Abstract
Brucella is a genus of Gram-negative intracellular pathogens that cause animal and human diseases. Brucella survival and replication inside immune cells is critical for the establishment of chronic infections. Protein modifications by small ubiquitin-related modifier proteins and the NF-κB pathway are involved in many cellular activities, playing major roles in regulating protein function that is essential for pathogenic bacteria during infection. However, the relationship between them in the intracellular survival of Brucella is still largely unknown. We demonstrated that Brucella abortus 2308 infection can activate the expression of small ubiquitin-related modifier-2 proteins in a time-dependent manner. We found the production of Th1 cytokines (IFN-γ and TNF-α) and the transcription of NF-κB/p65 were promoted by overexpression and inhibited by interference of small ubiquitin-related modifier-2. In addition, we showed that small ubiquitin-related modifier-2 can inhibit intracellular survival of Brucella abortus 2308 by regulating activation of the NF-κB pathway. Taken together, this work shows that small ubiquitin-related modifier-2 modification of NF-κB2/p65 is essential for the survival of Brucella abortus 2308 inside macrophages. This work may help to unravel the pathogenic mechanisms of Brucella infections.
Introduction
Brucella is a genus of Gram-negative, facultative intracellular pathogens that cause the serious human and animal disease brucellosis. 1 , 2 In animals, Brucella infection causes epididymitis in males and abortions in pregnant animals, resulting in serious economic and animal husbandry losses. 3 In humans, Brucella infections cause joint swelling and undulant fevers. At present, the mechanisms by which Brucella escapes removal by the immune system, invades host cells and replicates in their own compartment in host cells remain unknown. Small ubiquitin-related modifiers (SUMOs) are a group of proteins known to greatly influence various cellular biological processes, 4 such as the cell cycle, signal transduction, immune recognition, cell apoptosis, cell proliferation and differentiation, protein transport, organ origin, inflammation, Ag presentation, endoplasmic reticulum regulation, DNA repair and the stress response. 5 , 6 This protein family includes SUMO-1, SUMO-2 and SUMO-3. As previous studies have mainly focused on SUMO-1, less is known about SUMO-2 and SUMO-3.
SUMO-2 and SUMO-3 have very similar amino acid sequences (95% similarity), both of which are mainly distributed in the nucleus. SUMOs 1–3 can covalently combine with different target proteins to exert different effects in specific cells or tissues. 7 Brucella infections have been shown to inhibit SUMO-1 expression, thereby effecting apoptosis of the host cell. 8 There is great interest in the interactions of Brucella with SUMO-2-modified proteins during macrophage infection.
NF-κB is a type of nuclear protein factor with multidirectional transcriptional regulation, playing a major role in the regulation of gene transcription at the beginning of many crucial biological processes, such as the immune response, inflammation, oxidative stress and apoptosis. 9 Participation in the immune response is the most important function of the NF-κB pathway during infections of pathogenic microorganisms. 10 Previous studies have shown the NF-κB pathway is differentially activated by Brucella 11 and that SUMO-1 modification is an important requirement for stimuli-induced NF-κB/p100 processing. 12 However, the relationship between SUMO modifications and NF-κB in the intracellular survival of Brucella is still unclear.
To investigate whether SUMO-2-mediated NF-κB pathway activation affects intracellular survival of Brucella abortus 2308, we constructed a SUMO-2 interference and overexpression cell line and used Western blot analysis, ELISA, and reverse transcription PCR (RT-PCR) to detect SUMO-2, NF-κB/p65 expression and the secretion of IFN-γ and TNF-α. We also evaluated the role of SUMO-2-mediated NF-κB pathway activation in the intracellular reproduction of B. abortus 2308 in macrophages.
Materials and methods
Growth conditions of bacterial strains and cells
B. abortus 2308 (Chinese Center for Disease Control and Prevention; Beijing, China) was cultured in tryptic soy agar or tryptic soy broth medium (Oxoid, UK) in a 37°C shaker. Escherichia coli strain DH5α (Chinese Center for Disease Control and Prevention; Beijing, China) was cultured with Luria-Bertani culture medium containing antibiotics (50 mg/ml of ampicillin, Table 1). HEK-293FT cells and mouse macrophages RAW264.7 (Shanghai Institute of Cell Biology, Chinese Academy of Sciences, Shanghai, China) (Table 1) were cultured in DMEM medium containing 10% FBS (Gibco, USA) and set in a 37°C 5% CO2 incubator.
Primers for amplifying and real-time quantitative PCR.
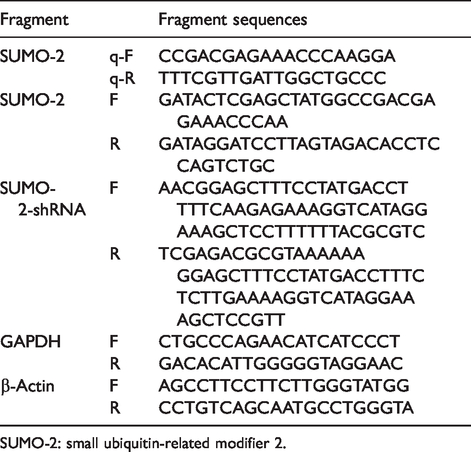
SUMO-2: small ubiquitin-related modifier 2.
