Abstract
Gegen Qinlian decoction (GQD) is a traditional Chinese medicine that is used to treat non-alcoholic fatty liver disease (NAFLD) in the clinic. The pharmacodynamics and cellular pathways governing the effects of GQD on NAFLD remain undefined. In this study, we investigated GQD pharmacology through assessment of its chemical constituents and evaluated and screened its components using drug likeness, pharmacokinetic characteristics (absorption, distribution, metabolism, excretion, and toxicity), and appropriate compensation mechanisms. We performed predictions of the active GQD ingredients based on reverse pharmacophore matching and compared multiple NAFLD-related genes to determine potential GQD targets. Molecular docking experiments of the active components were performed to reveal cellular targets. Annotation analysis of both target genes and related pathways were assessed through the DAVID database. Cytoscape software was used to construct a “component-target-path” network for the treatment of NAFLD by GQD. Through data analysis, 9 active GQD substances and 10 targets related to NAFLD encompassing 4 cellular pathways were identified. Data were verified through enzyme-linked immunosorbent assay and Western blot analysis. These findings provide new references for the network pharmacology of Chinese medicinal compounds and NAFLD treatment.
Keywords
Gegen Qinlian decoction (GQD) is a traditional prescription in Zhang Zhongjing’s “Treatise on Febrile Diseases” and is composed of Puerariae Radix, Scutellariae Radix, Coptidis Rhizoma, and Licorice. GQD treats non-alcoholic fatty liver disease (NAFLD) through its action on the peroxisome proliferator-activated receptor gamma (PPARγ) pathway. 1 Guo and co-workers 2 showed that GQD can affect lipid metabolism and reduce inflammation by regulating the intestinal flora balance and can ultimately play a role in treating NAFLD/non-alcoholic steatohepatitis (NASH). Xia and co-workers 3,4 found that GQD mainly improves glucose and lipid metabolism by regulating adipokine content, which can be used in the clinical treatment of type 2 diabetes mellitus with NAFLD. Guo and co-workers 5 showed that GQD treatment may be a potent strategy for managing NAFLD by managing lipid metabolism and inflammatory and histological abnormalities by triggering the Sirt1 pathway. Despite this knowledge, the active ingredients of GQD used in NAFLD treatment remain unclear and the major cellular targets governing its activity remain undefined. In order to address this, this study employed network pharmacology analysis, visual network software, existing databases, and “multialgorithm-multicomponent-multitarget” integrated analysis, to study the active components of GQD and its mechanism(s) of action during NAFLD treatment.
Materials and Methods
This study assessed the chemical composition of GQD through literature databases, drug likeness (DL), and absorption, distribution, metabolism, excretion, and toxicity (ADMET) assessments, to predict both pharmacology and toxicology. The identified compounds underwent reverse pharmacophore matching to obtain potential targets. NAFLD-related genes were assessed in Malacard and GeneCard databases and compared with GQD targets to obtain potential mechanisms of action for GQD during NAFLD therapy. Using autodock software, GQD active ingredients were docked with potential targets in the reverse molecule, and the targets of active GQD constituents during NAFLD treatment were predicted. Gene targets were analyzed using the DAVID database, and related GQD pathways for NAFLD were predicted. The “component-target-path” network for GQD treatment of NAFLD was constructed using Cytoscape software. We completed network pharmacological studies on the components and mechanisms of GQD activity during NAFLD treatment. The specific parameters were as follows.
Acquisition of Chemical Constituents
The chemical composition of GQD was inferred from literature and traditional Chinese medicine systems pharmacology (TCMSP) databases, and specific structural formulas were identified in Pubchem and Chemspider. When a specific structural formula was not identified, original papers were sourced. Structures were compiled using ChemBioDraw 14.0 software and files were maintained in the mol2 format. 6 -9
Screening of Chemical Constituents
The obtained chemical components were sequentially subjected to DL and ADMET screening. The DL screening tool was provided by the molsoft and SymMap 10 website, and the mol format file was inputted into DL predictions. The active ingredients of GQD were screened at a DL ≥0.18 as a reference for drug-like screening. We imported compound files obtained from DL screening into PreADMET and performed ADME and toxicity predictions. For ADME, we selected Caco-2 permeability (Caco2), oral bioavailability (OB), and plasma protein binding (PPB) as indicators, and set the thresholds as Caco2 ≥0.4, OB ≥30%, and PPB ≥20%. For toxicity predictions, Ames Carcino Mouse, and hERG inhibition tests were selected as indicators totaling 10 points. The Ames test comprised 4 points, the Carcino Mouse test accounted for 3 points, and hERG inhibition tests accounted for 3 points. Scores ≥6 were selected. A total of 97 compounds were finally screened. Scoring criteria are shown in Table 1.
