrence of intestinal mucosal inflammation. 39 TNF-α induces the expression of cell adhesion molecules in vascular endothelial cells. It can also be a stimulating factor for the local release of angiogenic substances, which can stimulate the release of VEGF and promote the proliferation and migration of endothelial cells. 59 Interleukin-17(IL-17) is a pro-inflammatory cytokine that can exert its effects through a variety of cytokines (eg, IL-1, IL-8, IL-6, and macrophage inflammatory proteins) and pathways (eg, NF-κB, MAPK, and JNK/c-jun). IL-17 can simultaneously activate and increase the expression of VEGF and then aggravate inflammation response. 60 The IL-17 signaling pathway is related to intestinal flora and plays a key role in regulating intestinal and autoimmune diseases. 61 In addition, multiple cancer pathways ranked high in the KEGG enrichment analysis. This may be related to the susceptibility to inflammation-cancer transformation of UC. Chronic inflammation is closely related to the occurrence of tumors. In recent years, numerous studies have shown that PD also has antitumor effects. The above results reveal the potential mechanism of MPD in the treatment of UC.
Subsequently, we conducted a molecular-docking simulation of the active ingredient and the core target. The results showed that quercetin, β-sitosterol, luteolin, kaempferol, and stigmasterol all docked well with core targets. These core targets are related to inflammation, angiogenesis, cell proliferation, apoptosis, and carcinogenesis. The above active ingredients are expected to become potential drugs for the treatment of UC and will become the focus of future research.
To validate the prediction results by network pharmacology analysis and molecular docking analysis, we established a UC model induced by DSS. We observed colon inflammation and injury by electron microscopy and conducted Western blot to detect the role of the MAPK signaling pathway in UC. The rats in the model group exhibited more severe injury and colitis, but these abnormalities were ameliorated by MPD administration. We found that the expression levels of p-p38MAPK and p-MLC in the colon tissue of the model group were significantly increased. However, they decreased after being treated with MPD. The results indicated that the p38 MAPK/MLCK signaling pathway was activated abnormally in the model group and that MPD had a protective effect in the treatment of UC. These results partly support our network pharmacology analysis results of MPD.
Conclusions
This study investigated the effective active ingredients and molecular mechanisms of MPD in the treatment of UC from the perspective of network pharmacology. The active ingredients of MPD in UC treatment comprise of 51 compounds. Quercetin, β-sitosterol, luteolin, kaempferol, and stigmasterol are the important active ingredients. There are 141 target genes involved in the treatment of UC by MPD, among which MAPK1, MAPK8, AKT1, VEGFA, JUN, and IL-6 are the key target genes. The signaling pathways of MPD in the treatment of UC primarily include the AGE-RAGE signaling pathway, the IL-17 signaling pathway, the MAPK signaling pathway, and the TNF signaling pathway. In vivo experiment indicated MPD could effectively ameliorate DSS-induced colitis, and the effect of MPD was associated with the MAPK signaling pathway.
This study has some limitations. We can conduct relevant clinical and basic experiments to verify further the accuracy of the results.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X221098850 - Supplemental material for Using Network Pharmacology and Molecular Docking Technology to Explore the Mechanism of Modified Pulsatilla Decoction in the Treatment of Ulcerative Colitis
Supplemental material, sj-docx-1-npx-10.1177_1934578X221098850 for Using Network Pharmacology and Molecular Docking Technology to Explore the Mechanism of Modified Pulsatilla Decoction in the Treatment of Ulcerative Colitis by Tingting Wu, Xin Yang, Bo Xu, Huiping Zhu, Jinwei Guo, Yu Zhou, Guoqiang Liang and Hongwen Sun in Natural Product Communications
Supplemental Material
sj-xlsx-2-npx-10.1177_1934578X221098850 - Supplemental material for Using Network Pharmacology and Molecular Docking Technology to Explore the Mechanism of Modified Pulsatilla Decoction in the Treatment of Ulcerative Colitis
Supplemental material, sj-xlsx-2-npx-10.1177_1934578X221098850 for Using Network Pharmacology and Molecular Docking Technology to Explore the Mechanism of Modified Pulsatilla Decoction in the Treatment of Ulcerative Colitis by Tingting Wu, Xin Yang, Bo Xu, Huiping Zhu, Jinwei Guo, Yu Zhou, Guoqiang Liang and Hongwen Sun in Natural Product Communications
Supplemental Material
sj-xlsx-3-npx-10.1177_1934578X221098850 - Supplemental material for Using Network Pharmacology and Molecular Docking Technology to Explore the Mechanism of Modified Pulsatilla Decoction in the Treatment of Ulcerative Colitis
Supplemental material, sj-xlsx-3-npx-10.1177_1934578X221098850 for Using Network Pharmacology and Molecular Docking Technology to Explore the Mechanism of Modified Pulsatilla Decoction in the Treatment of Ulcerative Colitis by Tingting Wu, Xin Yang, Bo Xu, Huiping Zhu, Jinwei Guo, Yu Zhou, Guoqiang Liang and Hongwen Sun in Natural Product Communications
Supplemental Material
sj-xlsx-4-npx-10.1177_1934578X221098850 - Supplemental material for Using Network Pharmacology and Molecular Docking Technology to Explore the Mechanism of Modified Pulsatilla Decoction in the Treatment of Ulcerative Colitis
Supplemental material, sj-xlsx-4-npx-10.1177_1934578X221098850 for Using Network Pharmacology and Molecular Docking Technology to Explore the Mechanism of Modified Pulsatilla Decoction in the Treatment of Ulcerative Colitis by Tingting Wu, Xin Yang, Bo Xu, Huiping Zhu, Jinwei Guo, Yu Zhou, Guoqiang Liang and Hongwen Sun in Natural Product Communications
Supplemental Material
sj-xlsx-5-npx-10.1177_1934578X221098850 - Supplemental material for Using Network Pharmacology and Molecular Docking Technology to Explore the Mechanism of Modified Pulsatilla Decoction in the Treatment of Ulcerative Colitis
Supplemental material, sj-xlsx-5-npx-10.1177_1934578X221098850 for Using Network Pharmacology and Molecular Docking Technology to Explore the Mechanism of Modified Pulsatilla Decoction in the Treatment of Ulcerative Colitis by Tingting Wu, Xin Yang, Bo Xu, Huiping Zhu, Jinwei Guo, Yu Zhou, Guoqiang Liang and Hongwen Sun in Natural Product Communications
Footnotes
Author Contributions
Huiping Zhu, Jinwei Guo, and Hongwen Sun designed the study. Tingting Wu and Bo Xu wrote the manuscript. Xin Yang and Guoqiang Liang performed the experiments. Yu Zhou analyzed the data. All authors have read and approved the final manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Science Foundation of Jiangsu Province (grant number BK20201179).
Ethical Approval
The study received ethical approval from the Ethics Committee of Suzhou Hospital of TCM (ECMUC2019005AO).
Informed Consent
Not applicable.
Trial Registration
Not applicable, because this article does not contain any clinical trials.
Statement of Animal Rights
We confirm that guidelines on animal rights and treatment have been met and any details of approval obtained are indicated within the text of the submitted manuscript.
Availability of Data and Materials
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
