Abstract
In the present study, we used microRNA (miRNA) sequencing to discover and explore the expression profiles of known and novel miRNAs in 1000 ng/ml LPS stimulated for 8 h vis-à-vis non-stimulated (i.e. control) PBMCs isolated from the blood of healthy pigs. A total of 291 known miRNAs were bio-computationally identified in porcine PBMCs, and 228 novel miRNAs (not enlisted in the swine mirBase) were identified. Among these miRNAs, ssc-miR-148a-3p, ssc-let-7g, ssc-let-7f, 3_8760, ssc-miR-26a, ssc-miR-451, ssc-miR-21, ssc-miR-30d, ssc-miR-99a and ssc-miR-103 were the top 10 most abundant miRNAs in porcine PBMCs. Through miRNA differential analysis combined with quantitative PCR, we found the expressions of ssc-miR-122, ssc-miR-129b, ssc-miR-17-5p and ssc-miR-152 were significantly changed in porcine PBMCs after LPS stimulation. Furthermore, targets prediction and function analysis indicated a significant enrichment in gene ontology functional categories related to diseases, immunity and inflammation. In conclusion, this study on profiling of miRNAs expressed in LPS-stimulated PBMCs provides an important reference point for future studies on regulatory roles of miRNAs in porcine immune system.
Introduction
In swine production, Gram-negative bacterial infection is one of the most important causes of severe inflammation, which may result in economic losses due to mortality, morbidity and decreased growth rate.1,2 Endotoxin/LPS, hosted in the outer membrane of Gram-negative bacteria, is a potent activator of the innate immune system. 3 LPS induces macrophage/monocyte activation and pro-inflammatory cytokine (TNF-α, IL-1β and IL-6) production by activating TLR4 signalling. 4 The TLR4/MyD88/NF-κB pathway and the TLR4/TRIF-NLRP3 inflammasome axis are key regulators involved in inflammatory processes.5,6 However, considering the complexity in the LPS-induced inflammatory response, other levels of regulation may also be involved.
MicroRNAs (miRNAs) represent a class of small non-coding RNAs (17–25 nt) that regulate gene expression through translational inhibition or transcript degradation. 7 Recent studies have shown that miRNAs play a key regulatory role in inflammation. For example, miR-223 reduced LPS-induced inflammation and suppressed the NLRP3 inflammasome and TLR4/NF-κB signalling pathway via targeting rho-related GTP-binding protein RhoB (RHOB). 8 MiR-9 increased LPS-induced inflammatory response of macrophages by targeting silent information regulator 1 (SIRT1). 9 Moreover, a number of miRNAs regulate elements of the TLR-signalling cascade or target cytokines directly. MiR-146b directly targets IL-1 receptor-associated kinase 1 (IRAK1) and TNF receptor-associated factor 6 (TRAF6), key adaptor proteins of the TLR4/NF-κB signalling cascade, resulting in inhibition of NF-κB activation and inflammatory cytokine production. 10 MiR-322 inhibits inflammatory cytokine expression in LPS-stimulated murine macrophages by targeting NF-κB1. 11 MiR-93-3p alleviates LPS-induced inflammation by inhibiting TLR4. 12 MiR-27a regulates the inflammatory response of macrophages by targeting IL-10, a potent anti-inflammatory cytokine. 13 However, miR-155 and miR-125b can target the 3′ untranslated region of TNF-α mRNA directly. 14
Considering the role of miRNAs in the innate immune response, the present study was designed to discover and explore the miRNA profile of porcine PBMCs in response to LPS. To date, miRNA expression in the LPS-induced inflammation in porcine blood cells has not been well characterised. This analysis of miRNA expression in porcine PBMCs challenged with LPS contributes significantly to the current knowledge of miRNAs playing roles in bacterial infection disease pathogenesis.
Materials and methods
Cell isolation, culture and stimulation
The PBMCs from the Duroc×Large White×Landrace (DLW) crossbred piglets (∼15 kg, ∼8 wk old) were isolated by Ficoll-Hypaque density gradient centrifugation at room temperature (25°C). Briefly, heparinised blood was layered on an equal volume of D-Han’s solution (pH 7.2), and then an equal volume of Ficoll-Hypaque medium was added in a conical centrifuge tube. The suspension was centrifuged at 472
Quantitative RT-PCR
Total RNA was isolated from cell samples using Trizol (Invitrogen, Carlsbad, CA) according to the manufacturer’s protocol. Expression of TNF-α, IL-1β and IL-6 mRNA was analysed using the Applied Biosystems 7500 Real-Time PCR system (Applied Biosystems, Foster City, CA). cDNA synthesis and quantitative RT-qPCR were carried out as previously described. 15 GAPDH was used as an internal normalisation control. Sequences of specific primers are shown in Supplemental Table S1. miRNA expression was analysed using a Hairpin-it™ miRNA qPCR Quantitation Kit (GenePharma, Shanghai, PR China) and the Applied Biosystems 7500 Real-Time PCR system according to the manufacturer. U6 was used as a normalisation gene. All data were analysed using the 2–ΔΔCT method. 16
Cytokine TNF-α and IL-1β measurements
The concentrations of TNF-α and IL-1β in supernatants of PBMCs were measured using commercially available porcine ELISA kits (R&D Systems, Inc., Minneapolis, MN) according to the manufacturer’s instructions.
