Abstract
Keywords
Introduction
Respiratory tract infections (RTI) are a prevalent and seriously damaging infectious disease. The specific mechanism is that sickness develops when pathogenic bacteria enter the respiratory system and grow on the mucosal surfaces. Based on where the infection occurs, it can be divided into upper and lower RTIs, with frequent attacks and chronic symptoms for recurrent RTIs. The majority of clinical symptoms include fever, cough, sore throat, etc. Nevertheless, if these symptoms are not treated quickly and accurately, they may become exacerbated by rhinitis, pharyngitis, pneumonia, etc. 1 The new coronavirus infection that causes Corona Virus Disease 2019 (COVID-19) spreads downhill through the upper respiratory tract. As air pollution worsens, the human body is continually exposed to an environment that is ripe for infection. As a consequence, the cellular immune system becomes fatigued, human immune globulin consumption rises, the number of immune cells and their activity significantly drop, and the prevalence of infectious diseases of the respiratory tract gradually rises. 2 To increase the body’s resistance to disease, it is therefore crucial to improve immune function throughout the entire body, including the respiratory system, in addition to the basic antibacterial and antiviral treatments. 3 Due to its low toxicity, Artemisia has special advantages in treating RTIs.
Artemisia belongs to the dicotyledonous plant family Compositae, which has a variety of resources and is extensively distributed in temperate, tropical, highland, steppe, desert, high altitude, and highly arid environments. 4 Several species of Artemisia, including Artemisia annua L. (common name Qinghao), Artemisia argyi L. (common name Aiye), and Artemisia capillaris Thunb. (common name Yinchen), have antibacterial, antiinflammatory, and immune-regulating properties. 5 Polysaccharides, organic acids, terpenoids, flavonoids, and volatile oils are among their secondary metabolites, which have considerable scientific value. Asteraceae family member Qinghao has been used safely for centuries to treat a range of fevers, most notably “intermittent fevers” and diseases connected to chills, including RTIs.6-8 Qinghao flavonoids have both antiinflammatory and antibacterial properties. They exert antiinflammatory effects by controlling the nuclear factor kappa-B (NF-κB) and mitogen-activated protein kinase (MAPK) signaling pathways, and they exhibit antibacterial properties by preventing Staphylococcus aureus membrane growth and mycoflavin production.7,9,10 Additionally, Qinghao and its derivatives have also demonstrated antiinflammatory properties, including inhibition of interleukin-6 (IL-6) that plays a key role in the development of severe COVID-19.6,11
Streptococcus pneumoniae, S. aureus, and respiratory syncytial virus, which cause RTIs, are inhibited by the volatile oil of Aiye. 8 The common component of Aiye and Yinchen, 6, 7-dimethoxy-coumarin, can regulate the activity of related enzymes in the phenylalanine metabolic pathway, thus interfering with the alcohol-induced primary hepatocyte damage. Yinchen extract (HACE) can protect the kidneys of diabetic rats, so Aiye and Yinchen can protect the kidneys and liver in the process of RTI. 12 Sesquiterpene lactones are mainly isolated from Asteraceae species, which were found in seven Artemisia species collected from Tajikistan, and exhibit the biological effects of inhibiting severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2).13,14 As a whole, Artemisia species displays pharmacological effects in the prevention and treatment of RTI, but more research is needed to determine the underlying pharmacological processes.
Artemisia species have complex components, many targets, and a wide range of therapeutic effects and signaling pathways. 15 Network pharmacology integrates diseases and drugs into biomolecular networks to predict their effective components, possible target pathways, and mechanisms of action.16,17 It has the advantages of a high success rate, fewer side effects, multiple genes, and multiple targets. It can systematically analyze the interaction network among drug components, protein targets, diseases, and genes to provide a more basic and scientific explanation for the research of traditional Chinese medicine. In this study, network pharmacology was used through data mining and analysis to construct a multilayered and multiangle biological network system of “drug-target-disease,” interpret the complex relationships between the networks, revealing multiple targets and signaling pathways regulated by the active ingredients of traditional Chinese medicine, and subsequently predict the mechanism of action of the active ingredients in the treatment of diseases. The mechanism of illness treatment is also revealed via molecular docking at the same time. In order to serve as a guide and foundation for future research on RTI-related diseases as well as clinical treatment, this research analyzed the active components, major targets, and signaling pathways of Artemisia species in the treatment of RTI using network pharmacology and molecular docking technology (Figure 1).

