Abstract
Objective
To explore the anti-COVID-19 active components and mechanism of Compound Houttuynia mixture by using network pharmacology and molecular docking.
Methods
First, the main chemical components of Compound Houttuynia mixture were obtained by using the TCMSP database and referring to relevant chemical composition literature. The components were screened for OB ≥30% and DL ≥0.18 as the threshold values. Then Swiss Target Prediction database was used to predict the target of the active components and map the targets of COVID-19 obtained through GeneCards database to obtain the gene pool of the potential target of COVID-19 resistance of the active components of Compound Houttuynia mixture. Next, DAVID database was used for GO enrichment and KEGG pathway annotation of targets function. Cytoscape 3.8.0 software was used to construct a “components-targets-pathways” network. Then String database was used to construct a “protein-protein interaction” network. Finally, the core targets, SARS-COV-2 3 Cl, ACE2 and the core active components of Compound Houttuyna Mixture were imported into the Discovery Studio 2016 Client database for molecular docking verification.
Results
Eighty-two active compounds, including Xylostosidine, Arctiin, ZINC12153652 and ZINC338038, were screened from Compound Houttuyniae mixture. The key targets involved 128 targets, including MAPK1, MAPK3, MAPK8, MAPK14, TP53, TNF, and IL6. The HIF-1 signaling, VEGF signaling, TNF signaling and another 127 signaling pathways associated with COVID-19 were affected (P < 0.05). From the results of molecular docking, the binding ability between the selected active components and the core targets was strong.
Conclusion
Through the combination of network pharmacology and molecular docking technology, this study revealed that the therapeutic effect of Compound Houttuynia mixture on COVID-19 was realized through multiple components, multiple targets and multiple pathways, which provided a certain scientific basis of the clinical application of Compound Houttuynia mixture.
Since December 2019, there has been a global outbreak of COVID-19 caused by the SARS-COV-2 virus, with rapid disease progression and high rates of infection and fatality. After the patient was infected with SARS-COV-2, the first symptoms were mostly fever, fatigue and dry cough. 1 Respiratory symptoms are common, and a few digestive ones such as diarrhea and nausea can be seen. 2 Severe cases may suffer from dyspnea, respiratory distress syndrome and septic shock. 3 Currently, no specific drug is available. 4 The novel Coronavirus infection diagnosis and treatment plan (trial) constantly updated by the National Health Commission has always emphasized the combination of Chinese and Western treatment and the antiviral effect of traditional Chinese medicine should be brought into full play. 5
Compound Houttuynia mixture (CHM) is prepared from Herba Houttuyniae, Radix Scutellariae, Radix Isatidis seu Baphicacanthi, Fructus Forsythiae, and Flos Lonicerae by extraction, concentration, alcohol precipitation and preparation. It has the effect of clearing away heat and detoxifying, and is used for throat pain caused by exogenous wind-heat, as well as acute pharyngitis and tonsillitis accompanied by wind-heat syndrome. 6 Modern clinical studies have shown that CHM can be used for the treatment of children with herpangina, 7 mycoplasma pneumonia, 8 and acute upper respiratory infection, 9 and not only for the treatment of hand-foot-mouth disease. 10 Currently, it has been included in the “Novel Coronavirus Prevention and Control Technical Guide of Traditional Chinese Medicine in Sichuan Province” issued by The Administration of Traditional Chinese Medicine of Sichuan Province, which is applicable to the treatment of the syndrome of wind-heat invading lung and excess heat obstructing lung.
Network pharmacology is based on the rapid development of omics and big data. It integrates systems biology, bioinformatics and other emerging cross-disciplines to understand the relationship between drugs and the body from the perspective of improving or restoring the balance of the biological network, and to analyze the biological system network as a whole. The application of network pharmacology to the study of TCM components has unique advantages, because on the 1 hoursand, the integrity and systematic characteristics of network pharmacology are consistent with the theory of “overall concept” and “treatment based on syndrome differentiation.” On the other hand, its research model has changed from “single drug and single target” to “disease-gene-target-drug,” which is similar to the synergistic characteristics of “multiple components, multiple pathways and multiple targets” of TCM components. In addition, molecular docking techniques can study intermolecular interactions (such as ligands and receptors) and predict their binding patterns and affinity. This study aims to explore the chemical composition and mechanism of CHM for the treatment of COVID-19 from the molecular level with the help of network pharmacology and molecular docking technology, so as to provide a scientific basis for the use of CHM in the treatment of COVID-19.
