Abstract
Purpose
Fufang Banlangen Keli (FBK) has been recommended for its clinical treatment of Coronavirus disease 2019 (COVID-19) and severe acute respiratory syndrome (SARS), but the mechanism of action is unclear. So, using network pharmacology and molecular docking, we studied the active components and mechanism of FBK in the treatment of COVID-19 and SARS.
Methods
The Encyclopedia of Traditional Chinese Medicine and Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform were used to screen the active components by oral bioactivity and drug likeness. Then, PharmMapper and SwissTargetPrediction databases were used to screen potential target genes of active components; the related target genes of COVID-19 and SARS were obtained from the GeneCards database. The intersection of the active components and disease-related targets was performed by the Venny2.1.0 database. The DAVID6.8 database and KOBAS3.0 database were used to get gene ontology (GO) function enrichment and Kyoto Encyclopedia of Genes and Genomes pathway annotation of gene targets. The “components-targets-pathways (C-T-P)” network of FBK was conducted by Cytoscape3.6.1 software. The top active components, angiotensin-converting enzyme 2 (ACE2) and SARS-CoV-2 3 Cl, were imported into AutoDock and PyMOL for molecular docking.
Results
From the FBK, a total of 28 active components and 73 gene targets were screened through network pharmacology. Twenty pathways were analyzed, including pathways in cancer, nod-like receptor signaling pathway, and pancreatic cancer, etc. (P < 0.05). A total of 337 items were obtained by GO functional enrichment analysis (P < 0.05), including 257 items for biological process, 38 items for cell composition, and 42 items for molecular function. Furthermore, molecular docking studies were performed to study potential binding between the key gene targets and selected active components.
Conclusion
Based on network pharmacology and molecular docking technology, qingdainone, (2Z)-2-(2-oxoindolin-3-ylidene) indolin-3-one, sinensetin, and acacetin in FBK were verified to bind to ACE2 and SARS-COV-2 3 Cl, so as to treat COVID-19 and SARS.
Since December 2019, a pneumonia caused by novel coronavirus has swept the world extremely rapidly, and the World Health Organization (WHO) named it Coronavirus disease 2019 (COVID-19). COVID-19 and severe acute respiratory syndrome (SARS) are both infectious diseases caused by coronaviruses, and their early symptoms are similar, including fever, cough, sore throat, and dyspnea. The critical patients even appear dyspnea, hypoxemia, and acute respiratory distress syndrome. 1 By October 2020, the COVID-19 has rebounded again in Europe and other regions, posing a great threat to human life, health, and property safety. Researchers and medical staff around the world are urgently seeking to develop specific drugs and vaccines for COVID-19.
Fufang Banlangen Keli (FBK) has the effect of clearing heat and detoxifying, cooling blood. It is composed of Isatidis radix and Isatidis folium. 2 Research found that FBK was used to prevent and treat the virus during the SARS epidemic, and it has antiviral and immunological effects. 3 Some reports also showed that FBK could resist novel coronavirus in vitro experiments. Therefore, it is of great significance to study the active components and mechanism of action of FBK in the treatment of COVID-19 and SARS.
Network pharmacology is a method that integrates system biology, multidirection pharmacology, and computer analysis technology. By exploring the targets of drugs and disease, the “components-targets-pathways (C-T-P)” network was established to analyze the interaction between drugs and targets so as to obtain active components and the mechanism of drugs systematically and holistically. 4 The concept of network pharmacology is quite consistent with the characteristics of traditional Chinese medicine (TCM) in the treatment of diseases through multicomponent, multitarget, and multipathway. 5,6 Molecular docking technology can simulate the docking activity between protein and drugs and describe the functional groups and protein residues. Molecular docking has become an important means to research and develop new drugs and is a validation and supplement to network pharmacology.
Furthermore, the “C-T-P” network of FBK in treating COVID-19 and SARS was constructed to screen the key components, targets and pathways by network pharmacology, and molecular docking technique. It provides the train of thought and the basis for its later research. The workflow of this study is shown in Figure 1.
