Abstract
Berberine (BBR) is a natural active principle with potential antitumor activity. The compound targets multiple cell signaling pathways, including proliferation, differentiation, and epithelial–mesenchymal transition. The aim of this study was to elucidate the mechanisms behind the anticancer activity of BBR by comparing the effects of purified BBR with those of the extract of
Introduction
The anticancer activity of natural origin compounds has been recently focused by several studies. 1 In this context, plants used in the traditional medicine systems can be considered a valuable source of active compounds and important supplements for chemotherapy/treatment. 2
Among the secondary metabolites produced by this plant, isoquinoline alkaloids such as berberine (BBR) are claimed to be responsible for most of the beneficial effects attributed to the plant source. For instance, BBR has been used to treat different types of diarrhea,4,5 and more recently, it has been found effective in the treatment of gastroenteritis, diabetes, hyperlipidemia, cardiovascular diseases, inflammatory conditions, hyperglycemia, and obesity.6–9 Moreover, this compound is endowed with anticancer activity on different cell lines. 10
In particular, regarding colon cancer cells, previous studies have shown the ability of BBR to induce cells death through different mechanisms, such as: activation of JNK/p38 apoptotic pathway, suppression of the Wnt/β-catenin signaling pathway and inhibition of growth/proliferation processes for example by inducing G1/S and G2/M cell cycle phase arrest.11,12
Nevertheless, only few studies have been focused on the gene expression effects of BBR in colon cancer cells. Specifically, Hu et al. 13 reported a strong dose-dependent antiproliferative BBR activity on human colon cancer cells (HT-29), together with a gradual increasing of apoptotic cells proportion associated with a reduction of the expression of the survivin gene. Moreover, BBR has been reported to increase nonsteroidal anti-inflammatory drug-activated gene-1 (NAG-1) expression in HCT-116 and CaCo-2 cells. This gene has been investigated for its proapoptotic and anti-tumorigenic activity, since it is highly expressed in colon but significantly reduced in most human tumor biopsies. 14
With the aim to explore the potential chemopreventive activity of
This evaluation could lead to the identification of genes and cellular pathways involved in colon cancer facilitating the identification of possible therapeutic targets.
Materials and methods
Plant material and preparation of extracts
Dried and powdered
Extraction was performed by treating 300 g of plant with 2 L of MeOH/H2O (1:1) and sonication for 1 h. The extract was filtered on Büchner funnel and the solvents were evaporated under vacuum at 40°C. The procedure was repeated for three times, obtaining 28.4 g of crude extract.
BBR analysis
The screening data obtained by UV and HPLC-UV were subsequently confirmed by MS and LC-MS (Waters, Milford, USA). The triple quadrupole mass analyzer was operating in positive mode electrospray ionization (ESI+), and acquisitions were performed in multiple reaction monitoring (MRM).
Both HPLC-UV and liquid chromatography–mass spectrometry (LC-MS) methods were original, developed, and validated
The title of BBR in the extract of
Cell culture
Human colon adenocarcinoma (HCA-7) cells line, kindly provided by Dr Mariella Chiricolo (University of Bologna, Italy) were initially grown in standard Dulbecco’s Modified Eagle’s Medium (DMEM) (Sigma-Aldrich, Inc) supplemented with 10% heat-inactivated fetal bovine serum (FBS), 0.6 mg/mL glutamine, and 200 U/mL penicillin/streptomycin (Sigma-Aldrich, Inc).
Cell cultures were replicated for subsequent experiments and maintained in water saturated atmosphere at 37°C and 5% CO2.
Cell viability test
A stock solution of
Further dilutions were made with the culture medium to the desired concentrations just before use.
HCA-7 cells were seeded into 96-well plates at a density of 104 cells per well, in 100 µL of cell culture medium, and incubated for 24 h to allow cell adherence.
Serial dilutions of
Cells treatment
Once determined, the most appropriate concentration for the treatments, HCA-7 cells were seeded at a density of 105 cells/mL into 9 cm2 (3 mL) wells. Cells were washed two times with phosphate buffered serum (PBS) and incubated for 18 h at 37°C with serum-free DMEM.
After serum starvation, cells were treated with the following solutions: (a) 30.9 and 92.7 µg/mL of
All treatments were performed for three biological replicates in DMEM supplemented with 2% FBS, antibiotics, and amino acids.
After the end of the exposure time (24 and 48 h), cells were trypsinized and processed for RNA extraction.