Scoring Criteria and Toxicity Predictions.

Chemical Constituents
Since the exclusion of compounds was inevitable, we established a compensatory mechanism to identify compounds with definitive activity in GQD and analyzed the factors that led to their removal. As an example, the parameters of baicalin were: DL = 0.75; Caco2 = −0.85; OB = 40.12, and PPB = 75.5%. The Ames test was positive, as were the Carcino Mouse scores, while hERG inhibition assays deemed the compounds low risk (total score = 5 points). According to these parameters, baicalin was screened out in “Caco2” and “toxicity predictions”. However, a comprehensive analysis of the compound suggested it should retain selection as its higher “OB” value compensates for the low “Caco2” values observed. According to the current literature, baicalin showed no toxicity and we therefore included this compound as an active ingredient. A total of 11 compounds were selected through such supplementary mechanisms.
Acquisition of NAFLD Targets
The results of searching keywords including “non-alcoholic fatty liver disease”, “non-alcoholic steatohepatitis”, and “simple fatty liver” on GeneCards and MalaCards were downloaded and comprehensively analyzed, leaving 84 relevant NAFLD targets. The active ingredients were uploaded onto the GQD “SystemsDock” website, and potential GQD targets based on reverse pharmacophore mapping technology were obtained. Through comparison of the target genes, 17 repetitive targets were obtained, that is, 17 potential target genes for the treatment of NAFLD by GQD.
Molecular Docking
Pymol software and AutoDock Tools 1.5.6 were used to modify proteins and active ingredients, respectively, which were introduced into AutoDockTools 1.5.6 for molecular docking. From the docking analysis, clusters with the highest number of conformations and highest absolute binding energies were selected, and maximum absolute values for the binding energy between the compound and the target protein were obtained. Drugbank and literature searches were used to identify drugs with definitive effects on the target proteins. Structures were downloaded using molecular docking approaches as positive controls. Binding energies were obtained as a screening threshold and used to assess the binding energy between compounds and proteins. Active ingredients were identified that bind to the target proteins.
Pathway Enrichment Analysis
A total of 17 target genes were integrated and uploaded to the DAVID database. We selected “Identifier” as “OFFICIAL_GENE_SYMBOL”, and “Homo sapiens” was selected for “species” for KEGG pathway annotation. KEGG pathways were analyzed, and those enriched in the target genes were deemed important regulatory networks for the GQD treatment of NAFLD.
Constructing “Active Component-–Target-Pathway” Networks
Active ingredients, target genes, and enrichment pathways were introduced into Cytoscape 3.6.1 software to construct active ingredient-target gene networks and target gene-enrichment pathway networks, respectively. Active components and target genes within the enrichment pathways were labeled as “nodes”, and if an association between two nodes was identified, it was represented as an “edge”. Two groups of networks were merged to obtain “Active Component-Target-Pathway" networks. In this network structure, “nodes” with a high “Degree” value (compounds, targets, pathways) were deemed to be of significance.
Preliminary Verification
Data were verified in HepG2 cells. Adiponectin receptor 2 (ADIPOR2) was selected as the target and the adiponectin content of cell supernatants was determined by enzyme-linked immunosorbent assay (ELISA). We investigated whether baicalin, worenine, wogonoside, glabridin, glabrene, and purerarin act through ADIPOQ/ADIPOR2. ADIPOR2 expression and adenosine monophosphate-activated protein kinase (AMPK) activity downstream of the key mediator APPL1 were assessed by Western blot.
Cells
HepG2 cells were purchased from Shanghai enzymology Biotechnology Co., Ltd. The cell source was ATCC hb-8065, generation P3.
