Abstract
Background
Curcumae Radix (CR), derived from the dry roots of Curcuma longa L., family Zingiberaceae, is widely used to treat depression. However, the ingredients and mechanisms of CR are still unclear. The purpose of this study was to solve this problem using network pharmacology and molecular docking.
Methods
The active ingredients of CR were screened through TCMSP, and the depression-related genes were obtained through the Genetic Association, GeneCards, and OMIM databases. Then, DisGeNET score was performed to evaluate the correlation between co-genes and depression. Topological analysis was conducted to screen hub genes and proteins, molecular docking was performed to evaluate the binding ability of the hub protein with active ingredients, and gene ontology (Go) function analysis, gene tissue localization, and KEGG pathway analysis were conducted to explore the function and location of genes, as well as the mechanism of CR for treating depression.
Results
Eight ingredients of CR were screened based on pharmacokinetic properties, five of which are closely related to depression, including (E)−5-hydroxy-7-(4-hydroxyphenyl)−1-phenyl-1-heptene, (E)−1,7-diphenyl-3-hydroxy-1-hepten-5-one, oxycurcumenol, β-sitosterol, and sitosterol. They interacted with 45 co-genes and co-proteins with a DisGeNET score ≥0.3. AR, NOS2, PTGS2, and TYK2 were pivot genes. EGFR, PTGS2, HSP90AA1, MAPK8, and ESR1 were hub proteins. PTGS2 was found to have good binding potential with oxycurcumenol, (E)−1,7-diphenyl-3-hydroxy-1-hepten-5-one and (E)−5-hydroxy-7-(4-hydroxyphenyl)−1-phenyl-1-heptene. Go functional analysis indicated that co-genes involved complex biological processes, cellular components and molecular functions. PER2, P2RX7, GRM1, TACR1, MAPK8, HCRTR1, EGFR, and TYK2 were highly expressed in the prefrontal cortex. The potential pathways for CR to exert antidepressant effects were calcium, estrogen, PI3K-Akt and ErbB signaling pathways.
Conclusions
This study revealed the ingredients, effective targets and mechanisms of CR in the treatment of depression, which provides a new perspective for the development of new antidepressants.
Keywords
Depression is a common worldwide mental disorder characterized by the persistence of negative emotions and the diminishment of positive emotions, which could cause serious functional and social disorders. 1 Previous studies have also found that depression is a major cause of global disability and is related to a 10-year reduction in life expectancy. 2 In addition, a recent study found that 50.4% of health care workers exposed to COVID-19 in Wuhan reported depressive symptoms, 3 and Vietnamese patients with COVID-19 were found to have a higher risk of depression (OR 2.88) and lower quality of life than those without COVID-19. 4 This is undoubtedly a heavy physical and mental burden for more than 150 million patients diagnosed with COVID-19 worldwide. What is worse, a recent network meta-analysis which comprehensively summarized the clinical effectiveness of antidepressants found that the benefits of all antidepressants were small compared to placebo, 5 especially in adolescents and children. 6 Therefore, seeking reliable, effective and safe antidepressant drugs has become an urgent requirement for doctors and patients with depression.
Chinese herbal medicine, as one kind of complementary and alternative medicine, has been shown to be superior to placebo in reducing the HAMD-17 score [mean difference (MD) = −4.53, 95% confidence interval (CI):−5.69‐−3.77, P < 0.00001], and the total effective rate is better than that of Western conventional medications alone when combined with Western conventional medications [Risk Ratio (RR): 1.16, 95% CI: 1.07‐1.27, P = 0.0004], and with less adverse reaction. 7 CR, a Chinese herbal medicine derived from the dry roots of Curcuma longa L., family Zingiberaceae, has been found not only to treat COVID-19, 8 but also to have antidepressant effects. 9 For example, a clinical trial showed that CR plays a role in improving the negative emotions and psychological stress of patients with post-stroke depression and improves their quality of life. 10 Animal experiments have also proved that CR can shorten the time of forced swimming and tail suspension in a mice model of depression, and antagonize the hypothermia induced by reserpine. 11 However, no study has been conducted to explore the active ingredients, antidepressant targets and systemic pharmacological mechanisms of CR.
Chinese herbal medicine holds the characteristics of multi-ingredient, multi-target, multi-pathway and synergistic effects, leading to an uncertain material basis and unclear pharmacological mechanism. 12 Therefore, it is difficult to understand the underlying molecular mechanism of Chinese herbal medicines through routine experimental research. In order to address this thorny scientific problem, innovative strategies and methods are urgently needed to comprehensively and systematically elucidate the pharmacological mechanisms of their specific therapeutic efficacy. Network pharmacology is a new field of pharmacology and pharmacodynamics, which integrates multi-pharmacology, systems biology, and computational biology to explain the complex regulation of Chinese herbal medicines on human organisms. 13 It investigates the synergistic effects and potential mechanisms of multiple ingredients by constructing an ingredient-target network, protein-protein target network and drug-disease network. 12 For example, Liu et al. identified 121 active ingredients of Chaihu Shugan powder and 15 targets related to depression through network pharmacology, and predicted its antidepressant mechanism. 14 Lin et al. identified 17 active components, 144 gene targets and 143 protein targets, as well as four important signaling pathways of Drynariae Rhizoma in fracture repair through network pharmacology, reflecting the characteristics of multiple components, multiple targets and multiple pathways of Chinese herbal medicine in promoting fracture healing. 15
In this study, a comprehensive network pharmacology approach was conducted, through drug active ingredient screening, drug and disease target screening, active ingredient-gene network construction, protein-protein interaction network construction, Go functional analysis, KEGG pathway enrichment and gene tissue localization, to explore the active ingredients and complex pharmacological mechanisms of CR for depression (Figure 1). Besides, the DisGeNET score was used to assess the correlation between the common targets and depression, and molecular docking was performed to assess the binding potential of the active ingredients to the corresponding hub proteins. This study provides a valuable reference for exploring the active ingredients of CR and clarifying the mechanism of CR for treating depression, which will be beneficial to patients with depression, especially those affected by COVID-19.