RNA isolation, reverse transcription, and quantitative RT-PCR
Total RNA was isolated from cells using GenElute mammalian total RNA purification mini prep kit (Sigma-Aldrich, Inc) according to manufacturer’s instructions. Total RNA concentration and quality were measured using a NanoDrop 2000 spectrophotometer (Thermo Scientific, Waltham, MA, USA). Complementary cDNA was obtained using PrimeScript RT Master Mix (Takara Bio Inc, Shiga, Japan) starting from 500 ng of total RNA. cDNA was amplified by real-time quantitative polymerase chain reaction (PCR) using the Power SYBR® Green Master Mix™ (Life Technologies, Foster City, CA, USA) and specific forward and reverse pre-designed assays (Sigma-Aldrich, Inc). The selected genes grouped by functional pathway are listed in Table 1.
Selected genes used in real-time PCR grouped by functional pathway.

PCR reactions were performed in 20 µL of final volume using the ABI PRISM 7500 (Applied Biosystems, Foster City, CA, USA). Each reaction contained 10 µL of 2× Power SYBR Green Master Mix, 400 nM concentration of each primer and 300 nM of cDNA.
After an initial denaturing step at 95°C for 10 min, the amplification proceeded with 40 cycles of a two-step profile of 15 s at 95°C and 60 s at 60°C. As final step, a melt curve dissociation analysis was performed. All experiments included non-template controls in order to check for the presence of contamination.