Materials and Methods
Collection, Screening and Targets Acquisition of Active Components
Information on the active components in CHM was collected through the systematic pharmacology analysis platform database of Chinese medicine (TCMSP, http://lsp.nwu.edu.cn/tcmsp.php), and related chemical components were collected in combination with literature as a supplement. Then, the active components were screened based on the threshold of oral bioavailability (OB) ≥30% and drug likeness (DL) ≥0.18. 11 Finally, the SMILE structural formula of each active component was collected on Pubchem (https://pubchem.ncbi.nlm.nih.gov/) website, and imported into SwissTargetPrediction (http://www.swisstargetprediction.ch/) website for acquiring its targets.
Acquisition of Disease Targets and Potential Targets of Anti-COVID-19 in CHM
In the GeneCards database (https://www.genecards.org/), “novel coronavirus” and “novel coronavirus pneumonia” were used as keywords to retrieve COVID-19-related genes. Then the potential targets of active components after deduplication point genes were compared to obtain the potential targets of CHM against COVID-19.
Gene Ontology and Pathway Enrichment Analysis
The potential targets obtained were uploaded to the DAVID database (https://david.ncifcrf.gov/) for GO enrichment and KEGG pathway annotation. The pathways for the enrichment of active components targets were considered to be important regulatory pathways for CHM in the treatment of COVID-19. The top 20 pathways and go enrichment results obtained by P < 0.05 were further analyzed on the omicshare platform.
Construction of “Components-Targets-Pathways” Network
The active components, targets and enrichment pathways were imported into Cytoscape 3.8.0 software to construct the “components-targets” network and “targets-pathways” network, respectively. Among them, different nodes were used to represent the above three, and edges were used to represent the relationship between the different nodes. Then the “components-targets-pathways” network was constructed by using the merge function. Finally, the components with high degree in the network were used as important active components.
“Protein-Protein Interaction (PPI)” Network Construction and Molecular Docking
The targets were imported into the STRING database to obtain the targets interaction network by setting the species as “Homo sapiens,” and the TSV format file of the network was downloaded. Then it was imported into Cytoscape 3.8.0 software for analysis by using the network analysis function. For the targets with a higher degree value in the network, the corresponding 3D structure was downloaded in the PDB database. Finally, the targets and active components’ structures were imported into Discovery Studio 2016 Client for molecular docking verification.
Result
Collection, Screening and Targets Acquisition of Active Components
Through the TCMSP database and related literature, we collected and screened 128 active components in CHM; 7 of them were from Herba Houttuyniae, 36 from Radix Scutellariae, 39 from Radix Isatidis seu Baphicacanthi, 23 from Fructus Forsythiae, and 23 from Lonicerae Japonicae Flos. After keeping the only components and removing those lacking target prediction data, 82 active components were obtained (The basic information on some active components is shown in Table 1). A total of 1031 targets were obtained from CHM after removing the duplicates through the SwissTargetPrediction database.
Basic Information of Some Active Components in CHM.

Potential Targets of Anti-COVID-19 in CHM
In total, 791 potential targets of COVID-19 were obtained by searching, summarizing and removing duplicates in the GeneCards database. Then the Venn diagram was used to map them with the 1031 targets of the active components. For the treatment of COVID-19, 128 potential targets of CHM were obtained, as shown in Figure 1. See Table 2 for some targets’ information.

The relationship between targets of active components in CHM and COVID-19 related targets.
Some Potential Targets for the Treatment of COVID-19 in CHM.

Targets’ Function and Pathway Annotation
GO enrichment analysis included biological process (BP), cell component (CC), and molecular function (MF). The top ones in BP were protein phosphorylation, peptidyl-serine phosphorylation and platelet activation (Figure 2(A)). The top in CC were cytosol, cytoplasm and extracellular exosome (Figure 2(B)), and in MF, protein binding, protein serine/threonine kinase activity and protein phosphatase binding (Figure 2(C)).