Reagents
Baicalin (high-performance liquid chromatography [HPLC] ≥98.0%, NO. 110715-201821), wogonoside (HPLC ≥98.0%, NO. 112002-201702), and puerarin (HPLC ≥98.0%, NO. 110752-201816) were purchased from the Chinese National Institutes for food and drug control. Worenine (HPLC ≥98.0%, NO. 20190315), glabridin (HPLC ≥98.0%, NO. 20190508), and glabrene (HPLC ≥98.0%, NO. 20190109) were purchased from Chengdu push biotechnology.
Dulbecco’s modified Eagle medium, fetal bovine serum, human adiponectin ELISA, human APPL1 antibody, human p-AMPKα antibody, human AMPKα antibody, and other Western blot reagents were purchased from Thermo Fisher Scientific (China) Co., Ltd.
Experimental assays
HepG2 cells in the logarithmic growth stage were seeded onto 96-well plates. Cells were either untreated (control) or treated with baicalin, worenine, wogonoside, glabridin, glabrene, or puerarin, 24 hours postincubation, at 37°C in 5% carbon dioxide; the control groups were treated with serum-free culture media while the administration group was administered 0.1 μmol/mL serum-free culture media for 30 minutes. ELISA was performed to assess the adiponectin levels of the supernatants.
For Western blot analysis, treated HepG2 cells were lysed and total protein content assessed via bicinchoninic acid assay. Samples were separated on 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis gels and transferred to polyvinylidene fluoride membranes. Membranes were blocked in 5% milk for 1 hour. Membranes were probed with antibodies against either p-AMPKα (1:1000), AMPKα (1:1000), APPL1 (1:1000), or glyceraldehyde 3-phosphate dehydrogenase (1:2000) overnight at 4°C. Membranes were washed in tris buffered saline-T and labeled with the appropriate horseradish peroxidase-conjugated secondary antibodies at room temperature for 2 hours. Proteins were visualized using the ECL system.
Results
Active Compound Screening
According to both screening and compensatory mechanisms, 108 compounds were identified, including 9 from Pueraria, 28 from Scutellaria, 10 from Coptidis, and 61 from Licorice. Most of the 108 compounds were flavonoids and their glycosides, followed by saponins and alkaloids. DL and ADME values of the compounds were screened for drug-like properties and listed. The results of these analyses are shown in Table 2.
Partial Active Ingredient Screening.

OB, oral bioavailability; PPB, plasma protein binding; DL, drug likeness.
Screening of Potential Targets
A total of 84 NAFLD-related genes were obtained from Malacards and GeneCards databases and compared with the results of reverse pharmacophore matching. A total of 17 potential NAFLD targets for GQD therapy were obtained. These targets were expressed as their gene symbols, as shown in Table 3.
Screening of Potential Targets.

Molecular Docking
The mol2 format of GQD active ingredients was downloaded from the TCMSP database and stored after conversion into the pdbqt format in batches. Protein X-ray diffraction structures were downloaded from the PDB database and saved in the PDB format. AutoDock Tools were used to deligand, desolvent, and hydrogenate the obtained protein structures, which were saved in the pdbdt format. Target structures and active ingredients were introduced into AutoDock Tools 1.5.6 for molecular docking. Gridbox parameters refer to the average coordinates obtained when removing protein ligands. This was performed 5 times for each protein and clusters with the largest conformations and absolute binding energies were selected, entailing the binding energy between the compound and the target protein. Through literature analysis, combined with current research hotspots, positive controls of each protein target were selected, and molecular docking was performed under identical conditions. Binding energies were obtained and used as a threshold for compound screening. For controversial positive control drugs, the first 30% (the first 32 compounds) were selected according to the binding energies of each compound/protein. 11 -21 Parameters of the positive drugs and gridboxes are shown in Table 4.
Positive Drugs and Gridbox Parameters for Each Target Protein.

Target Path Annotations
All potential target genes were integrated and uploaded onto the DAVID database for KEGG pathway annotation. The threshold value was set to P ≤ 0.05 and either higher pathway or gene functions were analyzed. Graphpad Prism 6 was used to plot the first ten items, with results shown in Figure 1.

