Abstract
Ulcerative colitis (UC) is a chronic idiopathic inflammatory bowel disorder of the colon that causes continuous mucosal inflammation extending from the rectum to the more proximal colon, to variable extents. 1 UC is characterized by a relapsing and remitting course, and it affects millions of people around the world and is now considered a global disease. 2 If it is not treated in a timely and appropriate manner, UC will transform into colorectal cancer. Treatments for UC include 5-aminosalicylic acid, steroids, and immunosuppressants. 3 However, various adverse reactions occur, drug withdrawal as well as side effects lead to relapse due to unstable efficacy, and the treatment course is difficult for patients to accept. 4 Recently, herbal medicines have been considered as potential treatments for UC based on syndrome differentiation, safety, effectiveness, and low recurrence rates. 5
With the growing popularity and promise of Traditional Chinese Medicine (TCM), it is necessary to uncover active components, pharmacological mechanisms, and potential drug efficacies. Traditional research on the active ingredients of TCM has mainly included separation and identification, after which biological activity screening is employed, which is time-consuming, labor-intensive, and cannot reflect the characteristics of multiple components, multiple targets, and multiple pathways. 12 In order to evaluate the pharmacological effects of TCM comprehensively, network pharmacology is used to discover pharmacologically active substances, explore their mechanisms of action, and elucidate their biological activities. 13 -15 Specifically, network pharmacology focuses on the whole system of interactions among components, targets, and diseases, and has transformed TCM research at the molecular level from “one target, one drug” to “network target, multi-component therapy.” 16 Network pharmacology can also be used as a way to screen for active ingredients of TCM and study their potential mechanisms in disease therapy, which can greatly reduce the workload of subsequent experimental research.
The aim of the present study was to discover potential drug candidates for the treatment of UC from the bioactive compounds isolated and characterized from the roots of ECH. For structural elucidation of the extracted compounds, detailed spectroscopic data analyses were performed, including 1D NMR and MS techniques, and spectroscopic data comparisons with related compounds. To examine further the mechanisms of compounds extracted from ECH on UC, the ingredients using network pharmacology were analyzed to obtain the main functional components, targets, biological processes, and pathways. This work deepens on our understanding of the effective substances and mechanisms of ECH in the treatment of UC and contributes to the development and application of TCM in disease treatment. The detailed procedures of the research protocol are shown in Figure 1.

Flowchart of a network pharmacology-based strategy to investigate the phytochemical and pharmacologic mechanisms of
Materials and Methods
Plant Material
The roots of
Reagents and Materials
The NMR spectroscopic data were obtained using Bruker-600 and Bruker-400 MHz spectrometers (Bruker BioSpin GmbH, Germany). HRESIMS were acquired with a Dionex UltiMate 3000 UHPLC equipped with a Thermo Scientific Q Exactive Focus Mass spectrometer (Thermo Fisher Scientific, USA). Column chromatography (CC) was performed using normal phase silica gel (200, 300 mesh, Qingdao Marine Chemical, Inc.). TLC was carried out on silica gel sheets (Qingdao Marine Chemical, Inc.), and Sephadex LH-20 (GE Healthcare, Uppsala, Sweden). Preparative HPLC was performed on an Agilent 1260 Chromatograph equipped with a pump, photodiode array detector at 274 nm and a ZORBAX SB-C18 ODS column (5 µm, 9.4 × 250 mm Agilent). Visualization: UV at 254 nm.
