Abstract
Alcoholic liver disease (ALD) progresses from steatosis to alcoholic hepatitis to fibrosis and cirrhosis. Liver biopsy is considered as the gold standard method for diagnosis of liver cirrhosis and provides useful information about damaging process which is an invasive procedure with complications. Existing biomarkers in clinical practice have narrow applicability due to lack of specificity and lack of sensitivity. The objective of this article is to identify proteomic biomarker candidates for alcoholic liver cirrhosis by differential expression analysis between alcoholic liver cirrhotic and healthy subjects. Blood samples were collected from 20 subjects (10 alcoholic liver cirrhosis and 10 healthy) from R. L. Jalapa Hospital and Research Centre, Kolar, Karnataka, India. Differential protein analysis was carried out by two-dimensional electrophoresis after albumin depletion, followed by liquid chromatography–mass spectrometry. The image analysis found 46 spots in cirrhotic gel and 69 spots in healthy gel, of which 14 spots were identified with significant altered expression levels. Based on the protein score and clinical significance, among 14 spots, a total of 28 protein biomarker candidates were identified: 13 with increased expression and 15 with decreased expression were categorized in alcoholic liver cirrhosis compared to healthy subjects. Protein biomarker candidates identified by “-omics” approach based on differential expression between alcoholic liver cirrhotic subjects and healthy subjects may give better insights for diagnosis of ALD. Prioritization of candidates identified is a prerequisite for validation regimen. Biomarker candidates require verification that demonstrates the differential expression will remain detectable by assay to be used for validation.
Keywords
Introduction
Cirrhosis of the liver is the histological development of regenerative nodules surrounded by fibrous bands in response to chronic liver injury, leading to portal hypertension and end-stage liver disease. Causes of cirrhosis of the liver are multifactorial. Despite varied etiology, the pathological characteristics which result in liver dysfunction are common. 1 In recent years, alcohol consumption has correlated with deaths from asymptomatic and self-limited fatty liver to cirrhosis of the liver. Alcoholic liver disease (ALD) progresses from steatosis to alcoholic hepatitis to fibrosis and frank cirrhosis (micronodular, occasionally mixed micronodular, and macronodular) and often occurs acutely against background chronic liver disease. 2 Multifactorial pathogenesis plays a role in progression of the disease. Accumulation of triacylglycerol in liver is an early and reversible effect of alcohol which increases peripheral lipolysis and alters liver redox potential leading to fatty acid synthesis. In ALD, generated prooxidants enhance antioxidant system results in lipid peroxidation. 3
Acetaldehyde generated from alcohol is highly reactive and toxic; it binds to phospholipids and amino acids, resulting in the formation of abnormal folding of proteins in endoplasmic reticulum (ER) leading to ER stress. 3 Proliferation and activation of hepatic stellate cells (HSCs) in ALD are induced by Kupffer cells and hepatocytes. Chief mitogen for the activation of HSCs is platelet-derived growth factor, which is produced by Kupffer cells. Kupffer cells induce collagen synthesis through the production of transforming growth factor β, tumor necrotic factor α (TNF-α), and reactive oxygen species (ROS). Activated HSCs migrate and accumulate at tissue repair sites and secrete large amounts of extracellular matrix (ECM). HSCs collagen synthesis is regulated at transcription and posttranscriptional levels. Dysregulation of cytokine metabolism and activity is vital for alcohol-induced liver damage. TNF-α, a pro-inflammatory cytokine, is one of the key factors for pathophysiology of ALD. Hepatocytes induce fibrosis through the production of ROS or apoptosis followed by regenerative nodular formation. 3,4
Diagnosing the degree of disease is a crucial step for successful management of ALD. Despite the development of potential diagnostic tests for the past 50 years, liver biopsy is considered as the gold standard method for diagnosis of liver cirrhosis and provides useful information about damaging process, namely, steatosis, lobular inflammation, periportal fibrosis, Mallory bodies, nuclear vacuolation, bile ductal proliferation, and fibrosis/cirrhosis. 2,5 Limitation of liver biopsy is highly invasive; poor sample quality and tissue size make biopsy nonreproducible, and it depends on the experience of the pathologist leading to interobserver variations. Risk allied for liver biopsy range from pain (84%) and hypertension, bleeding (0.5%), and damage to biliary system with approximately 0.01% mortality rate. 6 No single biomarker can establish alcohol to be the etiology of liver disease; existing biomarkers in clinical practice have narrow applicability due to lack of specificity and lack of sensitivity (distinguish intermediate stages) which prevent reliance on any single biomarker. 2,7
An ideal biomarker should be organ specific, a sensitive indicator for active damage, easily accessible in peripheral tissue, and cost-effective and should give insights for diagnosis, monitor the activity of disease, and assess therapeutic response. The determination of biomarkers could be an easy, noninvasive, and inexpensive method to monitor the progression of liver disease. This leads to urgency in the progression of biomarker discovery for cirrhosis of the liver with the help of technological advancement in “-omics” approach. Discovery of biomarker candidates should be a simplified, unbiased, semiquantitative binary comparison between diseased and normal. 8 Although individual sample analysis is recommended, pooling strategy with definite selection criteria from multiple individuals reduces sample number and cost. 9 Alterations in protein domain due to ALD which enters into circulation hold good for discovery of biomarker candidates. In the present study, we tried to discover protein biomarker candidates for alcoholic liver cirrhosis whose concentration may be altered due to changes in translation, posttranslational modifications, and/or degradation using two-dimensional electrophoresis (2DE) after albumin depletion, followed by liquid chromatography–mass spectrometry (LC-MS).