Plasmids
Lentiviral overexpression vector pLEX-MCS, pLEX-GFP and enveloping and packaging vectors (pSPAX2 and pMD2.G) were purchased from Xinjiang Academy of Animal Science (Xinjiang, China). Lentiviral RNAi vector pLL3.7-GFP and helper plasmids PRSV-REV and PCMV-VSVG were purchased from Xinjiang Academy of Animal Science (Xinjiang, China). pMD19-T cloning vector was purchased from Takara (Dalian, China).
Construction of SUMO-2 lentivirus interference vector
We used the GenBank SUMO-2 (NM_133354.2) gene sequences to design the specific interference fragments (Table 1). Interference fragments and pLL3.7 plasmids were double enzyme by Hpa I and Xho I, then connected and transformed into E. coli DH5α. PLL3.7-shRNA plasmid was extracted by a plasmid DNA Extraction Kit (Qiagen, China).
Construction of SUMO-2 lentivirus overexpression vector
We used the GenBank SUMO-2 (NM_133354.2) mRNA sequences to design the PCR amplification primers (Table 1). We then extracted mouse macrophage RAW264.7 total RNA (CW Biotech, China) and through RNA reverse transcription synthesis of cDNA (cDNA HiFi-Script first chain synthesis kit, CWbiotech), SUMO-2 genes were amplified with cDNA as the template. We connected to the pLEX-MCS and then pLEX-SUMO-2 were obtained, using a Plasmid Extraction Kit (Qiagen, China).
Identification of SUMO-2 stable interference and overexpression of cell lines
HEK-293FT and RAW264.7 cells were transfected with the lentivirus interference vector and overexpression vector, with a ratio of 5 ml of plasmids for 2 × 105 cells. Plasmids and cells were thoroughly mixed and placed in a 37°C 5% CO2 incubator. SUMO-2 stable cell interference (I-S group) and SUMO-2 stably overexpressing cells (O-S group) were generated. After incubating for 48 h, the cell lines were observed with a fluorescence microscope (Nikon, Japan). In addition, the total RNA was extracted (RNA extraction kit, CWbiotech, China) from different groups of culturing murine macrophages RAW264.7 and reverse transcribed to cDNA, with β-actin as a reference gene. Real-time PCR was conducted on the Light-Cycler 480 (Roche Applied Science) with the SYBR Premix Ex Taq™ reagent kit (TaKaRa, Japan) using 2−ΔΔCt analysis to calculate the relative gene expression (Table 1).
A cell model of B. abortus 2308 infection is established
To establish a cell model of B. abortus 2308 infection, cells were infected with B. abortus 2308 at the logarithmic growth phase. The MOI (bacteria:cells) was 100:1 and all cells were cultured in DMEM medium containing 10% FBS at 37°C and 5% CO2. After 1 h, gentamicin (30 g/ml, Sigma-Aldrich) was added to the cell medium for 30 min to kill the extracellular B. abortus 2308. Cells received different treatments at 0, 2, 4, 8, 12 and 24 h.
ELISA assay
At 4, 12 and 24 h after Brucella infection, the supernatant from differently treated cells culture was collected and filtered with a 0.22 μm membrane filter. After centrifugation at 400 g for 20 min, the cytokines TNF-α and IFN-γ were detected by ELISA (R&D, USA).
RT-PCR detection
According to the B. abortus 2308 infection models, at 4, 8, 12 and 24 h after infection, the total RNA of macrophages was extracted and reverse transcribed to cDNA, with GAPDH as a control gene. The real-time PCR was conducted on the Light-Cycler 480 (Roche Applied Science, Switzerland) with the SYBR Premix Ex Taq™ reagent kit (TaKaRa, Dalian, China), and the p65 transcription level was calculated using 2−ΔΔCt analysis.
Immuno-coprecipitation
Brucella M5-infected cells were lysed in a buffer containing 0.01 M Tris HCl (pH 7.4), 0.15 M NaCl, 1% sodium deoxycholate, 1% Triton X-100, 0.1% NaDodSO4 and 1 mM phenylmethyl sulfonyl fluoride. Immuno-coprecipitations were performed with Ab (p65, SUMO-2, IgG) (Santa Cruz Biotechnology), which were crosslinked to protein A agarose beads for 12 h at 4°C. After washing the beads, bound proteins were eluted and subjected to Western blotting.
Ability of Brucella to survive in different groups of cells
Using the model of the Brucella infection, at 0, 2, 4, 8, 12 and 24 h different groups of cells were collected and cleaved, followed by gradient dilution, producing concentrations of 101, 102, 103, 104, 105 and 106. The bacteria were then evenly coated on the Brucella solid medium; we incubated it at 37°C for 72 h then counted the number of colonies.