Extraction and Isolation
The dried and powdered roots of ECH (12 kg) were extracted with 75% ethanol (10 L × 3) at room temperature for 24 hours. The total extraction was evaporated to dryness and the resultant residue (230.0 g) was subjected to silica gel CC using gradient elution with light petroleum-EtOAc and CH2Cl2-MeOH (50:1 to 10:1 and 20:1 to 0:1). Seven fractions (A-G) were obtained according to TLC. Fr. A (4.1 g) was separated by silica gel CC eluting with ether-EtOAc 50:1 to 10:1, giving three pooled fractions (A1-A3) and compound 1 (62.0 mg). Fr. A3 (1.2 g) was purified by silica gel CC eluting with ether-EtOAc (30:1), to afford compounds 2 (10.5 mg) and 3 (7.4 mg). Fr. A1 (0.3 g) was purified by silica gel CC eluting with ether-EtOAc (50:1), to obtain compound 4 (6.5 mg). Fr. B (7.8 g) was fractionated by CC on silica gel eluting with ether-EtOAc 30:1 to obtain compound 5 (41.6 mg), and fractions (B1-B3). Compounds 6 (8.7 mg) and 7 (4.9 mg) were isolated from Fr. B2 by CC on silica gel eluted with ether-EtOAc (20:1), and Fr. C (6.1 g) was separated by silica gel CC eluting with ether-EtOAc (20:1 to 5:1), to give 4 pooled fractions (C1-C4). Fr. C1 (1.4 g) was further purified by CC on silica gel eluting with ether-EtOAc (20:1) to give compounds 8 (12.0 mg) and 9 (7.7 mg). Fr. C3 (1.7 g) was separated by CC on Sephadex LH-20 (CH2Cl2-MeOH, 1:1) and silica gel (ether-EtOAc, 15:1) to obtain compounds 10 (7.5 mg) and 11 (8.1 mg). Fr. D (11.0 g) was purified by silica gel CC eluting with CH2Cl2-MeOH (20:1), to obtain compounds 12 (210.5 mg) and 13 (177.4 mg). Fr. E (46.0 g) was separated by silica gel CC eluting with CH2Cl2-MeOH (20:1 to 5:1), to give compounds 14 (127.0 mg), 15 (98.0 mg) and pooled fractions (E1-E5). Fr. E2 (4.2 g) was fractionated by CC on Sephadex LH-20 (CH2Cl2-MeOH, 1:1) and further separated by HPLC (MeCN-H2O, 45:55) to obtain 16 (7.6 mg, tR: 23.2 minutes) and 17 (5.8 mg, tR: 28.6 minutes). Fr. F (61.0 g) was purified on a C18 reversed-phase column eluting with MeOH-H2O (40:60 to 90:10), to obtain 8 pooled fractions (F1-F8). Fr. F6 (3.3 g) was further separated by semi-preparative HPLC , using MeCN-H2O at 35:65 to obtain compounds 18 (10.0 mg, tR: 15.4 minutes) and 19 (5.6 mg, tR: 17.8 minutes). Fr. F5 was purified by semi-preparative HPLC (MeCN-H2O, 25:75) to obtain compound 20 (8.4 mg, tR: 12.7 minutes). Fr. F4 was purified by semi-preparative HPLC (MeCN-H2O, 22:78) to obtain compound 21 (6.4 mg, tR: 14.5 minutes). All the reagents, drugs and solvents used were analytically pure. The NMR data and spectra of the compounds are provided in the Supporting Information.
Identification of Active Compounds Potential Targets
Importing 12 candidate compounds into the DrugBank database (https:/www.drugbank.ca/) identified targets for the active compounds. The respective target protein gene names were queried via Uniprot database (https:/www.uniprot.org/), and target proteins were standardized. Fifty targets were obtained after the duplications were removed.
UC-Related Targets
In Genecards (http://www.genecards.org/) and OMIM (https://data.omim.org/) in the database for retrieval to “Ulcerative Colitis” as the keyword search and search criteria were set to “Gene” and “Homo sapiens.”
Network Construction
Venny2.1 was used to map the potential drug target proteins and UC gene, and the target of ECH in treating UC was screened out. The ECH potential therapeutic target obtained was imported into the STRING database (https://string-db.org/) and selected as “Homo sapiens” to create a protein–protein interaction (PPI) network. Cytoscape 3.8.0 software was used to analyze the data and map the drug-active component–disease–target interaction network.