Small-RNA library construction and deep sequencing
The RNA samples from LPS-stimulated and unstimulated PBMCs of three pigs were used to create three experimental small RNA libraries and three control libraries for biological replicates. For each small RNA library construction, 1 μg total RNA was ligated to a 5′ and a 3′ adapter sequentially according to the protocol (NEBNext® Multiplex Small RNA Library Prep Set for Illumina). Briefly, 1 µg RNA was mixed with 1 µl 3′ SR adaptor and added nuclease-free water up to 7 µl total volume. Then, the mix was incubated in a preheated thermal cycler for 2 min at 70°C. The tube was transferred to ice, and the following components were added: 10 µl 3′ ligation reaction buffer (2×) mix and 3 µl of 3′ ligation enzyme mix. The mix (total volume 20 µl) was then incubated for 1 h at 25°C in a thermal cycler. Next, 4.5 µl nuclease-free water and 1 µl SR RT Primer for Illumina were added to the ligation mixture, and the total volume was 25.5 µl. The samples were heated for 5 min at 75°C, 15 min at 37°C and 15 min at 25°C. Next, 1.1 µl 5′ SR adaptor was aliquoted into a separate nuclease-free 200 µl PCR tube. The adaptor was incubated in a thermal cycler at 70°C for 2 min, and then the tube was immediately placed on ice. The following components were added to the 3′ adaptor ligation mixture and mixed well: 1 µl 5′ SR adaptor, 1 µl 5′ ligation reaction buffer (10×) and 2.5 µl 5′ ligation enzyme mix. The mixture was incubated for 1 h at 25°C in a thermal cycler. The 5′ adapter sequence was 5′-rGrUrUrCrArGrArGrUrUrC rUrArCrArGrUrCrCrGrArCrGrArUrC-3′. The 3′ adapter sequence was 5′-rAppAGATCGGAAGAGC ACACGTCT-NH2-3′. Then, reverse transcription and PCR amplification with adapter-specific primers were performed according to the protocol. Next, the 145–160 bp amplification products were isolated on a 6% polyacrylamide gel. Briefly, load wells with 25 μl of mixed Amplified cDNA Construct and loading dye on the 6% polyacrylamide gel. A total volume of 50 μl was loaded on the gel. The gel was run for 60 min at 145 V. Using a razor blade, the bands corresponding to adapter-ligated constructs derived from the 21 and 30 nucleotide RNA fragments, respectively, were cut out. Then, 300 μl ultra-pure water was added to the gel debris. The DNA was eluted by rotating or shaking the tube at room temperature for at least 2 h. The eluate and the gel debris were transferred to the top of a 5 μm filter. Then, 2 μl glycogen, 30 μl 3M NaOAc, 2 μl 0.1× pellet paint and 975 μl 100% pre-chilled ethanol (–25°C to –15°C) were added to the eluate. The eluate was immediately centrifuged at 20,000
Basic data processing
The raw sequence reads produced by deep sequencing were pre-processed using the FASTX-Toolkit (http://hannonlab.cshl.edu/fastx_toolkit/). After adapter trimming, low-quality reads and reads shorter than 15 nt were removed to obtain clean reads. All clean reads were further annotated and classified by aligning to the pig genome mRAN (ftp://ftp.ensemble.org/pub/release-90/fasta/), miRBase (http://www.mirbase.org/), Rfam (http://rfam.xfam.org//) and RepBase (https://www.girinst.org/repbase/) databases. After alignment to Rfam and Genebank, non-miRNA reads (rRNA, tRNA, snoRNA and snRNA) were discarded, and the remaining reads (clean miRNA reads) were analysed by miRDeep2 (2.0.0.5) to identify known and novel miRNAs. 17
MiRNA differential expression analysis
To determine miRNA differential expression between the LPS-stimulated and unstimulated groups, the expression of each miRNA was normalised to the total number of reads in the sample using the following formula: normalised expression = (actual miRNA read count/total clean read count) × 106. 18 The fold change in miRNA reads was presented as log 2 transformation using the following formula: fold change=log 2 (LPS/control).
Target prediction and functional analysis
The potential target genes of the differentially expressed miRNAs were predicted using MiRanda (http://www.microrna.org/microrna/getDownloads.do). We applied DAVID 6.8 to perform gene ontology (GO) functional classification and pathway enrichment analysis of the target genes.
Statistical analyses
The data are shown as means ±
Results
The up-regulated expressions of TNF-α, IL-1β and IL-6 in LPS-stimulated PBMCs
In order to identify principal LPS-responsive miRNAs, PBMCs isolated from the whole blood of the three healthy pigs were stimulated with LPS for 8 h to induce inflammation. We applied ELISA to determine the protein levels of TNF-α and IL-1β. As shown in Figure 1a, the concentration of TNF-α and IL-1β in supernatants of the LPS-stimulated PBMCs were obviously increased compared to the control (
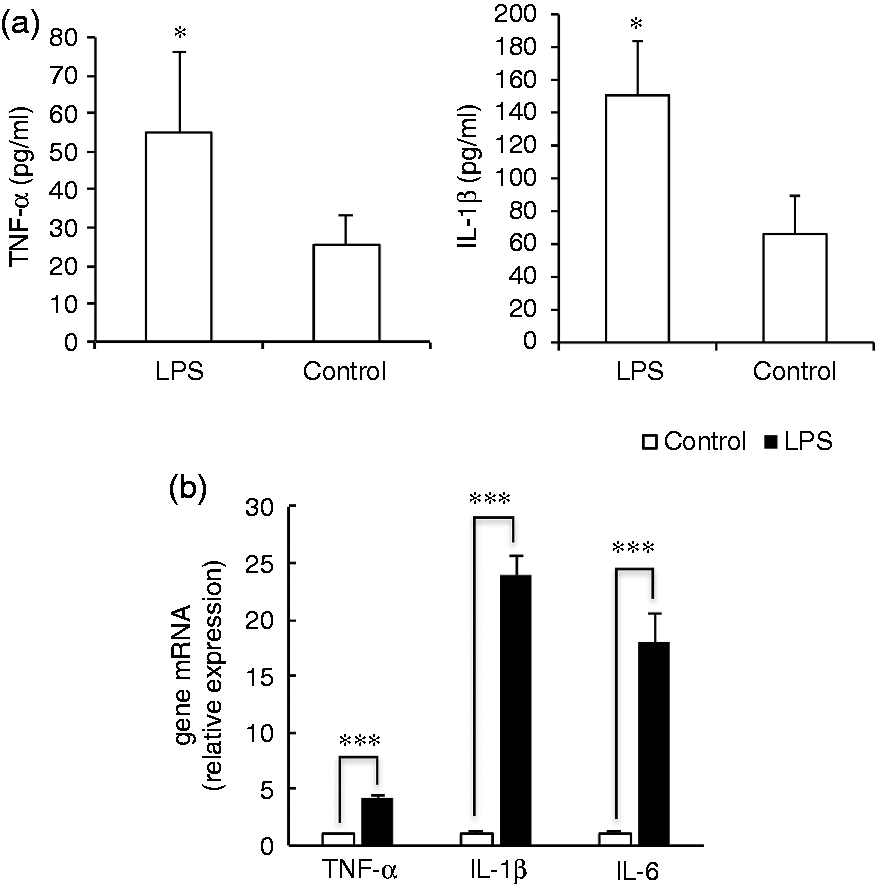
LPS-induced acute inflammation in porcine PBMCs. (a) The concentration of TNF-α in supernatants of the PBMCs were measured 8 h after treatment with or without LPS. (b) TNF-α, IL-1β and IL-6 mRNA expression were determined by quantitative PCR in porcine PBMCs 8 h after treatment with or without LPS. The data represent the mean ± SD.
