Abstract
Severe acute respiratory syndrome (SARS) once caused great harm in China, but now it is the coronavirus disease 2019 (COVID-19) pandemic that has become a huge threat to global health, which raises urgent demand for developing effective treatment strategies to avoid the recurrence of tragedies. Yinqiao powder, combined with modified Sangju decoction (YPCMSD), has been clinically proven to have a good therapeutic effect on COVID-19 in China. This study aimed to analyze the common mechanism of YPCMSD in the treatment of SARS and COVID-19 through network pharmacology and molecular docking and further explore the potential application value of YPCMSD in the treatment of coronavirus infections. Firstly, the active components were collected from the literature and Traditional Chinese Medicine Systems Pharmacology database platform. The COVID-19 and SARS associated targets of the active components were forecasted by the SwissTargetPrediction database and GeneCards. A protein–protein-interaction network was drawn and the core targets were obtained by selecting the targets larger than the average degree. By importing the core targets into database for annotation, visualization, and integrated discovery, enrichment analysis of gene ontology, and construction of a Kyoto Encyclopedia of genes and genomes pathway was conducted. Cytoscape 3.6.1 software was used to construct a “components–targets–pathways” network. Active components were selected to dock with acute respiratory syndrome coronavirus type 2 (SARS-COV-2) 3CL and angiotensin-converting enzyme 2 (ACE2) through Discovery Studio 2016 software. A network of “components–targets–pathways” was successfully constructed, with key targets involving mitogen-activated protein kinase 1, caspase-3 (CASP3), tumor necrosis factor (TNF), and interleukin 6. Major metabolic pathways affected were those in cancer, the hypoxia-inducible factor 1 signaling pathway, the TNF signaling pathway, the Toll-like receptor signaling pathway, and the PI3K-Akt signaling pathway. The core components, such as arctiin, scopolin, linarin, and isovitexin, showed a strong binding ability with SARS-COV-2 3CL and ACE2. We predicted that the mechanism of action of this prescription in the treatment of COVID-19 and SARS might be associated with multicomponents that bind to SARS-COV-2 3CL and ACE2, thereby regulating targets that coexpressed with them and pathways related to inflammation and the immune system.
Keywords
Introduction
Application of traditional Chinese medicine (TCM) in the treatment of the novel coronavirus acute respiratory syndrome coronavirus type 2 (SARS-CoV-2) has been largely inspired by the treatment of severe acute respiratory syndrome (SARS) caused by the outbreak of SARS-CoV in late 2002 in Guangdong Province, China; this spread rapidly during 2003, with the cumulative number worldwide being over 8000. 1 In late December, 2019, acute pneumonia that had been raging for over 2 months and caused by a novel coronavirus was observed in Wuhan, Hubei Province, China. Coronavirus Disease 2019 (COVID-19), which was caused by the novel coronavirus SARS-CoV-2, has become a serious threat for public health worldwide. 2 According to statistics, as of April 19, 2021, 141 922 191 individuals had been infected with COVID-19 worldwide, of which 3 026 088 had died. The novel coronavirus was mainly spread by respiratory droplets in crowds, but was also less often spread through aerosols, fecal-oral transmission, and other channels.3,4 The virus is characterized by strong infection, fast transmission, long incubation period, and general susceptibility. The main symptoms of patients after infection are fever and fatigue. In severe cases, shock and multiple organ dysfunction may occur in a short period, which poses a serious threat to the life and health of patients.5,6
At present, it is considered that controlling the imbalance of inflammatory factors, improving the immune system, and inhibiting cell apoptosis are the main directions for the treatment of COVID-19. 7 In the absence of the availability of specific antiviral drugs or vaccines, a few Asian countries, such as China, Thailand, and India, have been relying on the use of traditional medicines. In TCM theory, Yinqiao powder combined with modified Sangju decoction (YPCMSD) is often used for treating acute upper respiratory tract infections, sore throat, headache, fever, and cough. 8 YPCMSD is included in the “Novel Coronavirus Prevention and Control Technical Guide of Traditional Chinese Medicine in Sichuan Province” issued by The Administration of TCM of Sichuan Province, which was applicable to the treatment of the novel coronavirus. YPCMSD consists of Lonicera japonica Thunb. (Jinyinhua), Forsythia suspensa (Thunb.) Vahl (Lianqiao), Morus alba L. (Sangye), Dendranthema morifolium (Ramat.) Tzvel. (Juhua), Platycodon grandiflorus (Jacq.) A. DC. (Jiegeng), Arctium lappa L. (Niubangzi), Lophatherum gracile Brongn. (Danzhuye), Phragmites communis Trin. (Lugen), Mentha haplocalyx Briq. (Bohe) and Glycyrrhiza uralensis Fisch. (Gancao). At present, the efficacy of this prescription has been verified in China, so it is necessary to explore further its active components and mechanism of action, so as to increase the popularization and use of this prescription. 9