Materials and methods
Samples
Blood samples were collected from 20 subjects: 10 clinically and diagnostically proven alcoholic liver cirrhotic subjects with varying degree and age- and gender-matched 10 healthy subjects (Table 1) from R. L. Jalappa Hospital and Research Centre, attached to Sri Devaraj Urs Medical College, a constituent of Sri Devaraj Urs Academy of Higher Education, Kolar, Karnataka, India. Individuals diagnosed with cirrhosis of the liver caused by ALD based on clinical history and symptoms, namely, ascites, encephalopathy, jaundice, and altered biochemical parameters, were included in the study. Individuals with diabetes and/or its complications, myocardial infarction, acute and chronic renal failure, pneumonia, and cancer were excluded from the study. Collection of blood samples from cirrhotic liver subjects and healthy subjects was carried out after obtaining informed consent, and the study is approved by Institutional Ethical Committee (DMC/KLR/IEC/61/2016-17).
Details of 20 blood samples (10 alcoholic liver cirrhotic subjects and age- and gender-matched 10 healthy subjects) used for discovery of biomarker candidates by proteomic approach.
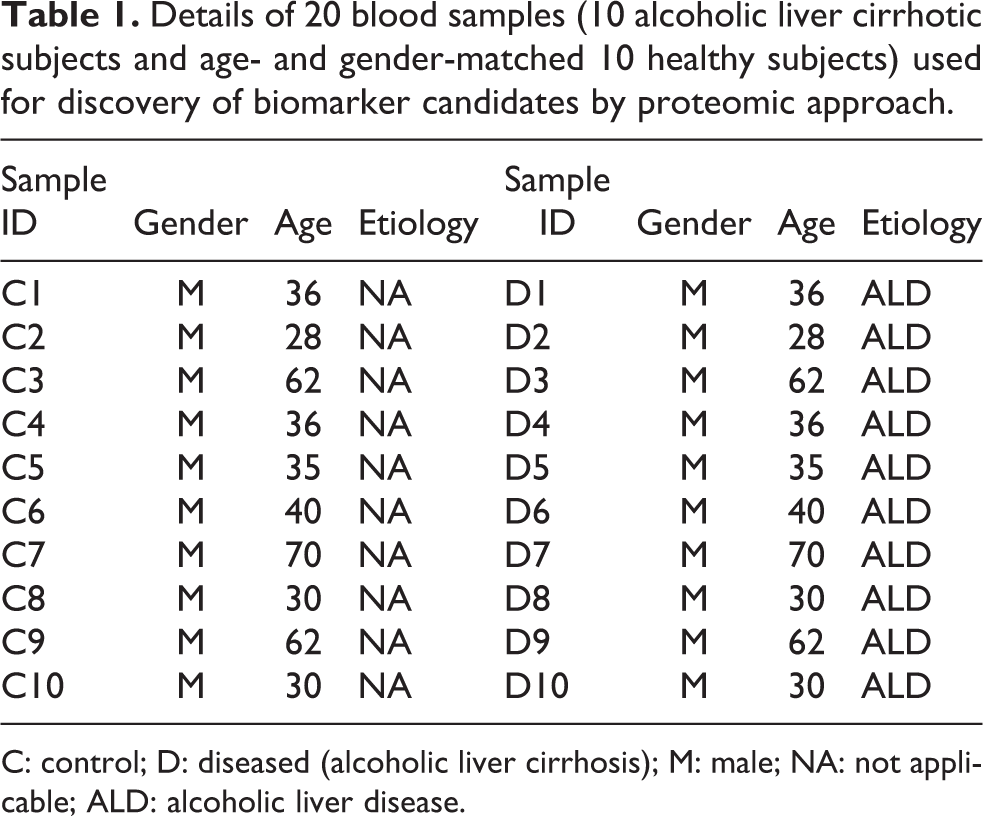
C: control; D: diseased (alcoholic liver cirrhosis); M: male; NA: not applicable; ALD: alcoholic liver disease.
Serum separation
Serum was collected from clotted blood using serum separator tubes centrifuged at 4000 r/min for 10 min. Serum was stored at −20°C till further analysis. All samples were used for discovery of protein biomarker candidates after depletion of abundant albumin. Desalting was carried out by acetone precipitation. 2DE was carried out to find differentially expressed proteins between cirrhotic and healthy subjects. Identified spots were characterized by LC-MS after in-gel trypsin digestion.
Reagents
Dye-based (Cibacron blue) prefractionation albumin depletion kit was procured from Thermo Fisher Scientific (Waltham, Massachusetts, USA). Precast gels and other chemicals of analytical grade for sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE), 2DE, in-gel trypsin digestion, and LC-MS were procured from Bio-Rad (Hercules, California, USA) and Sigma Aldrich (St Louis, Missouri, USA).