Western blotting
According to the Brucella infection models, at 0, 2, 4, 8, 12 and 24 h after infection, protein lysate samples were separated by 12% SDS-PAGE gel and transferred to the Nitrocellulose membrane (NC) membrane for 40 min (100 mM of Tris-HCl, 150 Mm of NaCl, 0.05% Tween 20 and pH 7.2). The NC membrane was blotted with 5% non-fat milk in TBST for 1 h, followed by incubation with a primary Ab diluted with TBST for 2 h at 37°C. The membrane was washed with TBST three times and incubated with a secondary Ab at 37°C for 2 h. After washing with PBS three times, the NC membrane was stained using a HRP-DAB kit (Zhongshan Golden Bridge, China).
Abs used herein are as follows: primary Abs were rabbit anti-SUMO-2 mAb (Abcam, UK), rabbit anti-β-actin mAb (Jackson ImmunoResearch, USA); the secondary Ab was HRP-conjugated mouse anti-rabbit IgG (Bioworld, USA).
Statistical analysis
SPSS Statistics 17.0 software (IBM, USA) was used for analysis. All experimental data were denoted as mean ± standard deviation (x̅ ± SD). Multiple group comparisons were performed using the Student’s t test.
Results
B. abortus 2308 infection induces SUMO-2 protein expression
The expression of SUMO-2 was assessed by Western blot analysis in B. abortus 2308-infected RAW264.7 cells at 0, 2, 4, 8, 12 and 24 h post-infection. We found that B. abortus 2308 infection caused SUMO-2 protein levels to gradually decrease over the 24 h period, indicating that B. abortus 2308 inhibits the expression of SUMO-2 in a time-dependent manner (Figure 1).
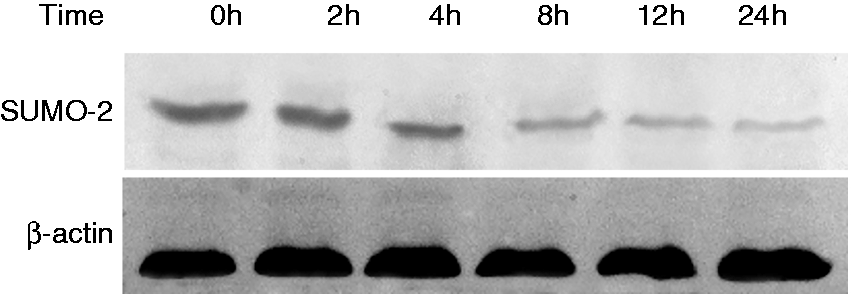
Western blot analysis of small ubiquitin-related modifier 2 (SUMO-2) protein expression in RAW264.7 cells 0, 2, 4, 8, 12 and 24 h after B. abortus 2308 infection. β-Actin levels were used as a control.