Flowchart of experimental designs.
Results
Screening the Active Ingredients and Targets of Artemisia
According to the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP), 126, 135, and 53 active ingredients of qinghao, aiye, and yinchen were retrieved, respectively, and the corresponding targets of each active ingredient were screened. According to oral bioavailability (OB) > 30% and drug-likeness (DL) > 0.18, 44 active ingredients (Table 1) and 206 drug targets were finally obtained.
Active Components of Qinghao, Aiye, and Yinchen.

OB, oral bioavailability; DL, drug-likeness.
The Active Components of Artemisia and RTI Share Common Targets
The Gene Cards database was used to retrieve 2418 disease-related targets, and the intersection of the active ingredient targets and RTI targets was selected as 133 key targets (Figure 2). The intersection targets are the key targets of Artemisia plants in the treatment of RTI diseases.

Wayne diagram of the intersection of drug targets and disease targets (yellow represents the active ingredient target of Artemisia, pink represents the target related to RTI). RTI, respiratory tract infection.
PPI Network Construction and Topology Analysis
The protein–protein interaction (PPI) network of 133 key targets was constructed, and after the discrete proteins were removed, the interaction network of 123 proteins was obtained, including 123 nodes and 601 sides, indicating the main interaction between Artemisia and RTI. The Cytoscape software visualizes the protein network map (Figure 3A). Combined with Analyze Network to analyze the topological characteristics of the PPI network, the degree of node genes is obtained, and the degree is sequenced. The genes of TOP5 are shown in Table 2. It can be seen from the table that TP53, AKT1, JUN, HSP90AA1, and RELA as hub genes play an important role in the occurrence and development of RTI. Five active components corresponding to hub genes were extracted, and a drug-active component-disease key target gene network was constructed. Visualization using Cytoscape is shown in Figure 3B, which contains 3 species of Artemisia, 23 drug active components, 5 key targets, and a total of 70 relationship pairs. The results show that the same active component of the same Artemisia plant acts on different targets, while the same target will be affected by different active components. It reflects the characteristics of the multicomponent and multitarget treatment of Artemisia.
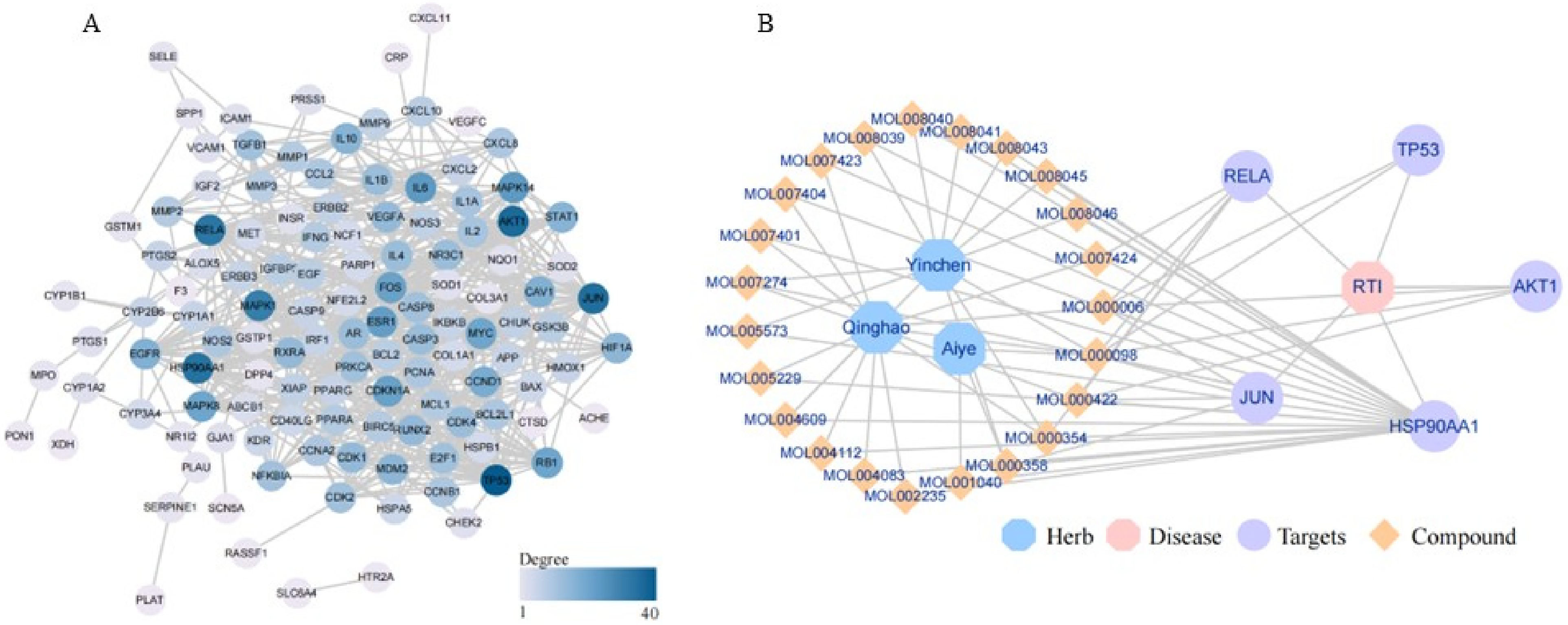
PPI network and drug-active ingredient-target network. (A) The lines represent the interaction between proteins; the color represents the degree value, and the darker the color, the higher the degree value, and the more in the core position. (B) Artemisia is represented by the dark blue octagonal shape; the purple circle represents the hub gene; the yellow diamond represents the drug's active ingredient; and the pink octagon represents disease. PPI, protein–protein interaction.
Topological Properties of the TOP5 Hub Gene.