Network pharmacology was introduced by Hopkins in 2008 using a visualization network software and a variety of algorithms combined with integrated analysis of existing related databases. This method breaks the previous single component/single target/disease concept, and it is also consistent with the TCM theory of multiple components/multiple targets/more complicated diseases. 9 The molecular docking technique can simulate interactions between receptors and drug molecules. It is usually used to study the active sites of drugs and plays an important role in the study of natural products. In this study, we predicted the core components and core targets of YPCMSD and the signaling pathways for treatment of SARS and COVID-19 through network pharmacology and molecular docking, so as to prove ideas and the basis for its further research. The flow chart of this study is shown in Figure 1.

Flow chart of this study.
Materials and Methods
Collection of Potential Active Components
The active components of YPCMSD were identified from literature and the Traditional Chinese Medicine Systems Pharmacology database and analysis platform (TCMSP, https://tcmspw.com/tcmsp.php). The platform provides pharmacokinetic properties of natural compounds such as oral bioavailability (OB) and drug-likeness (DL) for new drug discovery.
Screening of Active Components and Targets
OB is an important pharmacokinetic parameter in drug absorption, distribution, metabolism, and excretion. It indicates the speed and degree at which the active components or active groups of the oral drug are absorbed into the systemic circulation. Further, DL refers to the similarity of a component with a known drug and the potential of this class of components to become drugs. Therefore, active components were selected according to their OB and DL characteristics (OB ≥ 30% and DL ≥ 0.18). The active components were imported into SwissTargetPrediction (http://www.swisstargetprediction.ch/) to collect the relevant targets. The targets related to viral pneumonia were obtained by searching “COVID-19” and “SARS” as the keywords in the GeneCards database (https://www.genecards.org/). All intersections with the predicted targets of the obtained components were used to obtain the potential targets of YPCMSD in the treatment of COVID-19 and SARS.
Gene Ontology and Kyoto Encyclopedia of Genes and Genomes Pathway Enrichment
The potential targets were sorted out and uploaded to the database for annotation, visualization, and integrated discovery (DAVID, https://david.ncifcrf.gov/) for gene ontology (GO), and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway annotation. The pathways of targets enrichment of active components were considered to be an important regulatory pathway of YPCMSD against COVID-19 and SARS. The top 20 relevant results from the KEGG pathway and GO annotation analysis were plotted using the OmicShare tool (http://www.omicshare.com) as bubble plots.
Construction of “Components–Targets–Pathways” Network
Cytoscape 3.6.1 software (https://cytoscape.org/download.html) was used to construct a “components–targets–pathways” network. Different nodes represent the active components, targets, pathways, and edges used to indicate the correlation between the 2 nodes. Topological analysis of the constructed pharmacological network was performed to screen the core active components. In the “components–targets–pathways” network, the components with a high degree were the most important active components.
Construction of Protein–Protein–Interaction Network
The STRING database (https://string-db.org/) collects, scores, and integrates all publicly available protein–protein–interaction (PPI) information, including direct (physical) and indirect (functional) interactions. The targets were imported into the STRING database to obtain the PPI network. The TSV file was downloaded and imported into the software of Cytoscape 3.6.1 and analyzed by using the function of network analysis.