Bioinformatic Analysis
To study further the synergistic effect of the potential target of ECH for the treatment of UC, the clusterProfiler toolkit of R was used to convert gene symbols into gene ID, and gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) metabolic pathway enrichment analysis of the intersection targets was conducted respectively. Visualization was achieved using the ggplot2.R package. The GO and KEGG pathway items with
Molecular Docking
A start was made by downloading the active ingredient molecular structures at ZINC database (http://zinc.docking.org/). The mol2 file of the active components was entered into structural optimization software SYBYL (Version 2.0). The Tripos Force field was used to select the Gasteiger-Huckel load type and the Powell iteration algorithm was used to set the maximum repeat rate to 1000 steps, and the Gradient was 0.005. Color option was Force. The ligand (mol2 format) was converted by importing Auto Dock Tool software into PDBQT format. The target gene protein conformation was then retrieved in the PDB database (http:/www.rcsb.org/), and the following screening conditions were applied: the protein structure obtained by X-ray crystal diffraction; protein crystal resolution was less than 3 Å; and proteins with well-defined typing. To remove the water molecule and its original ligand, the receptor protein (PDB format) was imported into Pymol 2.3.4 software (Warren Lyford DeLano), and then entered into Auto Dock Tool software to pretreat the protein with polar hydrogen. The docking pocket of the receptor protein was determined based on the location of its original ligand. Finally, we used Auto Dock Vina software for molecular docking, setting up the energy range for 10, exhaustiveness at 20, number modes at 100, and scoring the affinity of each docking.
Results
Identification of Compounds in Eomecon chionantha Hance
Twenty-one alkaloids were extracted, isolated and purified from the roots of ECH; 13 compounds (

Chemical structures of compounds 1
Screening of Effective Anti-ulcerative Colitis Ingredients in Eomecon chionantha Hance
All alkaloids identified in ECH were searched from the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP, http://tcmspw.com), which provides the pharmacokinetic properties of natural compounds such as oral bioavailability (OB), drug-likeness (DL), and blood brain barrier (BBB). Early assessment of the ADME properties of candidate active ingredients has become an indispensable process in modern drug discovery. Thus, a variety of researchers have recommended that molecules with OB ≥30%, DL ≥0.18 and BBB ≥ −0.3 will be considered to have better pharmacological effects. 28 In this research, all alkaloids that met the screening criteria were considered as candidate ingredients. Twelve components with effective pharmacological activities were identified after screening ADME parameters (Table 1).