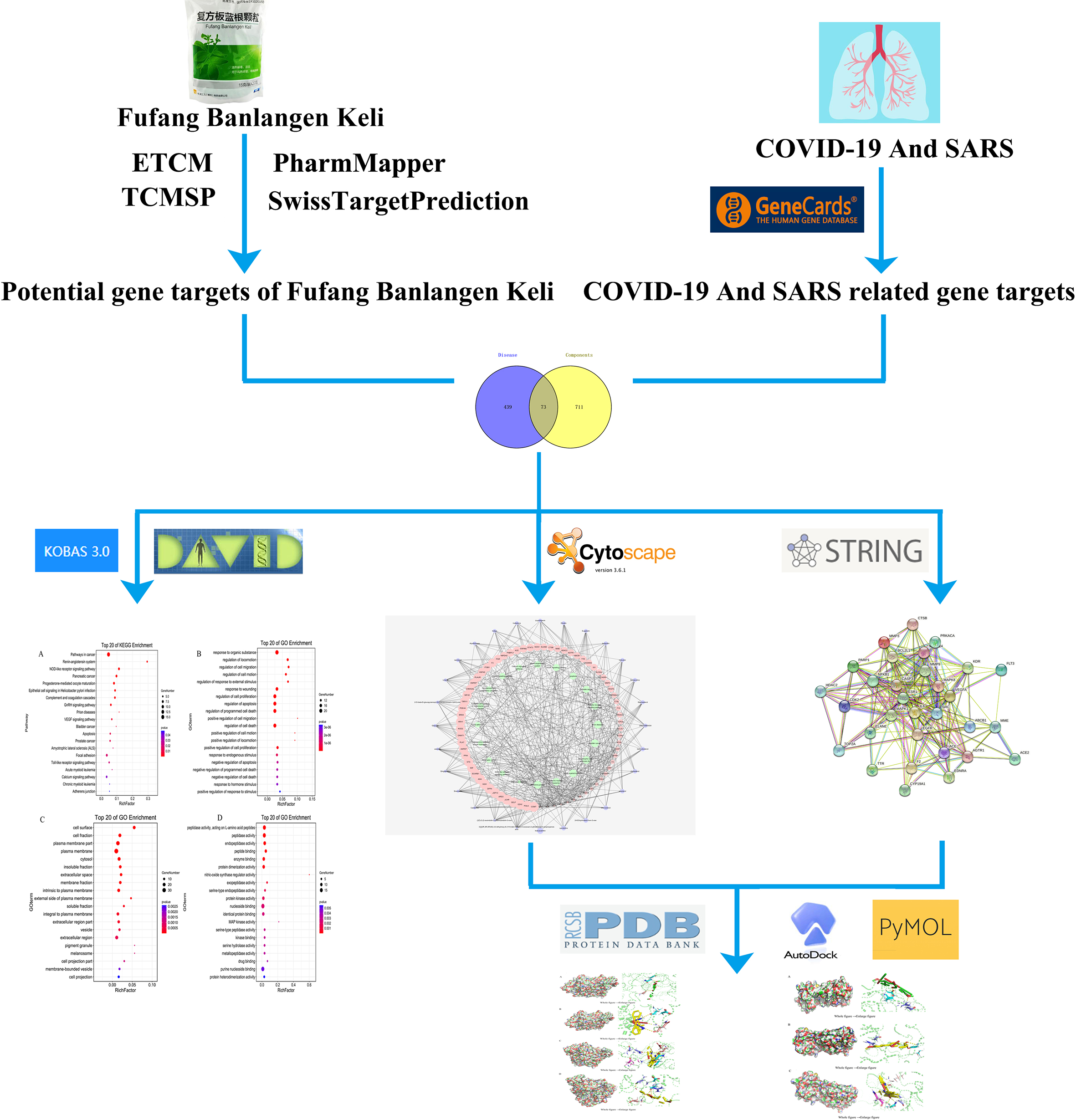
The workflow of network pharmacological study of Fufang Banlangen Keli against COVID-19. COVID-19, coronavirus disease 2019; ETCM, Encyclopedia of Traditional Chinese Medicine; SARS, severe acute respiratory syndrome; TCMSP, Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform.