The process of systematically elucidating the mechanism of Curcumae Radix in treating depression.
Materials and Methods
Screening Active Ingredients and Chemical Structures
Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform 16 (TCMSP, http://tcmspw.com/, Version 2.3) is a systematic pharmacology platform that can identify the relationship between Chinese herbal medicines, targets and diseases, and is used for new drug discovery. The platform also provides pharmacokinetic properties of natural compounds such as oral bioavailability (OB), drug-likeness (DL), aqueous solubility, and potential to cross the blood-brain barrier (BBB). A compound with a BBB from −0.3 to +0.3 is considered to be moderately penetrating, and greater than 0.3 indicates strong penetrating power. The potential active ingredients of CR were obtained from the TCMSP according to the pharmacokinetic properties of CR, including OB ≥30%, DL ≥0.18 and BBB ≥ −0.3. Then, the chemical structures and their Pubchem Cid of the potential active ingredients of CR were downloaded and stored in the mol2 format.
Protein and Gene Targets of CR’s Potential Active Ingredients
According to Pubchem Cid of the active ingredients, the corresponding SMILES were retrieved in the Pubchem database (https://pubchem.ncbi.nlm.nih.gov/). If there were no canonical SMILES in the Pubchem database, the SMILES of the active ingredients were obtained by searching for the chemical name in Zinc15 (http://zinc.docking.org/substances/home/). Next, the SMILES of the CR’s active ingredients were used to search for the relevant protein targets of the active ingredients in the Swiss Target Prediction (http://www.swisstargetprediction.ch/). The protein targets with a probability ≥0 were included, then the duplicate data were deleted, and the name of the protein targets and their Uniprot IDs were saved. Finally, the UniProt KB search function (http://www.uniprot.org/uniprot/) was used to obtain the gene targets of the CR’s active ingredient.
Screening of Co-Genes for Curcumae Radix and Depression
The following keywords were used to search for genes associated with depression in the Genetic Association Database (GAD, https://geneticassociationdb.nih.gov/), GeneCards database (http://www.genecards.org/), and OMIM database (http://www.ncbi.nlm.nih.gov/omim), such as depression, depressive, antidepressant, and depressed; duplicate data and false positive genes were removed, and then the gene targets of CR were matched to obtain the co-gene targets of CR’s active ingredients and depression, as potential gene targets for the treatment of depression with CR’s active ingredients.
DisGeNET Score of Co-Genes and Depression
The DisGeNET database 17 (http://www.disgenet.org/search, Version 5.0) is one of the platforms that contains the largest available genes and variants associated with human disease. In the database, disease information related to the input gene can be obtained, and genetic information related to the input disease can also be received. A confidence score (DisGeNET score) was performed to navigate the more than 400,000 gene-disease associations in the DisGeNET database, which reflects the recurrence of gene-disease associations across all data sources. 17 In the DisGeNET database, we used co-gene targets of CR’s active ingredients and depression to search, extracted the association results of depression in Mental Disorders, and screened genes with a DisGeNET score ≥0.3.
Construction of Active Ingredient-Gene Network
Cytoscape Version 3.6.0 is an open source software platform that integrates data integration, analysis and visualization and is now widely used to visualize molecular interaction networks and biological pathways. 18 We introduced the gene targets with DisGeNET score ≥0.3 and the active ingredients of CR into Cytoscape Version 3.6.0 software to construct a CR active ingredient-gene target network. Then, we performed a topology analysis to obtain the gene targets with the highest degree values, betweenness centrality and closeness centrality at the same time, as the pivot gene.
Construction of Protein-Protein Interaction (PPI) Networks
The String database (version 11.0, https://string-db.org/, updated January 19, 2019) collects, scores, and integrates all publicly available PPI information, including direct (physical) and indirect (functional) interactions. 19 The co-gene targets with DisGeNET score ≥0.3 were imported into the String database, the species was defined as ‘human’, and the PPI relationship was obtained. The medium confidence was set to 0.4 as the minimum required interaction score, and the updated result was saved in TSV format. Node1, node2 and the combined score were imported into Cytoscape software to draw the PPI network, and topology analysis was performed to obtain the protein targets with the top 5 degrees. The Generate style from statistics tool was used to set the protein targets format, including setting the node size and color to reflect the degree value, and setting the color of the edge to reflect the combined score, and then the PPI network was uploaded.
Molecular Docking
CB-Dock (http://cao.labshare.cn/cb-dock/) integrates the popular docking program Autodock Vina to predict the binding of the target protein to the compound, with an accuracy rate of 70%. 20 CB-Dock can automatically identify the protein-ligand binding site, calculate the center and size, customize the size of the docking box according to the query ligand, and then perform molecular docking with AutoDock Vina. The Uniprot ID of the five highest-degree proteins recognized by the PPI network was inputted into the UniProt database (https://www.uniprot.org/), then the corresponding PDB ID with the minimum resolution (Å) in the 3D structure databases was selected, and its protein structure downloaded in PDB format. Finally, CB-Dock was used to dock the CR active ingredients in mol2 format with the corresponding protein targets, and Schrodinger Suites (Version 2018‐1, Materials) was used to show the 3D and 2D map of the corresponding molecular docking.