Ingredients of ECH.

Compound-Target Network Analysis
We finally selected 12 active compounds of ECH. These identified 4356 UC-related targets from the Genecards and OMIM databases. The drug-ingredients-targets-disease (D–I–T–D) network was constructed as shown in Figure 3. The network contained 64 nodes (12 compounds in ECH and 23 intersection targets) and 243 edges which indicated the D–I–T–D interactions. According to the value of degree centrality (DC), the important compounds included C
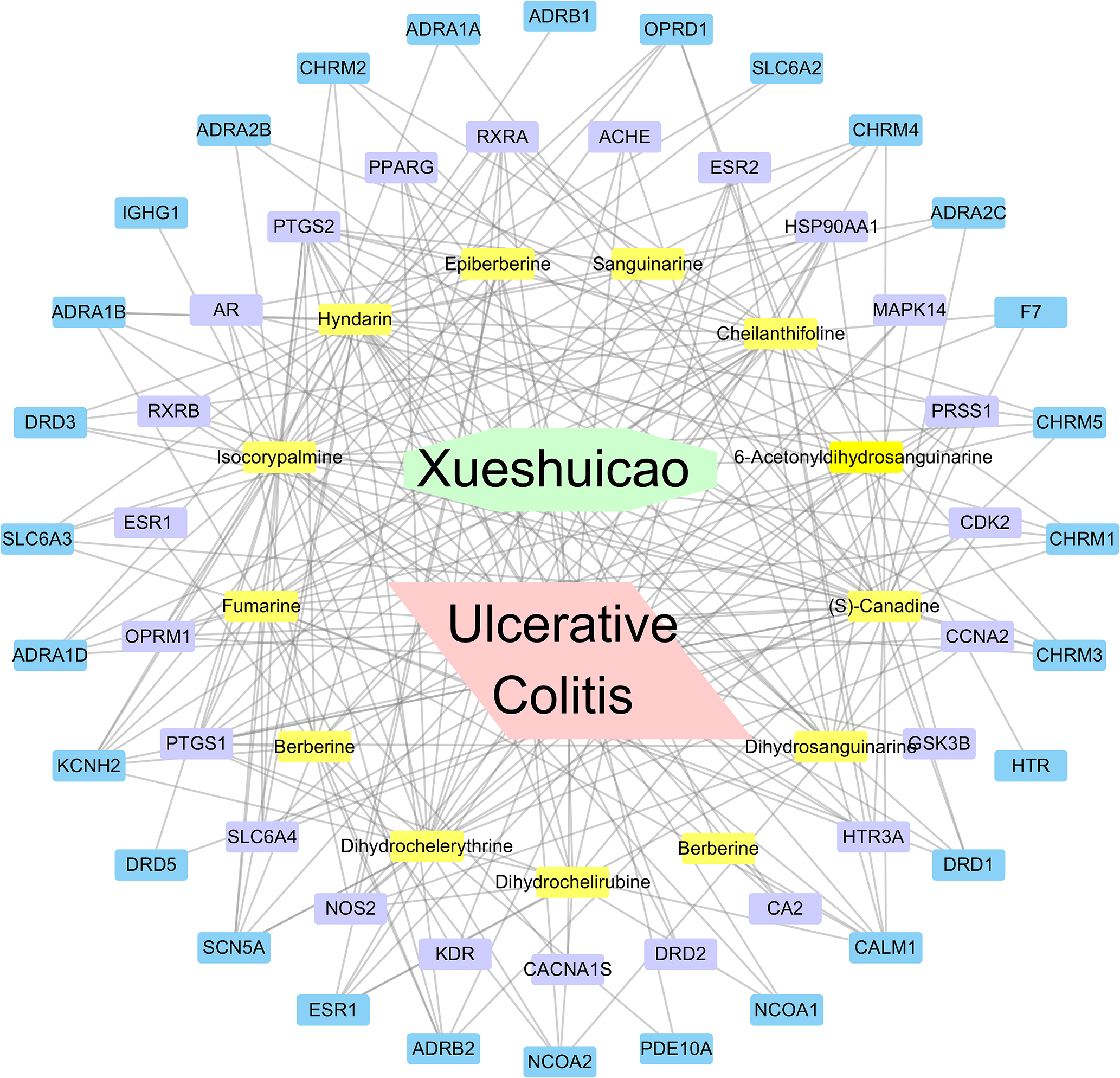
The drug-ingredients-targets-disease network of
Protein–Pprotein interaction Networks Construction
After the intersection of drug targets and genes of UC, 23 common targets were obtained. At the same time, the screened disease-drug common targets were entered into the STRING platform to create the PPI network model, as shown in Figure 4.

Protein–protein interaction (PPI) network analysis.
Gene Ontology and Kyoto Encyclopedia of Genes and Genomes Pathway Enrichment Analysis
In the GO functional enrichment analysis of 23 intersection targets, 401 results were obtained, mainly involving biological processes (BP) (345 items), cellular components (CC) (23 items) and molecular functions (MF) (41 items). The analysis condition was set as

Twenty main terms of geneontology (GO) enrichment.
The KEGG pathway results (Table 2) showed that 22 significantly enriched pathways were related to the mechanism of ECH in treating UC, mainly involving small cell lung cancer, serotonergic synapse, IL-17 signaling pathway, Th17 cell differentiation, estrogen signaling pathway, transcriptional misregulation in cancer, and PI3K-Akt signaling pathway; these are presented in Figure 6, in which the area of color represents the number of genes, and the color represents the adjusted

The 20 main pathways enriched by major hubs. Area of the color represented the number of genes and color represented of
KEGC Pathway Results of 23 Intersection Targets.

Gene-Pathway Network Analysis
The prominently enriched pathways and genes that regulate these pathways were used to create the gene-pathway network, as shown in Figure 7. The topological analysis of 19 pathways and 15 genes was carried out with DC. The V-shapes represent pathways and the rectangles represent target genes in Figure 7, and it can be seen from the network diagram that MAPK14 had the maximum DC, which is the most core target gene. The DC of several other genes is also large, such as RXRA, GSK3B, CDK2, RXRB, HSP90AA1, PTGS2, and ESR1, which might be key target genes for ECH against UC.