Depletion of albumin
Resuspended resin (200 μL aliquot of resin) was transferred into spin column (column volume: 900 μL; 10-μm pore-size polyethylene filter). Bottom of the column was twisted off and placed in a 1.5-mL collecting tube. Then, the sample is centrifuged at 12,000 × g for 1 min, the flow-through is discarded, and finally the spin column back is placed into the same collection tube. Around 200 μL of binding/wash buffer was added to the spin column. Then, the sample is centrifuged at 12,000 × g for 1 min, the flow-through is discarded, and, finally, the spin column back is placed into the new collection tube. About 50 μL of pooled serum sample (cirrhotic and healthy in separate columns) was added into resin and incubated for 2 min at room temperature. Then, the sample is centrifuged at 12,000 × g for 1 min, flow-through is reapplied to spin column, and incubated for 2 min at room temperature. Again, the sample is centrifuged at 12,000 × g for 1 min and flow-through is retained. Spin column was placed in a new collection tube. Resin was washed to release unbound proteins by adding 50 μL of binding/wash buffer for each 200 μL of the resin used. Retained fractions of cirrhotic and healthy samples were run in SDS-PAGE for confirmation of albumin depletion. 10
Sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE)
SDS gel was prepared according to the standard protocol. Samples from cirrhotic liver subjects and healthy subjects after depletion of albumin along with un-depleted samples were loaded in gel, and SDS-PAGE was carried out at 25 mA in 1× SDS running buffer for confirmation of depletion along with molecular weight marker. After electrophoresis, gel was incubated in a fixing solution (40% methanol, 10% acetic acid) at room temperature for 20 min. Gel was subjected for staining with sliver stain (0.1% silver nitrate, 36% formaldehyde) at room temperature for 20 min. Excess staining solution was removed, and the gel was washed with 5% acetic acid. 11 –13
Acetone precipitation
Acetone precipitation was carried out to remove excess salts which interferes electrophoretic run. Protein samples after depletion of albumin (cirrhotic and healthy separately) were placed in acetone-compatible tubes. Four times the sample volume of cold acetone (−20°C) was added into both tubes. The tubes were vortexed and incubated for 60 min at −20°C, followed by centrifugation at 13,000 × g for 10 min. Then the supernatant was disposed carefully for the retention of protein pellet. 14
Two-dimensional polyacrylamide gel electrophoresis
Albumin depleted and desalted protein pellet (200 μg) from cirrhotic and healthy subjects were diluted with rehydration buffer (8 M urea, 2% 3-((3-cholamidopropyl) dimethylammonio)-1-propanesulfonate, 50 mM dithiothreitol (DTT), 0.2% w/v Bio-Lyte 3/10 ampholyte, bromophenol blue) and used separately for 2DE with 7 cm (pH 4–7) nonlinear immobilized pH gradient dry strips (Bio-Rad). Samples were left overnight for rehydration on 7 cm (pH 4–7) dry strips. Isoelectric focusing was carried out at 250 V for 20 min, followed by 4000 V for 5 h at 20°C. Proteins were separated by 8–16% precast polyacrylamide gels at 200 V for 40 min. After electrophoresis, gels were subjected to staining with silver stain (0.1% silver nitrate, 36% formaldehyde) at room temperature for 20 min. Excess staining solution was removed, and the gel was washed with 5% acetic acid. 15
Image analysis
For image analysis, scanned gels were processed using PDQuest 2-D analysis software (Bio-Rad). For differential analysis, the cirrhotic gel was compared with that of the healthy gel. Differential expression of proteins present in both cirrhotic and healthy gels was considered significant when the fold change was least 2 and p ≤ 0.05 with 95% confidence interval with the application of rank-sum test.
In-gel digestion and peptide extraction
Excised spots were cut into cubes and transferred into a microcentrifuge tube, and 100 μL of destaining solution (100 mM ammonium bicarbonate/acetonitrile (1:1 vol/vol)) was added and incubated for 30 min; 500 μL of neat acetonitrile was added and the tubes were incubated for 10 min until gel pieces shrink; 50 μL of DTT solution (10 mM DTT in 100 mM ammonium bicarbonate buffer) was added to cover the gel pieces and incubated in 56°C thermostat for 30 min; and 500 μL of acetonitrile was added to the tubes and further incubated for 10 min. All the liquid was removed from the tube. Following DTT treatment, to get reduction and alkylation of cystines and cysteines in the protein, 50 μL of iodoacetamide solution (55 mM iodoacetamide in 100 mM ammonium bicarbonate solution) was added to the tubes and incubated for 20 min at room temperature in dark. The gel pieces were again treated with acetonitrile for 10 min, and the entire liquid was removed from the tube. The gel pieces were saturated with trypsin buffer (13 ng/µL of trypsin in 10 mM ammonium bicarbonate in 10% acetonitrile) for 30 min. Cold trypsin (20 µg of trypsin in 1.5 mL of ice-cold 1 mM hydrochloric acid) was added to the tubes and incubated overnight at 37°C. Tubes were cooled to room temperature, gel pieces were centrifuged at 10,000 r/min for 1 min, and peptides were extracted in 100 µL of extraction buffer (1:2, 5% formic acid/acetonitrile) by incubating for 15 min at 37°C shaker, and the supernatant was withdrawn directly for LC-MS analysis. 16
Mass spectrometric analysis
Mass spectrometric analysis of the extracted peptides was performed using Nano LCMS-LTQ-Orbitrap Discovery (Thermo Scientific) coupled to Nano LC (Agilent 1200). The samples were reconstituted in 0.1% formic acid prior to injection; 70-min gradient run was set up using acetonitrile and water with formic acid as the mobile phase. LTQ Orbitrap Discovery is a hybrid-type MS system with the ability to determine accurate m/z of intact precursors. The raw files post-MS run was analyzed using Proteome Discover software and MASCOT as search engine against human database. 17
Results
SDS-PAGE analysis for depletion of albumin
Immobilized resin form of Cibacron blue was effective in binding abundant albumin from plasma/serum samples for depletion of significant amount. Human serum albumin (HSA) from pooled serum samples of cirrhotic and healthy subjects was depleted using Cibacron blue dye-loaded resin columns. Proteins in the flow-through were analyzed by SDS-PAGE (Figure 1) along with prestained molecular weight marker to investigate efficient depletion of HSA. Sensitive staining, silver stain, was helpful for the detection of low nanogram proteins when compared with Coomassie Brilliant Blue. Silver-stained gel demonstrated significant amount of abundant albumin depletion from serum samples of both cirrhotic and healthy subjects.