Target Gene Enrichment Analysis
Gene ontology (GO) enrichment analysis results are shown in Figure 4A: a total of 2269 terms were obtained in the biological process (BP), which was significantly correlated with drug response and bacterial-derived molecule response. In terms of molecular function (MF), a total of 164 DNAs were obtained, and the key target genes were significantly related to receptor ligand activity, signal receptor activator activity, cytokine activity, and cytokine receptor binding. In terms of cell composition (CC), a total of 61 terms were obtained, and the key target genes were significantly related to plasma membrane raft, vesicular cavity, membrane region, membrane microdomain, membrane raft, and so on.

Gene enrichment analysis of key target genes of Artemisia in the treatment of RTI. (A) In the GO enrichment bar chart of the key target genes, the ordinate indicates the enriched GO term, and the bar length indicates the number of enriched GO terms for the key targets. (B) Enrichment of the KEGG pathway of key target genes, the vertical axis of the bar chart indicates the KEGG pathway, and the length of the bar indicates the number of genes in the pathway. GO, gene ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; RTI, respiratory tract infection.
Sixty-two related pathways were obtained by the Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis of 133 key target genes, and the TOP15 pathway was displayed by drawing a bar graph using the R language function enrichplot (Figure 4B). It can be seen from the diagram that the KEGG enrichment results include the phosphatidylinositol 3-kinase (PI3K)-protein kinase B (Akt) signaling pathway, the tumor necrosis factor (TNF) signaling pathway, and the interleukin-17 (IL-17) signaling pathway, indicating that the key target genes are significantly related to the PI3K-Akt signaling pathway and the TNF signaling pathway.
Key Target-Function-Pathway Regulatory Network
Key targets-the BP network is shown in Figure 5A, which contains 10 terms and 89 key targets with a total of 357 relationship pairs. Key targets-the MF network is shown in Figure 5B, which contains 10 terms and 67 key targets with a total of 177 relationship pairs. Key targets-the CC network is shown in Figure 5C, which contains 10 terms and 44 key targets with a total of 149 relationship pairs. Key target pathways are shown in Figure 5D, including 20 KEGG pathways, 100 key targets, and a total of 533 relationship pairs.