Gene Ontology Function, KEGG Pathway, and Tissue Localization Analysis of Genes
The Database for Annotation, Visualization and Integrated Discovery (DAVID, https://david. ncifcrf.gov/, Version 6.8) provides systematic, comprehensive biofunctional annotation information for a large number of genes and proteins, enabling the identification of the most significantly enriched biological annotations. 21 We introduced depression-related genes with a DisGeNET score ≥0.3 into the DAVID database, set the “Select Identifier” to “official gene symbol,” set the “list type” to “gene list,” and defined the species as “homo sapiens” in the “background” and “list.” Then, we performed Go function, KEGG pathway and tissue localization analysis on the gene targets. The most important biological process (BP), cellular component (CC), molecular function (MF), KEGG pathway and tissue localization results were screened and stored according to a threshold P < 0.05. The BP, CC, and MF was designed using GraphPad Prism 7.0 software. The advanced bubble chart for the KEGG pathway was created with Omicshare Tools (http://omicshare.com/tools/Home/Soft/getsoft/type/index).
Brain-Related Gene Localization Analysis
Founded in 2008, BioGPS (http://biogps.org/) is a centralized gene annotation portal with a “Gene expression/activity chart” plugin that contains approximately 6,000 datasets, enabling researchers to access distributed gene annotation resources. 22 The gene targets identified above in the brain were input into BioGPS for specific distribution analysis, the raw data were downloaded, the expression value of each gene in each part was calculated, and the gene expression heat map was drawn using GraphPad Prism 7.0 software.
Integrated Signal Pathway
We imported the UniprotID of genes with DisGeNET score ≥0.3 into the KEGG mapper of the KEGG database (http://www.kegg.jp/), and set the species to humans, thus obtaining the signaling pathways of the genes. Finally, screening and integrating was carried out of the signaling pathways related to depression as the potential mechanism for CR treatment of depression.
Results
Screening of Active Ingredients of Curcumae Radix
There were 222 ingredients of CR in the TCMSP database. According to the pharmacokinetic properties of CR, including OB ≥30%, DL ≥0.18, and BBB ≥ −0.3, eight potential active ingredients were selected, as shown in Table 1 (β-sitosterol, sitosterol, (4aR,5R,8R,8aR)−5,8-dihydroxy-3,5,8a-trimethyl-6,7,8,9-tetrahydro-4aH-benzo[f]benzofuran-4-one, (E)−1,7-diphenyl-3-hydroxy-1-hepten-5-one, (E)−5-hydroxy-7-(4-hydroxyphenyl)−1-phenyl-1-heptene, oxycurcumenol, zedoalactone E, and 1,7-diphenyl-3-acetoxy-6(E)-hepten).
Main Active Ingredients in Curcumae Radix.

Co-Genes of Curcumae Radix and Depression
Except for (4aR,5R,8R,8aR)−5,8-dihydroxy-3,5,8a-trimethyl-6,7,8,9-tetrahydro-4aH-benzo- [f]benzofuran-4-one and 1,7-diphenyl-3-acetoxy-6(E)-hepten, six active ingredients of CR were able to be used to retrieve the corresponding protein targets in the Swiss Target Prediction database. A total of 239 proteins, and 239 corresponding genes were retrieved by UniProt database.
In terms of disease targets, 235 genes related to depression were retrieved from the GAD database, 24435 from the GeneCards database, and 256 from the OMIM database. Excluding the duplicate data, 7920 genes related to depression were obtained, and then matched with the gene targets of CR’s active ingredients (Supplemental Figure S1); 203 potential co-genes for CR treatment of depression were screened, as shown in Table 2.
Potential Gene Targets of Curcumae Radix in the Treatment of Depression.