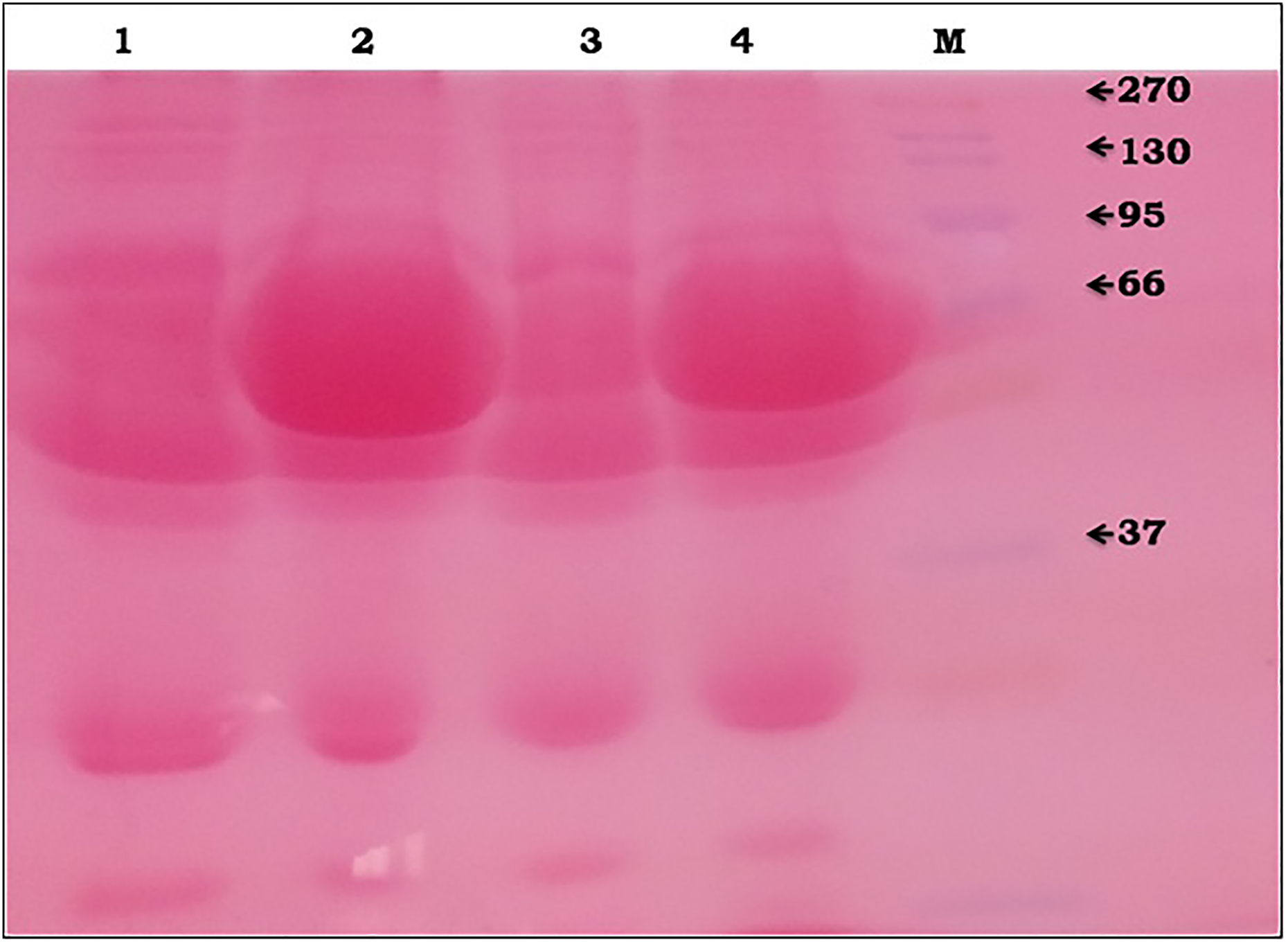
SDS-PAGE analysis for confirmation of albumin depletion (silver-stained gel). 1: Normal pooled albumin depleted serum. 2: Normal pooled serum. 3: Cirrhotic liver pooled albumin depleted serum. 4: Cirrhotic liver pooled serum. M: pre-stained molecular weight marker. SDS-PAGE: sodium dodecyl sulfate polyacrylamide gel electrophoresis.