Gene-Pathway network of
Molecular Docking
The key active components of ECH selected above were verified through molecular docking with their potential targets associated with UC, and the results are shown in Table 3. It can be seen that the key active components are well docked with the genes MAPK14, RXRA, GSK3B, CDK2, HSP90AA1, PTGS2, and ESR1. Some of the molecular docking simulation details are shown in Figure 8. The specifics show that the small molecules bind to the protein’s active pocket and match well with the active pocket by hydrogen bonding. Figure 8 shows that isocorypalmine interacts with CDK2’s active sites HIS82 and HIS84 to form three hoursydrogen bonds, which are the main force that promotes its binding to the active site.
Affinity of Active Compound With Key Targets (kcal/mol).

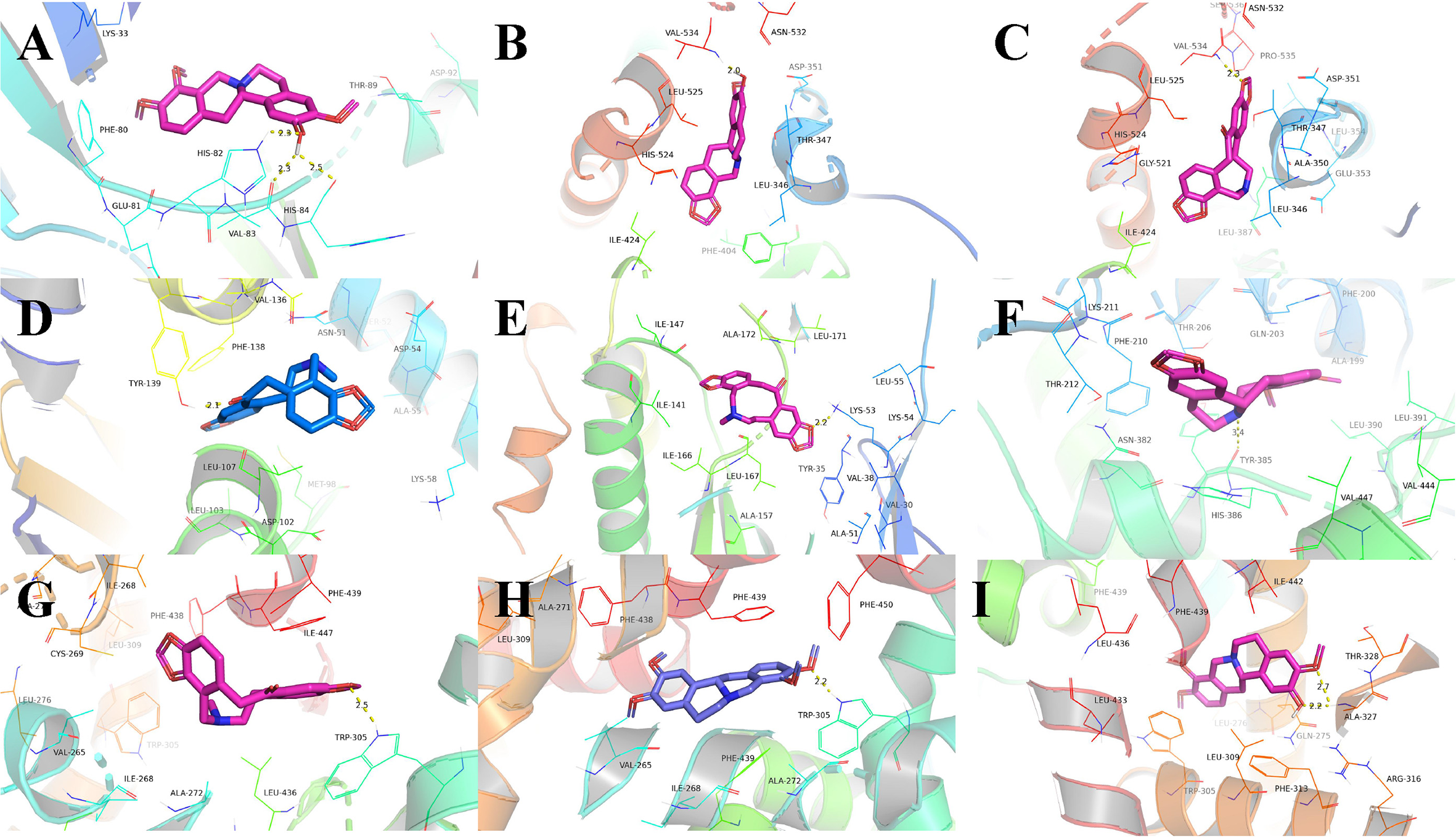
Binding studies of selected ingredients–targets interactions. (A) C20 with CDK2; (B) C21 with ESR1; (C) C15 with ESR1; (D) C15 with HSP90AA1; (E) C15 with MAPK14; (F) C19 with PTGS2; (G) C15 with RXRA; (H) C18 with RXRA; (I) C20 with RXRA. Molecules are represented by a ball-and-stick model, the hydrogen bonds are represented by a dotted line, and the distance is in angstroms. Atoms C, O, and N are purple, red, and blue, respectively.