DisGeNET Score Between Co-Genes and Depression
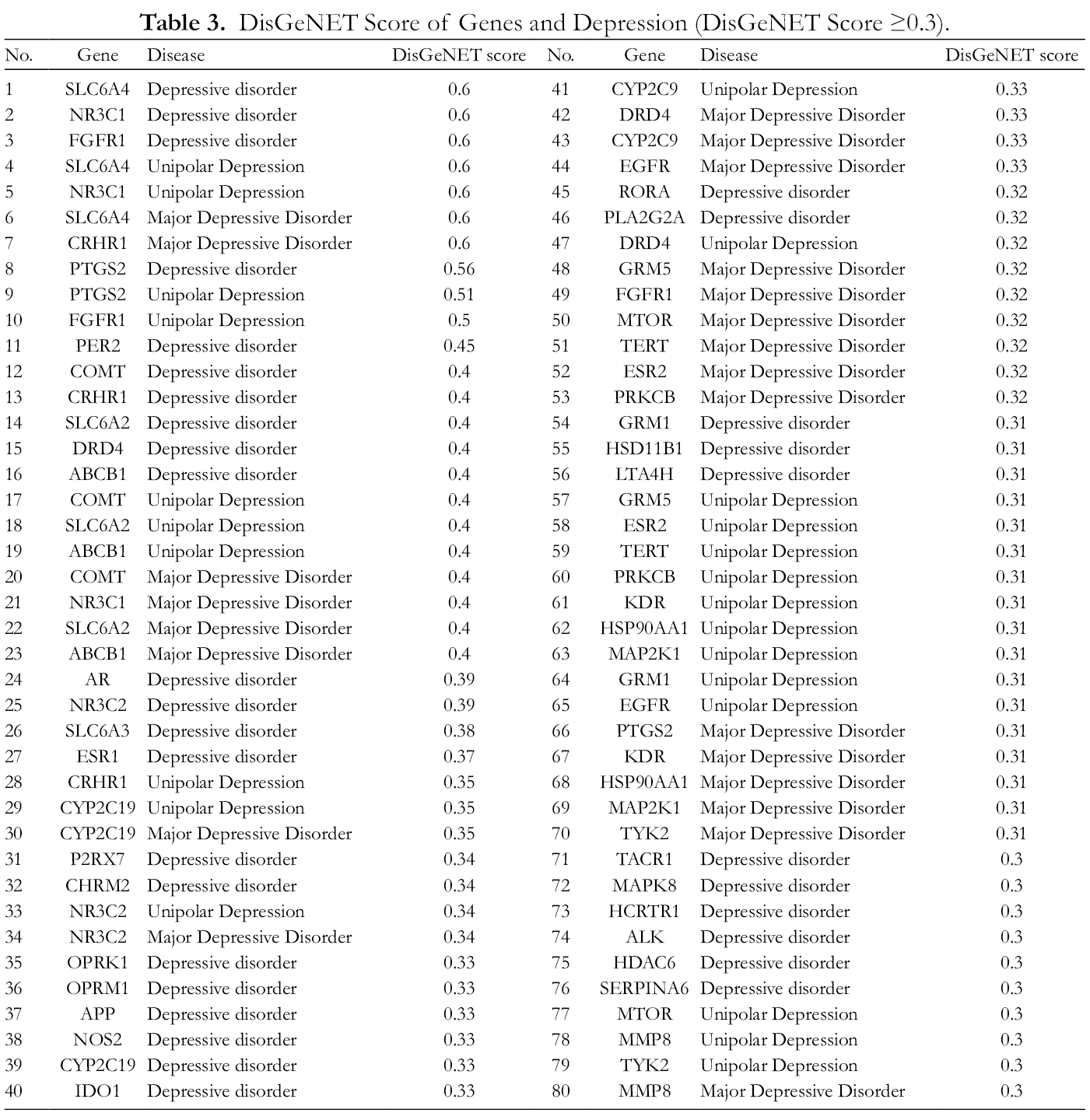
In order to identify genes strongly related to different depressions, we conducted a DisGeNET score to assess the correlation between co-genes and depression. In the DisGeNET database, 36,617 correlations between co-gene targets and depression were obtained, and 138 genes were associated with depressive disorder, unipolar depression, and major depressive disorder. Among them, 45 genes had a DisGeNET score ≥0.3, which may be those strongly related to depression. The detailed DisGeNET scores of these genes and different types of depression are shown in Table 3. There were seven genes with a DisGeNET score ≥0.6, of which three were related to depressive disorder: SLC6A4, NR3C1, FGFR1; two were associated with unipolar depression: SLC6A4, NR3C1, and two were related to major depressive disorder: SLC6A4, CRHR1.
DisGeNET Score of Genes and Depression (DisGeNET Score ≥0.3).
Ingredient-Gene Network of Curcumae Radix
In order to identify the pivotal genes and corresponding active ingredients that are strongly related to depression, we constructed an active ingredient-gene network using Cytoscape software, which is shown in Figure 2. The active ingredient-gene network contains 50 nodes and 80 lines. Only five active ingredients in CR were associated with the 45 genes related to depression. (E)−5-Hydroxy-7-(4-hydroxyphenyl)−1-phenyl-1-heptene was linked to 24 genes, (E)−1,7-diphenyl-3-hydroxy-1-hepten-5-one to 19, oxycurcumenol to 13, and β-sitosterol and sitosterol to 12. Topology analysis showed that there were four genes with the top degree values, betweenness centrality and closeness centrality at the same time: AR, NOS2, PTGS2, TYK2 (Table 4).
