Abstract
Introduction
Stroke is a common disease with high incidence, high mortality, and disability, which imposes a significant burden on individuals, their families, and the society. Stroke is ranked in the second place among all the disabling diseases 1 . Cerebral ischemic stroke (CIS) is the most common type of stroke, accounting for 80% of all cases 2 . A survey of incidence on more than 480 000 people has found that the current stroke incidence (247 of 100,000 person-years) and mortality rates (115 of 100,000 person-years) in China appear to be the highest in the world, and found that the stroke incidence has continued increasing in the past 30 years 3 . Furthermore, the mortality rate from stroke has been increasing steeply in China, rising faster than the other countries over the past 3 decades. In addition, the prevalence and incidence of stroke in China have risen faster than the other countries 4 . In 2004, the average fee for stroke admission was 6356 RMB ($1000.94), which was 2 times the annual income of rural residents 5 . Therefore, safer and more effective drugs for the treatment of CIS are urgently needed.
Thrombolytic therapy is an effective therapeutic approach for CIS, but its application in clinical practice is restricted by some factors, such as the strict contraindications to thrombolysis, narrow therapeutic time window, and serious side effects 6 . Numerous experimental and clinical observations have clearly suggested that traditional Chinese medicine (TCM) has unique advantages in the treatment of CIS. Various traditional Chinese prescriptions, single traditional Chinese medicine, and bioactive compounds of TCM have all demonstrated the neuroprotective effects against CIS7-11.
The Chuanhong Stroke Capsule (CHSC) prescription is a traditional Chinese herbal formulation created by Professor Ji-xue Ren, who is a national famous Chinese physician. CHSC is currently used as an in-hospital preparation at First Affiliated Hospital of Changchun University of Traditional Chinese Medicine for more than several decades. Dr Ren believes that Qi stagnation and blood stagnation is the basic pathogenesis of CIS, and the therapeutic principle is blood-activating and stasis-removing. Clinical observations have found that CHSC could significantly improve capacity of patients to undertake daily activities, enhance the quality of life, and lessen the rate of disability or mortality 12 . CHSC consists of the following 9 herbs: Radix Paeoniae Rubra (Chishao), Ligusticum chuanxiong Hort (Chuanxiong), Typhae pollen (Puhuang), Pueraria Radix (Gegen), Flos Sophorae (Huaihua), Panax notoginseng (Sanqi), Carthamus tinctorius (Honghua), Geosaurus (Dilong), and Herba Siegesbeckiae (Xixiancao). Each ingredient of CHSC has its own specific functions. For example, Adix Paeoniae Rubra, Ligusticum chuanxiong Hort, Pueraria Radix, P notoginseng, and C tinctorius have already shown the function of anti-inflammation and anti-apoptosis 13-18; Typhae pollen has demonstrated the ability to attenuate cell injury through various functions against neuronal apoptosis and oxidative stress 19 ; Geosaurus could repress inflammatory responses, inhibit apoptosis, and ameliorate ischemic brain injury 20 ; Siegesbeckiae can reduce brain injury through reducing platelet adhesion and anti-oxidative stress 21 . Due to its complex components, it is difficult to conduct studies on the curative mechanisms of CHSC in previous studies.
At present, network pharmacology or system biology provides new methods for the understanding of pharmacological mechanisms of complex Chinese medicine. Milla Kibble et al argued that a network pharmacology approach would naturally lend itself as an unbiased strategy for uncovering the overall mode of action of multi-targeted natural products and their mixtures, and it enables one not only to explore the therapeutic potential of natural products but also to better understand their potential adverse effects prior to the actual clinical trials, thereby de-risking and speeding up the drug development process 22 . As one of the core technologies in virtual screening (VS), molecular docking has been extensively used in network pharmacology. Computational docking can be used to predict bound conformations and free energies of binding for small-molecule ligands to macromolecular targets. Docking is widely used for the study of biomolecular interactions and mechanisms, and is applied to structure-based drug design. The methods are fast enough to allow VS of ligand libraries containing tens of thousands of compounds 23 . Molecular docking helps to comprehensively characterize the underlying mechanism of a drug candidate and to interpret its cascading effects, improving the prediction of drug efficacy and safety 24 . Thus, network pharmacology is a feasible method to investigate TCM-related issues with a holistic perspective.
In this study, a network pharmacology study of CHSC was established through molecular docking and network analysis. The detailed workflow of the study was shown in Figure 1.First, we obtained the active components of CHSC and potential targets against CIS. Then, the protein–protein interaction (PPI) of potential targets against CIS was constructed. Next, a systematic analysis of potential targets and biofunctional modules was conducted. Meanwhile, the interactions between the components of CHSC and key targets were confirmed using molecular docking. Collectively, we aimed to explore the potential mechanism of action of CHSC and provide a theoretical basis for further pharmacological research and clinical application of this prescription.

The workflow of this study.
Materials and Methods
Screen of Active Ingredients of CHSC
The Traditional Chinese Medicine System Pharmacological Database and Analysis Platform (TCMSP, http://lsp.nwu.edu.cn/tcmsp.php) was used to collect the active ingredients of CHSC. These main active ingredients were screened according to the absorption, distribution, metabolism, and excretion (ADME) rules reported in the literature (oral bioavailability [OB] ≥30%, drug likeness [DL] ≥0.18) 25 . Since Geosaurus was not in the TCMSP database, the chemical components of Geosaurus were searched using the following bibliographic databases: China National Knowledge Infrastructure (CNKI) Database (https://www.cnki.net/), and PubMed database (https://pubmed.ncbi.nlm.nih.gov/). Then, the chemical composition of Geosaurus was analyzed with the SwissADME database (http://www.swissadme.ch/) which was used to calculate the molecular properties of the collected compounds by importing the simplified molecular-input line-entry specification (SMILES) information. Based on the yielded properties, the compounds were selected using standard screening criteria which involve gastrointestinal absorption, and DL rules 26 .