Discussion
In the present study, 21 alkaloids were extracted, isolated, and purified from ECH roots, among which 12 active ingredients, including dihydrosanguinarine, dihydrochelirubine, and isocorypalmine, have different pharmacological activities. D–I–T–D network analysis of ECH for UC was conducted using the 12 compounds and 50 compound targets. The pivotal active compounds with pleiotropic effects of ECH are likely to be C
PPI network analysis of the intersection targets and GO and KEGG enrichment analysis were conducted to explain the possible molecular mechanisms of ECH in the treatment of UC. GO enrichment results showed that ECH mainly regulated biological processes in response to alkaloids, response to xenobiotic stimulus, transcription initiation from RNA polymerase II promoter, regulation of neurotransmitter levels, “regulation of synaptic transmission, dopaminergic,” response to steroid hormones, and steroid hormone-mediated signaling pathway. This result was similar to the findings of previous pathological studies of UC, indicating that ECH can intervene in the occurrence and development of UC at different levels and in multiple ways.
29
For example, previous research showed that UC is caused by a reactive exogenous metabolite that binds before excretion into bile.
29
Xenobiotics may reduce disease-related gut mucosal alterations and clinical symptoms. Magro et al. pointed out that significant reductions in tissue levels of dopamine, norepinephrine, and 5-hydroxytryptamine are associated with symptoms in UC,
30
and Ushijima et al. reported that the activity of RNA polymerase II in intestinal mucosal epithelial cells is connected with
Estrogen exerts its physiological effects by binding to estrogen receptors (ESRs), which include the classical nuclear ESRs and the membrane ESRs. Nuclear receptors including ESR alpha (ESR1) and beta (ESR2) are located in the nuclei of cells and react with ligands by regulating the transcription of downstream target genes. 32 Since UC increases the risk of colorectal carcinoma development, Khalili et al. suggested that estrogen-related pathways might mediate the pathogenesis of UC. 33 An increasing number of reports have highlighted the relationship between estrogen signaling and inflammation, suggesting that ESR1 and ESR2 are vital mediators of intestinal inflammation, according to epidemiological and experimental data. 34 A number of scholars have proposed that the ESR1/ESR2 signaling ratio affects colitis in males and females. Principi et al. claimed that the alpha/beta balance of the ESRs appears to have a direct influence on colorectal carcinogenesis, and the resulting apoptosis indicates that ESR2 has an anticarcinogenic effect. 35 In addition, Principi et al. reviewed the ESR anti-inflammatory and anti-tumor effects in the colon, showing the potential to prevent and/or treat UC at the translational level. Jacenik et al. described dysregulation of ESRs in the intestinal mucosa of patients with Crohn’s disease and UC, indicating that estrogen signaling may play a role in the local immune response and maintain epithelial homeostasis in a gender- and age-dependent manner. In the classical mechanisms of action, estrogen-bound ESRs are located in the cytoplasm and undergo conformational changes to dissociate from scaffolding heat shock protein (Hsp90 and Hsp70) dimers and translocate to the nucleus, where they can bind to specific DNA sequences in the promoter regions of target genes to modulate their expression levels.
A growing body of evidence also suggests that endoplasmic reticulum stress (ERS) plays a key role in the pathogenesis of UC. Originating from endoplasmic reticulum lumen misfolding or abnormal protein folding, ERS induces the protein folding reaction (unfolded protein response, UPR) to resolve the stress, recover the endoplasmic reticulum, promote cell survival, and adapt to the signaling network. UPR defects in animal models can induce colitis susceptibility. In contrast, over activation of the UPR can promote inflammation and induce apoptosis associated with intestinal inflammation, as well as the destruction of the UC bowel epithelial barrier. 36 The estrogen signaling pathway can help protect the intestinal epithelial barrier of UC patients by inhibiting ERS and inflammatory factor generation.