Identification of biomarker candidates
Synthetic gel image representative of all features in the differential analysis comparing samples from cirrhotic and healthy is shown in Figure 2. The image analysis software and statistical analysis found 46 spots in cirrhotic gel and 69 spots in control gel, of which 14 spots were identified with significant altered expression levels between cirrhotic and healthy based on quantitative ratio. These spots were excised, digested with trypsin, analyzed by LC-MS, and identified by MASCOT database. These spots contain more than one protein; among 14 spots, a total of 68 proteins were identified. Many of the proteins were identified as the same protein in different locations on the gels and so among 68 proteins, we identified 46 candidate biomarkers for alcoholic liver cirrhosis. Among 46 candidate biomarkers, 28 were identified based on protein score and clinical significance (Table 2).
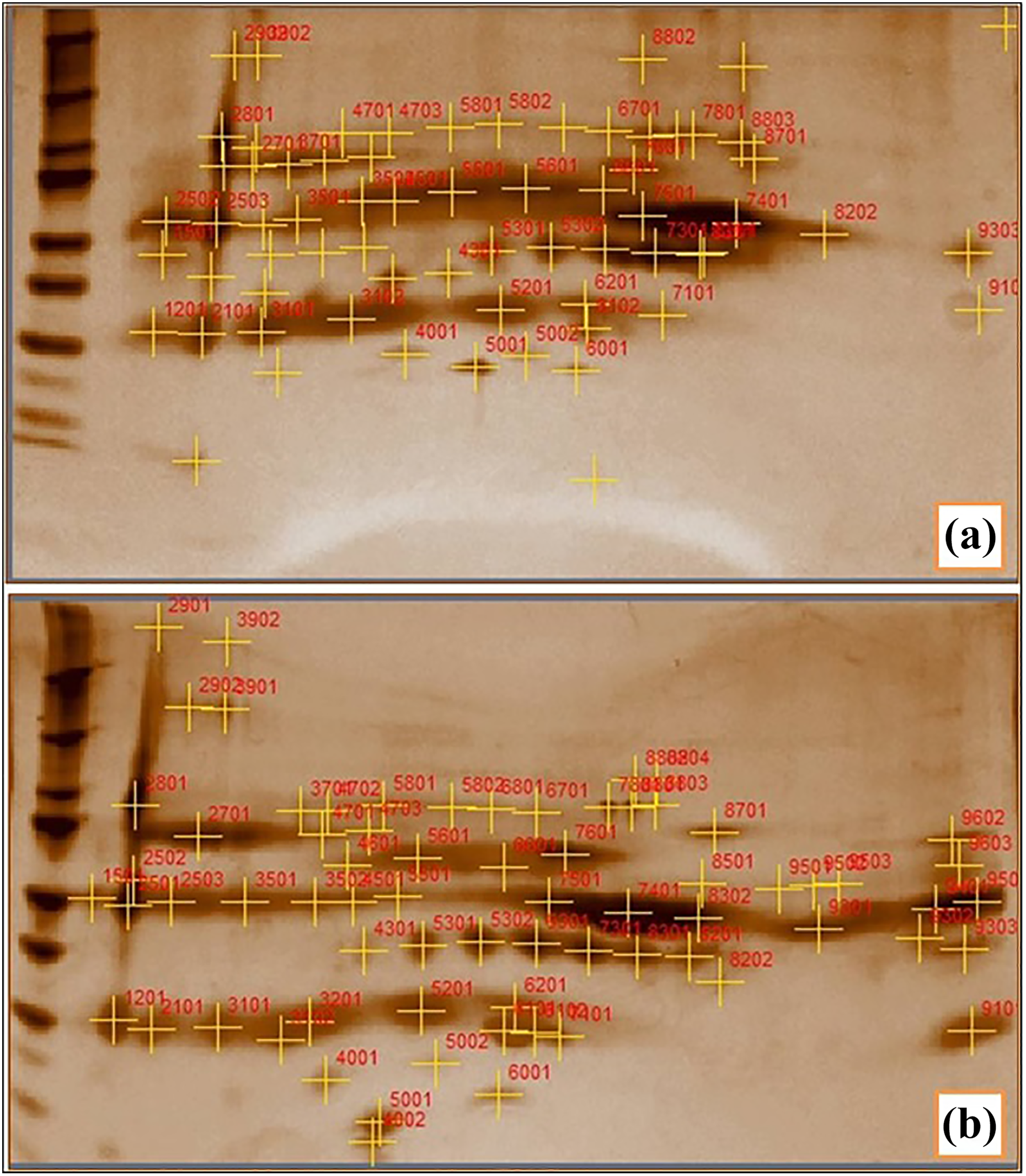
Comparison of 2DE gel images representative of all features in differential analysis from alcoholic cirrhotic and healthy subjects. (a) 2DE gel image of alcoholic liver cirrhotic subjects. (b) 2DE gel image of healthy subjects. 2DE: two-dimensional electrophoresis.
Proteomic biomarker candidates identified by 2DE followed by LC-MS for alcoholic liver cirrhosis.