Molecular Docking
The core components selected from the YPCMSD were docked with SARS-COV-2 3CL hydrolase and angiotensin-converting enzyme 2 (ACE2). Subsequently, the structures of related proteins were downloaded from the RSCB PDB database (https://www.rcsb.org/). The structures of components were downloaded from the Pubchem database (https://pubchem.ncbi.nlm.nih.gov/). DiscoveryStudio software was used for solvent molecules and ligand removal, hydrogenation, electron addition, and other operations. Molecular docking was then performed (Discovery Studio 2016; BIOVIA). It is generally believed that LiDockscore ≥ 90 has a strong binding ability when docking with the processed protein. 12
Result
Active Components of YPCMSD
The TCMSP database was used to screen the components of YPCMSD. With OB ≥ 30% and DL ≥ 0.18, 223 active components were selected, of which 16 were from Lianqiao, 24 from Sangye, 16 from Jinyinhua, 18 from Juhua, 4 from Jiegeng, 7 from Niubangzi, 1 from Lugen, 9 from Bohe, 76 from Gancao, and 5 from Danzhuye. There were 17 components contained in more than 2 herbs. The basic information of some components in YPCMSD is shown in Table 1. All collected components information is shown in Supplemental Table S1.
Basic Information of Some Active Components in YPCMSD.

Abbreviations: OB, oral bioavailability; DL, drug-likeness; YPCMSD, Yinqiao powder, combined with modified sangju decoction.
Potential Targets of YPCMSD
A total of 1138 potential targets of active components were collected from SwissTargetPrediction database. In addition, we screened the top 500 core targets related to COVID-19 and SARS from the GeneCard database and intersected them with the possible targets of YPCMSD to obtain 75 coincidence targets (Figure 2).

Venn diagram of coincidence targets.
Target Biological Function Analysis
GO functional enrichment analysis in DAVID yielded 213 GO entries (P < .05), including 124 biological process (BP) entries, 35 cell composition entries, and 54 molecular functional entries, accounting for 58.2%, 16.4%, and 25.4%, respectively (shown in Figure 3A to C).

Bubble chart of the results of KEGG pathway enrichment and GO functional enrichment. (A) The top 20 significantly enriched entries in BP; (B) the top 20 significantly enriched entries in CC; (C) the top 20 significantly enriched terms in MF; and (D) the top 20 KEGG pathways enrichment.
The KEGG pathway annotation of DAVID involved 113 pathways (P < .05) and the top ones were pathways in cancer, hypoxia-inducible factor 1 (HIF-1) signaling pathway, tumor necrosis factor (TNF) signaling pathway, vascular endothelial growth factor (VEGF) signaling pathway, Toll-like receptor signaling pathway, and PI3K-Akt signaling pathway (Figure 3D). Among them, pathways in cancer involved 26 genes including epidermal growth factor receptor (EGFR), phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma (PIK3CG), CASP3, mitogen-activated protein kinase 1 (MAPK1), tumor protein p53 (TP53), and interleukin 6 (IL6). The HIF-1 signaling pathway involved 14 genes, including PIK3CG, IL6, and B-cell lymphoma-2 (BCL2). The TNF signaling pathway involved 14 genes including TNF, MAPK8, CC motif chemokine 2 (CCL2), and CC motif chemokine 5 (CCL5). Enrichment analysis of the key KEGG pathways showed that YPCMSD was mainly involved in tumorigenesis pathways, inflammatory response signaling pathways, immune regulation pathways, and bacterial infections to achieve anti-CoV effects.