Lentiviral packaging and infection
pLL3.7-SUMO-2 interference (Figure 2a) and pLEX-SUMO-2 overexpression (Figure 2b) lentiviral vectors harbouring EGFP genes were constructed and recombinant viruses were transfected into RAW264.7 cells. At 48 h post-infection, cells with green fluorescence were observed, indicating the successful transfection of the SUMO-2 gene.
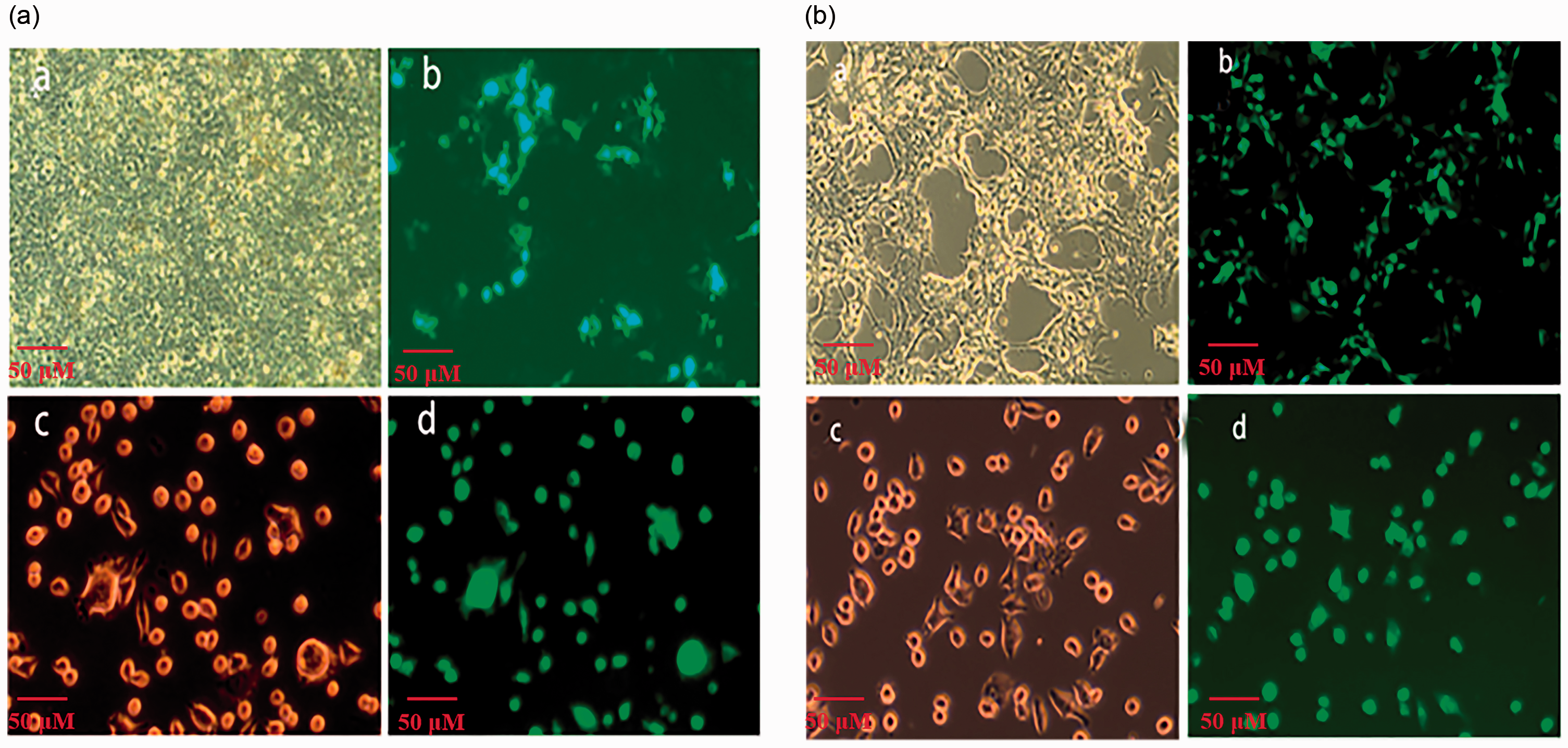
Identification of the small ubiquitin-related modifier 2 (SUMO-2) interference and overexpression vectors. The constructed vectors pLL3.7-SUMO-2 and pLEX-SUMO-2 were transfected into HEK-293FT cells following the instructions in the Highly Efficient Transfection kit (HET). Recombinant viruses were constructed and RAW264.7 cells infected. Packaging and infection process of (a) pLL3.7-SUMO-2 and (b) pLEX-SUMO-2. (a) 293T cells; (b) lentivirus package; (c) RAW264.7 cells; (d) lentivirus transfection.
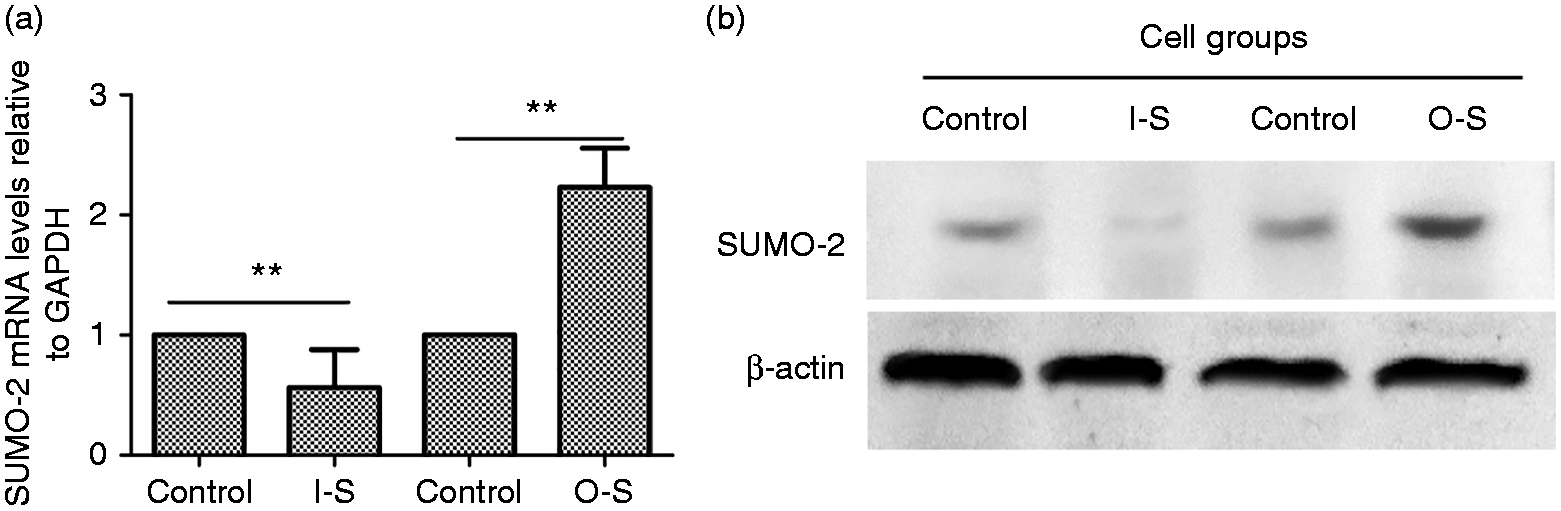
Relative expression of small ubiquitin-related modifier 2 (SUMO-2) at the mRNA and protein level in differently treated cells. RAW264.7 cells were infected with SUMO-2 lentivirus vectors. (a) After 48 h, total RNA was extracted and reverse transcribed to cDNA, with β-actin as a reference gene. Real-time PCR was conducted to detect the level of SUMO-2 mRNA. 2−ΔΔCt analysis was used to calculate relative gene expression. (b) Western blot detected protein expression of SUMO-2 in interference (I-S) cells and in overexpressing (O-S) cells. All treatments were repeated three times with n = 3 per time point. Statistical significance is indicated by a double asterisk (**P < 0.01).