The TCMSP and Swiss Target Prediction (http://www.swisstargetprediction.ch) were used to identify the potential target genes of the active compounds in CHSC. We queried each target protein name (protein ID) through the protein database (UniProt), selected the species as “Homo sapiens”, obtained the target gene name (gene ID) corresponding to each target protein. Then the resulting data is the predicted target of the active ingredients.
Prediction of Potential Targets of CIS
Microarray data of differentially expressed mRNAs in CIS compared with the normal group were obtained from the GEO database (https://www.ncbi.nlm.nih.gov/geo/). Series: GSE158312 Sva and Limma of Rmur3.6.1 were used to carry out the joint analysis of multiple chips and correct data batches. The 2 software packages can be obtained from (https://www.bioconductor.org/). The genes with an adjusted P <0 .05 and log2 (fold change) >1 or log2 (fold change) <−1 were considered as significantly differentially expressed genes and CIS-related targets. In addition, CIS-related targets were integrated in the database, including the Genecards (https://www.drugbank.ca/), and OMIM databases (https://www.disgenet.org/web/DisGeNET/menu/home), using “CIS” as the keyword.
Construction of the PPI Interaction Network
String11.0 (https://string-db.org/) is a database for storing known and predicted protein interactions, including direct and indirect protein interactions, and it provides scores for each protein interaction. The selected cross targets were introduced into String database for protein interaction analysis, and a protein interaction network with species limited to “Homo sapiens” and confidence>0.9 was obtained. Protein interaction data was further imported into Cytoscape 3.7.2 (https://cytoscape.org/) to build the PPI network.
GO and Pathway Enrichment Analysis
The clusterProfiler package installed in R offers the group gene ontology (GO) and KEGG methods to classify genes. The two software packages can be obtained from Bioconductor (https://www.bioconductor.org/) acquisition. Also, GO and KEGG enrichment analyzes the potential biological pathways and functions associated with the target. Pathways with the adjusted P less than 0.05 (P < 0.05) were considered as significant.
Construction of the Herb-Target-Pathway Network
All the networks can be created by the Cytoscape3.7.2 software. Cytoscape supplies a basic set of features for data integration, analysis, and visualization of complicated networks. In the network diagram, nodes and edges are the two important elements of a network, “node” represents the active component and target in CHSC, and “edge” represents the relationship between the active component and target 27 . The “degree” is an important parameter for the network pharmacology approach, which was used to evaluate the potential targets.
Active Components-Targets Docking
Eleven components were selected among the core components of CHSC and docked with 5 proteins selected from the core targets to verify the accuracy of the main components and predicted targets. The 2D structure of the candidate composition and the target were downloaded from the PubChem database and RCSB protein data (https://www.pdb.org/), respectively. The latter preferably selects a model with ligand binding smaller than 3A, and then imports the 2D structure into the Pymol 1.7.2.1 Software (https://pymol.org/2/) for dehydration, hydrogenation, and separation of ligands, then imports into AutoDockTools 1.5.6 to construct the docking grid box for each target. Docking was completed by Autodock Vina 1.1.2 software, and the molecules with the lowest binding energy in the docking conformation were selected to observe the binding effect by matching with the original ligands and intermolecular interactions (such as hydrophobicity, hydrogen bonding).
Results
Identification of Active Compounds of CHSC
Through the retrieval of TCMSP and SwissADME databases, the active chemical components of CHSC were screened. After deleting the duplicate data, a total of 81 active chemical components were obtained, including 13 from Adix Paeoniae Rubra, 6 from Ligusticum chuanxiong Hort, 4 from Pueraria Radix, 17 from C tinctorius, 6 from Flos Sophorae, 6 from Typhae pollen, 7 from P notoginseng, 9 from Herba Siegesbeckiae, and 13 from Geosaurus. The details of candidate compounds were described in Table 1. Radix Paeoniae Rubra, Ligusticum chuanxiong Hort, Typhae pollen, Pueraria Radix, Flos Sophorae, P notoginseng, C tinctorius, Geosaurus, and Herba Siegesbeckiae flos were simplified as CS, CX, PH, GG, HJ, SQ, HH, DL, and XXC, respectively, in the following paragraphs.
Active Compounds of CHSC and Their Parameters.