Construction of “Components–Targets–Pathways” Network
The constructed network included 201 nodes (128 components, 53 targets, and 20 pathways), with green representing the components, blue representing the targets, and yellow representing the KEGG pathways (shown in Figure 4). By analyzing between the centrality, closeness centrality, and degree between the components and the targets, we found that the top 10 components were MOL001484: inermine, MOL000522: arctiin, MOL002311: glycyrol, MOL003656: lupiwighteone, MOL004857: gancaonin B, MOL003370: onjixanthone I, MOL001790: linarin, MOL002218: scopolin, MOL002322: isovitexin, and MOL006070: robinin. The top 10 targets were found to be cyclin-dependent kinase 2 (CDK2), EGFR, phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (PIK3CA), prostaglandin G/H synthase 2 (PTGS2), glycogen synthase kinase 3 beta (GSK3B), PIK3CG, prostaglandin G/H synthase 1 (PTGS1), MAPK14, poly(ADP-ribose)polymerase-1 (PARP1), and peroxisome proliferative activated receptor gamma (PPARG). It is suggested that the prescription for YPCMSD for the treatment of COVID-19 and SARS might work through multiple components which regulate genes coexpressed with ACE2 and SARS-CoV-2 3CL.

Components–targets–pathways network: The green, blue, and yellow nodes, respectively, represented the active components, targets, and pathways.
PPI Network Construction
The potential targets of YPCMSD in the treatment of COVID-19 and SARS were recorded on the STRING website (https://string-db.org/cgi/input.pl) and the PPI network of the potential targets was obtained. We further found that TP53, MAPK1, CASP3, STAT3, TNF, and IL6 were the links between ACE2 and other targets, as shown in Figure 5. This suggested that the mechanism of action of YPCMSD in the treatment of the COVID-19 and SARS might be related to the regulation of genes coexpressed with ACE2.
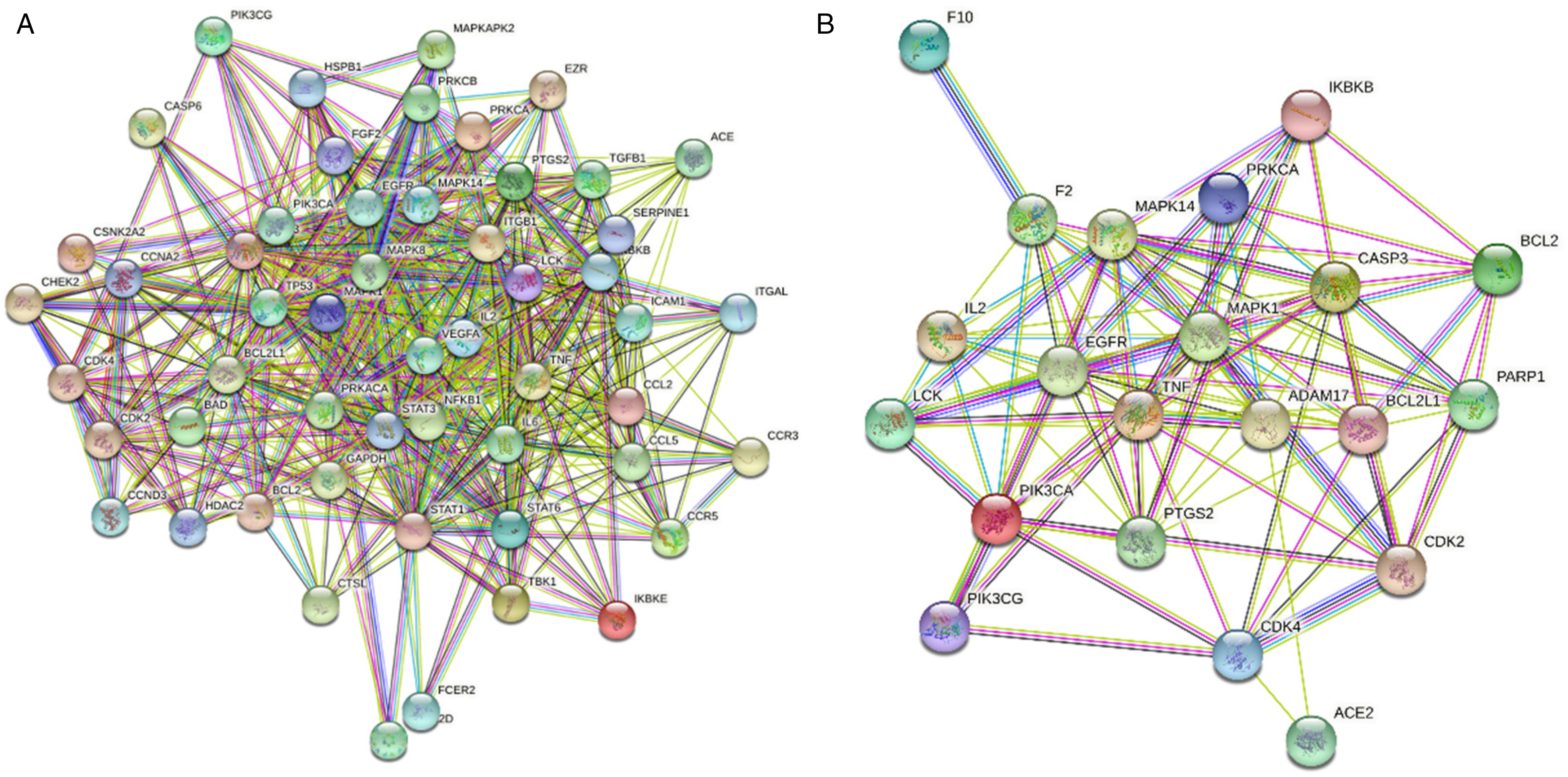
PPI interaction network. (A) Interaction of potential targets of YPCMSD for the treatment of COVID-19 and SARS and (B) The top 20 targets that interacted with ACE2.
Molecular Docking and Analysis
It is a general belief that the lower the energy of ligand and receptor binding, the higher the LiDock score, the greater the possibility of interaction, and the stronger the potential activity of the components. Until now, there was no clinical therapeutic drug that could be used as a positive drug. Thus, we directly docked the components with the targets. The results of the docking were analyzed with LiDockscore ≥ 90 as the screening criterion. The 4 core components (arctiin, scopolin, linarin, and isovitexin) mentioned in topology analysis network were docked with SARS-COV-2 3CL and ACE2. The details are shown in Figures 6 and 7, and Table 2. The result showed that all core components had strong affinity for ACE2 protein and SARS-CoV-2 3 Cl protein. Compared with other core components, the docking scores of scopolin and inermine with these 2 proteins were not too high.

Results of molecular docking of each component with ACE2. (A) arctiin, (B) linarin, (C) scopolin, and (D) isovitexin.