General analysis of small RNAs
Three independent RNA libraries were constructed for both the LPS-stimulated and unstimulated PBMCs, and then deep sequenced using the Illumina Hiseq2500 system. The six small RNA libraries from LPS treatment groups L1–L3 and control groups N1–N3 yielded a total of 30,002,256, 26,862,983, 25,945,596, 28,820,384, 25,815,123 and 22,992,993 raw reads, respectively. After eliminating adaptor and low-quality reads, a total of 25,785,759, 23,072,789, 21,888,402, 24,466,303, 21,924,186 and 19,793,933 clean reads were obtained in the L1–L3 and N1–N3 libraries, respectively (Table 1). All clean reads were then aligned to the pig genome databases (i.e. miRBase, Rfam, RepBase and mRNA/EST; Table 1). The sequence length distribution in the six libraries exhibited wide variation, ranging from 14 to 41 nt. Most of the small RNAs were 20–3 nt in length, predominantly 22 nt, which is the typical length of Dicer-derived products (Figure 2).
Preliminary analysis of high-throughput sequencing data.
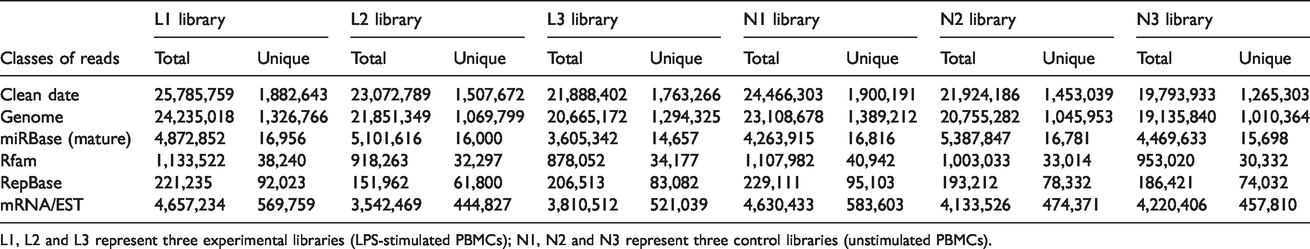
L1, L2 and L3 represent three experimental libraries (LPS-stimulated PBMCs); N1, N2 and N3 represent three control libraries (unstimulated PBMCs).
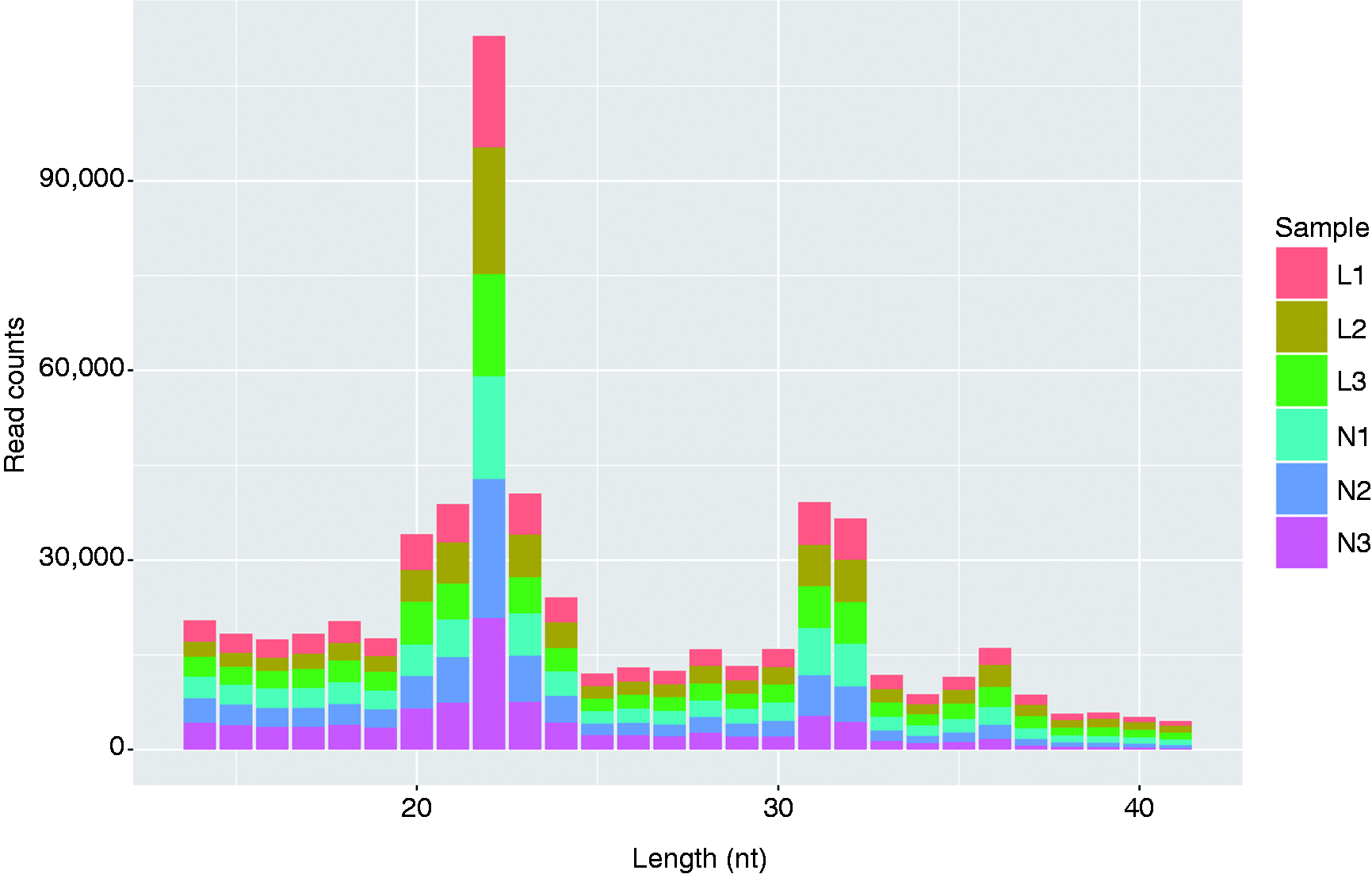