Materials and Methods
Screening of Potential Active Ingredients
Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP) (http://lsp.nwu.edu.cn/tcmsp.php) and Encyclopedia of Traditional Chinese Medicine (ETCM) (https://pubchem.ncbi.nlm.nih.gov/) were used to collect all the chemical components of FBK. The common name and PubChem CID were collected from the PubChem database to establish the chemical components database of FBK. Meanwhile, according to the ADME principle, the active components in FBK were screened by using the index of oral bioactivity (OB) ≥30% and drug likeness (DL) ≥0.18. 6,7
Screening Potential Targets of Active Components Associated With Disease
The gene targets corresponding to the chemical components were obtained from the PharmMapper database (http://lilab.ecust.edu.cn/pharmmapper/index) and the SwissTargetPredict database (http://www.swisstargetprediction.ch/index.php). Using the Retrieve/ID mapping tool and setting the species as “human”, the Gene Official Symbol format of the corresponding genes of active components were obtained by UniProt. Then GeneCards (https://www.genecards.org/) were searched for keywords “COVID-19” and “SARS” to get disease-related gene targets. Finally, using Venny2.1.0 database and tabular tool to collect the intersection target genes of the active components in FBK with the 2 diseases.
Gene Ontology and KEGG Pathway Enrichment
Both the DAVID 6.8 database (https://david.ncifcrf.gol/) and the KOBAS 3.0 system (http://kobas.cbi.pku.edu.cn/anno_iden.php) were used for gene ontology (GO) enrichment and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis. For GO enrichment and KEGG pathway annotations, the potential target genes were imported into the 2 databases, respectively, selecting Identifier for GENE_OFFICIAL_SYMBOL, species as “Homo sapiens”. Using a threshold value of P < 0.05, the top 20 pathways and GO enrichment were plotted by the Omicshare database (http://www.omicshare.com) as bubble plots.
Construction of “C-T-P” Network
Cytoscape 3.6.2 database is a bioinformatics analysis software for integrating modular network and bioscience network diagrams. The 28 active components, 73 gene targets, and the top 20 KEGG pathways were imported into Cytoscape 3.6.2 database. Different nodes were used to represent the active components, gene targets, and enrichment pathways, respectively, and edges were used to represent the correlation between the two nodes. The “C-T-P” network was obtained by merging the components-targets network and the targets-pathways network. With the degree value greater than twice the median as the standard, components with high degree value in the topographic parameters were screened and analyzed as important active components in the later study.
Constructing Protein-Protein Interaction Network
STRING database (https://string-db.org/) is a part of the ELIXIR infrastructure and is one of ELIXIR’s Core Data Resources. It contains 5090 species, 24 584 628 proteins, 3 123 056 667 interactions. The plug-in in Cytoscape 3.6.2 software was used to analyze the topology parameters and chose 30 targets with the highest degree value. Then, those 30 targets and angiotensin-converting enzyme 2 (ACE2) were screened into the STRING database by setting the species as “human”. The interaction between gene targets was analyzed, and the protein-protein interaction (PPI) network diagram was downloaded after ranking.
Molecular Docking
RSCBPDB (https://www.rcsb.org/) is a database that has tens of thousands of gene targets. AutoDockTools-1.5.6 software is used for molecular docking to obtain the binding energy value.
The reported 3-dimensional (3D) structure of SARS-COV-2 3 Cl and ACE2 were downloaded from RSCBPDB in the PDB format. The 3D structures of the components downloaded from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/) were minimized by Chemdraw software, and imported into AutDockTools-1.5.6 to be hydrogenated, charged, and then the rotatable bond number was calculated. At the same time, the ligands and nonprotein molecules (such as water molecules) in the gene targets were removed by PyMol software and added all hydrogen to balance charges, calculated the total charge in AutoDockTools-1.5.6. It is generally believed that the binding capacity is stronger when the binding energy is lower than −5 Kcal/mol. The docking results were imported into PyMol software for visualization processing to show the functional groups and protein residues at the binding sites of components and proteins.
Results
Collection of Potential Active Ingredients
FBK is composed of Isatidis radix and Isatidis folium. Using the TCMSP database to search herb names of Isatidis radix and Isatidis folium, respectively, 48 chemical components were obtained by selecting OB ≥30% and DL ≥0.18. The chemical components of the 2 were analyzed and compared by using the Venny2.1.0 database, and 28 potential effective components were obtained by removing the repetitive chemical components and the noncorresponding target components, as shown in Figure 2(A). The specific components’ information is shown in Table 1.