SUMO-2 gene expression levels in differently treated cells
Relative gene expression of SUMO-2 was calculated using the 2−ΔΔCt method (Table 1). In the I-S group, SUMO-2 expression was significantly decreased (P < 0.01) when compared with the control group. The highest interference efficiency was 65%. In the O-S group, SUMO-2 expression was significantly increased after lentiviral pLEX-SUMO-2 infection (P < 0.01) (Figure 3a). Additionally, the Western blot test also showed similar results (Figure 3b). These results confirmed successful construction of SUMO-2 interference, overexpression and rescue cell lines.
ELISA detection of TNF-α and IFN-γ levels in B. abortus 2308-infected cells
We measured TNF-α and IFN-γ levels in supernatants of 2308-infected RAW264.7 cells in the presence of SUMO-2 interference or overexpression. O-S cells produced significantly higher levels of TNF-α and IFN-γ at 4, 12 and 24 h post-infection compared to the control group (P < 0.01) and I-S cells produced significantly lower levels of TNF-α and IFN-γ at 2 and 24 h post-infection (P < 0.05; Figure 4). These results indicate the secretion of inflammatory cytokines is closely related to SUMO-2 protein expression.
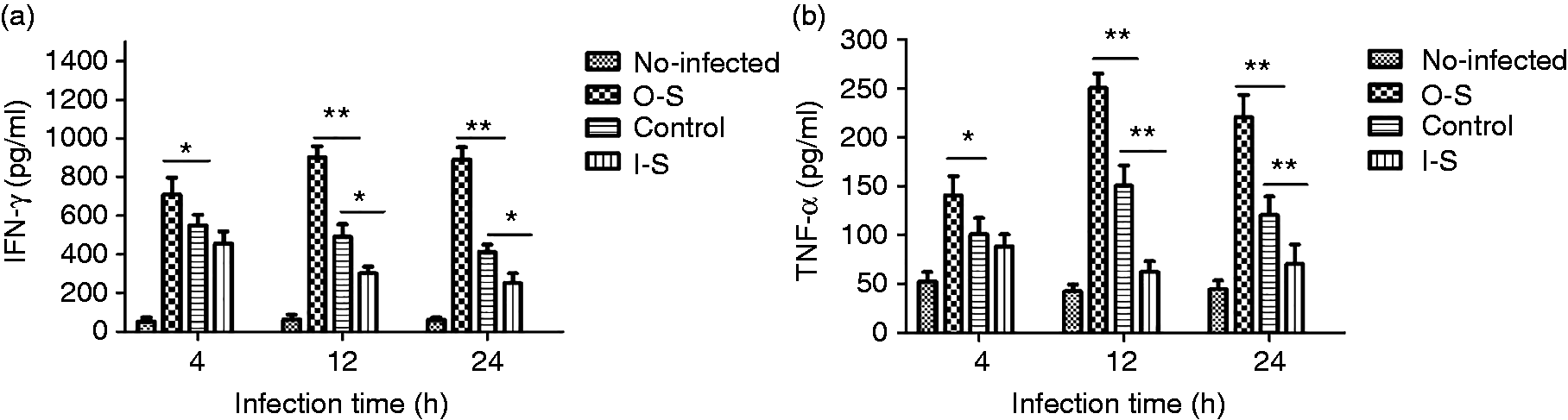
ELISA analysis of IFN-γ (a) and TNF-α (b) production from the supernatants of B. abortus 2308-infected RAW264.7 cells in the presence of small ubiquitin-related modifier 2 (SUMO-2) overexpression or interference. Significant differences when compared with uninfected cells are indicated by a single asterisk (*P < 0.05) and a double asterisk (**P < 0.01).