Length distribution of sequencing reads in the six libraries.
Identification of known and novel miRNAs
By performing BLASTN searches against miRBase v21.0, a total of 291 known miRNAs were identified (Supplemental Table S2). The relative number of normalised sequence reads of each miRNA was considered as an indication of its abundance. The top 10 most abundant miRNAs (ssc-miR-148a-3p, ssc-let-7g, ssc-let-7f, 3_8760, ssc-miR-26a, ssc-miR-451, ssc-miR-21, ssc-miR-30d, ssc-miR-99a and ssc-miR-103) in the two groups were ordered by the average proportion of each miRNA read relative to the total normalised miRNA reads (Figure 3). Then, we identified novel miRNAs in our small RNA deep-sequencing libraries by miRDeep2, which predicted 274 potential novel miRNAs at the relatively stringent score cut-off of 5 and signal-to-noise ratio of 17.1 (Table 2). The predicted novel miRNAs were then filtered by removal of RNA without hairpin structure, which reduced the list to 228 novel miRNAs with significant randfold
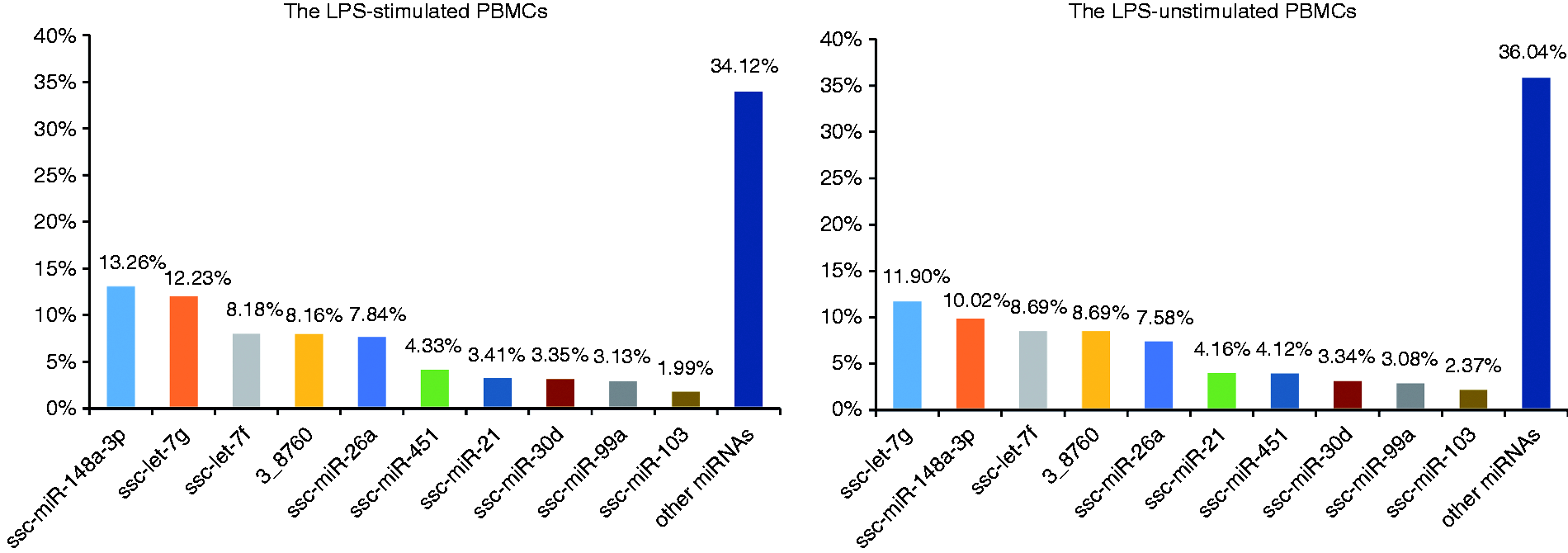
Top 10 most abundant miRNAs in the LPS-stimulated and unstimulated porcine PBMCs ordered by the average proportion of each miRNA read relative to total normalised miRNA reads.