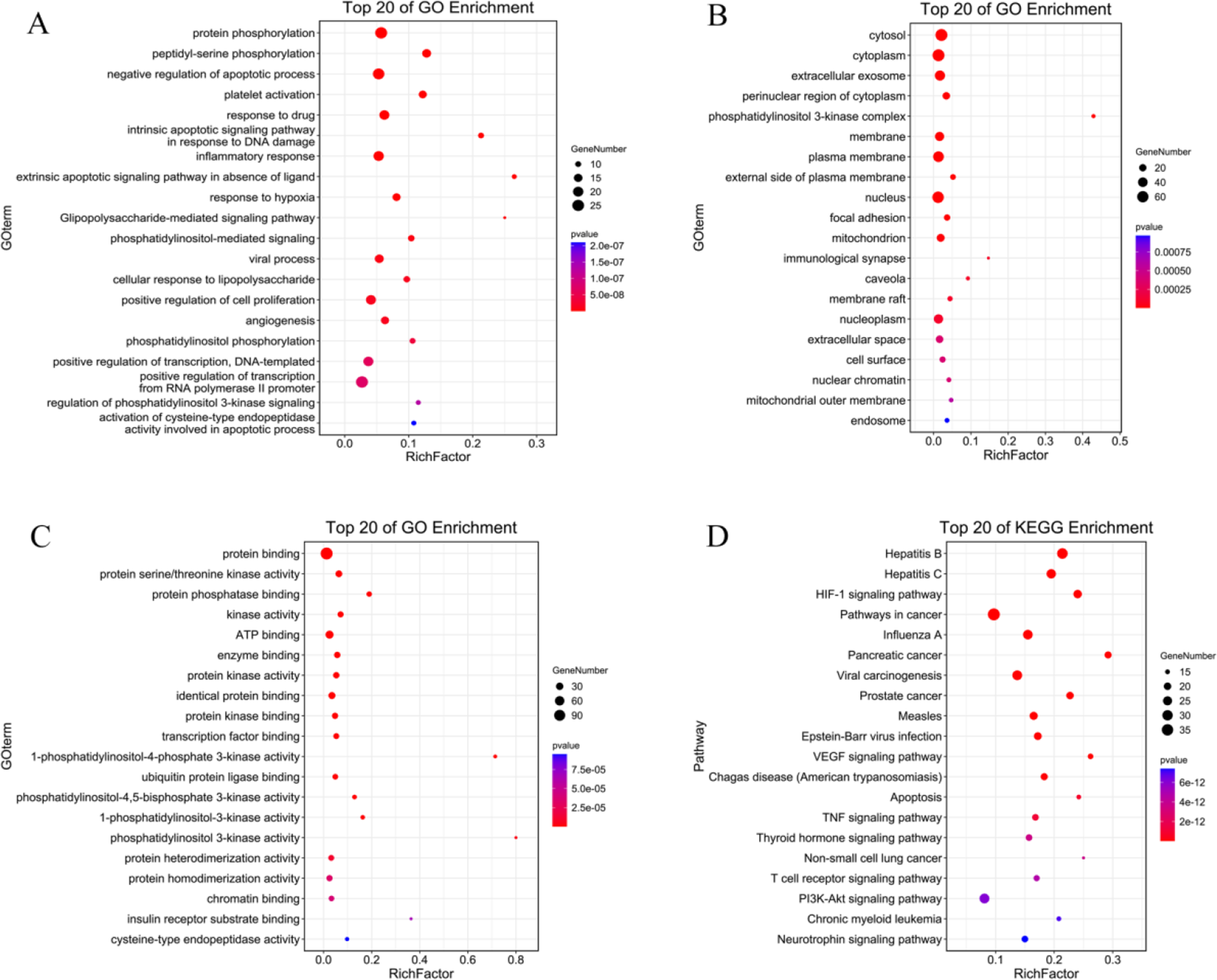
The top 20 ones of GO and KEGG enrichment analysis of CHM in COVID-19 treatment. (
KEGG pathway annotation results showed that 128 potential targets were involved in 132 pathways, of which 127 were significantly related to targets (P < 0.05). The top 20 pathways were associated with the HIF-1 signaling, VEGF signaling and TNF signaling pathways (Figure 2(D)).