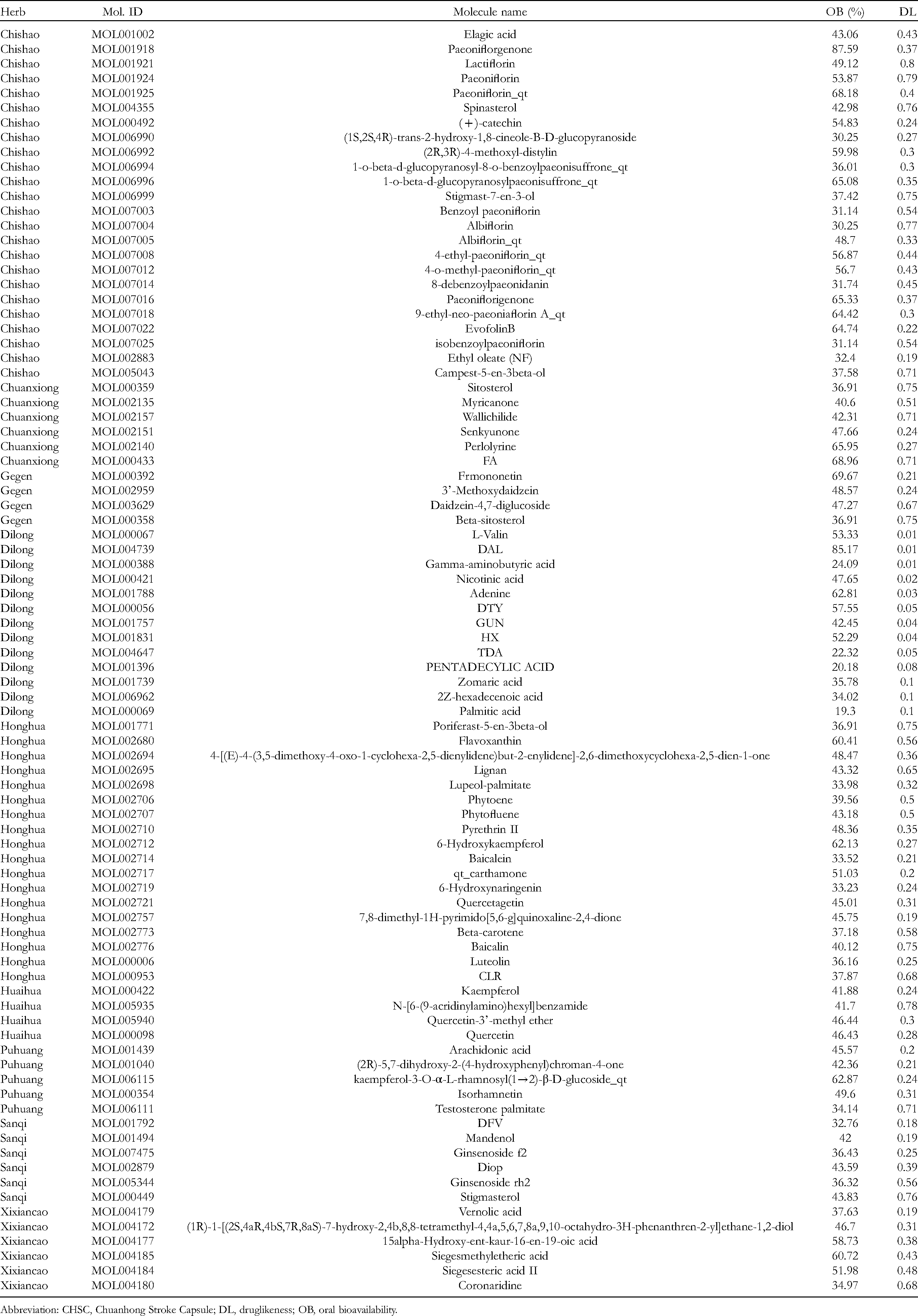
Abbreviation: CHSC, Chuanhong Stroke Capsule; DL, druglikeness; OB, oral bioavailability.
Targets of CHSC and CIS
Joint analysis of a gene chip in the GEO database (GSE158312) has identified 414 differentially expressed genes related to CIS, which were used to build a volcano map (Figure 2). A total of 293 active ingredient targets in CHSC were obtained by TCMSP and Swiss Target Prediction databases. A total of 3053 CIS-related targets were obtained from Genecards, OMIM disease databases, and GEO chip analysis database. Finally, 209 target genes were obtained for both CHSC and CIS through VENNY2.1 (https://bioinfogp.cnb.csic.es/tools/venny/), as shown in Figure 3.

Volcano map of the differential genes jointly analyzed by GEO chips from normal group and CIS group.

Intersection targets of CHSC and CIS. The blue zone indicates targets for active components of CIS; the yellow zone indicates targets for CHSC; and the overlap zone indicates the intersection targets of both CHSC and CIS.
CHSC-CIS PPI Network
The PPI network diagram was constructed using the CytoNCA plug-in in Cytoscape, as shown in Figure 4. According to the topological characteristics of the network, the four most important parameters were selected to screen the core targets of CHSC: degree centrality (DC), closeness centrality (CC), eigenvector centrality (EC), and betweenness centrality (BC). Using the CytoNCA plug-in, we set the DC, BC, EC, and CC should be greater than the median value, screened the protein interaction with high enrichment, and built PPI core network diagram. The nodes closing to the center and in dark color represent that they play a key role in the whole network. The results showed that RELA, JUN, AR, NR3C1, CTNNB1, FOS, TP53, RB1, MAPK1, MYC, HIF1A, Mapk14, AKT1, CDKN1A, HSP90AA1, and ESR1 ranked high. This indicated that these proteins play key roles in the whole network, and the targets corresponding to these protein genes play an important role in the treatment of CIS by CHSC, suggesting that these proteins may be the key targets of CHSC in the treatment of CIS. Table 2 shows the specific information of the 16 core targets. At the same time, we also constructed the network diagram of core targets and noncore targets (Figure 5).

Topological analysis of the protein–protein interaction (PPI) network. According to the topological analysis, the rightmost represents the target of the core.