Statistical analysis
A preliminary test was performed to select the reference gene among three housekeeping genes. RPL13 expression appeared most consistent with the amount of RNA input. Gene expression quantification was conducted with the delta/delta Ct calculation method, 15 and RPL13 was used as reference gene to normalize the gene expression levels.
After normalization, the delta Ct of treated cells, and control were compared by a paired sample t-test. Mean expression levels of treated cells were calculated as fold changes relative to the expression of untreated cells with the delta/delta calculation method.
Results
The assay based on vital dye PrestoBlue showed that both
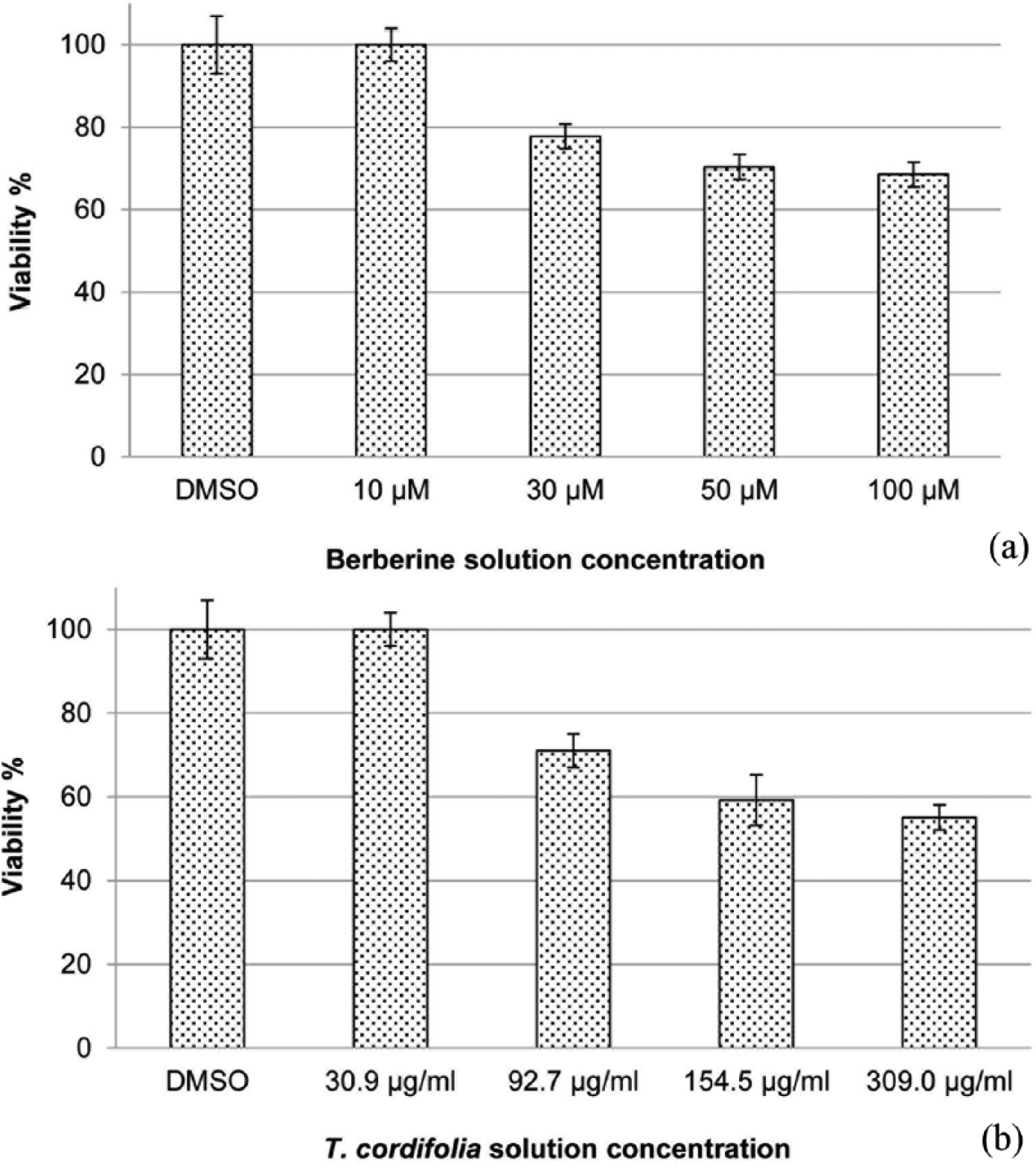
(a) Cell viability of berberine solution in HCA-7 cell line assessed by PrestoBlue reagent protocol. Cells were treated with different concentrations of drug (10, 30, 50, and 100 µM) for 48 h and (b) cell viability of
Gene expression of 44 genes belonging to different pathways involved in tumor progression and cell differentiation was investigated in HCA-7 cell line treated with
Significantly de-regulated genes after 24 h of treatment in HCA-7 cell line.
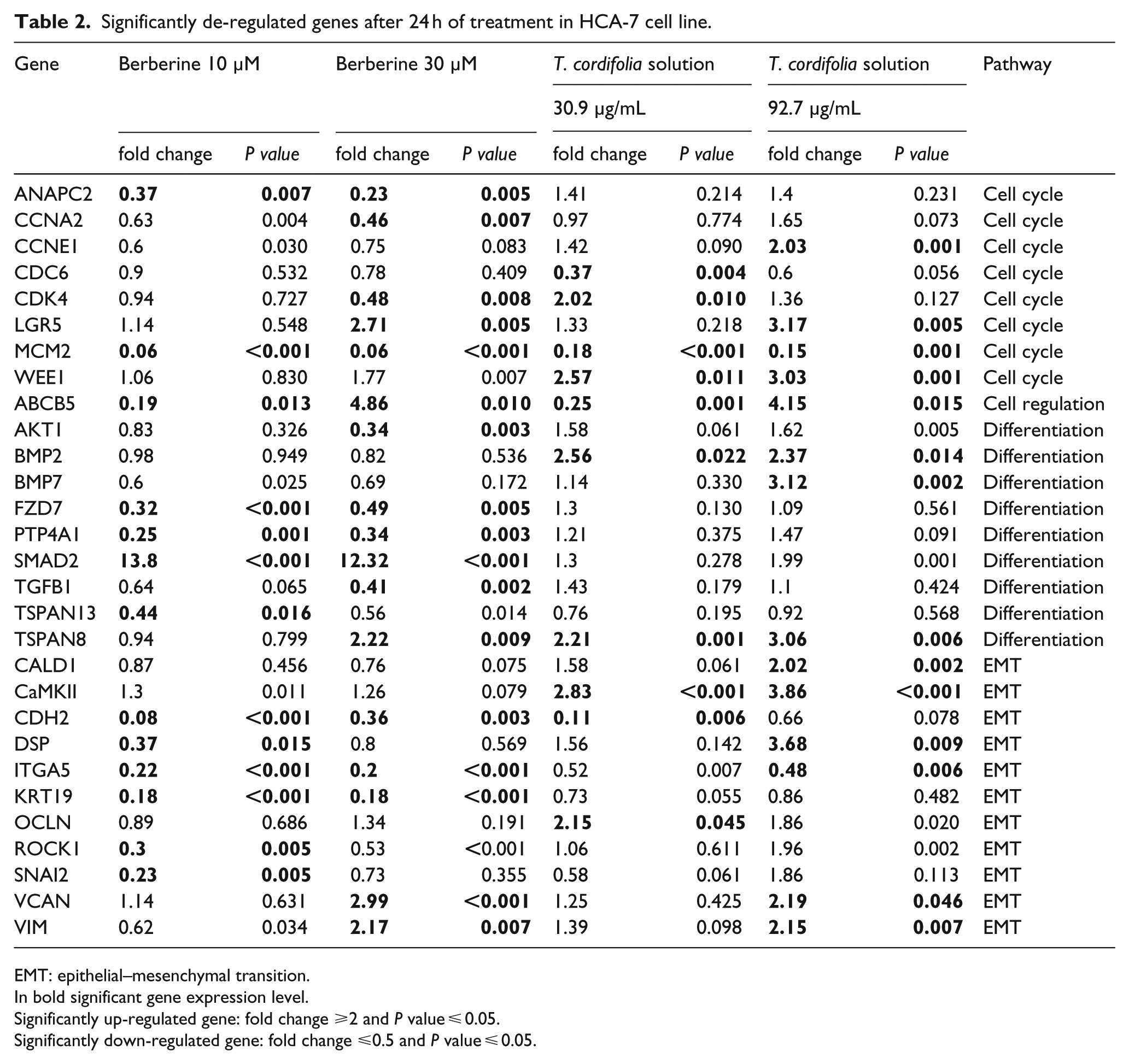
EMT: epithelial–mesenchymal transition.