Results of molecular docking of each component with SARS-COV-2 3CL. (A) arctiin, (B) linarin, (C) scopolin, and (D) isovitexin.
Results of Molecular Docking.

Discussion
TCM has a long clinical history for the prevention and treatment of this kind of acute infectious diseases. According to the principle of TCM, COVID-19 is a disease caused by the evil of wet poison, because wet poison attacks the lung. At present, China has issued many prescriptions for the treatment of COVID-19 and the clinical results show that they have significant efficacy. The prescription of YPCMSD is composed of Yinqiao San and Sanju Yin, which is a treatment plan against COVID-19 issued by the general office of the National Health Commission and the office of the State Administration of TCM. 13
In this study, the mechanism underlying the treatment of CoV infection with YPCMSD was studied by network pharmacology and 75 effective components, 53 effective targets, and 113 signal pathways were screened out. We used the network pharmacology method and molecular docking technique to screen the core components in YPCMSD and predicted its possible action mechanism in COVID-19 and SARS treatment. The pathways most relevant to the lungs obtained by KEGG analysis were the HIF-1 signaling pathway, the TNF signaling pathway, the VEGF signaling pathway, the Toll-like receptor signaling pathway, the PI3K-Akt signaling pathway, small cell lung cancer, and pathways in cancer, all of which are mostly involved with IL6, CASP3, IL2, STAT3, TP53, PIK3CG, PIK3CA, MAPK14, and CCL2 genes. These genes are mostly targets of arctiin, inermine, scopolin, robinin, linarin, isovitexin, and some other components. It has been reported that the anti-inflammatory effect of isovitexin is by inhibiting the production of TNF-α, IL-1 β, and IL-6. In addition, it can inhibit the production of malondialdehyde (MDA) and reactive oxygen species (ROS) to enhance the antioxidant effect and reduce the production of pro-inflammatory cytokines in a concentration-dependent manner. 14 Arctiin has a wide range of pharmacological activities such as antitumor, immunomodulation, and anti-inflammatory effects, especially its powerful anti-inflammatory effect. Arctiin can significantly inhibit the release of nitric oxide, prostaglandin E2, TNF-α, IL-6, and IL-2 from macrophages activated by lipopolysaccharide 1β and other inflammatory active substances and down-regulate the activation of NF-κB, AP-1, and other transcription factors in the nucleus to exert anti-inflammatory effects, 15 which are related to the anti-SARS and anti-COVID effects. Scopolin can reduce IL-6, VEGF, and FGF-2 expressions in rat synovial tissues. 16 “COVID-19 clinical protocols, seventh edition” pointed out that severe and critically ill patients often had elevated inflammatory factors and clinical manifestations of “cytokine storm.” Regulating the expression of these genes could further reduce inflammation and reduce the damage of inflammatory response to the body. 17