Construction of “Components-Targets-Pathways” Network
A network diagram of "Components-Targets-Pathways" was constructed for the screened 82 active components, 128 targets and the first 20 pathways sorted by the P value in ascending order, as shown in Figure 3. There were 230 nodes (82 active components, 128 targets, and 20 pathways) and 1519 edges in this network diagram. It showed that CHM exerted its anti-COVID-19 effect through multiple components, multiple targets, and multiple pathways coordinated regulation. Xylostosidine, Arctiin, ZINC12153652, and ZINC338038 with higher degree values obtained from the network diagram would be used for molecular docking verification. Their structural formulas are shown in Figure 4.
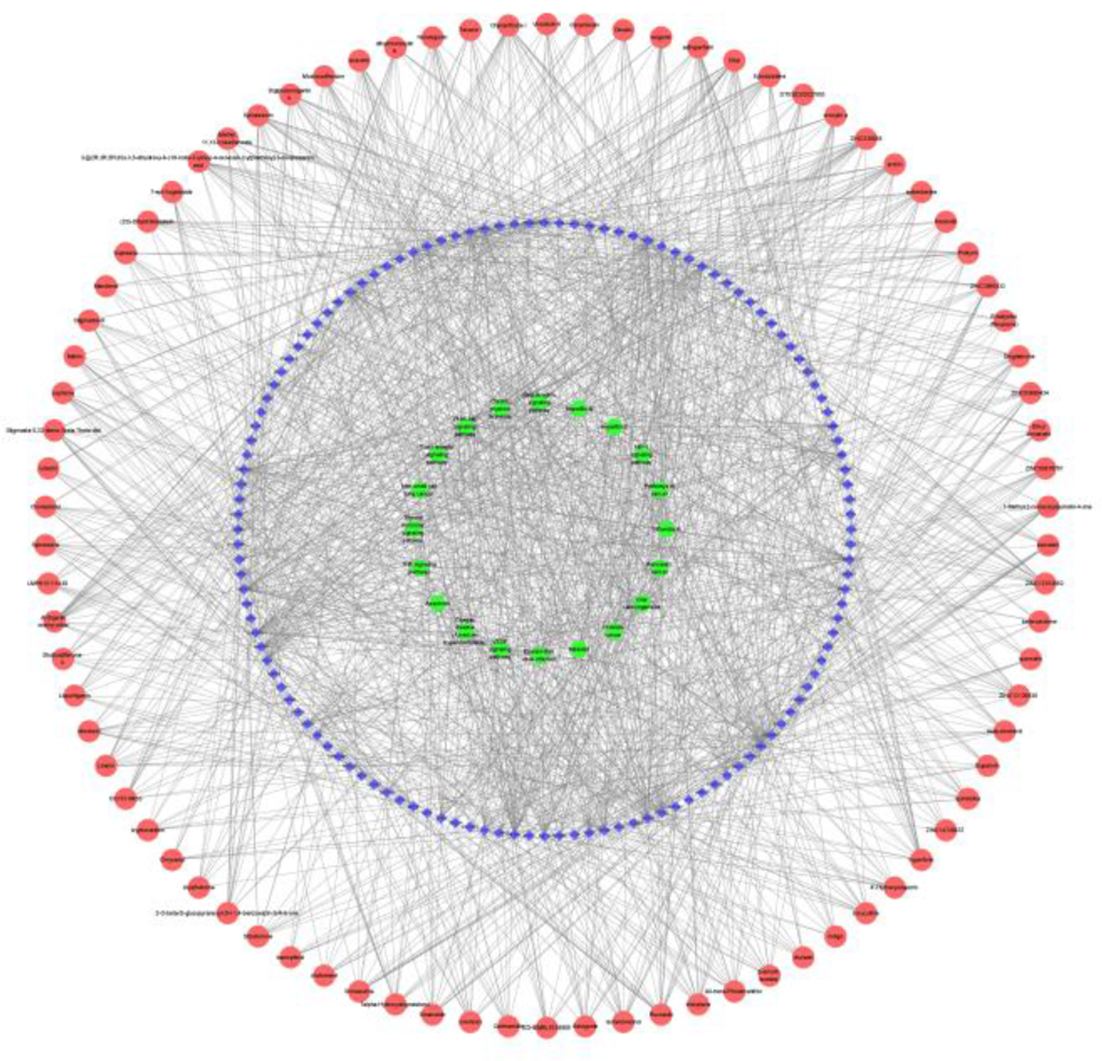
“Components–targets–pathways” network. The red, blue, and green nodes respectively represent the active components, targets and pathways.