In bold significant gene expression level.
Significantly up-regulated gene: fold change ⩾2 and
Significantly down-regulated gene: fold change ⩽0.5 and
Significantly de-regulated genes after 48 h of treatment in HCA-7 cell line.

EMT: epithelial–mesenchymal transition.
In bold significant gene expression level.
Significantly up-regulated gene: fold change ⩾2 and
Significantly down-regulated gene: fold change ⩽0.5 and
The most relevant effects were observed after 48 h (Figures 2 and 3).

Gene expression profile of HCA-7-treated cells (30.9 µg/mL of
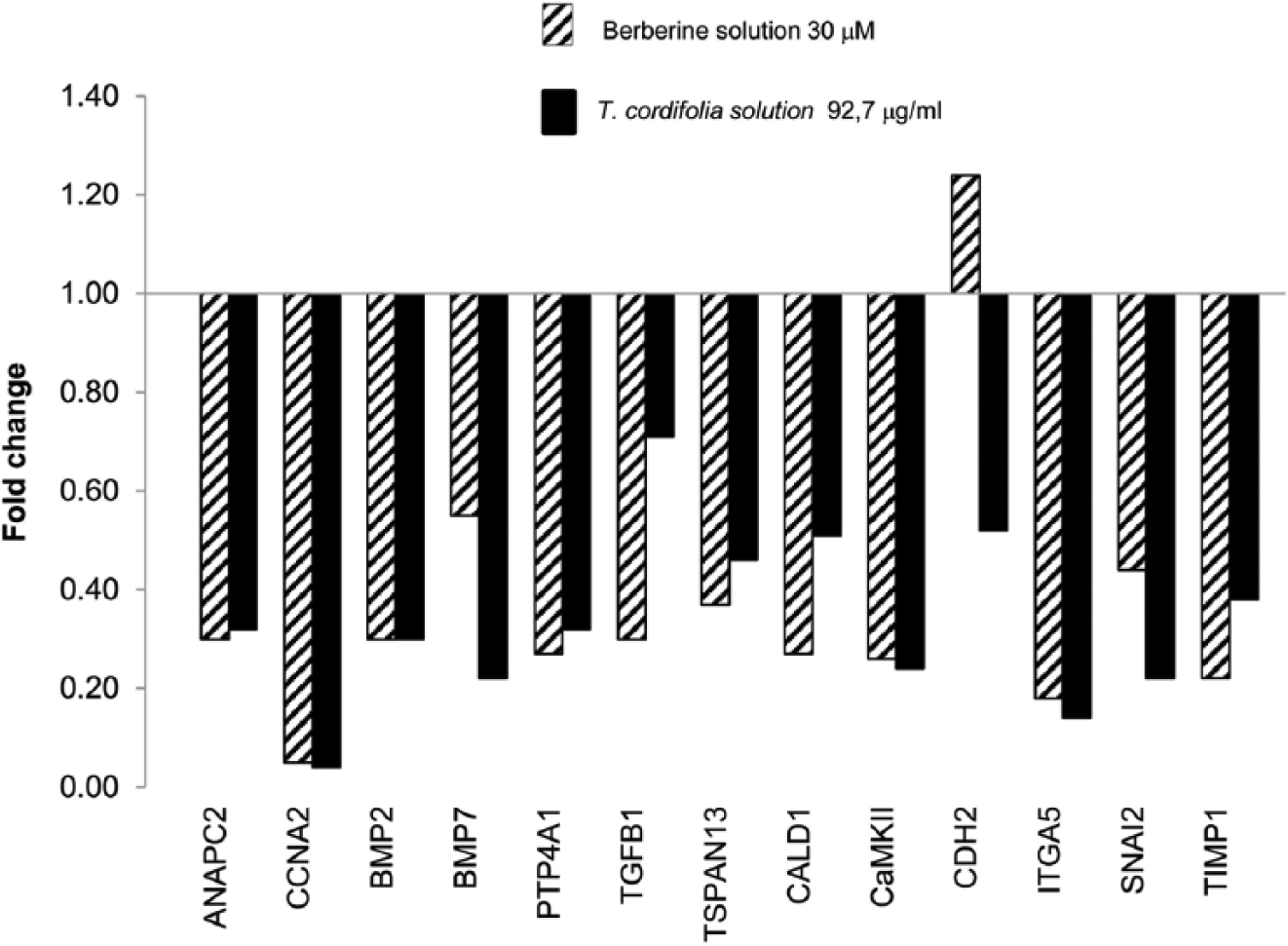
Gene expression profile of HCA-7-treated cells (92.7 µg/mL of
A total of 13 genes of the 44 studied were significantly over- and down-regulated and belong essentially to three pathways: cell cycle and regulation, differentiation, epithelial–mesenchymal transition (EMT).