Survey by miRDeep2 analysis showing the number of novel miRNAs under different score cut-offs ranging from 10 to 1.
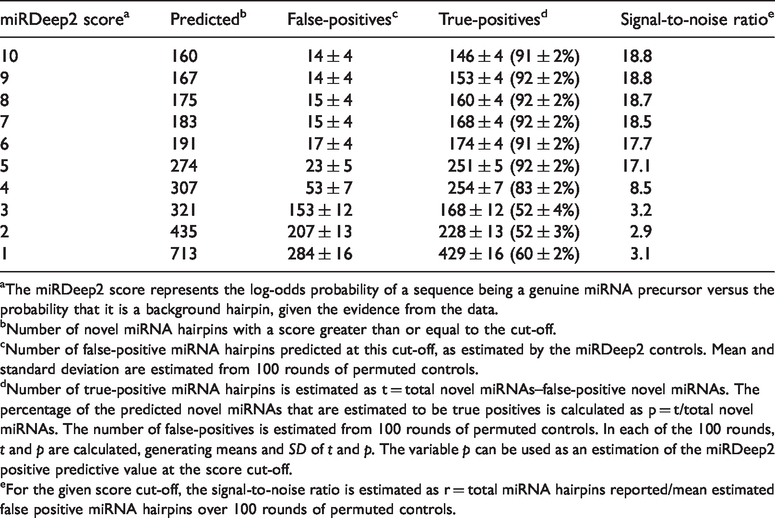
aThe miRDeep2 score represents the log-odds probability of a sequence being a genuine miRNA precursor versus the probability that it is a background hairpin, given the evidence from the data.
bNumber of novel miRNA hairpins with a score greater than or equal to the cut-off.
cNumber of false-positive miRNA hairpins predicted at this cut-off, as estimated by the miRDeep2 controls. Mean and standard deviation are estimated from 100 rounds of permuted controls.
dNumber of true-positive miRNA hairpins is estimated as t = total novel miRNAs–false-positive novel miRNAs. The percentage of the predicted novel miRNAs that are estimated to be true positives is calculated as p = t/total novel miRNAs. The number of false-positives is estimated from 100 rounds of permuted controls. In each of the 100 rounds,
eFor the given score cut-off, the signal-to-noise ratio is estimated as r = total miRNA hairpins reported/mean estimated false positive miRNA hairpins over 100 rounds of permuted controls.
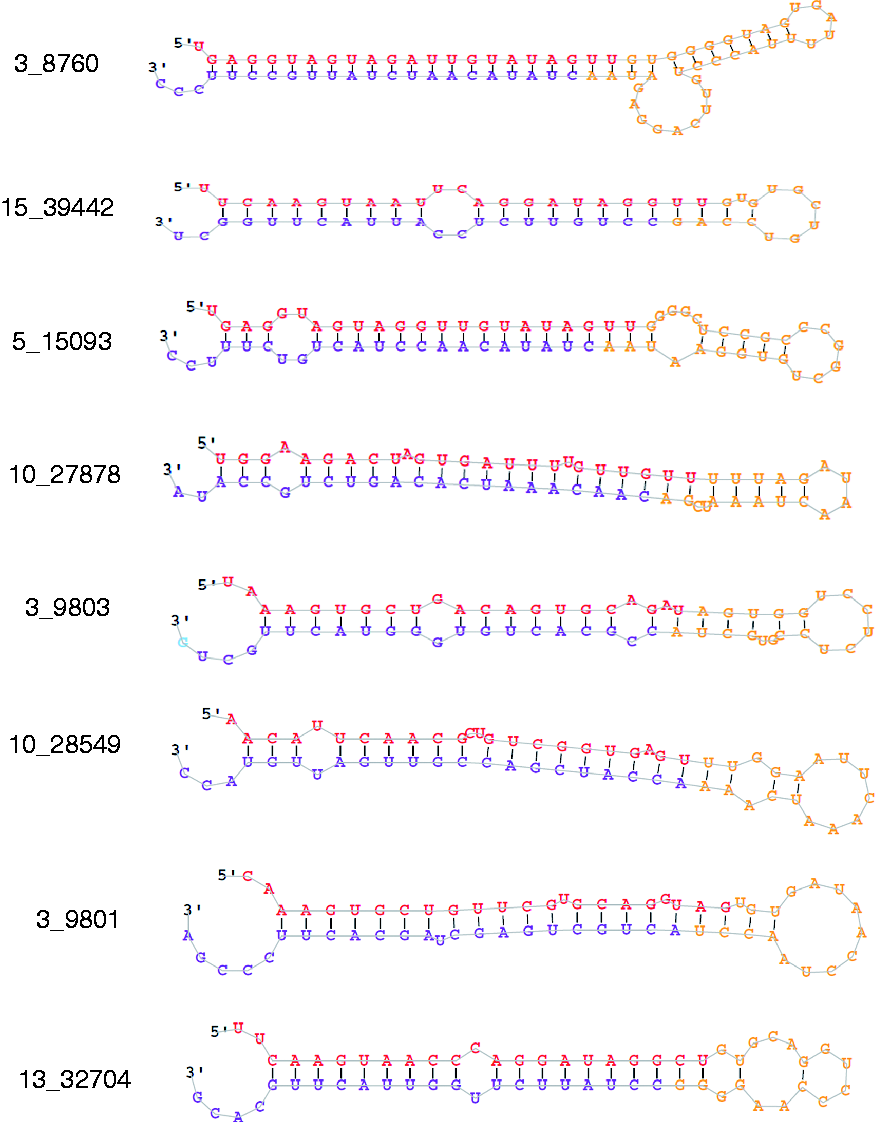
Predicted stem-loop secondary structures of eight candidate novel pig miRNAs. Red: mature sequence; purple: star sequence with no reads cover; yellow: stem sequence.