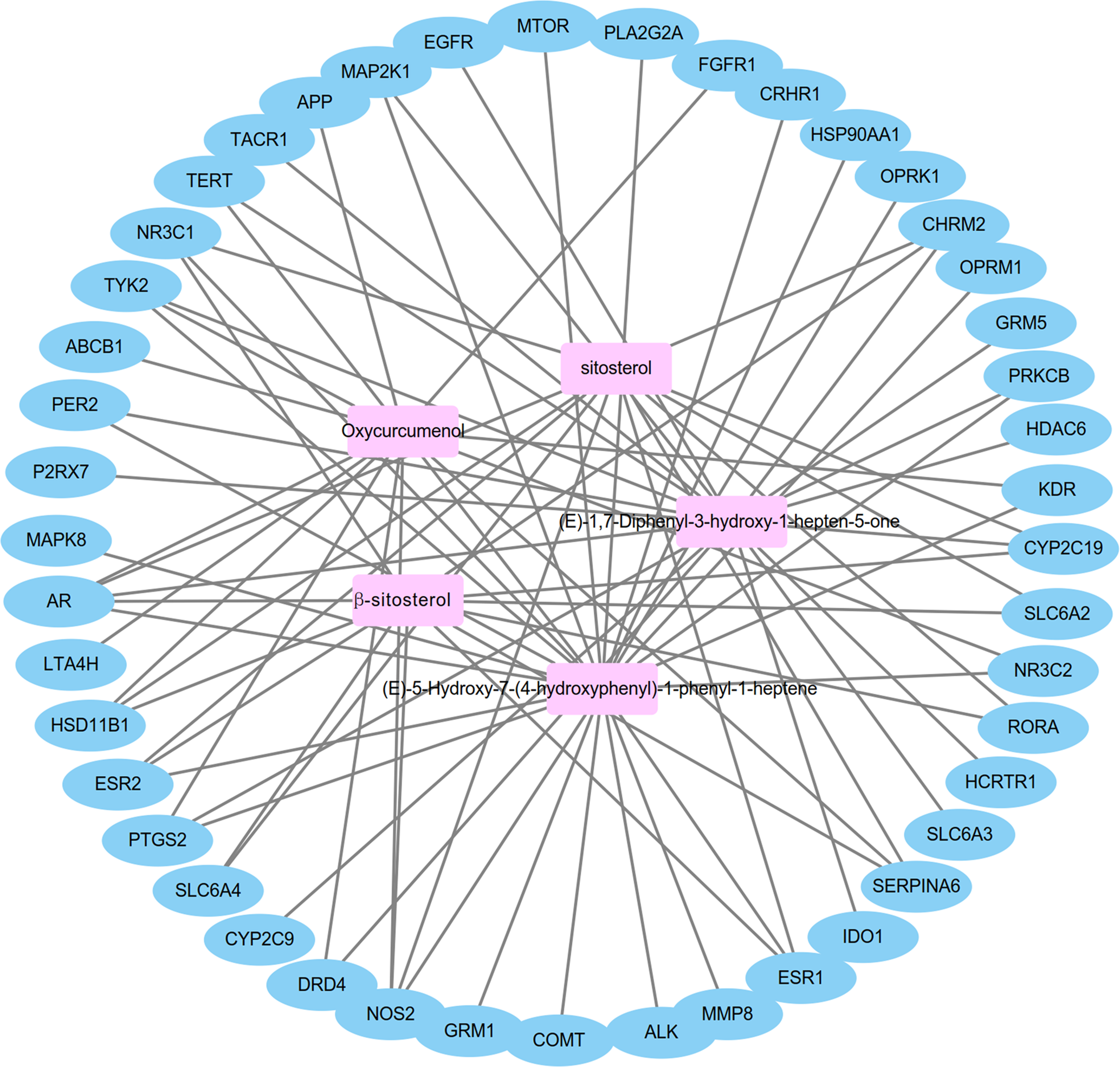
Ingredient-gene network of Curcumae Radix. The pink rectangle nodes (■) are the main active ingredients of Curcumae Radix, the light blue oval (●) is a potential gene for treating depression with Curcumae Radix. The line represents the correlation between the active ingredients and the genes.
Genes With the Top 4 Degree Values, Betweenness Centrality and Closeness Centrality.

Protein-Protein Interaction Network
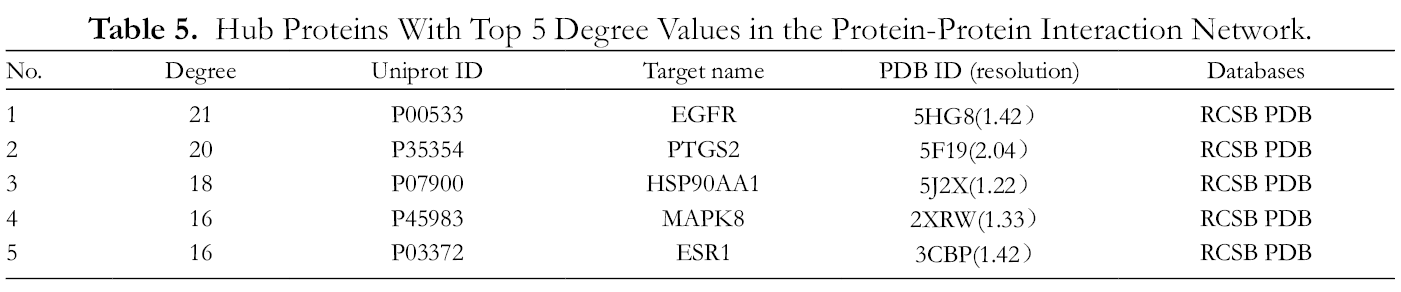
The PPI network of CR shown in Figure 3 involves 45 nodes and 189 lines. The average node degree of the PPI network was 8.4. We performed topological analysis to screen for proteins with top 5 degree values in the PPI network, which is shown in Table 5. The results showed that EGFR had the highest degree of 21; PTGS2 had the second highest degree of 20; HSP90AA1 with a degree of 18; and MAPK8 and ESR1 with a degree of 16; these were considered to be hub proteins in the PPI network.
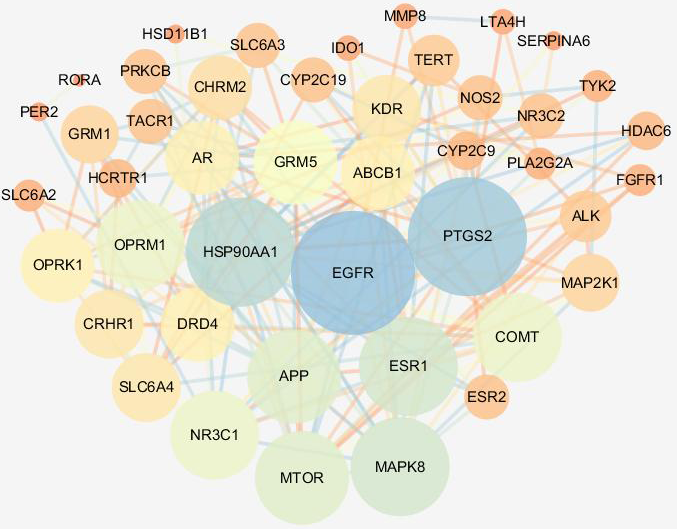
Protein-protein interaction network of Curcumae Radix. Nodes represent proteins and lines represent associations between proteins; the size and the color of the node represent the value of the degree, and the color of the line indicates the value of the combined score (low values to bright colors, low values to small sizes).