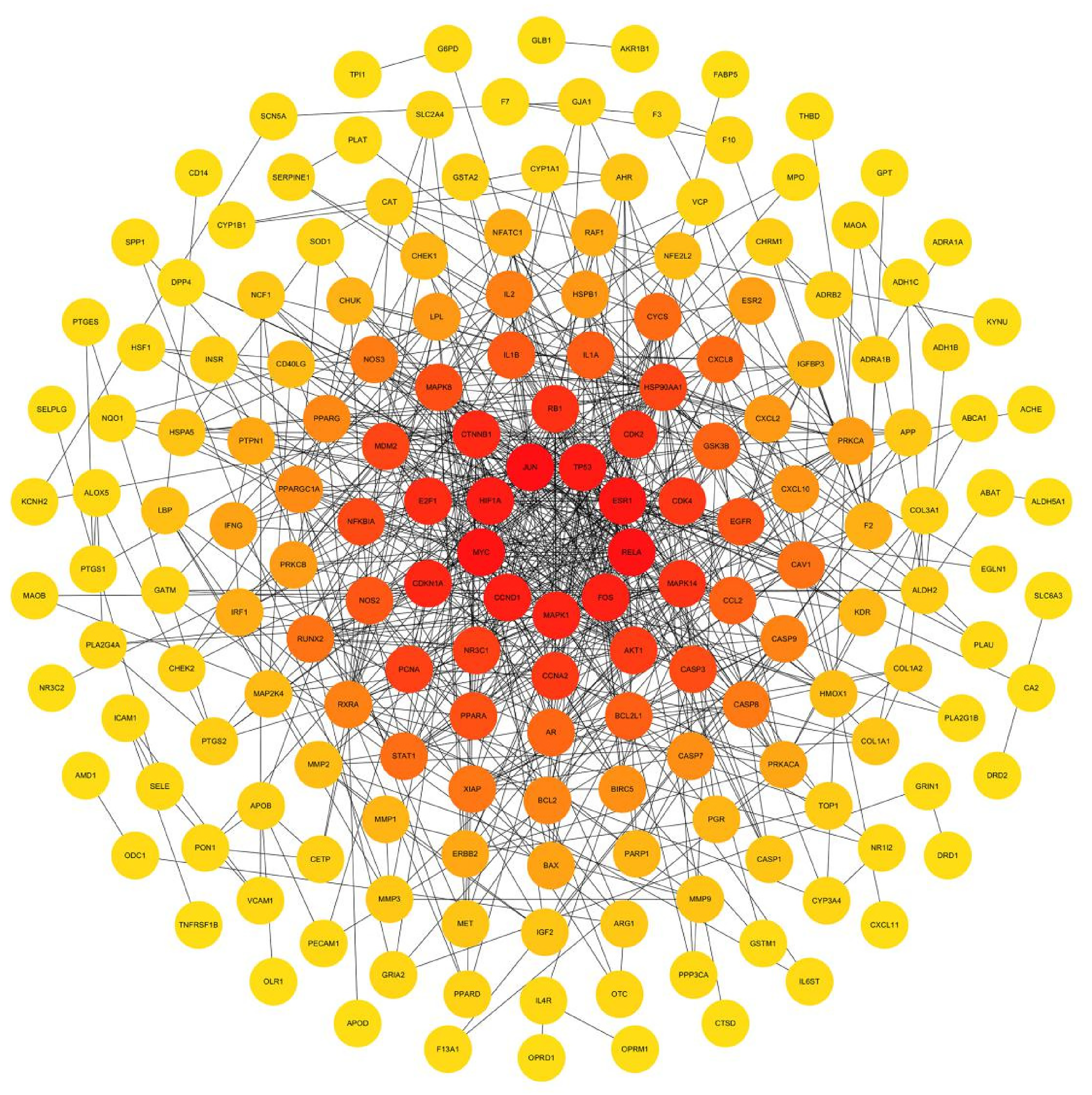
The network diagram of core targets and noncore targets. The nodes close to the center and dark color represent that they play a key role in the whole network.
The Specific Information of the 16 Core Targets from Topological Analysis of 209 Targets.

Abbreviation: DC, degree centrality.
GO and KEGG Enrichment Analysis
GO Enrichment
GO analysis of 209 candidate targets for CHSC against CIS was performed using the clusterProfiler. R data package was used to understand the relationship between functional units and their underlying significance in the biological system networks. The results were divided into three parts including biological processes (MF), cellular component (CC), and molecular function (MF) as shown in Figure 6. For MF ontology, some of the top GO terms associated with CIS were protein domain specific binding, kinase binding, protein homodimerization activity, DNA-binding transcription factor binding, oxidoreductase activity, amide binding, phosphatase binding, serine hydrolase activity, protein heterodimerization activity, and cytokine receptor binding. For BP ontology, some of the top 10 GO terms associated with CIS were in response to inorganic substance, cellular response to nitrogen compound, cellular response to organic cyclic compound, response to drug, response to wounding, response to lipopolysaccharide, circulatory system process, response to growth factor, apoptotic signaling pathway, and response to radiation. For CC ontology, top-scoring GO terms were membrane raft, perinuclear region of cytoplasm, neuronal cell body, axon, receptor complex, side of membrane, mitochondrial envelope, vesicle lumen, extracellular matrix, and transcription regulator complex.

GO enrichment analysis of CHSC in treating CIS. GO items and gene count are represented by the x-axis and y-axis, respectively.
Pathway Enrichment
Through clusterProfiler R data package analysis, we obtained all the pathways that illustrate the role of CHSC in the treatment of CIS. Top 10 KEGG signaling pathways of CHSC were obtained and constructed based on the correlation of pathway to CIS, as shown in Figure 7. From the KEGG enrichment analysis, the top 10 KEGG pathways were IL-17 signaling pathway, TNF signaling pathway, Toll-like receptor signaling pathway, Th17 cell differentiation, NF-kappa B signaling pathway, MAPK signaling pathway, PI3K-Akt signaling pathway, HIF-1 signaling pathway, VEGF signaling pathway, and p53 signaling pathway.