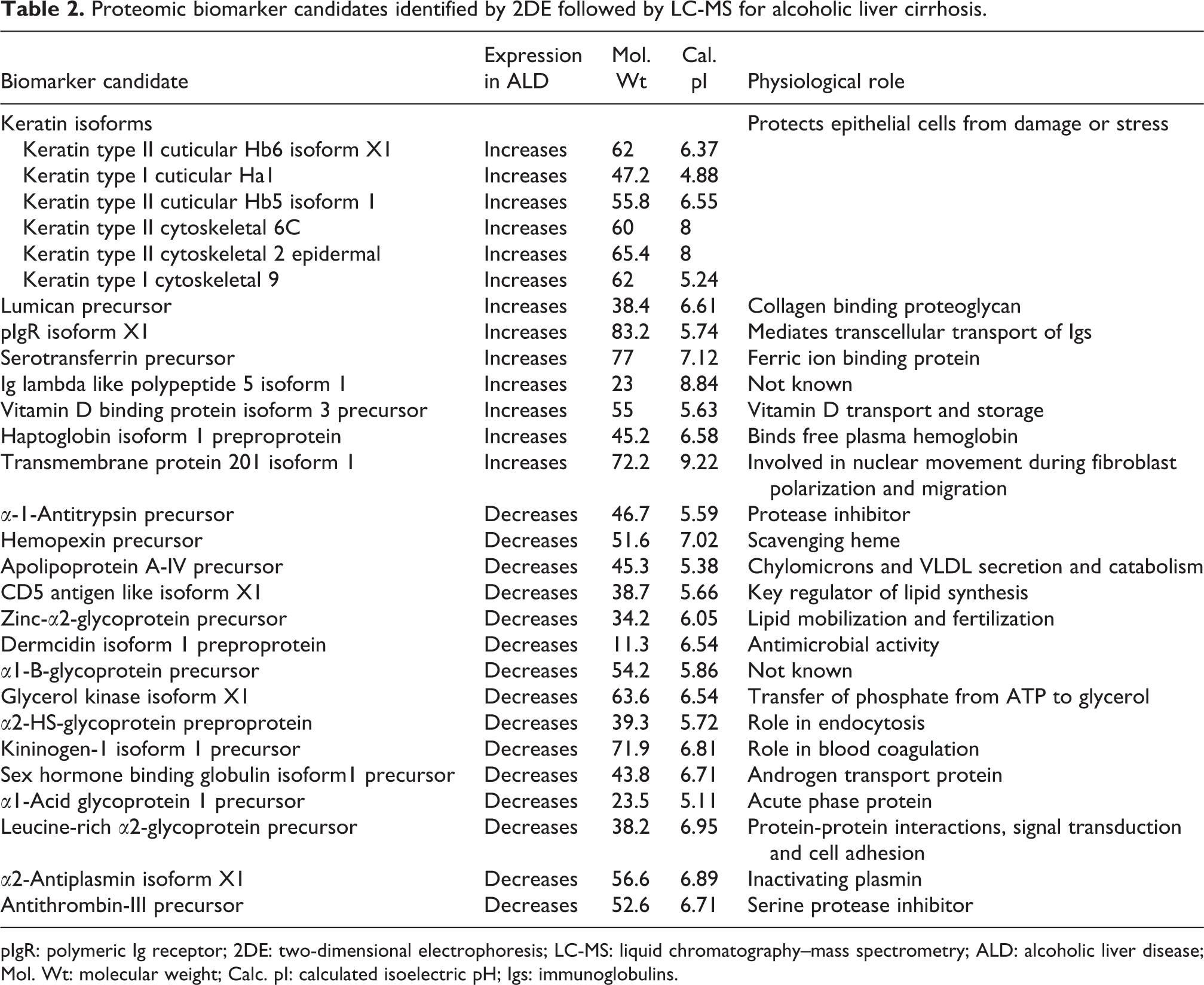
pIgR: polymeric Ig receptor; 2DE: two-dimensional electrophoresis; LC-MS: liquid chromatography–mass spectrometry; ALD: alcoholic liver disease; Mol. Wt: molecular weight; Calc. pI: calculated isoelectric pH; Igs: immunoglobulins.
Among 28 protein biomarker candidates, 13 with increased expression and 15 with decreased expression were identified in alcoholic liver cirrhotic when compared to healthy subjects. Serum concentrations of keratin isoforms were found to increase in alcoholic cirrhosis. Immunoglobulins (Igs), namely, polymeric immunoglobulin receptor isoform X1 and IgGFc-binding protein precursor, were increased in cirrhotic liver when compared to healthy subjects but with low protein scores. Along with 13 features whose expression is increased in cirrhosis of the liver, angiotensinogen preproprotein, α2-macroglobulin isoform X1, were found to increase compared to healthy subjects. Serum albumin preproprotein, α-1-antitrypsin precursor and α-1-antichymotrypsin precursor showed decreased expression in alcoholic liver cirrhosis. Concentrations of glycerol kinase isoform X1 and kininogen-1 isoform 1 precursor were decreased in cirrhotic liver compared to healthy subjects.