Top 10 Pathways and Value Diagrams
KEGG pathway annotation showed that all 17 target genes were involved in pathway enrichment (100%), involving 12 pathways, 11 of which significantly correlated with the target genes (P < 0.05). The leading pathways included AMPK signaling (11, 64.7%), NAFLD (10, 58.8%), adipocytokine signaling (7, 41.2%), PPAR signaling (6, 35.3%), insulin resistance (5, 29.4%), and type II diabetes mellitus (5, 29.4%). Further comparative analysis of the 10 KEGG pathways with the strongest correlation revealed that among the 17 target genes, the associated genes included MAPK8, TNF, INSR, PPARA, PPARG, ACACACA, ADIPOQ, FASN, SREBF1, and ADIPOR1. These genes are the likely targets of GQD during NAFLD treatment.
Construction of “Active Component-Target Gene-Enrichment Pathway” Networks
The 4 pathways with the highest number of target genes were obtained from the pathways significantly correlating with target genes. These pathways, 17 target genes, and 108 components were used to construct a network of active ingredient-target gene-enrichment pathways using Cytoscape software. According to the data, compounds with a degree >10 were selected for analysis. Among them, the number of flavonoids was the highest, followed by coumarins, while the number of alkaloids was relatively low. In these compounds, 14 originated from licorice, 3 from Coptidis, 2 from Scutellaria and 1 from Pueraria. Results are shown in Figure 2 and Table 5.
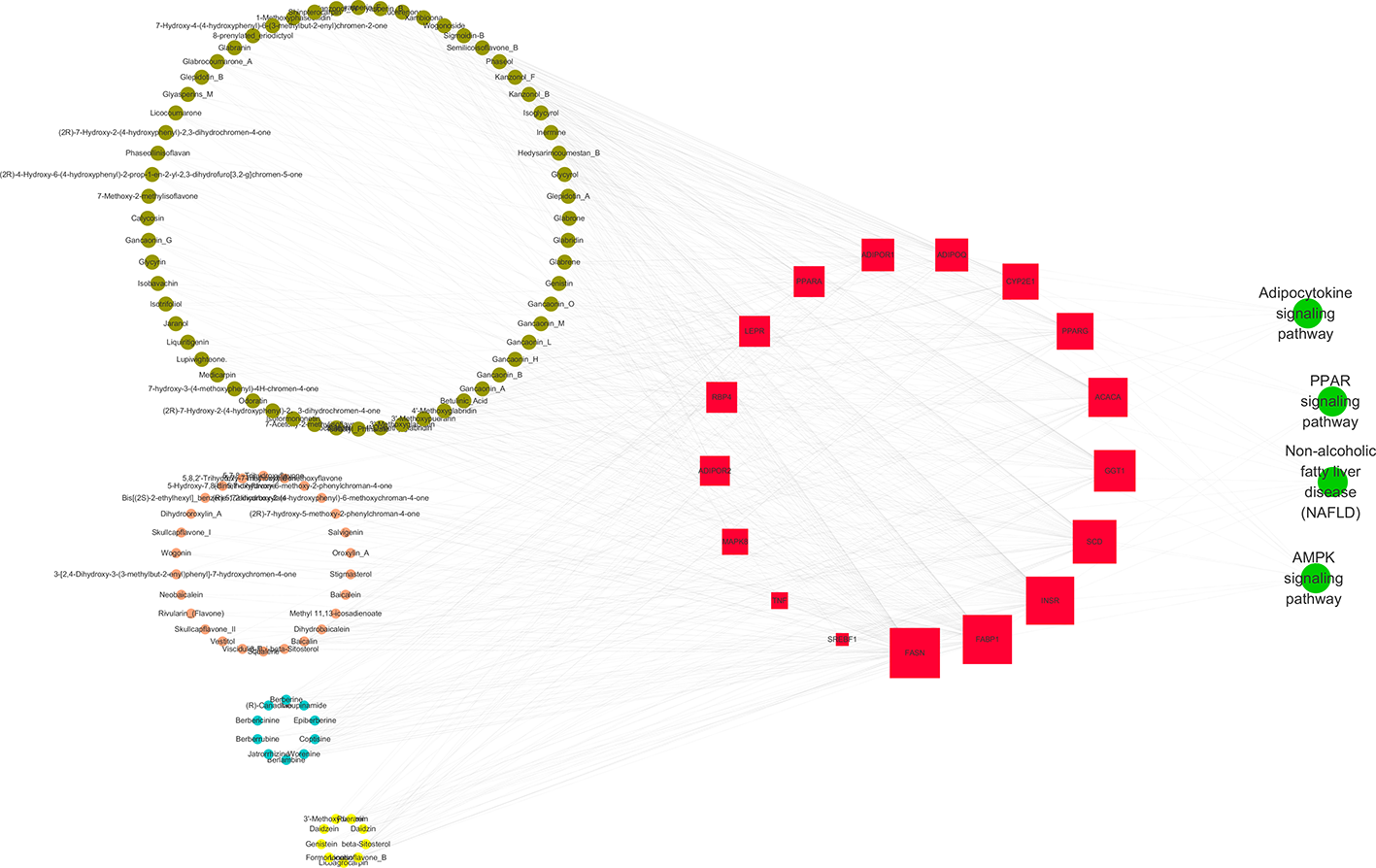

Gegen Qinlian decoction treatment of the Multidimensional Network
Partial Summary of the Cytoscape Data.