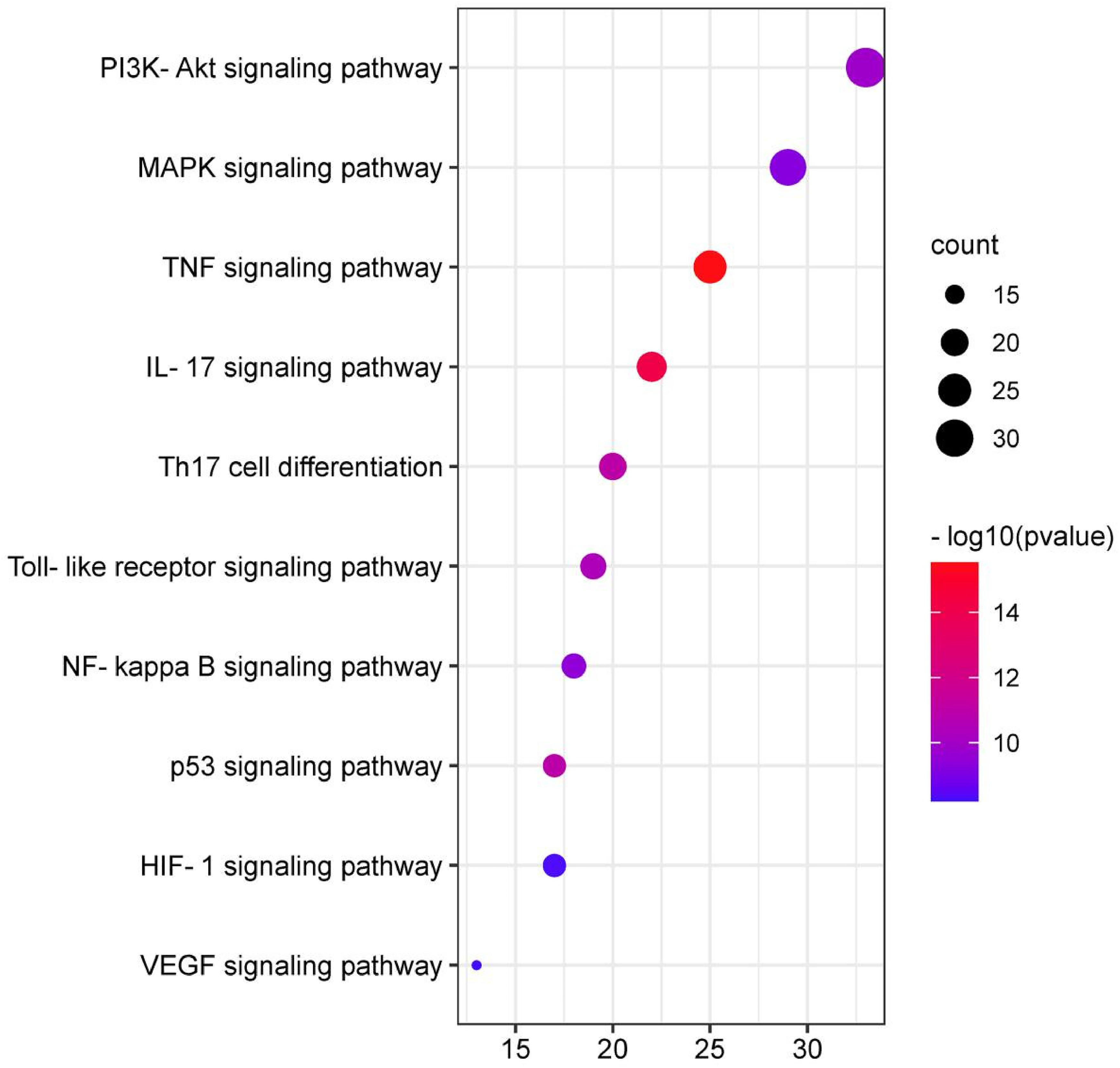
Top 10 enrichments of KEGG analysis with CHSC in treating CIS.
CHSC-CIS Network Analysis
Herb-Target Network
The active components and their targets were constructed to establish the compound-target network using Cytoscape. This network was composed of 298 nodes and 1374 edges. The herb-target network is shown in Figure 8.

Herb-compound-targets network of CHSC in treating CIS. The peripheral circles of different colors represent different compounds, including CS, CX, PH, GG, HJ, SQ, HH, DL, and XXC, and A1, A2, A3, A4, B1, B2, and C1 represent the repeated components of each herb, and the inner rectangle represents the targets.
Components-Target Network
The components-target network of CHSC was constructed using the “merge” function in Cytoscape software, as shown in Figure 9. In this network, we found that one target could be hit by several compounds (central nodes, such as PTGS2, PTGS1, PGR, HSP90AA1, PPARG, AR, and EAR1), but some were modulated by only one compound in this network. Furthermore, one potential active compound could correspond to multiple targets. The results showed that the top 11 key active components of CHSC in the treatment of CIS were quercetin, luteolin, kaempferol, beta-carotene, formononetin, beta-sitosterol, baicalein, arachidonic acid, 3'-methoxydidzein, myricanone, and isorhamnetin. These components play an important role in the treatment of ischemic stroke.