Key targets of Artemisia in the treatment of the RTI-function-pathway regulatory network (gray circle represents key targets). (A) Key targets-BP functional networks (blue hexagons represent GO-BP terms). (B) Key targets-MF functional networks (yellow hexagons represent GO-MF terms). (C) Key targets-CC functional networks (pink hexagons represent GO-CC terms). (D) Key target pathway regulatory network diagram (purple hexagons represent KEGG pathways). GO, gene ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; RTI, respiratory tract infection; BP, biological process; MF, molecular function; CC, cell composition.
Molecular Docking
The core gene of the PPI network, TOP5, was selected and the active compounds targeting the TOP5 gene were selected from the pharmacological network of traditional Chinese medicine (Table 3). Table 4 displays the docking energy parameters. Figure 6 shows the best docking image between receptor and ligand after visualization. The result of molecular docking between AKT1 (PDB ID: IUNP) and quercetin is shown in Figure 6A, in which the LYS-64 residue interacts with the quercetin by a hydrogen bond, and the docking affinity between the active molecule and protein is −6.8 kcal/mol. The results of molecular docking between HSP90AA1 (PDB ID: 1BYQ) and artemetin are shown in Figure 6B, in which GL Ymur137, PHE-138, and THR-184 residues interact with artemetin by hydrogen bonding, and the docking affinity between active molecules and proteins is −6.9 kcal/mol. The results of molecular docking between JUN (PDB ID: 1JUN) and β-sitosterol are shown in Figure 6C, in which THR-297, V AL-305, ALA-306, L YS-309, and ARG-302 residues interact with β-sitosterol by hydrogen bonding, and the docking affinity between active molecules and proteins is −5.9 kcal/mol. The results of molecular docking between RELA (PDB ID: 1NFI) and isorhamnetin are shown in Figure 6D, in which MET279, L YS-28, SER-51, GLN-266, and GLN-241 residues have hydrophobic 10-bond interactions with the isorhamnetin, and the docking affinity between the active molecule and protein is −7.8 kcal/mol. The result of molecular docking between TP53 (PDBID: 1A1U) and quercetin is shown in Figure 6E, in which the LYS-40 residue interacts with the quercetin by a hydrogen bond and the docking affinity between the active molecule and protein is −5.9 kcal/mol. The results of molecular docking show that these binding energies are all less than −5 kcal/mol, indicating that there is a high affinity between the active components and the target. Based on this, we predict that all these active components will play a key role in the RTI.

The main active component’s molecular docking with the target protein (the pink stick structure is an amino acid residue with a hydrogen bond interaction with the active component). Gray dotted lines are hydrophobic bonds formed between active components and amino acid residues, and each gray dotted line represents a hydrophobic bond. Each yellow dotted line represents a hydrogen bond between active components and amino acid residues. (A) The blue double-ring stick model is the active molecule quercetin. (B) The green double-ring stick model is the active molecule artemetin. (C) The active molecule β-sitosterol is represented by the green double-ring stick model. (D) The green double-ring stick model is the active molecule isorhamnetin. (E) The green double-ring stick model is the active molecule quercetin.
Molecule Components of Hub Gene Docking.

oral bioavailability; DL, drug-likeness.
Energy Parameters of Molecular Docking.

Antiviral Effects of Lactones on the Target Proteins of the SARS-CoV-2 Virus
The average value of molecular docking evaluation for selected sesquiterpene lactones and remdesivir with all target proteins of the SARS-CoV-2 virus are shown in Table 5. The average value of molecular docking evaluation for selected sesquiterpene lactones with all target proteins of SARS-CoV-2 virus ranged from −6.979 kcal/mol to −9.5966 kcal/mol. The average score of molecular docking for anabsintine, absintine and lactucopicrin were −9.5966 kcal/mol; −9.5824 kcal/mol, and −8.8395 kcal/mol, respectively. However, the docking score of remdesivir was −8.4096 kcal/mol.
Name, Chemical Structure and Average Docking Score of Selected Sesquiterpene Lactones.

Cell Viability Assay
A. annua extract was added to the MDCK cell culture medium at different concentrations, and the cell survival rate was determined using the CCK-8 method after 24 h culture. As illustrated in Figure 7A, the concentration that yielded a cell survival rate exceeding 90% was selected as the maximum nontoxic concentration of cells (TC0), and the concentration range of A. annua extract was 25–200 μg/mL. These findings indicate that A. annua extract has a very low impact on cells and high safety.