Differential expression analysis
To investigate the changes in miRNA expression in porcine PBMCs after
miRNAs significantly up- or down-regulated in LPS stimulated PBMCs.
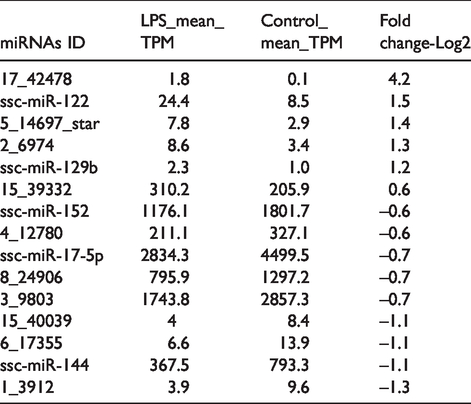
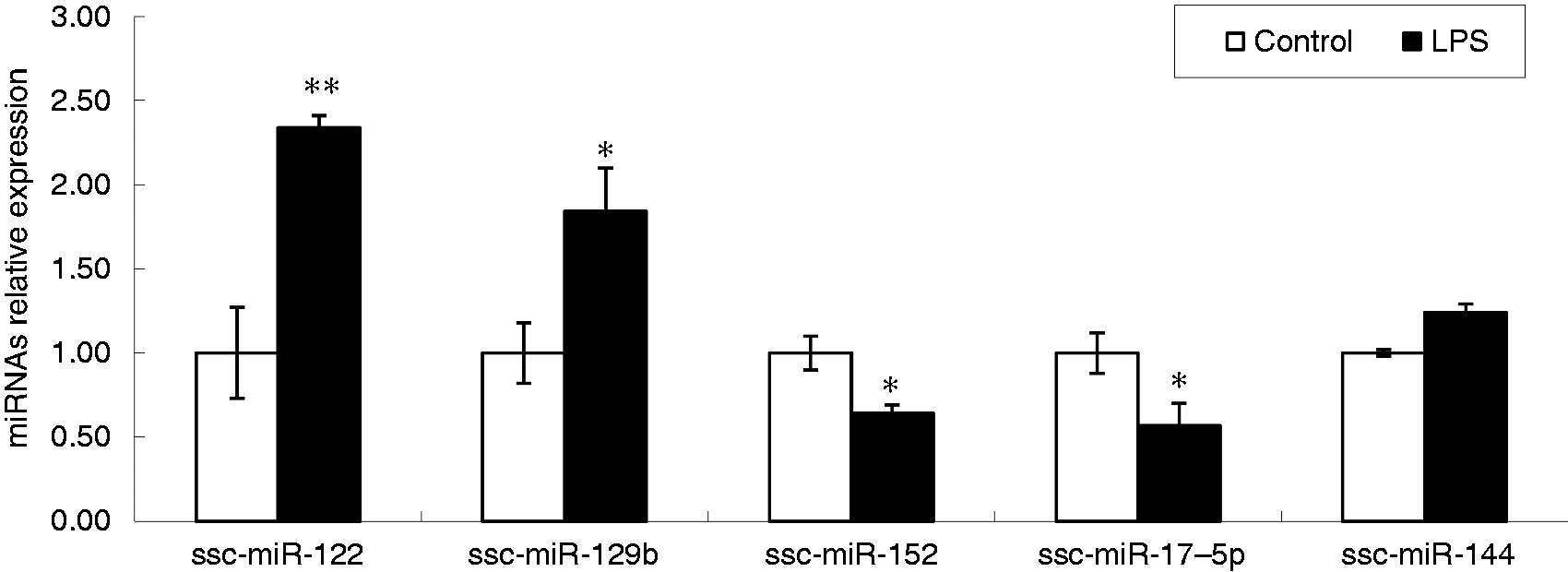
Quantitative PCR was performed to verify expressions of the five miRNAs (ssc-miR-122, ssc-miR-129b, ssc-miR-152, ssc-miR-17-5p and ssc-miR-144) in RNA sequencing. Expression was normalised by U6. The data represent the mean ± SD. n = 3. *
GO and functional classification
The potential targets of the novel differentially expressed miRNAs are listed in Supplemental Table S4. The genes associated with the significantly enriched GO terms and KEGG pathways are shown in Figures 6 and 7, respectively. The GO annotation was classified into three functional categories: 40 catalogues of biological process (BP), 16 catalogues of cellular component (CC) and seven catalogues of molecular function (MF). In the biological process category, most of the annotated targets were involved in the positive regulation of transcription from RNA polymerase II promoter, the negative regulation of cell proliferation and the intracellular protein transport. In the cellular component category, most targets were assigned to the cytoplasm and the extracellular exosome. In the molecular function category, most targets were associated with the thiol-dependent ubiquitin-specific protease activity, the cysteine-type endopeptidase activity and the protein heterodimerisation activity.