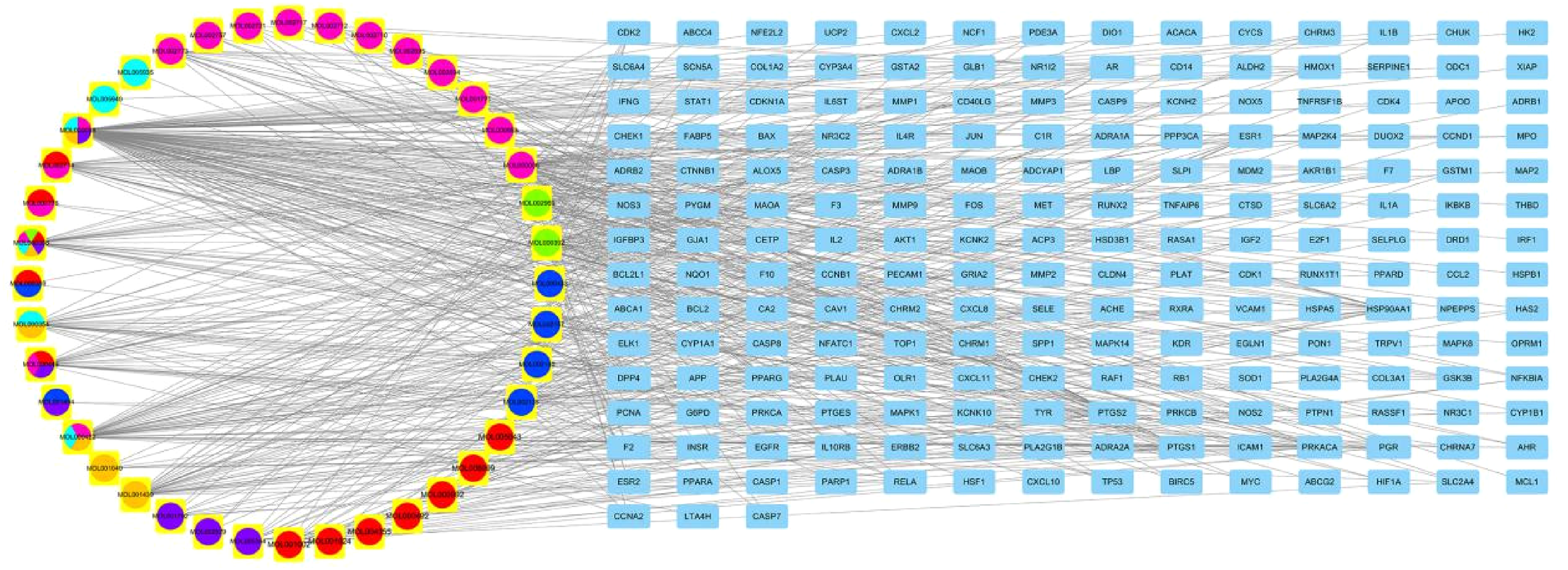
The network analysis of the drug-CHSC-potential target of CIS. The circle on the left represents the active components of the herbs, and the rectangle on the right represents the targets.
Herb-Components-Pathway Network
After the analysis of herb-components-pathway, we found that CIS was closely related to the PI3K-Akt signaling pathway, MAPK signaling pathway, and TNF signaling pathway. The herb-components-pathway network was shown in Figure 10 (Sankey diagram). The PI3K-Akt signaling pathway map of CHSC in treating CIS was obtained from KEGG pathway database, as shown in Figure 11. The targets related to pathways were also listed in Table 3.

Sankey diagram of herb-components-pathway. The band between the 2 bars represents a targeting relationship.
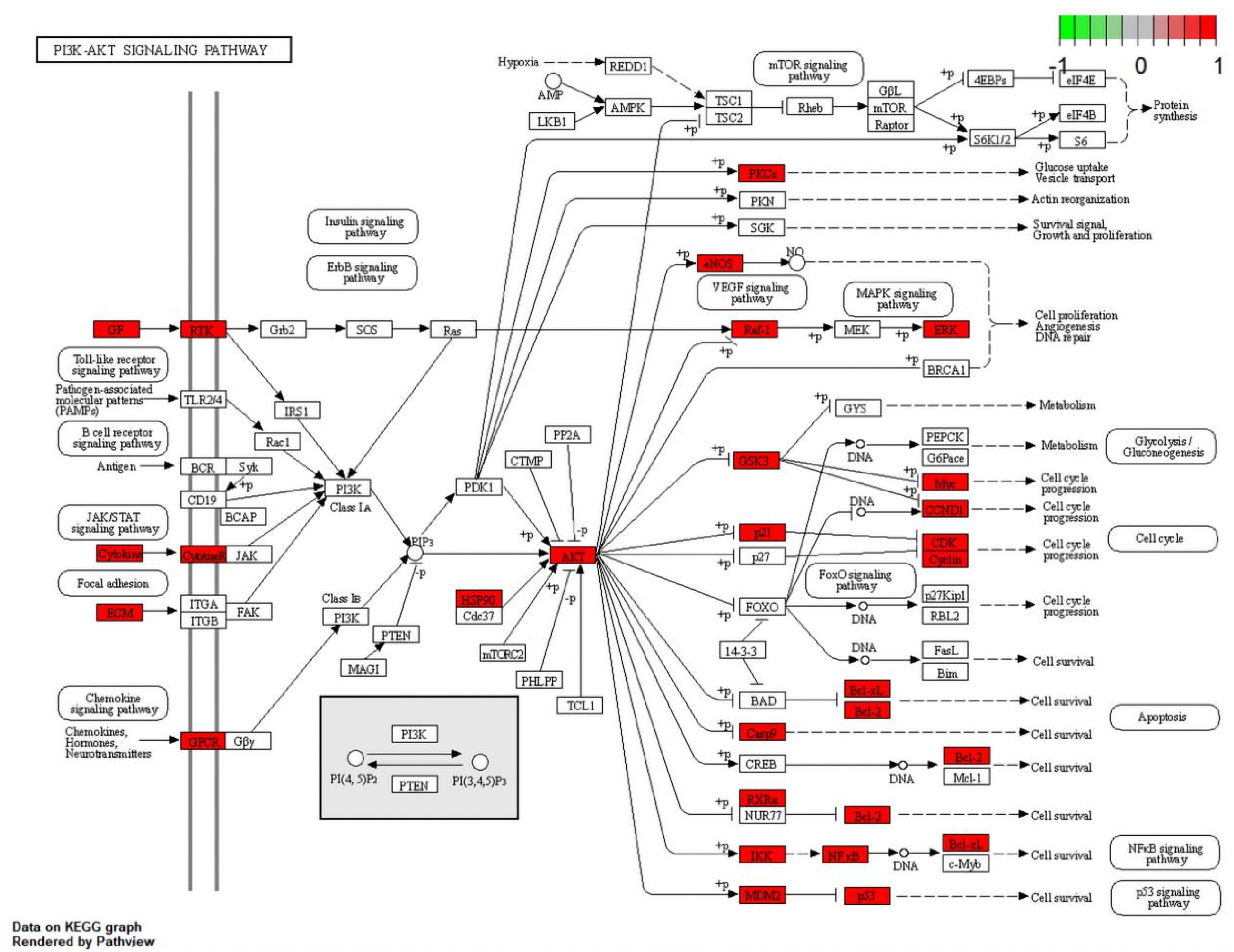
Pathway map of CHSC against CIS. The key targets of CHSC in the treatment of CIS were shown as rose red in the PI3K-Akt signal pathway.