The structural formulas of the anti-COVID-19 active components screened from CHM and used for molecular docking verification. (
PPI Network Construction and Molecular Docking
The interaction network of the corresponding targets was obtained by importing 128 targets into the STRING database (Figure 5(A)), and it was further analyzed through Cytoscape 3.8.0 software. Finally, MAPK1 and TNF with higher degree were selected as the targets for molecular docking verification. In addition, it is reported in the literature that the two targets of ACE2 and SARS-COV-2 3 Cl are closely related to the new coronary pneumonia, so these two proteins and active components were also tested for molecular docking. 12 Figure 5(B) shows the interaction network between ACE2 and the top 20 targets in the PPI network.

“Protein–protein interaction” network. (
The 3D structures of the corresponding proteins were downloaded in the PDB database. Then the 3D structure of the proteins and the components were imported into Discovery Studio 2016 Client software for molecular docking. Before docking, the water molecules, hydrogen bonds and ligands in the protein structure needed to be removed. Then the binding site between the components and the proteins were also needed to be found and the sphere radius were set to 10. The docking information is shown in Table 3. The 3D and 2D views of components and proteins docking are shown in Figures 6 and 7.
Partial Results of Molecular Docking.


Molecular docking results of xylostosidine and arctiin with each protein. (

Molecular docking results of ZINC12153652 and ZINC338038 with each protein. (
When the docking score between the components and the proteins was greater than 100, it is believed that they have a better binding ability. 13 The higher the docking score, the stronger the binding ability. The docking scores between these 4 components and 4 proteins were all greater than 100, and the binding score between Arctiin and SARS-COV-2 3 Cl was the highest, reaching 139.483. Besides, the most important chemical bonds between the components and the proteins were conventional hydrogen bonds and carbon hydrogen bonds (Table 3). These results indicated that the predicted active components were likely to bind to the predicted key targets, and indirectly proved the reliability of the predicted results of network pharmacology.
Discussion
CHM is made of Herba Houttuyniae, Radix Scutellariae, Radix Isatidis seu Baphicacanthi, Fructus Forsythiae, and Flos Lonicerae. Among them, Herba Houttuyniae is used as the emperor medicine, which has the effects of clearing heat and detoxification, diuresis and dehumidification, clearing heat and stopping diarrhea. Radix Scutellariae is taken as a minister medicine, which has functions of clearing away heat and dampness, purging fire and detoxification. The rest are adjuvants, which together help to exert their effects. 14 Modern pharmacological research shows that CHM has antibacterial, antiviral, improving immunity, diuresis and other effects. 15 In TCM theory, COVID-19 belongs to the category of epidemic disease. The disease is located in the lungs. The basic pathogenesis is characterized by “dampness, heat, poison, and blood stasis.” TCM has unique advantages in dialectical treatment. In this study, network pharmacology and molecular docking were used to analyze systematically the potential treatment mechanism of CHM on COVID-19.
As a result, we screened out 82 active components in CHM through the TCMSP database and performed target predictions. By intersecting with COVID-19 related targets, 128 targets of CHM for the treatment of COVID-19 were obtained. Go enrichment and KEGG annotation were performed on them to obtain enrichment pathways of targets, and then the “components-targets-pathways” network diagram was analyzed to obtain the core chemical components of anti-COVID-19, such as Xylostosidine, Arctiin, ZINC12153652 and ZINC338038, which were used for subsequent molecular docking verification. One study showed that a compound cocktail composed of arctiin, daidzein, glycyrrhizic acid, and liquiritin inhibited mouse pneumonia resulting from a PR8 viral infection and caused a weight gain after oral administration. 16 This also verified the reliability of network pharmacology from the side.