Further analysis revealed 24 compounds corresponding to 10 targets. Of the compounds, 9 out of 24 had a degree ≥10 including xambioona, baicalin, phaseo, worenine, wogonoside, inermine, glabrene, puerarin, and glabridin. These compounds are key components of GQD for the treatment of NAFLD. Their potential targets included FASN, FABP1, INSR, SCD, GGT1, ACA, PPARG, CYP2E1, ADIPOQ, and ADIPOR1.
Enzyme-Linked Immunosorbent Assay
ELISA was performed to assess the content of adiponectin in the supernatants of HepG2 cells. The 6 drug groups determined are shown in Figure 3.
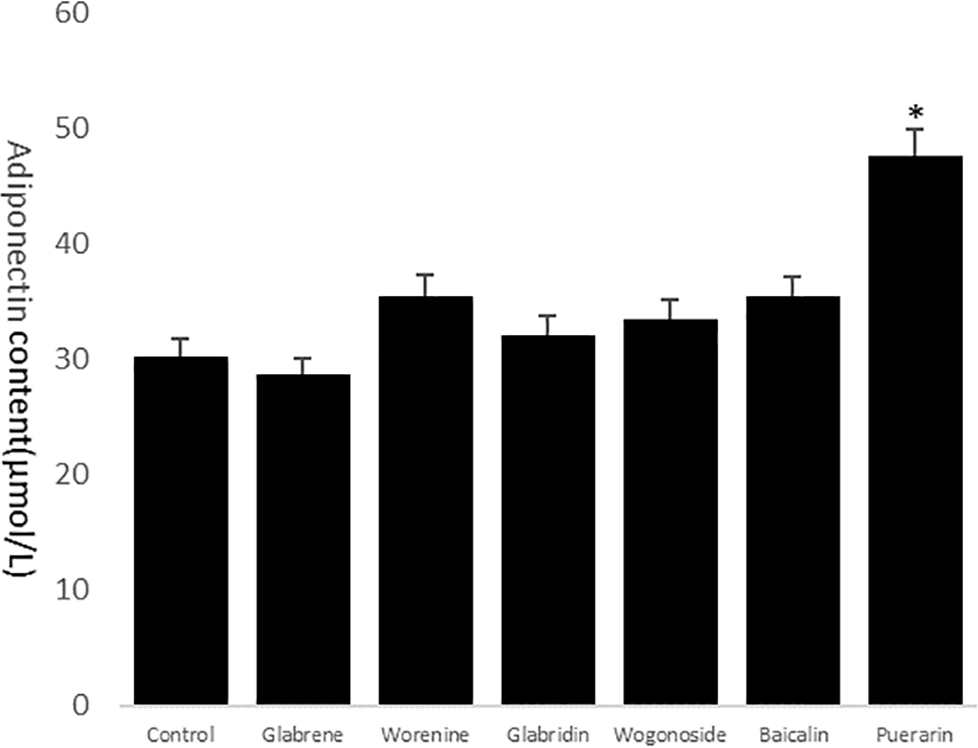
Adiponectin ELISA assays.