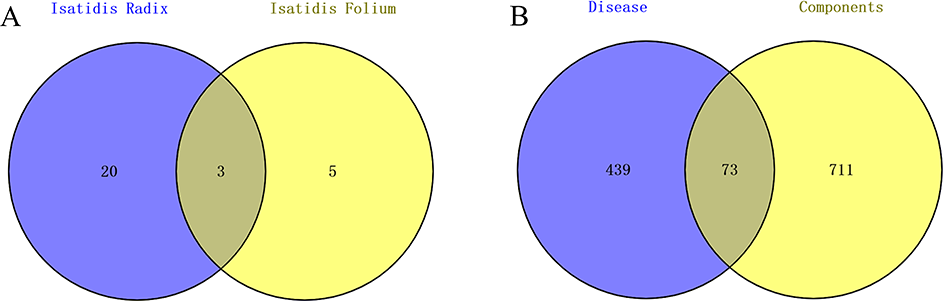
Venn diagram of (A) chemical components and (B) coincidence targets.
Potential Active Components of Fufang Banlangen Keli.

Abbreviations: DL, drug likeness; OB, oral bioactivity.
Collection of Potential Targets
The targets of the selected active components were compared with COVID-19 and SARS by Venny2.1.0 database, as shown in Figure 2(B). According to the analysis, 73 potential targets for the treatment of COVID-19 and SARS were identified and the details are shown in Table 2.
Potential Target Genes.

Target Biological Function Analysis
The potential targets were imported into DAVID and KOBAS 3.0 databases for enrichment analysis. The KEGG pathway annotation showed that pathways in cancer, NOD-like receptor signaling pathway, and pancreatic cancer were at the top of the list. And a total of 20 pathways (P < 0.05) were directly or indirectly related to COVID-19 and SARS, which were in line with the characteristics of multitarget and multipathway therapy for diseases with TCM, as shown in Figure 3(A). Under the condition of P < 0.05, 257 items of biological process (BP), 38 items of cell composition (CC), and 42 items of molecular function (MF) were obtained by GO enrichment analysis. The top 20 BP enrichments were response to organic substance, regulation of locomotion, and regulation of cell migration, etc, as shown in Figure 3(B). Figure 3(C) showed that the top 20 of CC enrichment are cell surface, cell fraction, plasma membrane part, etc. The top 20 MF enrichments are peptidase activity, acting on

(A) KEGG pathways annotation; (B) GO BP enrichment; (C) GO CC enrichment; and (D) GO MF enrichment. BP, biological process; CC, cell composition; GO, gene ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; MF, molecular function.
Construction of “C-T-P” Network