Phosphoinositide 3-kinase (PI3K) is widely involved in biological activities such as cell growth, metabolism, and proliferation. 37 As a downstream effector of PI3K, Akt can be activated by factors such as PI3K, inflammation, and DNA damage. Studies have shown that the PI3K/Akt signaling pathway is closely involved in the regulation and release of pro-inflammatory factors such as TNF-α, and that this pathway is one of the most important in the pathogenesis of UC. 38 Intervention in the PI3K/Akt signaling pathway is thus a relevant mechanism for effective treatments of UC by many drugs, such as oxymatrine and mesalazine. 39
Intestinal mucosal barrier defects are an important pathological feature of UC. In the occurrence of UC, a large number of pro-inflammatory factors are generated that form negative feedback regulation after repeated activation of inflammatory cells, resulting in continuous intestinal inflammation and sustained aggravation of the intestinal mucosal injury.
Prostaglandin G/H synthase 2 (PTSG2), also known as cyclooxygenase-2 (COX-2), is a key enzyme in the biosynthesis of prostaglandins and is thought to be involved in the development of UC. 44 After mechanical stimulation of the intestines (expansion/stretching) in mice, the mRNA levels of COX-2 in the smooth muscle of the colon increase, but inhibition of COX-2 can reduce the anaphylaxis of the gut after expansion. 45 Prostaglandin, a product of PTSG2, has positive effects on inflammation, pain, and fever. Clinical and preclinical studies have shown that celecoxib and rofecoxib, PTGS2 inhibitors, are effective in treating UC in both patients and animals. 46
Mitogen-activated protein kinase 14 (MAPK14), also named p38α, is the alpha subtype of p38, which coordinates the cellular responses to stress as well as inflammation and aids in the production of inflammatory mediators, in particular TNF-α and COX-2, and is closely correlated with UC. 47 The genes for p38α are localized in the IBD-susceptible region of chromosome 6. Waetzig et al. found that p38α is activated significantly in the inflamed colonic mucosa of IBD patients, with p38α exhibiting the strongest activation in UC and Crohn’s disease. 48 Furthermore, they incubated intestinal mucosal biopsy tissue culture medium of UC patients with the MAPK14 inhibitor SB203580, and found that TNF-α decreased significantly, which was positively correlated with the concentration of SB203580. More importantly, Grossi et al. suggested that the pain burden in newly diagnosed IBD patients is connected with the expression levels of PTGS2 and MAPK14, indicating potential therapeutic targets for managing pain in IBD. 49 Consequently, MAPK14 may play a major regulatory role in a variety of inflammatory mediators in diseases such as UC. MAPK inhibitors are potentially powerful drugs for UC, as they can treat not only inflammation but also the pain caused by inflammation.