Information of Related Target Gene Symbols in Pathway.

Components-Targets Docking Analysis
The molecular docking results were less than −5.0 kJ/mol, suggesting that these compounds could exert a strong binding effect with the key proteins (Figure 12), including JUN with quercetin (A), JUN with luteolin (B), JUN with kaempferol (C), MAPK1 with arachidonic acid (D), MAPK1 with quercetin (E), and MAPK1 with luteolin (F), as shown in Table 4.

The conformations of some important compounds and key targets in CHSC. (A) JUN with quercetin (binding energy = −8.2). (B) JUN with luteolin (binding energy = -8.2). (C) JUN with kaempferol (binding energy = −8.1). (D) MAPK1 with arachidonic acid (binding energy = −5.5). (E) MAPK1 with quercetin (binding energy = −8.4). (F) MAPK1 with luteolin (binding energy =−8.5).
Target Docking Parameters and Corresponding Calculation Results.

Discussion
In this study, we used a network pharmacology strategy to discover the molecular mechanism of the neuroprotective effect of CHSC on the treatment of ischemic stroke. The network pharmacology strategy helps to clarify the mechanism of action of TCM from a systematic perspective. At present, the application of network pharmacology to study the mechanism of TCM has become a research hotspot. In this study, we have found that 62 active components of CHSC can act on 209 information system-related targets. Further analysis has shown that CHSC can act on a variety of biological processes of ischemic stroke through PI3K-Akt, MAPK, and TNF signaling pathways, and affect the prognosis of stroke. We further confirmed that CHSC has the characteristics of multi-component, multichannel, and multi-target. The core components with the highest degree in compound target network are considered to be responsible for the neuroprotective function of CHSC, including quercetin, luteolin, kaempferol, beta-carotene, formononetin, beta-sitosterol, baicalein, arachidonic acid, 3'-methoxydidzein, myricanone, and isorhamnetin.
Quercetin performs a neuroprotective function against focal cerebral ischemia through the modulation of various proteins, which have demonstrated the neuroprotective, anti-oxidative, anti-inflammatory, and anti-apoptotic effects, or inhibitory effect on microglia-derived oxidative stress28-30. Luteolin can inhibit MMP9, and activate PI3K/Akt signaling pathway and SIRT3/AMPK/mTOR signaling pathway in the treatment of CIS31,32. Kaempferol can inhibit brain injury, neuroinflammation, and blood–brain barrier dysfunction to improve neurological deficits in CIS by activating NF-κB and STAT333,34. Formononetin is able to recover injured nerve functions by enhancing synaptic plasticity, downregulating of the Bax/Bcl-2 ratio, and upregulating PI3K/Akt signaling pathway in the treatment of CIS35,36. Baicalein exerts a neuroprotective effect against ischemic/reperfusion injury via alteration of NF-κB, LOX, and AMPK/Nrf2 signaling pathway 37 . Arachidonic acid can play a neuroprotective role after cerebral infarction by reducing oxidative stress 38 . It was found that 2-hydroxy arachidonic acid could reduce brain injury by inhibiting apoptosis, activating Nrf2/HO-1, inhibiting iNOS/NO, and reducing the protein levels of IL-1B, IL-6, and TNF-α 39 .
According to the node degree in the core component target network we constructed, RELA, JUN, AR, NR3C1, CTNNB1, FOS, TP53, RB1, MAPK1, MYC, HIF1A, MAPK14, AKT1, CDKN1A, HSP90AA1, and ESR1 were considered as the core targets of CHSC in the treatment of CIS. In addition, molecular docking was carried out to verify the correlation between core components and core targets. Enenstein et al reported that multiple control points in RELA signaling were associated with the risk of childhood stroke 40 . One study has found that resveratrol and valproate reduced RELA (K310) acetylation of neurons exposed to oxygen and glucose deprivation, and reduced infarct volume and neurological deficit 41 . Lanzillotta et al have shown that targeted acetylation of NF-κB/RELA and histones by epigenetic drugs could reduce ischemic brain injury 42 ]. It is reported that CTNNB1 was related to the pathological process of ischemic stroke 43 . Meng’s research has shown that increasing the expression of FOS and its downstream genes can improve the treatment of ischemic brain injury 44 . Zhai et al suggested that FOS may be associated with the development of ischemic stroke 45 . A study has found that TP53 may be considered as a genetic marker of predicting neurologic deterioration and poor functional outcome after stroke 46 . Wang et al believed that activation of TP53 could initiate the transcription of NOTCH1 and promote the process of neurogenesis 47 . Prestel’s findings suggested that RB1 was the main factor regulating vascular risk in atherosclerotic variables 48 . Recent research has found that neuronal injury can be reduced by regulating MAPK1 49 . Ma et al have found that MYC was involved in ischemia–hypoxia-induced astrocyte apoptosis 50 . Some studies have found that MYC downstream-regulated genes can protect the integrity of blood–brain barrier and cerebral edema following cerebral ischemia51,52. Ewida et al reported that HIF1A can improve the prognosis of stroke by activating angiogenesis53,54. Meanwhile, HIF1A can also be used as a diagnostic marker of ischemic stroke 55 . A recent study has demonstrated that MAPK14 can be used as a potential biomarker of cardioembolic stroke 56 . AKT1 is one of serine/threonine protein kinases (eg, AKT1, AKT2, and AKT3), and it participates in a variety of biological functions 57 ]. The activation of Akt is the main factor of neuronal survival after ischemic injury. Many neuroprotective agents can enhance Akt activity and protect brain injury, indicating that AKT is the main mechanism of cell repair after injury 58 . Gao’s findings have indicated that ESR1 genetic polymorphisms may contribute to the development of cerebral infarction, especially in the female population 59 ]. A recent study found that stroke was significantly associated with ESR1 rs2234693 polymorphism, especially ischemic stroke 60 . A study has found that JNK/JUN not only played an important role in promoting the proliferation and inhibiting the apoptosis of cerebral smooth muscle cells in vitro, but also played a role in reducing cerebral ischemia–reperfusion injury61,62.