The 28 active components, 73 gene targets, and the top 20 pathways were imported into Cytoscape3.6.2 software, and the “C-T-P” network was obtained as shown in Figure 4. Purple represents active components, pink represents potential targets, and green represents pathways. Through the software plug-in, the node is visualized by degree value, and the node size is proportional to degree value. According to the requirements of topological parameters, the key nodes were determined by degree values greater than twice the median so as to obtain potential active components for later molecular docking tests.

Active components-targets-Kyoto Encyclopedia of Genes and Genomes pathways network.
Construction of PPI Network
The top 30 potential targets obtained by Cytoscape 3.6.2 software were imported into the STRING database together with ACE2. The protein interaction network diagram was obtained through a plug-in, and the relationship between gene targets and ACE2 was analyzed, the results were shown in Figure 5.
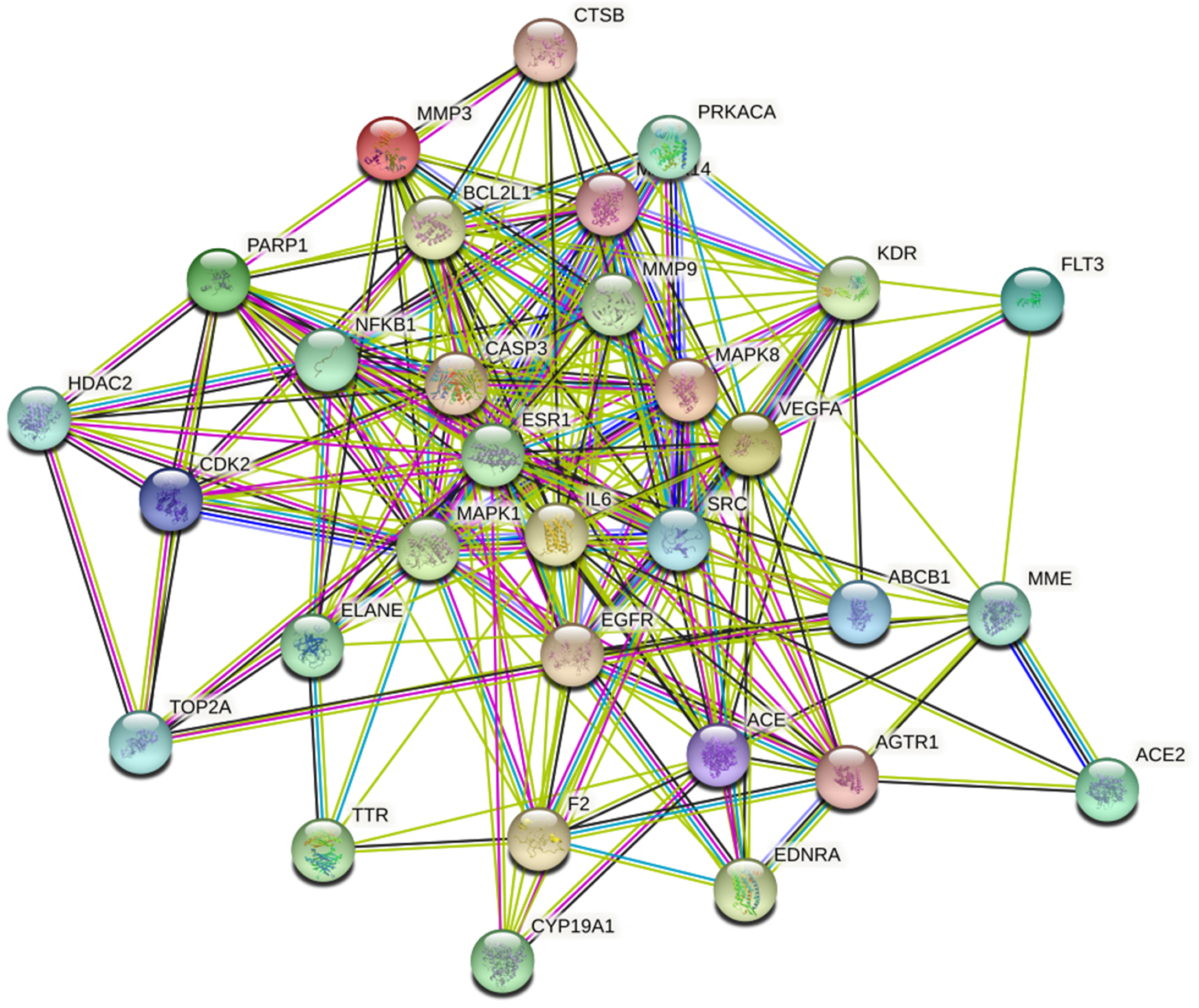
Protein-protein interaction network diagram.
Molecular Docking and Analysis
The components with a high degree were docked with SARS-COV-2 3 Cl and ACE2, respectively. It is generally believed that the energy required for the binding of the receptor to the ligand is inversely proportional to the stability of the binding, and the related docking results are shown in Table 3. Results showed that qingdainon, sinensetin, and acacetin had a strong binding ability with SARS-COV-2 3 Cl and ACE2, and (2Z)-2-(2-oxoindolin-3-ylidene) indolin-3-one had a strong binding ability with ACE2. And the docking mode, functional groups, and protein residues were displayed via PyMol software. The docking patterns of components with SARS-COV-2 3 Cl are shown in Figure 6, and the docking results with ACE2 are shown in Figure 7.