Through PPI network analysis, we found that MAPK1, MAPK3, MAPK8, TP53, TNF, IL6 and other targets were the main targets of CHM in the treatment of COVID-19. MAPK1, MAPK3, and MAPK8 belong to the family of mitogen-activated protein kinases (MAPK). After being activated, they participate in the biological processes of cellular immune regulation, 17 inflammation, 18 cell proliferation, 19 apoptosis, 20 and the signal transduction of various stress responses. 21 Tumor protein p53 (TP53) is a cell cycle-related gene. It mainly synthesizes cell cycle-related proteins through transcription and plays an important role in the regulation of cell proliferation and apoptosis. The normal p53 gene is wild-type (wt p53), which plays an important regulatory role in the normal growth of cells, 22 and obviously inhibits the transformation and growth of cells. Once the p53 gene is lost or mutated, it is called mutant (mt p53), which induces a variety of cancers. 23 -25 Tumor Necrosis Factor-α (TNF-α) is a pro-inflammatory factor mainly produced by activated monocytes and macrophages, and is a key factor in the process of inflammatory reaction. It can participate in the physical immune response by enhancing the immune activity of neutrophils and cytotoxic T cells, and plays an essential role in the body’s resistance to pathogen infection. 26,27 Interleukin-6 (IL6) is a multi-functional, multi-effect and multi-directional cytokine secreted by T cells, B cells, monocytes, macrophages, fibroblasts, vascular endothelial cells and some tumor cells. It participates in a variety of inflammatory and immune response processes, and plays a major role in the occurrence and development of tumors. 28 -30
In order to verify further the feasibility and reliability of network pharmacology, we molecularly docked the top 4 components with MAPK1, TNF, SARS-CoV-2 3 Cl, and ACE2. The results showed that the components and the targets both had better ability to combine, thus confirming this.
Besides, enrichment analysis, the KEGG pathway showed that the signal pathways of CHM in the treatment of COVID-19 include the HIF-1, VEGF, and TNF signal pathways. Hypoxia inducible factor (HIF) is composed of 2 subunits of HIF-1α and HIF-1β. Under hypoxia, it could activate the transcription of a variety of target genes, and plays an important role in the growth and development of cells, tissues, physiological stress and certain pathological processes. Studies have reported that the role of HIF-1 in the inflammatory response was complex and diverse, and had a significant impact on the inflammation-related diseases and progression of cancer in immune and non-immune cells. 31 Huang et al. found that the HIF-1 signaling pathway might be one of the key pathways for the treatment of COVID-19. 32 In addition, because of the immunomodulatory effects, HIF-1α inhibition through pharmacological strategies might provide a new approach to aid the treatment of patients affected with COVID-19. 33 Vascular endothelial growth factor (VEGF) is a highly specific cytokine that could promote the division of vascular endothelial cells, induce blood vessel formation and increase blood vessel permeability. In addition, reports have shown that VEGF plays a vital role in the process of lung inflammation. 34,35 Besides, VEGF was deemed a promising therapeutic target in suppressing inflammation during SARS-CoV-2 infection with neurological symptoms. 36 Tumor necrosis factor (TNF) is a cytokine with a wide range of biological activities produced by multinucleated giant cells. On the 1 hoursand, it has the ability to regulate the bodies’ immune function and could cause necrosis of certain tumor cells. 37 On the other hand, it mediated pathophysiological reactions such as inflammation, 38 tissue damage, 39 and shock. 40 One study proved a role for blockade of TNF-α in the treatment of COVID-19 inflammatory cascade. 41 Therefore, we speculate that CHM may treat COVID-19 by inhibiting inflammation and regulating the immune function of the body.
Conclusion
Taken together, this study screened out the active components and targets of CHM for the potential treatment of COVID-19 through the method of network pharmacology, and verified them with molecular docking technology, which provided a certain scientific basis for in-depth understanding of its treatment mechanism and clinical application.
Supplemental Material
Online supplementary file 1 - Supplemental material for Mechanism of Compound Houttuynia Mixture as an Anti-COVID-19 Drug Based on Network Pharmacology and Molecular Docking
Supplemental material, Online supplementary file 1, for Mechanism of Compound Houttuynia Mixture as an Anti-COVID-19 Drug Based on Network Pharmacology and Molecular Docking by Xing-Pan Wu, Tian-Shun Wang, Zi-Xin Yuan, Yan-Fang Yang and He-Zhen Wu in Natural Product Communications
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the esearch, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