Purerarin was found to enhance the expression of adiponectin in HepG2 cells, showing significant differences compared with the control group. No significant effects on adiponectin expression were observed in HepG2 cells.
Western Blot Analysis
After 30 minutes of incubation, the expression of AMPKα in HepG2 cells did not differ between glabrene, worenine, glabridin, wogonoside, baicalin, and puerarin. The expression of p-AMPKα significantly increased in worenine, wogonoside, baicalin, and puerarin groups, with no changes in total AMPKα levels, confirming AMPKα activation. Worenine, wogonoside, and baicalin significantly increased the expression of APPL1 after 30 minutes of treatment, suggesting that they act on ADIPOR2. These results are summarized in Figure 4. However, the mechanism of puerarin may differ, as it is known to enhance adiponectin expression, an activator of ADIPOR2. The mechanism of action of this compound requires further elucidation in future studies. 22 -24

Western Blot Analysis.
Discussion
NAFLD is caused by liver stress that is closely related to glycolipid metabolism disorders and is characterized by hepatocyte steatosis and liver triglyceride accumulation. Its clinical manifestations include NAFL, NASH, and non-alcoholic cirrhosis. The incidence of NAFLD in the general population is as high as 20%-33%. With improved human living standards, NAFLD is increasing on a yearly basis. It has been shown that approximately 10%-25% of NAFLD patients progressively develop NASH; 40% of NASH patients develop liver fibrosis, 10%-15% of NASH patients develop cirrhosis, and 9%-26% of NASH-related cirrhosis patients die within 4-10 years of follow-up. NAFLD therefore poses a serious threat to human health. Effective preventative and therapeutic NAFLD treatments can reduce the incidence of cirrhosis, liver cancer, and reduce atherosclerotic cardiovascular and cerebrovascular diseases. 24,25
In this study, 17 target genes were selected, which were closely related to 4 pathways (AMPK, NAFLD, adipocytokine signaling, and PPAR). The relationship between GQD and these pathways is mapped in Figure 5. Many signaling pathways are dysregulated during NAFLD, with GQDs likely to exert clinical effects through these targets. Nine compounds in GQDs closely related to NAFLD pathways were identified, including xambioona, baicalin, phaseo, worenine, wogonoside, inermine, glabrene, puerarin, and glabridin. Despite the knowledge that Puerariae, Scutellaria, Coptis, and Licorice display anti-NAFLD effects, their mechanism(s) of action are poorly defined. 26,27

Pathway Maps of GQD for NAFLD
ADIPOR2 is expressed in liver cells and plays an important role in glycolipid metabolism. When activated, the receptor recruits APPL1 to directly or indirectly modulate cell signaling pathways (including AMPK). ADIPOR2 activity can be assessed through APPL1 expression. The activation of this pathway is also dependent on AMPK signaling. AMPK consists of 3 subunits: α, β, and γ 28,29 ; Thr172 represents the active site of the α-catalytic subunit and the levels of p-AMPKα directly reflect the levels of AMPK activation. It is speculated that the compounds influence lipid metabolism in HepG2 cells through the ADIPOR2/APPL1/AMPK axis to alleviate the effects of NAFLD. 30 Specifically, worenine, wogonoside, and baicalin significantly increased the expression of APPL1 and p-AMPKα in HepG2 cells, suggesting that they act on ADIPOR2. These data verified the network pharmacology data. Of note, the expression of APPL1 did not change upon puerarin treatment, despite the activation of p-AMPKα. Therefore, puerarin may not activate AMPK through the ADIPOR2 receptor and its specific mechanisms require further investigation. 26
In summary, we have studied the pharmacodynamics and mechanisms of GQD action during the treatment of NAFLD through network pharmacology. Through a series of measurements, we screened 9 potentially active substances in GQD, 10 potential targets related to NAFLD, and 4 pathways likely to exert their pharmacodynamic effects. It was confirmed that worenine, wogonoside, and baicalin act upon their predicted targets and that ADIPOR2 activates AMPK signaling to regulate lipid metabolism. These data highlight the reliability of the network pharmacology data and support the treatment of NAFLD using Gegen Qinlian decoction.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