Hub Proteins With Top 5 Degree Values in the Protein-Protein Interaction Network.
Molecular Docking
Molecular docking was performed to evaluate the binding ability of the active ingredients with the corresponding hub-proteins (Table 6). The hub-proteins with the top 5 degree values were molecularly docked with the CR’s active ingredients, and 9 docking results were received and are represented in Figure 4. It is believed that the more negative the vina score, the more stable the binding of the active ingredients and the proteins. 20 The vina scores of the active ingredients and core protein targets of CR in the treatment of depression were negative and less than −5.9. The results show that oxycurcumenol, (E)−1,7-diphenyl-3-hydroxy-1-hepten-5-one and (E)−5-hydroxy-7-(4-hydroxyphenyl)−1-phenyl-1-heptene had good binding activities to PTGS2. Oxycurcumenol was fixed in the binding cavity of the protein PTGS2 through π-π bonds with residue ALA 527. (E)−1,7-Diphenyl-3-hydroxy-1-hepten-5-one was fixed in the binding cavity of the protein PTGS2 through π-π bonds with residues PHE 209 and TRP387. (E)−5-Hydroxy-7 -(4-hydroxyphenyl)−1-phenyl-1-heptene was fixed in the binding cavity of the protein PTGS2 through π-π bonds with residues PHE 518 and TRP 387.

Molecular docking of active ingredients with the corresponding hub proteins (Left: 3D structure, right: 2D structure; the green arrows indicate π-π bonds, and the pink arrows indicate H bonds).
Molecular Docking of Active Ingredients With the Corresponding Hub Proteins.

Gene Ontology Function, KEGG Pathway, and Tissue Localization Analysis
The Go enrichment analysis and tissue localization analysis of the potential genes for CR’s active ingredients in the treatment of depression are shown in Figure 5, including BP, CC, and MF. In the BP (Figure 5(A)), signal transduction involved 11 (24.4%) genes of depression, chemical synaptic transmission involved eight (17.8%), protein phosphorylation seven (15.6%), and positive regulation of nitric oxide biosynthetic process six (13.3%). In the CC (Figure 5(B)), plasma membrane involved 27 (60%) gene targets for treatment of depression, nucleus 22 (48.9%), the integral component of the membrane 20 (44.4%), and the integral component of the plasma membrane 17 (37.8%). In the MF (Figure 5(C)), protein binding involved 35 (77.8%) gene targets of depression, ATP binding 12 (26.7%), zinc ion binding 10 (22.2%), and enzyme binding nine (20%).

Enriched gene ontology terms of potential targets for treating depression from main active ingredients of Curcumae Radix. (
In the tissue localization analysis (Figure 5(D)), 30 depression-related genes were highly expressed in the brain, three in the breast, and 10 in the liver (Supplemental Table S1). Thirty brain-related genes were further analyzed for specific distribution using the BioGPS database (Figure 5(E)). HSP90AA1 was highly expressed in cingulate cortex, hypothalamus, medulla oblongata, occipital lobe, olfactory bulb, pineal (day), and pineal (night). COMT was highly expressed in hypothalamus, olfactory bulb, whole brain, pineal (day), and pineal (night). There were eight genes that were only highly expressed in the prefrontal cortex: PER2, P2RX7, GRM1, TACR1, MAPK8, HCRTR1, EGFR, and TYK2; APP was highly expressed in the cingulate cortex.
The KEGG pathway analysis revealed the potential mechanism of CR in the treatment of depression (Figure 5(F)). The calcium signaling pathway was identified as the most important one involving eight genes, accounting for 17.8%; the estrogen, Ras, and PI3K-Akt signaling pathways involved 7 gene targets, respectively, accounting for 15.6%; and the ErbB, HIF-1, Rap1, and MAPK signaling pathways involved 5 gene targets (11.1%).
Signaling Pathway Integration
Four important signaling pathways for treatment of depression with CR’s active ingredients are shown in Figure 6. The gene targets are marked in light blue, and the potential gene targets for CR active ingredients treatment of depression are marked in dark blue. There were 15 potential gene targets for the treatment of depression in the calcium, ERBB, PI3K-AKT, and estrogen signaling pathways, accounting for 33.3%, suggesting that CR may exerted antidepressant effects at these gene targets. PKC and MEK were recognized to play a role in multiple signaling pathways simultaneously, suggesting that they may be “hub genes.”

Anti-depression pathways of potential targets from main active ingredients of Curcumae Radix. (The color represents the signal pathway, the arrow “→” indicates the promoting effect, and T-arrow “┨” indicates the inhibition).