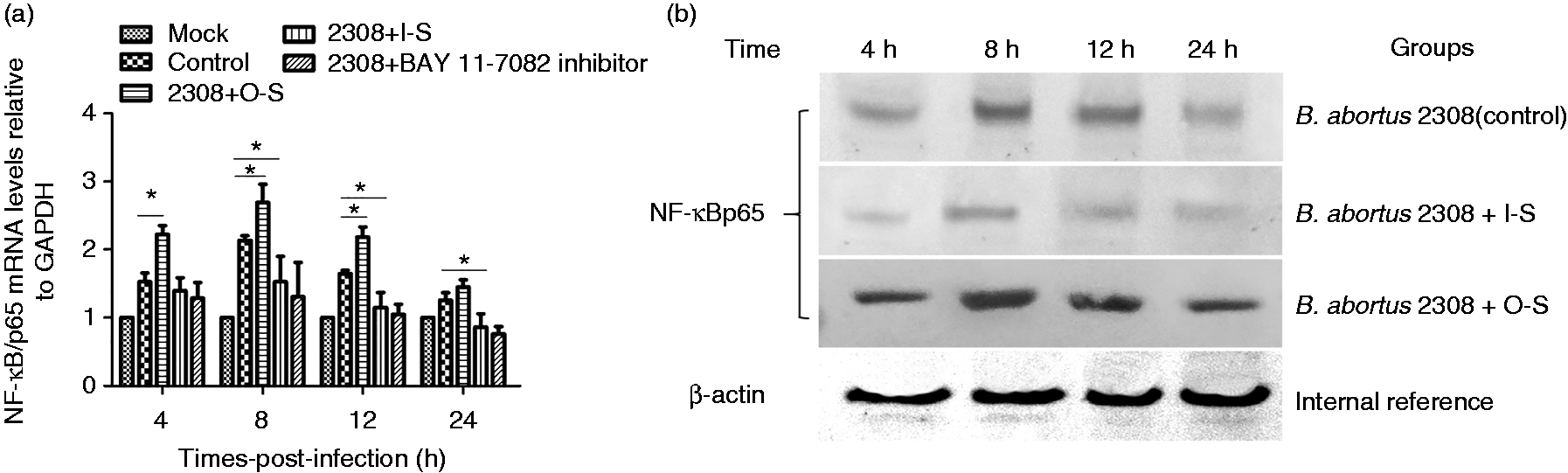
The levels of NF-κB/p65 gene expression in different B. abortus 2308-infected cells. Levels of NF-κB/p65 mRNA and protein expression in B. abortus 2308-infected macrophages were measured at 4, 8, 12 and 24 h post-infection by reverse transcription PCR (RT-PCR) (a) or Western blot (b). p65 mRNA levels of each group were normalized to expression of GAPDH to generate ΔCt. All treatments were repeated three times with n = 3 per time point. Statistical significance is indicated by a single asterisk (*P < 0.05).
NF-κB/p65 mRNA and protein levels in differently treated cells infected with B. abortus 2308
To explore whether NF-κB is involved in the SUMO-mediated immune response, we used RT-PCR (Figure 5a) and Western blot (Figure 5b) to detect p65 expression levels in differently treated cells infected with B. abortus 2308. The results show that B. abortus 2308 induced p65 mRNA transcription and protein expression in macrophages in a time-dependent manner. At 4, 8 and 12 h after B. abortus 2308 infection, NF-κB/p65 mRNA transcription in O-S cells was significantly higher than in control cells (P < 0.05), showing that SUMO-2 expression induces NF-κB pathway activation in B. abortus 2308-infected cells. At 8, 12 and 24 h after B. abortus 2308 infection, NF-κB/p65 mRNA transcription in I-S cells was lower than in the control group, indicating SUMO-2 gene silencing inhibited NF-κB pathway activation. Adding NF-κB inhibitors or interfering with the SUMO-2 gene also had the ability to inhibit p65 mRNA transcription. In addition, the results also showed the overexpression and interference of SUMO-2 in macrophages would affect NF-κB/p65 protein expression during B. abortus 2308 infection.
Direct interaction of NF-κB/p65 with SUMO-2 in RAW264.7 cells
In order to further explore the interaction of NF-κB/p65 and SUMO-2 in B. abortus 2308-infected cells, we detected the p65 and SUMO-2 expression in cell lysates by Immuno-coprecipitation, results showed that immune complexes precipitated with anti-NF-κB/p65 contained SUMO-2 (Figure 6), demonstrating an interaction between the endogenously expressed proteins in RAW264.7 cells.
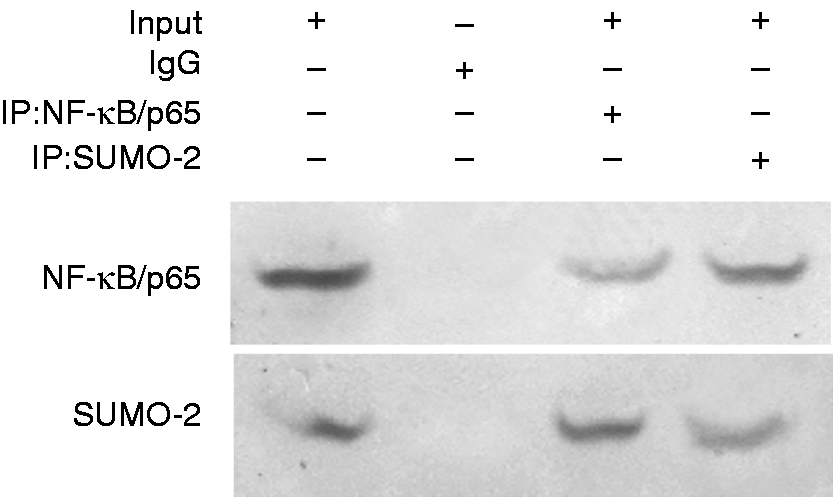
The interaction of NF-κB/p65 with small ubiquitin-related modifier 2 (SUMO-2). Lysates from RAW264.7 cells expressing p65 and SUMO-2 protein were immunoprecipitated with anti-SUMO-2 or control IgG. Cell lysates and eluted proteins were subjected to Western blotting with anti-NF-κB/p65 (upper) or anti-SUMO-2 (lower).