Interestingly, both the plant extract and the BBR showed the same effects on the expression profiling of treated cells. In particular, the genes that maintained the same expression trend following all the treatments were ANAPC2 and CCNA2, involved in cycle regulation; BMP7 and PTP4A1 belonging to cell differentiation pathway; CaMKII and ITGA5 involved in EMT. This set of genes was significantly down-regulated in treated cells with respect to the control.
Discussion
Plant-based drug discovery has played an important role in the development of anticancer drugs currently in clinical use. 1
Several medicinal plants used in different traditional medicine systems showed anti-tumorigenic activity.16–18 Specifically, plants containing BBR might be promising suppressing agents.
In this investigation, we observed the effect of
Considering the quality of the mRNA extracted from treated cells the highest concentration of
These observations let hypothesize the involvement of BBR in the activity showed by the
The transcriptional level of 44 genes known to be involved in specific pathways such as cell cycle, differentiation, or EMT was monitored.
Among the genes modulated by the treatments, a couple of genes of cell cycle pathway resulted significantly down-regulated: ANAPC2 and CCNA2. The first one codes for a subunit of the anaphase-promoting complex (APC), involved in the progression of the cell cycle. Recently, the chemopreventive and chemotherapeutic potential of curcumin, contained in the Asian spice
Alteration of epithelial cells homeostasis and acquisition of migratory mesenchymal phenotype are crucial steps in progression of epithelial cancers. In this study, it was shown that the expression level of several genes involved in differentiation and EMT was modulated by
Bone morphogenetic proteins (BMPs) are responsible for tumorigenesis and metastatic progression of many types of cancer, inducing EMT and activating the tumorigenic potential of cancer stem cells in human colorectal cancer. 21 The up-regulation of BMP7 in colorectal cancer determines a poor prognosis caused by the increased aggressiveness of the tumor and onset of liver metastases. 22 We found significantly down-regulated both BMP2 and BMP7 in our system.
The expression of PTP4A1 was found four times lower in treated cells compared to control. Its product is localized preferentially in the cytoplasm of colon adenocarcinoma cells, endothelial cells, and smooth muscle cells in proximity of vessels surrounding lymph node metastases.23,24
Another down-regulated gene in treated cells was TSPAN13 that encodes a transmembrane protein involved in cell motility. Up-regulation of this gene has been reported at the progression stage of many cancers such as ovarian adenocarcinoma, breast cancer, bladder carcinoma, and acute lymphoblastic leukemia. 25
Two Ca2+/calmodulin-dependent proteins, CALD1 and CAMKIIG, were down-regulated in HCA-7 cells after exposure to
TGFB1 increases the invasive properties of colon cancer cells increasing the expression of CDH2 coding for the N-cadherin, located at the tips of myofibroblast filopodia. 27 In our investigation, CDH2 was significantly down-regulated at the higher concentration of both treatments.
The integrin α5, ITGA5, was strongly down-regulated in treated cells compared to control with an expression decreased by 10 times. The altered expression of ITGA5 during tumor progression has been largely demonstrated in colon cancer cells line with highly invasive potential. 28
Two genes involved in EMT and down-regulated only at the higher concentration of
Conclusion
Our study demonstrated that extract of
Footnotes
Acknowledgements
The authors are grateful to Dr Paolo Scartezzini (Maharishi Ayurveda Product, Italy) for providing
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