Discussion
According to the pharmacokinetic properties of CR, including OB, DL, and BBB, eight CR active ingredients were selected in our study, five of which are closely related to depression, namely β-sitosterol, sitosterol, (E)−1,7-diphenyl-3-hydroxy-1-hepten-5-one, (E)−5-hydroxy-7-(4-hydroxyphenyl)−1-phenyl-1-heptene, and oxycurcumenol. The first two are terpenoids, and the latter three are volatile oil components. Mou et al. found that either terpenoids or volatile oils extracted from Shugan Hewei Decoction containing CR could reduce the resting time in the open-field test in depression model rats and increase the content of serotonin (5-HT), γ-aminobutyric acid (GABA), and glutamic acid in the rats’ hypothalamus. 23 Besides, it has been reported that the 95% ethanol, light petroleum and ethyl acetate extracts of CR have a good antidepressant effect, especially the light petroleum and ethyl acetate extracts. 24 In the active ingredient-gene network, these five active ingredients of CR have also been identified to be closely linked to depression-related genes. Therefore, the active ingredients of CR may have good antidepressant effects.
In our study, 45 co-genes of CR and depression with DisGeNET score ≥0.3 were associated with depressive disorder, unipolar depression, or major depressive disorder. Topology analysis showed that genes AR, NOS2, PTGS2 and TYK2 with the top 4 degree values, betweenness centrality and closeness centrality at the same time, suggested that AR, NOS2, PTGS2 and TYK2 were closely related to depression. These results are partially consistent with previously published studies. For example, Owens found that decreased AR mRNA expression levels may be associated with depressive symptoms in healthy men. 25 Galecki et al. observed that NOS2 transcription was increased in peripheral blood of patients with recurrent depression, 26 and that the polymorphism (−1026C/A) in the NOS2 promoter was associated with the risk of recurrent depressive disorder. 27 Brkic et al. found that lipopolysaccharide reduced the level of PTGS2 mRNA in the prefrontal cortex of male Wistar depression rats. 28 Kalkman suggested that inhibiting TYK2 could reduce depression by blocking IL-6-mediated inflammation, which could be a potential new treatment based on the cause of depression. 29 Therefore, CR may reduce the onset and development of depression by regulating these important gene targets.
In our study, 45 proteins and 189 interactions were identified in the PPI network. Topological analysis showed that EGFR, PTGS2, HSP90AA1, MAPK8, and ESR1 have the highest values, which may be the hub protein targets that regulate or affect the progression of depression. These results have been partially confirmed by previous studies. For example, Hu et al. observed by biotin-labeled protein chip technology that the expression of EGFR protein in the hippocampus of rats with chronic stress depression increased significantly. 30 Jacobs et al. also confirmed that patients with stage IV non-small cell lung cancer with EGFR mutations showed elevated pro-inflammatory marker TNF-a, but the severity of depression was lower. 31 Previous studies found that Shugan Hewei Decoction containing CR could increase the content of glutamate in the nucleus accumbens to inhibit depressive-like behavior, which may be related to glutamate promoting the expression of PTGS2 protein in nerve cells. 32,33 Wei et al. found that the expression of HSP90AA1 protein in patients with moderate or severe depression was significantly higher than that in patients with mild depression or without depression. 34 Mohammad et al. identified MAPK8 (also called JNK1) as a central repressor of neurogenesis and promoter of depressive-like behavior in the adult hippocampus. 35 Besides, Chai also found that MAPK8 protein was highly expressed in the hippocampus of diabetic rats with depression, and the drug containing the Curcuma ingredient could reverse this trend. 36 Kajta et al. observed that the depressive-like effects of prenatal exposure to dichlorodiphenyltrichloroethane impair GPER1/ESR1 protein. 37 In addition, our molecular docking experiments further confirmed that these hub protein targets have good binding activity with the corresponding CR active ingredients, especially PTGS2 was identified to bind well to oxycurcumenol, (E)−1,7-diphenyl-3-hydroxy-1-hepten-5-one and (E)−5-hydroxy-7-(4-hydroxyphenyl)−1-phenyl-1-heptene. Chen et al. found that cortical PTGS2 protein (also known as cyclooxygenase 2, COX2) is activated in rats with depression-like behavior induced by chronic unpredictable mild stress (CUMS). 38 In addition, Song also confirmed that the expression of COX-2 in the hippocampal CA1 area of CUMS-induced depressed rats increased, suggesting that the occurrence of neuroinflammation is related to the depression-like behavior of rats. 39 Although the direct effect of oxycurcumol with COX2 is still unclear, it has been confirmed that oxycurcumol could prevent cell damage induced by oxidative stress. 40 However, few studies have been reported on (E)−1,7-diphenyl-3-hydroxy-1-hepten-5-one and (E)−5-hydroxy-7-(4-hydroxyphenyl)−1-phenyl-1-heptene. In short, these results indicate that EGFR, PTGS2, HSP90AA1, MAPK8 and ESR1 proteins may be important targets for the active ingredients of CR to regulate depression, but further animal or cell experiments are needed to confirm this.