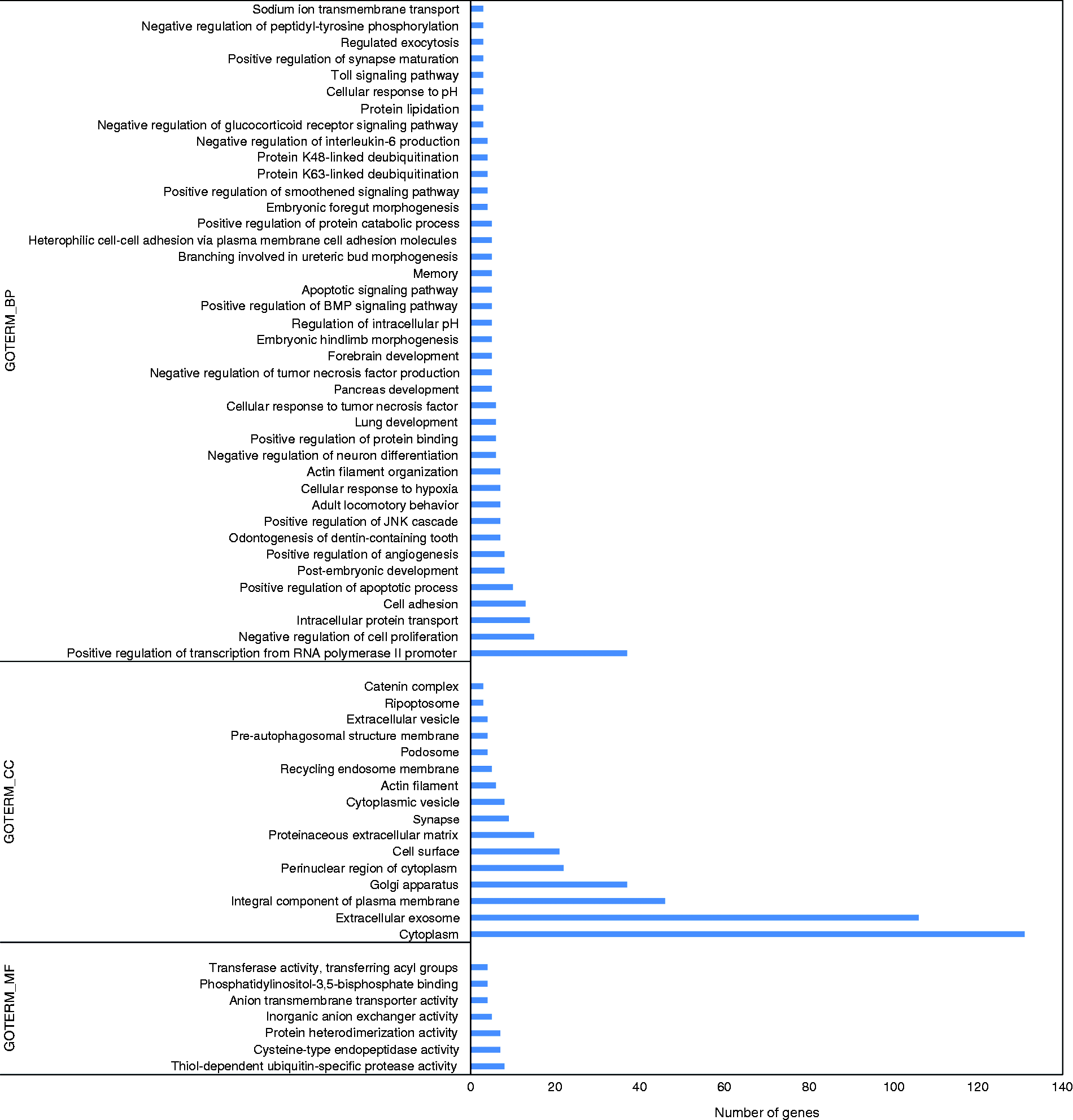
Gene ontology functional analysis of the potential targets of the novel differentially expressed miRNAs. We display the enriched pathway terms with a
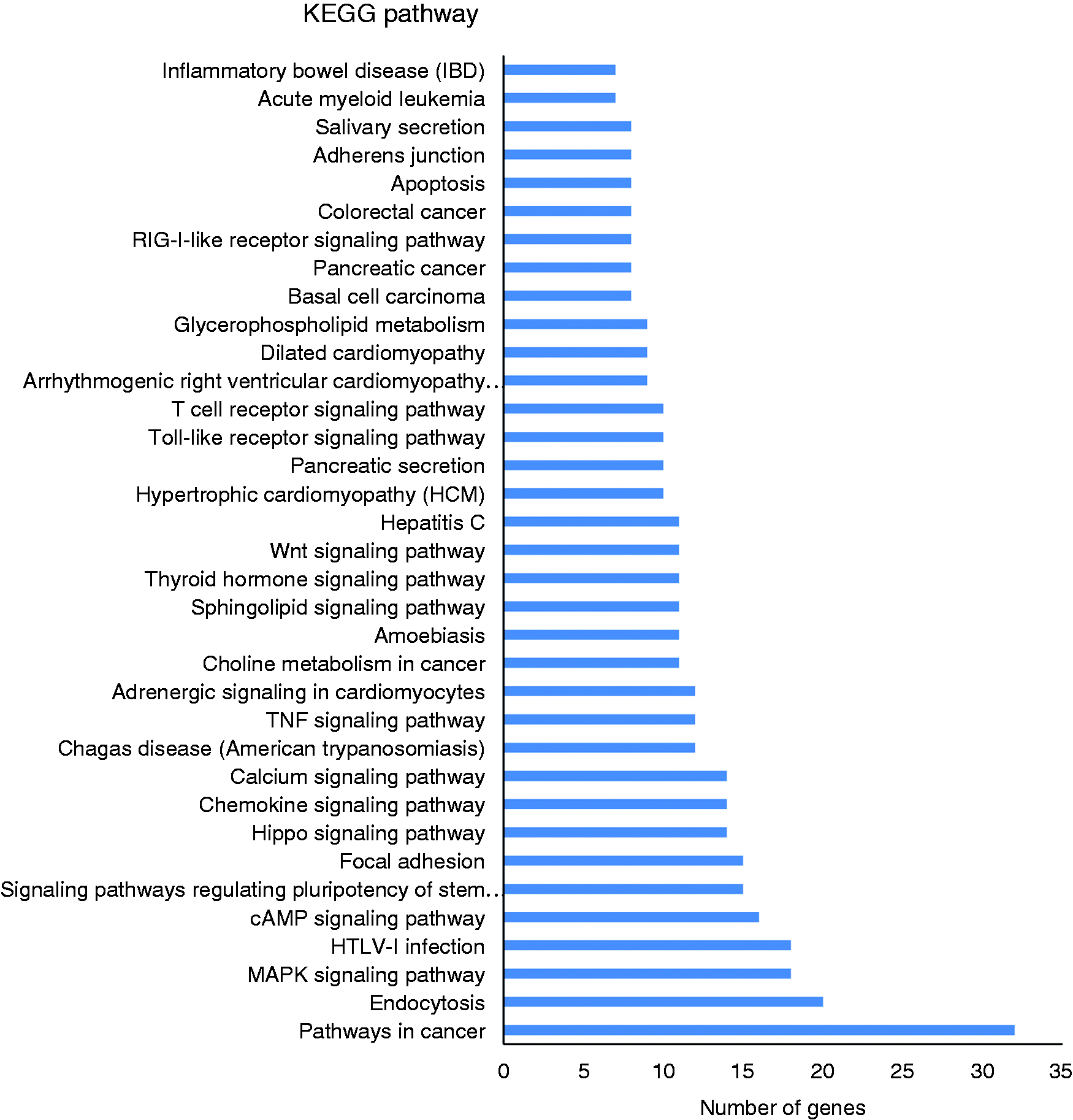
KEGG pathway analysis of the potential targets of the novel differentially expressed miRNAs. We display the enriched pathway terms with a
Furthermore, KEGG pathway analysis demonstrated that the target genes were mainly involved in immunity, such as the pathways in cancer, endocytosis, the chemokine signalling pathway, the TNF signalling pathway, the TLR signalling pathway and the T-cell receptor signalling pathway.