Further research on the main KEGG signaling pathways of YPCMSD found that the TNF signaling pathway and MAPK signaling pathway could reduce the excessive immune response and eliminate inflammation. The PI3K-Akt signaling pathway could recognize cell stimulation or toxic damage activation, thereby regulating basic cell functions. 18 The HIF-1 signaling pathway had the function of regulating the body's oxygen homeostasis. The Toll-like receptor signaling pathway has an antiviral effect by regulating immune response. 19 At the same time, the Toll-like receptor signaling pathway induced the secretion of a variety of inflammatory factors and participated in the inflammatory response. Many BP played a key role in the treatment of COVID-19 and SARS, including anti-inflammatory, antiviral, and immunomodulatory effects. The main genes involved in these pathways include CASP3, EGFR, IL6, MAPK1, IKBKB, PIK3CA, PIK3CG, CCL5, TNF, TP53, and VEGFA. The above genes could also confirm the results of the PPI network analysis. All the above results proved that the multiple targets of TCM could treat COVID-19 and SARS by regulating the KEGG signaling pathway.
However, there are still some limitations in network pharmacology and molecular docking technology. In the future, we need to use network pharmacology as a guide to conduct in-depth pharmacodynamics and pharmacokinetics experiments to provide a more specific pharmacological basis for YPCMSD to treat coronavirus-infected pneumonia.
Conclusion
In the summary, active components such as arctin, scopolin, linarin, and isovitexin, core targets such as MAPK1, CASP3, STAT3, TNF, and IL6, and potential signaling pathways such as the HIF-1 signaling pathway and TNF signaling pathway of YPCMSD for the treatment of SARS and COVID-19 were screened out for the first time through network pharmacology and molecular docking technology. It is predicted that YPCMSD can achieve an anticoronavirus-induced disease effect mainly through regulating tumorigenesis pathways, inflammatory response signaling pathways, immune regulation pathways, and bacterial infections. This study will provide valuable reference for the application of YPCMSD.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X211035290 - Supplemental material for A Network Pharmacology and Molecular Docking Approach to Investigate the Anticoronavirus-Induced Diseases Effect of Yinqiao Powder Combined With Modified Sangju Decoction
Supplemental material, sj-docx-1-npx-10.1177_1934578X211035290 for A Network Pharmacology and Molecular Docking Approach to Investigate the Anticoronavirus-Induced Diseases Effect of Yinqiao Powder Combined With Modified Sangju Decoction by Tian-Shun Wang, Xing-Pan Wu, Qiu-Yuan Jian, Yan-Fang Yang and Wu He-Zhen in Natural Product Communications
Footnotes
Authors' Note
Tianshun Wang designed the entire experiment and wrote the manuscript. Xingpan Wu also participated in the design of the experiment and search for literature databases.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (grant number [31570343]).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