Effect of the SUMO-2/NF-κB pathway on the intracellular survival of Brucella in differently treated macrophages
We measured the intracellular survival of B. abortus 2308 in SUMO-2 RAW264.7 cells after different treatments. Cells were collected at 0, 2, 4, 8, 12 and 24 h post-infection, lysed and the number of Brucella CFU (log CFU) was counted. As with the control group (RAW cells), log CFU values for the O-S group showed a decreasing trend at 12 and 24 h, although overall values were significantly lower in the O-S group (Figure 7a; P < 0.01). Log CFU values for the I-S group were significantly raised when compared to RAW cells at 8, 12 and 24 h post-infection (P < 0.05, P < 0.01). However, when the O-S group was pre-treated with NF-κB-inhibitor BAY11-7082, intracellular survival increased as reflected by significantly lower log CFU values for the O-S group than the BAY11+O-S group at 24 h post-infection (Figure 7b; P < 0.05). These results show that SUMO-2-mediated activation of NF-κB can inhibit intracellular Brucella survivability.
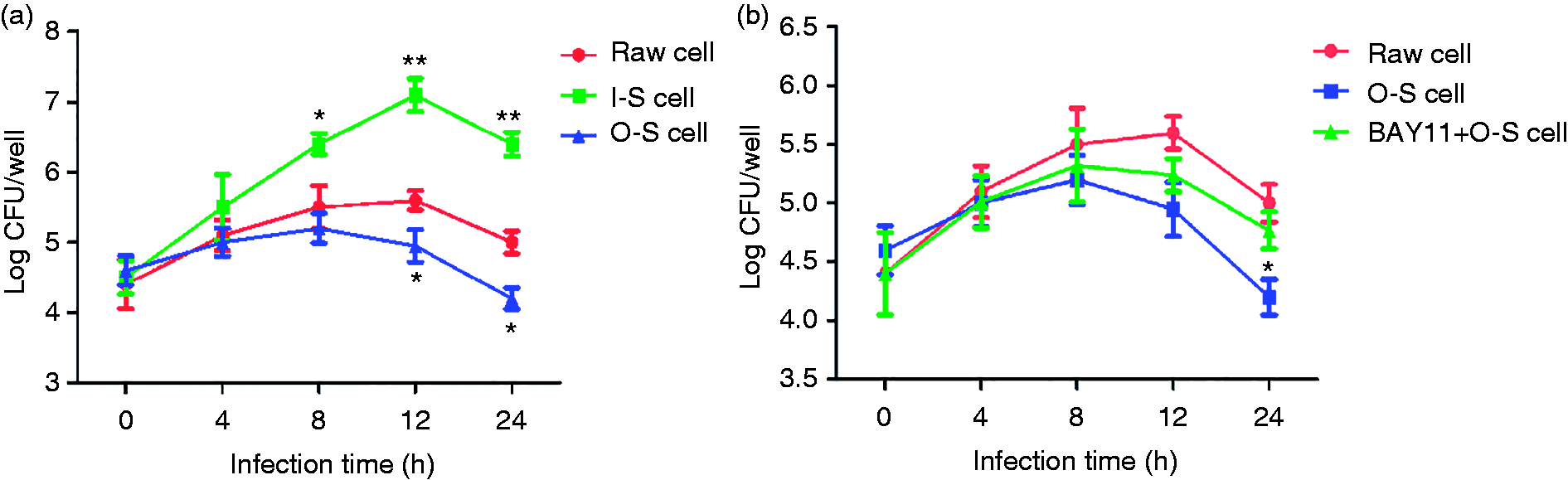
Evaluation of intracellular survival of B. abortus 2308 after infecting different groups of RAW264.7 cells. Small ubiquitin-related modifier 2 (SUMO-2) stable interference or overexpression cell lines (a) and cells pre-treated with NF-κB inhibitor (b) were lysed at 0, 2, 4, 8, 12 and 24 h post-infection and cellular extracts were coated on trypticase soy agar culture medium and cultured at 37°C for 72 h. The number of CFUs were counted and represented here on a log scale. All treatments were repeated three times with n = 3 per time point. Statistical significance is indicated a single asterisk (*P < 0.05) and a double asterisk (**P < 0.01).