(A) Toxic effects of Artemisia annua extract on MDCK cell. (B) Inhibitory effect of A. annua extract on influenza virus.
The Inhibition of the Drug on the Virus
As shown in Figure 7B, A. annua extract (25 μg/mL) had an inhibitory effect on the virus, and the inhibitory effect was the highest when the concentration was 200 μg/mL, indicating that the inhibitory effect of A. annua extract on the virus was dose-dependent. A. annua extract can treat RTI by inhibiting the influenza virus.
Discussion
The disease targets, active components, and drug targets of Qinghao, Aiye, and Yinchen were predicted by an online database, and the following results and conclusions were obtained by the method of network pharmacology: firstly, 44 active components and 206 drug targets were obtained by TCMSP database. The active ingredients include β-sitosterol, quercetin, isorhamnetin, artemisinin, and others. It has been shown that β-sitosterol regulates chemokine expression in cells and reduces excessive inflammatory responses. 18 β-sitosterol decreased the levels of proinflammatory cytokines IL-1 β and TNF-α, increased the levels of antiinflammatory IL-4 and IL-10, and reduced inflammatory response in rats with acute lung injury.19,20 Quercetin inhibited cisplatin-induced renal injury by downregulating the expression of NF-κB and iNOS proteins, which proved that quercetin has a renal protective effect.21-23 Quercetin, by upregulating miR-124 to inhibit NF-κB, alleviates the inflammatory response of lipopolysaccharide induction. 24 Isorhamnetin inhibits chronic bronchitis by reducing the inflow of inflammatory cells.25,26 Artemisinin and its derivatives show the highest potential for anti-SARS-COV-2 strains and could become potential drugs against COVID-19.7,13,27 Furthermore, PPI network analysis of key targets shows that the network contains 123 nodes and 601 edges, and according to the value of the Top5 hub genes, they are TP53, AKT1, JUN, HSP90AA1, and RELA, indicating that Artemisia plants use the above genes to achieve the role of treatment of respiratory infections.28-31 GO functional enrichment results show that the therapeutic effect of Artemisia on RTI is related to the binding of membrane and cytokine receptors, and most of the BPs take place in the membrane, which participates in the activation and binding of a series of cellular receptors and the cascade of downstream signal pathways. KEGG functional enrichment analysis showed that the key target genes were significantly associated with the PI3K-Akt signal pathway, the TNF signal pathway, the IL-17 signal pathway, and so on. The PI3K/AKT signal pathway, the TNF signal pathway, and the IL-17 signal pathway are all important pathways in the inflammatory response. The TNF signaling pathway plays its role through the expression of cytokines, chemokines, and cell adhesion molecules and is one of the main pathways involved in immune and inflammatory responses.32,33 It is speculated that the active components of Artemisia may inhibit inflammatory response and regulate immune function in the treatment of RTI by acting on these pathways. Finally, molecular docking results show that there are hydrogen bonds and hydrophobic bonds between proteins and small molecules, with binding energies indicating that isorhamnetin, artemetin, and quercetin have good binding affinity with the target. In silico study of the antiviral effects of selected sesquiterpene lactones on the target proteins of the SARS-CoV-2 virus shows that the performance of SARS-CoV-2 virus with sesquiterpene lactone absinine, anapsinine, and lactoside extracted from A. annua was better than that of the antiviral drug Redesivir.
In summary, using network pharmacology and molecular docking technology, the key elements, core targets, and signaling pathways of Artemisia in the treatment of RTI were discussed, providing a theoretical foundation and a point of reference for further research on the mechanism of treatment for RTI.
Materials and Methods
Identification and Screening of Active Compounds and Their Target Genes
Use of TCMSP (http://tcmspw.com/tcmsp.php) for Qinghao, Aiye, and Yinchen active ingredients and the corresponding targets. Screened active ingredients for OB > 30% and DL >0.18. 27
Intersection of Disease Target Genes and Compound Target Genes
Using “respiratory tract infection” as a keyword, RTI-related genes were screened in the Gene Cards database (https://www.genecards.org/). 27 The target genes predicted from the active ingredients in Artemisia campestris L. were intersected and mapped with the target genes predicted for the RTI disease to obtain the potential therapeutic targets of Artemisia species to treat RTI. Automatically retrieved the overlapping genes and drew the Venn diagram using the R language's Venn Diagram package.
PPI Network Construction and Topology Analysis
The PPI network was constructed by uploading the common drug-disease targets to the String database (https://string-db.org), and the confidence level was set at 0.9 (Confidence = 0.9). The Network Analyzer tool of the Cytoscape 3.8.2 software was used for topological analysis of the PPI network. Genes were sorted based on degree values, and genes with scores greater than the mean value were selected as the core targets. 34 The Hub gene was obtained through topological analysis, and genes were sorted based on degree values. The Top5 genes were selected as the core targets, and the corresponding active components of the hub gene were extracted to construct the pharmacology regulatory network of traditional Chinese medicine.