According to GO annotation analysis, the core targets of CHSC played an important role in protein homodimerization activity, protein domain specific binding, kinase binding, response to inorganic substance, cellular response to nitrogen compound, cellular response to organic cyclic compound, etc., KEGG pathway analysis showed that many pathways were closely related to the pathogenesis of CIS. Typically, PI3K-Akt signaling pathway, MAPK signaling pathway, TNF signaling pathway, IL-17 signaling pathway, and Th17 cell differentiation were the top 5 ranks of KEGG analysis. T cells play an important role in regulating stroke injury. Th-17 cell, a member of CD4 T cells, secretes IL-17 cytokine, which includes 6 members from IL-17A to IL-17F, and most of which have pro-inflammatory effects. Th17 cell has been proved to be the main source of neuroinflammatory cascade in acute ischemic brain injury. After stroke, IL-17A and IL-17F induced inflammatory factors (such as TNF-α) to promote the development of stroke, and IL-17B/IL-17c/IL-17d showed excessive pro-inflammatory effects. On the contrary, IL-17e plays an immunoprotective role in the setting of CIS. Therefore, IL-17 functions as a two-edged sword63-65. Previous studies have shown that PI3K/Akt signaling pathway and TNF-α are important for neural development. PI3K/Akt pathway can not only prevent nerve injury, but also promote neuronal survival. Its effect of repairing nerve injury was positively correlated to VEGF expression 66 . Lv et al have found that mice with ischemic stroke can promote the activation of glial cells and inhibit neuroinflammation by activating PI3K/Akt/NF-κB signaling pathway 67 . TNF-α is closely related to inflammation and is one of the key factors in the progression of stroke. It had a beneficial effect on poststroke functional recovery 68 . Yong et al have found that the apoptosis of cerebral microvascular endothelial cells in ischemic stroke could be inhibited by downregulating MAPK-related pathway 69 . There are lot of studies supporting that downregulation of MAPK-related pathways can reduce nerve injury70,71.
Finally, we used molecular docking to explore the mechanism of CHSC in the treatment of CIS. In this study, we found that the therapeutic effect of CHSC on CIS may be due to the active compounds of CHSC, especially myricanone, isorhamnetin, luteolin, baicalein, arachidonic acid, formononetin, etc., which may affect the regulation of PI3K-Akt, MAPK, and TNF signaling pathways. In addition, through PPI analysis, these key targets play an important role in the treatment of CIS, such as JUN, FOS, MAPK1, MAPK14, and AKT. The results of molecular docking indicated that there was a strong affinity between the above components and key targets.
Conclusions
In this study, network pharmacology and molecular docking were used to explore the biological mechanism of CHSC in the treatment of CIS. It was observed that CHSC achieved therapeutic effects through a complex network of multiple components, pathways, and targets. This study showed that the main active ingredients of CHSC were 3'-methoxydaidzein, arachidonic acid, baicalein, beta-carotene, beta-sitosterol, formononetin, isorhamnetin, kaempferol, luteolin, myricanone, and quercetin, which mainly target RELA, JUN, FOS, TP53, MAPK1, MAPK14, AKT1, ESR1, etc. The main regulatory pathways were PI3K-Akt, MAPK, and TNF signaling pathways. Our findings provided a theoretical basis for further study of the effective components and mechanism of CHSC in the treatment of CIS. However, our study also has some limitations. For example, the research of network pharmacology was based on data and the real-life situation may be slightly different. In addition, the efficacy and dose effects of additional components should be further examined. Finally, further studies using in vitro and in vivo experiments are required to corroborate our findings.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was supported by Jilin University (Start-up Grant to Jinhua Li), Bethune Research Project of Jilin University (2018B34), and Graduate Innovation Fund of Jilin University (101832020CX272).
Data Availability
The data for this study can be provided by the corresponding author (Jinhua Li: jinhua1@jlu.edu.cn).
Authors’ Contributions
Xu Wang contributed to conceptualization and writing original draft. Jun Wang and Xin Chen contributed to software and data curation. Zi-Qiao Yu and Mo-Xuan Han contributed to supervision, review and editing. Wei-sen Zhao contributed to methodology and software. Jinhua Li contributed to supervision, funding acquisition, and writing revision.
Ethical Approval
Not applicable, because this article does not contain any studies with human or animal subjects.
Informed Consent
Not applicable, because this article does not contain any studies with human or animal subjects.
Trial Registration
Not applicable, because this article does not contain any clinical trials.