Discussion
Brucella is a Gram-negative bacterium that invades the host after surviving in macrophages. 13 As the host tries to clear the infection, a few Brucella are transferred to the endoplasmic reticulum in the form of Brucella-containing vacuoles to escape host immune killing, resulting in chronic infection. 14 SUMOs modify proteins in cells to change their function and some pathogenic bacteria have evolved mechanisms to affect different stages of the SUMO modification system, intentionally or unintentionally, to avoid clearance by the host cell. 15 Ribet et al. indicated that the bacterial virulence factor Listeria hemolysin O can interfere with the SUMO modification process in Listeria-infected cells. 16 Shigella, an intracellular pathogen, can weaken the cell SUMO pathway by modifying SUMO proteins to reduce the binding stability of the SUMO E2-conjugation enzyme Ubc9 with substrate proteins. 17 , 18 Recent studies have confirmed that SUMO-1 plays an essential role in regulating B. abortus 2308 infection inside macrophages, and researchers speculate that the type IV secretion system may be involved in protein damage of SUMO-1. 19 Although both SUMO-1 and SUMO-2 belong to the SUMO gene family, 20 their amino acid sequence homology is only 50%. In addition, SUMO-1 and SUMO-2 have been shown to have distinct and overlapping sets of target proteins, suggesting that SUMO-1 and SUMO-2 may have different biological functions. 21 In this study, we explored whether SUMO-2 has a similar role in Brucella infections. Brucella-infecting macrophages is a gradual and complex process. Many researchers believed there were many critical time points in the process of Brucella infection, invasion and reproduction, such as 0, 2, 4, 8, 12 or 24 h. 22 , 23 Our results showed that B. abortus 2308 could induce SUMO-2 expression at 0, 2, 4, 8, 12 and 24 h time post-infection, but its expression decreased significantly over time. However, whether the activation of SUMO-2 affects the intracellular replication of Brucella needs further investigation.
Previously, Th1 immune responses characterized by production of IFN-γ have been shown to be associated with protective immunity to Brucella. 24 IFN-α; is an important cytokine required for the bactericidal activity of macrophages; IFN-α; elicits antibacterial responses and can therefore eliminate Brucella. 25 TNF-α is another important regulator of host immune responses to the invasion of pathogenic microorganisms. 26 In our study, we found the expression of IFN-α and TNF-α levels decreased in I-S cells infected with Brucella when compared with control cells. Brucella survives and causes persistent infections in hosts due to multiple virulence factors that alter cytokine production. The overexpression of SUMO-2 in RAW264.7 cells inhibited the survival of intracellular Brucella; conversely, after silencing SUMO-2, Brucella intracellular survival capacity significantly increased. These results show that B. abortus 2308 intracellular reproduction is inhibited by SUMO-2 overexpression and promoted by SUMO-2 interference. We speculate that the release of Th1 cytokines induced by SUMO-2 modification is a key factor in protecting against persistent Brucella infections.
Studies have shown that SUMOylation can affect the activation of the NF-κB pathway and SUMO ligase-induced NF-κB activation is essential for DNA damage response. 27 In this study, we discussed the relationship between SUMOylation and NF-κB in Brucella infections. Immuno-coprecipitation showed there was an interaction between SUMO-2 and NF-κB in Brucella-infected cells, RT-PCR and Western blot assays also indicated that SUMO-2 could directly affect p65 mRNA transcription and protein expression, which revealed that SUMO-2 may be an important modified protein that regulates the activation of the NF-κB signaling pathway. Transcriptional factor NF-κB is a key mediator of TNF-α signaling, and NF-κB-mediated TNF-α-induced host immunity is required to inhibit the intracellular survival of Brucella. 25 When NF-κB inhibitor BAY11-7082 was added, we found it could effectively inhibit the SUMO-2-mediated anti-Brucella effect. Therefore, we believe the activation of NF-κB is a key factor in the initiation of the anti-Brucella effect of SUMO-2. NF-κB can regulate secretion of the pro-inflammatory cytokines IL-1β, IL-6 and IL-12, amongst others. 28 We suggest that SUMO-2 may promote the release of Th1 cytokines by modifying NF-κB molecules, thereby inhibiting B. abortus 2308 intracellular reproduction.
In summary, the SUMO-2 protein plays an important role in Brucella survival in RAW264.7 cells. Inhibition of SUMO-2 can inhibit Th1 immune responses and permits intracellular survival of Brucella. Importantly, the activation of the NF-κB pathway is essential for SUMO-2 to exert its antibacterial effect. Although this study provides new insights into how Th1 develops anti-Brucella immunity in cells, the molecular mechanisms underlying the interaction between SUMO-2 and NF-κB transcription factors need to be explored further.
Footnotes
Acknowledgements
This work was supported by Tiankang Biological Co., Ltd. The authors would like to thank Professor Chuangfu Chen and Jihai Yi for their guidance.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (No. U1803236), National Natural Key Research and Development Plan (No. 2017YFD0500304), and Corps Major Scientific and Technological Projects (No. 2017AA003).