Retinoic acid receptors are ligand active transcription factors that consist of two receptor families, the retinol X receptor family (RXR) and the retinoic acid receptor family (RAR), each consisting of three subtypes (α, β, γ). 50 Retinoic acid signaling pathways regulate a variety of biological functions, such as cell differentiation, proliferation, apoptosis, and inflammatory responses. 51 RXR-A is closely related to bowel immunity and inflammation, and in the intestinal mucosa of vitamin-A deficient rats, the expression levels of inflammatory cytokines increase while the expression levels of RARA mRNA decrease. 52 Macrophages have vital functions in damaged intestines in UC patients and mice colitis models by secreting various inflammatory factors and bioactive substances. It has been shown that downregulation of RXR-A increases the activities of nuclear factor kappa-B (NF-κB) in macrophages, which mediates COX2 expression and PGE2 production. 53 In azoxymethane/dextran sulfate sodium-induced RXRA +/-mice, the expression of RXRA in the intestinal tissues of RXRA +/-mice decreases compared with wild-type mice, accompanied by upregulation of two genes, Snail1 and Snail2, which are transcription factors downstream of inflammatory mediators. 54 Additionally, although UC presents a high risk of developing colorectal cancer, RXRA control can be beneficial in accelerating colitis-related colon cancer by promoting IL-6/STAT3 signaling. 55
Glycogen synthase kinase-3 beta (GSK3B), originally identified as a regulator of glycogen metabolism, is a serine/threonine kinase in mammalian eukaryotic cells that acts on numerous signaling proteins and transcription factors such as c-Jun, AP-1, β-catenin, CREB, and NF-κB to regulate cell differentiation, proliferation, survival, apoptosis, and inflammation. 56 Relevant studies have shown that GSK3B can regulate inflammation in a variety of diseases, including type 1 auto-immune diabetes, experimental colitis, and asthma. 57 -59 Consequently, GSK3B has become a novel inflammatory target. Numerous studies have investigated the positive effects of GSK3B in the regulation of the inflammatory process via controlling NF-κB activity. NF-κB is a pro-inflammatory transcription factor involved in the pathogenesis of UC and is one of the most important regulators of pro-inflammatory cytokine gene transcription, including IL1α, IL2, IL2R, IFN-β, and TNF-α. 60 GSK3B inhibition has been shown to attenuate NF-κB activation by triggering epigenetic chromatin remodeling, stabilizing p105, and activating the Wnt/β-catenin signaling pathway, and has been shown to suppress the expression of NF-κB-mediated genes NOS, BCL-2, X-IAP (X-linked inhibitor of apoptosis protein), cyclin D1, COX-2, MMP-9, survivin, BCL-XL, BFL-1/A1, TRAF1 (TNF-receptor-associated factor-1), and IL-6. Furthermore, the inhibition of GSK3B has been shown to attenuate NF-κB function by activating the production of anti-inflammatory IL-10 cytokines and by triggering intracellular accumulation of CREB, which co-competes for the NF-κB transcriptional co-activator CREB-binding protein.
In this research, we also used virtual molecular docking to explore the binding energy of the key active components with the genes MAPK14, RXRA, GSK3B, CDK2, HSP90AA1, PTGS2, and ESR1 based on the above-mentioned screening content via network pharmacology. As shown in Table 3, the majority of the active components have a high affinity to the corresponding protein (binding energy is less than −7.0), and the primary amino acids function mainly by hydrogen bonding. Hydrogen bonding minimizes the energy of the small molecule-receptor complex and thus increases stability (Figure 8). The binding energy value is relatively small, indicating that the compound is able to bind to the receptor protein stably. The molecular docking data are thus consistent with the results of network pharmacology screening in this work, which further verifies the reliability of the network pharmacology prediction.
Conclusions
In conclusion, through a multi-pathway and multi-target model, various active components of ECH were found to play a likely biological role in the prevention and control of UC. According to the DC values of the nodes in the network, the pivotal active compounds with multiple effects in ECH were likely to be isocorypalmine, hyndarine, (s)-canadine, cheilanthifoline, fumarine, dihydrochelerythrine, and dihydrosanguinarine. On the basis of relevant literature and network pharmacological analysis, our work suggested that regulating inflammatory pathways, interfering with the occurrence and development of inflammation, regulating cell apoptosis, and inhibiting cell proliferation are the main mechanisms by which ECH treats UC. Target genes such as MAPK14, GSK3B, and HSP90AA1 can be used to study the molecular mechanisms of the prevention and treatment of UC and UC-related cancers clinically. This study addressed ECH’s mechanism of action against UC, presented new approaches and ideas for elucidating ECH’s mechanism of action, and also provided a theoretical reference for its application. Nevertheless, additional experiments are needed to verify the active components and the mechanism of action of the targets.
Supplemental Material
Supplementary Material 1 - Supplemental material for A Combined Phytochemical and Network Pharmacology Approach to Reveal the Effective Substances and Mechanism of Eomecon chionantha Hance for the Treatment of Ulcerative Colitis
Supplemental material, Supplementary Material 1, for A Combined Phytochemical and Network Pharmacology Approach to Reveal the Effective Substances and Mechanism of
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Scientific Research Fund of Hunan Provincial Education Department (19B402) and the innovation teams of laboratory medicine, School of Nursing, Hunan University of Medicine (2019-48).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