To identify further the role of genes in the regulation of depression, we performed Go functional analysis. In the BP, signal transduction, chemical synaptic transmission and protein phosphorylation were the most involved biological processes of the genes. Several lines of evidence also prove this finding. For example, a previous study reported that corticosterone-induced depression and anxiety-like behavior reduced intracellular signal transduction in the hippocampus. 41 Lagus et al. confirmed that most people with depressive disorder experience sleep disorders, which were associated with disruption of chemical synaptic transmission. 42 Rajamanickam et al. found that Akt could regulate the expression and phosphorylation of serotonin transporter, the target of antidepressants. 43 In the CC, the plasma membrane and nucleus were identified as sites of gene action. Bowman et al. found that monoamine transporters on the plasma membrane were involved in the steady-state regulation of serotonin in adolescent antidepressant therapy. 44 Pace et al. confirmed that blocking the transport of glucocorticoid receptors from the cytoplasm to the nucleus and disrupting glucocorticoid receptor-DNA binding may lead to major depression. 45 In the MF, protein binding and ATP binding were common functions of depression-related gene targets. A previous study found that overexpression of p11 promotes protein binding between P11 and 5-HT receptors and alleviates depression-like behavior. 46 ATP-binding related genes, such as ABCB1, were also identified as being associated with major depression. 47
In terms of the pathological part of depression, previous studies have reported that the dysregulated hypothalamus-pituitary-adrenocortical axis and the neurotrophin system of the brain are considered to be important in the pathophysiology of depression, which is closely related to the brain. 48 In our study, most co-genes were identified as being related to the brain. However, many brain regions are involved in the pathology and development of depression. In order to identify accurately high expression regions of genes, we used BioGPS for further analysis. The results revealed that PER2, P2RX7, GRM1, TACR1, MAPK8, HCRTR1, EGFR and TYK2 were highly expressed in the prefrontal cortex. Previous studies also suggested that PER2 and P2RX7 are associated with stress-induced depression. 49,50 GRM1 was identified as high expression in the prefrontal cortex in female patients with major depressive disorder. 51 However, the expression of TACR1, MAPK8, HCRTR1, EGFR, and TYK2 in the prefrontal cortex has not been verified in patients with depression.
KEGG pathway enrichment analysis indicated that multiple signaling pathways were involved in the pathophysiological processes of depression, such as the calcium, estrogen, Ras, PI3K-Akt, ErbB, and HIF-1 signaling pathways. Previous studies found that Jiawei Wendan decoction containing CR could exert antidepressant effects by inhibiting the massive influx of Ca2+ in hippocampal neurons and improving neuroplasticity, 52 as well as up-regulating Ras protein in the hippocampus to improve the spatial cognitive ability in rats with depression. 53 Besides, regulation of the estrogen signaling pathway has also been found to play an important role in the neuroprotection of postpartum depression. 54 Fan et al. found through network pharmacology that the mechanism of the drug combination Tatarinowii Rhizoma and CR in the treatment of depression is related to the secretion of sex hormones and hippocampal neuronal apoptosis, which is similar to our study. 55 However, its vague pictures, single data, and lack of in-depth analysis cannot provide more effective information for the treatment of depression. Guo et al. demonstrated that enhancing the PI3K/AKT/FoxO1 pathway to inhibit TLR4 expression could alleviate neuroinflammation and cause depression-like behavior. 56 Previous studies confirmed that the antidepressant effect of β-sitosterol is mediated by 5-HT, dopamine and GABA-ergic systems, while mGluR1-dependent long-term inhibition in brain dopamine neurons is regulated by the NRG1/ErbB signaling pathway and involves the ErbB2/ErbB4 receptor. 57,58 Chen et al. found that oxycurcumol protection of nerve cells from oxidative stress-induced cell damage may be related to the HIF1α signaling pathway. 40 Eyre et al. also believed that the onset of depression is associated with impaired neuroplasticity and that the HIF-1α signaling pathway is involved in this process in rodent depression models. 59
Conclusions
This study revealed the ingredients, effective targets and mechanisms of CR in the treatment of depression, as well as high expression areas of depression-related targets, which provide a new perspective for the development of new antidepressants.
Supplemental Material
Online supplementary file 1 - Supplemental material for Identification of the Ingredients and Mechanisms of Curcumae Radix for Depression Based on Network Pharmacology and Molecular Docking
Supplemental material, Online supplementary file 1, for Identification of the Ingredients and Mechanisms of Curcumae Radix for Depression Based on Network Pharmacology and Molecular Docking by Xiaotong Wang, Qiaoru Lin, Meiqing Shen, Haixiong Lin, Junjie Feng, Lulu Peng, Minling Huang, Xiaoxuan Zhan, Ziyin Chen and Tengfei Ma in Natural Product Communications
Supplemental Material
Figure S1 - Supplemental material for Identification of the Ingredients and Mechanisms of Curcumae Radix for Depression Based on Network Pharmacology and Molecular Docking
Supplemental material, Figure S1, for Identification of the Ingredients and Mechanisms of Curcumae Radix for Depression Based on Network Pharmacology and Molecular Docking by Xiaotong Wang, Qiaoru Lin, Meiqing Shen, Haixiong Lin, Junjie Feng, Lulu Peng, Minling Huang, Xiaoxuan Zhan, Ziyin Chen and Tengfei Ma in Natural Product Communications
Supplemental Material
Table S1 - Supplemental material for Identification of the Ingredients and Mechanisms of Curcumae Radix for Depression Based on Network Pharmacology and Molecular Docking
Supplemental material, Table S1, for Identification of the Ingredients and Mechanisms of Curcumae Radix for Depression Based on Network Pharmacology and Molecular Docking by Xiaotong Wang, Qiaoru Lin, Meiqing Shen, Haixiong Lin, Junjie Feng, Lulu Peng, Minling Huang, Xiaoxuan Zhan, Ziyin Chen and Tengfei Ma in Natural Product Communications
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported, in part, by the International Program for Postgraduates, Guangzhou University of Chinese Medicine (GZYXB[2021]52), Excellent Doctoral Dissertation Incubation Grant of Guangzhou University of Chinese Medicine (GZYXB2020-18), and Excellent Doctoral Dissertation Incubation Grant of First Clinical School of Guangzhou University of Chinese Medicine (YB201902).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