Discussion
Invasive liver biopsy is the gold standard diagnostic tool for liver fibrosis/cirrhosis with varied etiology and to distinguish between intermediate stages. Reliable noninvasive biomarker with sensitivity and specificity is needed for diagnosis/prognosis and effective management of the disease. 8 In the present study, we used 2DE followed by LC-MS for identification of biomarker candidates for alcoholic liver cirrhosis. For maximal detection of meaningful protein expression difference, cases and controls should differ absolutely in terms of disease of interest. Simplified, unbiased binary comparison between diseased and healthy avoids contamination by other diseases and confounding factors which may alter the expression of protein results in false discovery of biomarker candidates. 18
Discovery of biomarker candidates by proteomic approach is difficult, especially when the pH range is between 3 and 7, as abundant albumin interferes in identification and characterization of low abundant proteins by mass spectral and electrophoretic analysis. Accurate protein biomarker candidate discovery was achieved after depletion of albumin using dye-based affinity columns. Antibody-based immunoprecipitation is more robust for depletion of abundant proteins from plasma/serum and is suitable for identification of novel biomarker candidates. 19 –21 Depletion dilemma can be rectified using narrow pH (3–5.6) range and avoids interference of abundant proteins (albumin, transferrin, and Igs) but chance to miss proteins whose isoelectric pH is in the alkaline range. 22
Technological advancement in biomarker candidate discovery resulted in identification of protein biomarker candidates for chronic liver diseases (CLDs) with varied etiology (Table 3). Biomarker candidates identified require verification which demonstrates that the differential expression should remain detectable by assay to be used for validation. 18 Despite numerous biomarker candidates identified, verification may be done only for few qualified candidates in terms of marker performance and reagent availability. 27 Biomarker candidates that show significant expressional differences between diseased and healthy in discovery phase are prioritized. Proteins that are secreted and/or present on cell surface and that act in cellular pathways and deregulated in ALD should be considered for further validation. 9
Protein biomarker candidates identified by proteomic analysis for liver fibrosis.
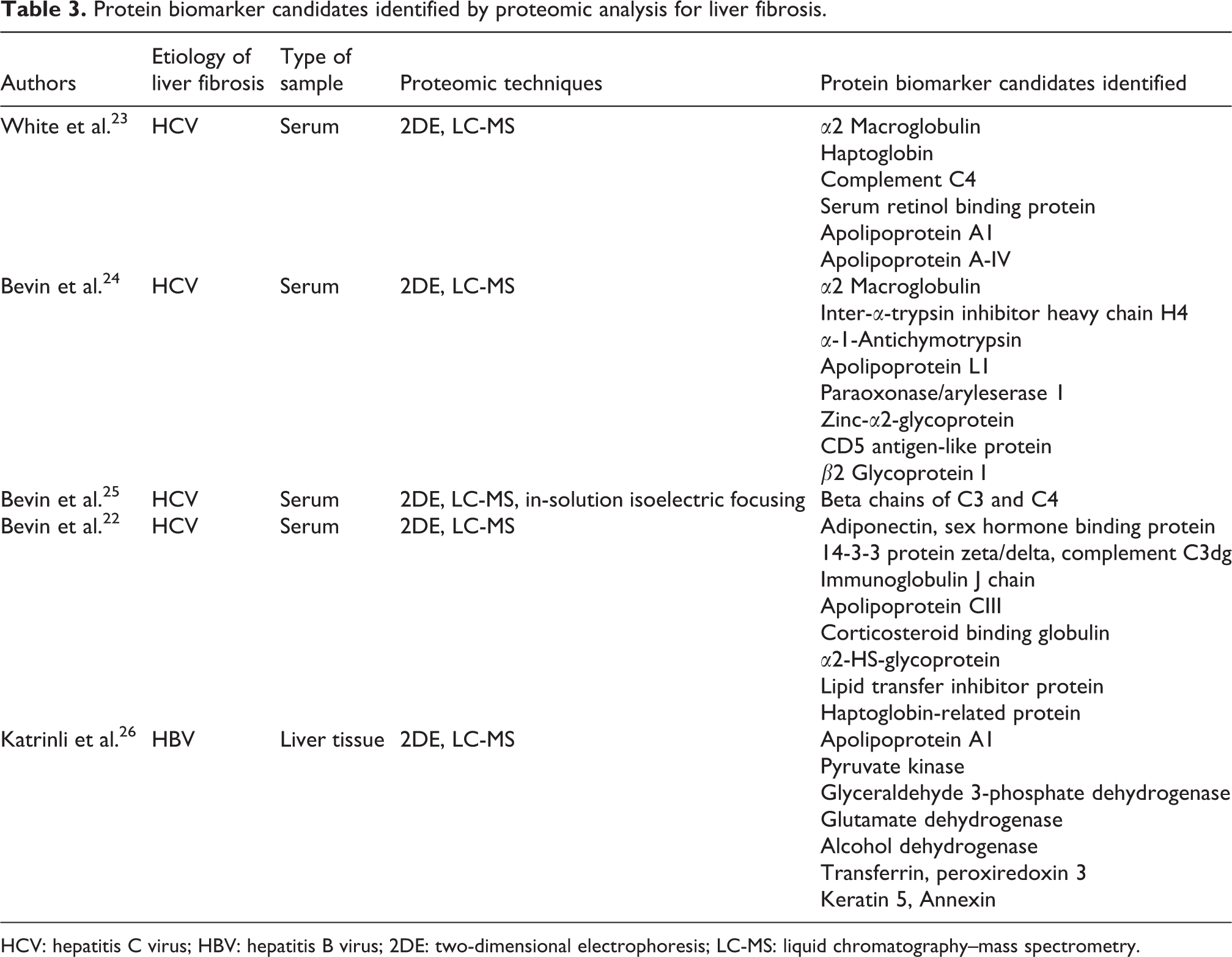
HCV: hepatitis C virus; HBV: hepatitis B virus; 2DE: two-dimensional electrophoresis; LC-MS: liquid chromatography–mass spectrometry.