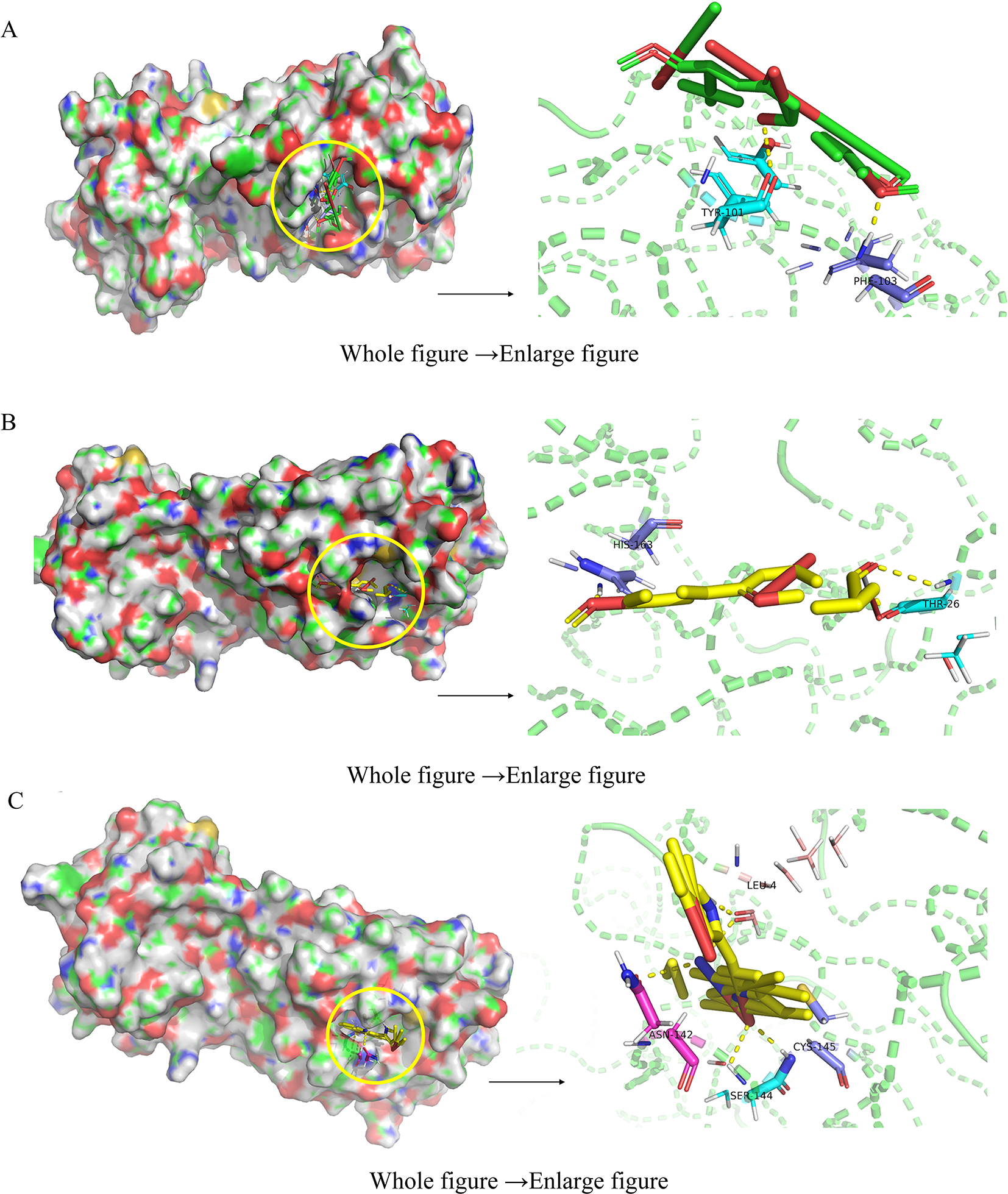
Three-dimensional docking mode diagram of SARS-CoV-2 3 Cl with (A) sinensetin; (B) acacetin; and (C) qingdainon.

(A) Three-dimensional docking mode diagram of angiotensin converting enzyme 2 (ACE2) with (A) sinensetin; (B) (2Z)-2-(2-oxoindolin-3-ylidene) indolin-3-one; (C) qingdainon; and (D) acacetin.
Binding Affinity of Core Components With SARS-CoV-2 3 Cl and ACE2.

Discussion
At the end of 2019, a novel coronavirus pneumonia caused by SARS-COV-2 swept the world, bringing unprecedented challenges to human survival and life. Patients with COVID-19 mainly have fever, cough, dyspnea, and other symptoms. Exudative edema may occur in the lungs, which leads to disturbance of pulmonary microcirculation 8 and imbalance of immune response. The overexpression of tumor necrosis factor-alpha, interleukin-6, and other inflammatory factors leads to “cytokine storm,” 9 which causes acute lung injury and damages other vital organs of the human body. FBK has the function of dispersing wind and heat, clearing lungs, and cooling blood. Isatidis radix and Isatidis folium have anti-inflammatory and antivirus effects; the former can also enhance the immune capacity of the body. FBK was used to prevent infection during the SARS outbreak, and it is now reported to have a significant inhibitory effect on SARS-CoV 3 Cl in vitro. 10 Therefore, it is of great significance to study the active components and mechanism of FBK in the treatment of COVID-19 and SARS.