GO and KEGG Enrichment Analysis
Active ingredient corresponding targets and RTI-related common targets were conducted by using the R Cluster Profiler package. BP, CC, and MF were chosen for GO analysis, P < .05 after correction and counts ≥ 2 were the screening conditions. In this study, enrichplot using the R language “Enrichplot” (Version 1.10.2) was used to create bar graphs to demonstrate the GO enrichment results. Using the KEGG database, all identified protein collections or screened differentially expressed proteins were annotated, and the most important metabolic and signal transduction pathways involved in these proteins or genes were analyzed. 34 KEGG pathway enrichment analysis of the key pathways linked to RTI and use R language “enrichplot” (Version 1.10.2) is used to create a bar graph to show the Top 15 pathways.
Construction of the “Key Target-Function-Pathway” Network Map of Artemisia for the Treatment of Respiratory Tract Infection
In order to observe the overall pharmacological effect of Artemisia plants on RTI, the TOP10 terms of GO-BP, GO-CC, and GO-MF and their corresponding key targets were selected, the KEGG Pathways TOP20 and their corresponding key targets were extracted, and targeted compounds of the key targets were selected. The regulatory network maps of the key target function and key target pathway were constructed respectively and visualized by Cytoscape software.
Molecular Docking
In order to further clarify the binding efficacy between the key active ingredients and targets of Artemisia plants in the treatment of RTI, the key active ingredients were selected as ligands and the core target as receptors. First, the protein structure of the key target was obtained from the PDB (https://www1.rcsb.org/) database, and the small molecules and water molecules originally contained in the target were removed. Complete protein hydrogenation and charge calculation with AutoDock Tools (Version 1.5.6). From the PubChem database (https://pubchem.ncbi.nlm.nih.gov/) to download active ingredient structure, and use AutoDock Tools to charge balance, small molecules that can rotate button to check. Then, the range of docking boxes is selected according to the receptor active center. Finally, AutoDock vina was used to calculate the receptor-ligand docking, and the structure with the lowest binding free energy (highest binding affinity) was selected in the output result. Finally, PyMol (Version 2.5) software was used for visualization and beautification. In addition, an in silico study of the antiviral effects of selected lactones on the 93 target proteins of the SARS-CoV-2 virus was carried out using D3 Pharma databases (D3 Targets-2019-nCoV).
Cell Assays
MDCK cells and strain A/Victoria/4897/2022 (H1N1) were all donated by Professor Fang Shisong, Shenzhen Center for Disease Control and Prevention. A. annua extract powder (Shaanxi Snow Biotechnology Co., LTD., Lot No. SMT231011).
Culture of Cells and Viruses
MDCK cells were cultured in DMEM medium containing 5% fetal bovine serum at 37 °C in an atmosphere of 5% CO2 and 95% air. When MDCK cells grow to a monolayer state, the medium in the culture dish is sucked, washed twice with PBS, and the maintenance solution is replaced (DMEM medium added with 0.25 μg/mL TPCK pancreatic enzyme). Add 10 μL of virus, culture in a 35 °C, 5% CO2 cell incubator, and then put into the refrigerator at −80 °C, freeze and thaw repeatedly, let the cells burst and release the virus particles.
Cell Viability Assay
The toxic effects of different concentrations of A. annua extract on MDCK cells were detected by the CCK8 method. MDCK cells were seeded in 96-well plates (104 cells/well) and cultured for 24 h with different concentrations of A. annua extract. After incubating, 10 μL of CCK-8 solution was added to each well for a further 1 h at 37 °C. The optical density at 450 nm was determined using a BioTek ELISA microplate reader.
The Inhibition of the Drug on the Virus
The extract of artemisinin was diluted into 4 concentrations of 200 μg/mL, 100 μg/mL, 50 μg/mL, and 25 μg/mL using DMEM medium. The 4 concentrations of drugs (1 mL) were mixed with the H1N1 virus (1 mL) and incubated at 37 °C for 5 h. Four concentrations of drugs (100 μL) were added to MDCK cells and incubated for 5. After 5 h, the drug solution was discarded and cleaned twice with PBS. The mixed solution of drugs and viruses was added to the cells and infected for 1 h. After 1 h, the disease venom was discarded, and the cells were cleaned twice with PBS. Add four concentrations of drugs (100 μL) and continue to culture for 24 h. At the same time, the virus control group (only virus, no drug solution) and the cell control group (no virus, no drug solution) were set up. After 24 h, 10 μL of CCK-8 reagent was added to each well and incubated for 1 h away from light. The optical density at 450 nm was determined using a BioTek ELISA microplate reader.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Ethical Approval is not applicable for this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Key Research and Development Program of “Inter-Governmental Cooperation in Science and Technology Innovation” (Grant No. 2023YFE0103200).
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