In the present study, keratin isoforms showed upregulation in alcoholic liver cirrhosis. Keratin is a fibrous structural protein that protects epithelial cells from damage and stress and regulates key cellular activities, namely, cell growth and protein synthesis. 28 Lumican, leucine-rich repeat proteoglycan, constitute an important fraction of noncollagenous ECM proteins. It plays a major role in tissue homeostasis and modulates cellular functions, namely, cell proliferation, migration, and differentiation. 29 Polymeric Ig receptor (pIgR) isoform X1 is a type-I transmembrane protein expressed from glandular epithelial cells of liver and breast. It mediates transcellular transport of polymeric Igs. Pro-inflammatory cytokines, namely, interferon-γ, TNF, and interleukin 1, which are the key regulators of pIgR expression, upregulate in ALD. 8,30,31
Vitamin D-binding protein, a multifunctional protein that belongs to the albumin gene family can bind various forms of vitamin D (ergocalciferol, cholecalciferol, and calcifediol) for the transport. It is synthesized by hepatic parenchymal cells. 32 Haptoglobin which is included in the existing noninvasive marker panel has showed increased expression in the present study as it is an acute phase protein. Liver is the major site for the synthesis of haptoglobin; hepatic expression will be stimulated by upregulated IL-6 in ALD. 23,33,34 Transmembrane protein 201 is involved in nuclear movement during fibroblast polarization and migration; actin-dependent nuclear movement is through association with transmembrane actin-associated nuclear lines. 35
Serine protease inhibitors, α-1-antitrypsin (SERPINA1) and α-1-antichymotrypsin (SERPINA3), produced primarily in liver hepatocytes and released directly into the blood stream showed downregulation in alcoholic liver cirrhosis compared to healthy subjects. 36 –38 Hemopexin which showed decreased expression in ALD is a single polypeptide chain of 439 amino acid residues with a molecular weight of 63 kDa is expressed from liver, and it acts as a heme-scavenging protein. 39 Apolipoprotein A-IV, even though not evident from liver, its expression, was also decreased in ALD. Downregulation of Apolipoprotein A-IV was reported in hepatic fibrosis in rat models. 23,40 CD5 antigen-like isoform X1, a key regulator of lipid synthesis, was downregulated in alcoholic liver cirrhosis, whereas upregulation was noted in liver cirrhotic patients in hepatitis C virus infection. 24 Zinc-α2-glycoprotein, adipokine, which plays an important role in fat catabolism and which reduces insulin resistance, was downregulated in ALD. 24,41
Glycerol kinase, a phosphotransferase and a key enzyme in the regulation of glycerol uptake and metabolism, is involved in triglyceride and glycerophospholipid synthesis. Glycerol kinase converts glycerol, a product of lipolysis to glucose in the liver, and shows downregulation in alcoholic liver cirrhosis. In the present study, α2-HS-glycoprotein has shown decreased expression in ALD. It is a secretory protein expressed from liver and key regulator in inhibition of vascular calcification, bone metabolism regulation, control of protease activity, insulin resistance, and breast tumor cell proliferative signaling. 42 α-1-Acid glycoprotein, an acute phase synthesized primarily in hepatocytes, which acts as a carrier of lipophilic compounds was down regulated in ALD. 43 An SERPINF2, α2-antiplasmin, synthesized in the liver as a single-chain glycoprotein with a molecular weight of 51 kDa inhibits plasmin and was decreased in liver cirrhotic patients. 44 Antithrombin III, a member of the serpin family and an inhibitor of proteinases, namely, thrombin and factor Xa, is primarily synthesized by hepatocytes and downregulated in ALD. 45 Our studies are also corroborated with the downregulation of α2-antiplasmin and antithrombin III in ALD.
Newly identified proteomic biomarker candidates for ALD need validation and clinical assay optimization which require measurement of thousands of patient samples with narrow measurement coefficient of variation values. 18 Concentration of proteins in serum/plasma ranges from picograms to nanograms per milliliter. Enzyme-linked immunosorbent assay is the best alternative for quantification of these protein candidates for ALD with high specificity and sensitivity capture and detection antibodies. Newly developed assay requires analytical validation before evaluating clinical utility in terms of performance characteristics, namely, outcome studies, clinical requirement, proficiency testing, and goals set by regulatory agencies. 46 Indicators of accuracy, precision, analytical measurement range, and reference intervals should be defined for newly discovered biomarker candidates. 47 After analytical validation of new methodology for protein of interest, biomarker candidate should confirm the performance characteristics in terms of consistency and accuracy in clinical evaluation to diagnose or predict the clinical outcome of ALD. The newly identified biomarker candidate should satisfy specificity and sensitivity. Evidence-based biomarker should fulfill regulatory requirements before introduced into clinical practice for ALD. 18,48
Conclusion
With the help of technological advancement in “-omics” approach, we identified 28 protein biomarker candidates (13 with increased expression and 15 with decreased expression) for alcoholic liver cirrhosis. Despite numerous biomarker candidates identified, verification may be done only for few qualified candidates that act in cellular pathways and deregulated in ALD. These differentially expressed proteins between alcoholic cirrhosis and healthy subjects need to be validated to get the same differential expression detectable by assay to be used for validation.
Footnotes
Acknowledgements
The authors thank Dr Kiranmayee P, SDUAHER for her timely suggestions to carry out this research work.
Declaration of Conflicting Interests
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