The structure of SARS-CoV-2 and SARS-CoV are highly similar, and they both attack the human body by binding human ACE2 protein. 11 The SARS-CoV-2 protein coat had spikes S1 and S2, spikes S1 mediated the binding of the virus to the antibody on the surface of the host cell, and S2 fused with the host cell membrane after the binding of S1 to the host cell membrane and entered the host cell. 12 SARS-CoV-2 is the binding of S1 and ACE2 receptors into cells, 13 negatively regulating Ang II levels, 14 thus upsetting the balance between Ang II and Ang 1‐7, and overactivating Ak1-α receptors, resulting in increased lung vascular permeability and pulmonary edema leading to the damage to the lungs. 15,16 SARS-CoV-2 3 Cl mediates the hydrolysis of replicase peptides into functional proteins, which plays an important role in viral replication. Therefore, using network pharmacology, we could screen out the effective components, the potential targets, and the KEGG pathways to construct the network of “C-P-T” and analyze the topological parameters to obtain the potential active components. Molecular docking of the screened components with SARS-COV-2 3 Cl and ACE2 were carried out to investigate their docking activities.
Using network pharmacology, 28 potential active ingredients and 73 potential targets for the treatment of COVID-19 and SARS were screened through the TCMSP database. Through DAVID and Kobars3.0 database analysis, a total of 20 signal pathways (P < 0.05) were directly or indirectly related to COVID-19 and SARS. Among them, pathways in cancer, NOD-like receptor signaling pathway, and pancreatic cancer are closely related to tumors, which was in line with the characteristics of Chinese medicine treating diseases through multiple targets and multiple pathways. At the same time, we found that the top-ranked compounds qingdainon, sinensetin, and acacetin had strong binding ability with SARS-CoV-2 3 Cl and ACE2 through the topological parameters and molecular docking results of the “C-T-P” network diagram; (2Z)-2-(2-oxoindolin-3-ylidene)indolin-3-one had a strong binding force with ACE2. Finally, we use PyMol software to show the docking mode, compound functional groups, and protein residues between them so as to provide data reference for later related research.
The main direction of treatment for COVID-19 is to block the entry of the virus into the cell and the biotransformation of the virus, 17 so it is predictable that qingdainone, sinensetin, acacetin, and (2Z)-2-(2-oxoindolin-3-ylidene) indolin-3-one blocks the entry of the virus into cells by binding to ACE2 hydrogen bonds. The combination of qingdainone, sinensetin, and acacetin with SARS-CoV 3 Cl inhibited the transformation of virus to cure the disease. Studies 18 have shown that sinensetin reduced the expression of nuclear factor kappa-light-chain-enhancer of activated B cells (NF-kB) by inactivating extracellular signal-regulated kinase 1/2 mitogen-activated protein kinase (MAPK), and P38 MAPK signal pathways. Sinensetin could bind to Toll-like receptors, so as to induce autophagic cell death by p53-related MAPK/mammalian target of rapamycin signal transduction. 19 It causes the cells attacked by the virus to die and keeps the body stable. According to the literature, acacetin can inhibit the activity of SARS-CoV 3CL 20 and block the signal of IL-6 to reduce the level of IL-6 to play the role of antivirus activity, so as to reduce the lung injury of COVID-19. 21 It is reported that Isatidis Radix can reduce the level of IL-6 in inflammation caused by endotoxin, which further proves that component Radix Isatidis granule can be used to treat COVID-19. 22 At present, there are few literatures about the pharmacology of qingdainone and (2Z)-2-(2-oxoindolin-3-ylidene) indolin-3-one and the mechanism of their activity needs further research and exploration.
Taken together, the potential active components and mechanisms of FBK in the treatment of COVID-19 and SRAS were predicted and analyzed from many aspects via the method of network pharmacology and molecular docking. But the results should be further verified by in vitro and in vivo experiments.
Conclusion
Both COVID-19 and SARS are a serious threat to people’s health and life. Using network pharmacology and molecular docking, we analyzed the active components and comprehended the pharmacological mechanism of FBK against COVID-19 and SARS. These findings provide a strong theoretical basis for further research related to the anti-COVID-19 and SARS action of FBK.
Footnotes
Acknowledgments
We sincerely thank all colleagues from Key Laboratory of Traditional Chinese Medicine Resources and Chemistry of Hubei Province, Wuhan, China, who advised on the process of writing the paper.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (No. 81573865).