Discussion
As a potent agonist of TLR4, LPS can activate TLR4 signalling and induces the expression of the inflammatory cytokines TNF-α, IL-1β and IL-6. 4 In this study, LPS stimulated the production of TNF-α, IL-1β and IL-6 in porcine PBMCs, indicating that acute inflammation was induced by LPS challenge.
To our knowledge, this study is the first to survey miRNA expression profiles in porcine PBMCs with LPS stimulation. A total of 291 known miRNAs and 228 novel miRNAs were identified in our sequencing. Zhang et al. identified the miRNA signature of human blood mononuclear cells and found 108 highly expressed miRNAs. 19 We found that there were common miRNAs with abundance expressions in both human and porcine PBMCs, such as miR-148, let-7, miR-26, miR-21, miR-99, miR-30 and miR-103. Singh et al. used miRNA sequencing to explore the miRNA expression profiles in TLR ligand-stimulated and non-stimulated PBMCs isolated from the blood of healthy Murrah buffaloes. They found that the abundantly expressed miRNAs include bta-miR-103, bta-miR-19b, -191 -29b, -15a, -19a, -30d, -30b-5p and members of the let family (let 7a-5p, let 7g and let 7f) in LPS- and CpG-treated and control samples, while bta-miR-21-5p was highly expressed in LPS-treated PBMCs only. 20 In our sequencing libraries, four miRNAs (ssc-let-7g, ssc-let-7f, ssc-miR-30d and ssc-miR-103) were also identified with high abundance, accounting for 26.78% and 26.30% of the total normalised miRNA reads in the LPS-treated and control groups, respectively. However, we found that ssc-miR-21 was abundantly expressed in both LPS-treated and control samples. The other most abundant miRNAs shared between the LPS-stimulated and unstimulated PBMCs were ssc-miR-148a-3p, 3_8760, ssc-miR-26a, ssc-miR-451 and ssc-miR-99a. We performed GO and KEGG analysis of the potential targets of the top nine most abundant known miRNAs (ssc-miR-148a-3p, ssc-let-7g, ssc-let-7f, 3_8760, ssc-miR-26a, ssc-miR-451, ssc-miR-21, ssc-miR-30d, ssc-miR-99a and ssc-miR-103). The results are shown in Supplemental Figures S1 and S2. In the biological process category, most of the annotated targets were involved in the regulation of transcription from RNA polymerase II promoter. In the cellular component category, most targets were assigned to the cytoplasm, nucleus and nucleoplasm. In the molecular function category, most targets were associated with zinc ion binding and ATP binding. The KEGG pathways predictions indicated a significant enrichment in GO functional categories related to the phosphatidylinositol signalling system, HTLV-I infection and protein processing in the endoplasmic reticulum. MiR-148a is implicated in a series of biological processes, including cellular proliferation, apoptosis, metastasis and invasion.21–23 MiR-26a has been reported to regulate important cellular processes such as cellular differentiation, cell growth, apoptosis and metastasis, thereby participating in the initiation and development of various diseases. 24 MiR-451 and miR-99a play role in a variety of tumours and are potential candidates for therapy.25–27 However, there is very little specific information on the expression and function of these four miRNA in PBMCs.
In the present study, four known miRNAs (ssc-miR-122, ssc-miR-129b, ssc-miR-152 and ssc-miR-17-5p) and 10 novel miRNAs showed expression changes on LPS stimulation of porcine PBMCs. MiR-122 is the most abundant miRNA in the liver, which can be involved in liver biology and disease.28,29 It can directly target cyclin G1, Bcl-w (anti-apoptosis gene), WNT1 and so on.30–32 A recent study reported that miR-122 plays a role in innate immunity by regulating TLR4 expression in hepatoma cells. 33 MiR-129 has been shown to be involved in the formation of many types of cancer and attenuates cell proliferation by targeting high-mobility group box 1 (HMGB1).34,35 As a ubiquitous nuclear protein, HMGB1 acts as a danger signal when released extracellularly after cellular activation, stress, damage or death. 36 Extracellular HMGB1 is a mediator implicated in the pathogenesis of many inflammatory conditions and diseases. 37 Therefore, further studies have to be carried out in porcine PBMCs to identify the potential functions of ssc-miR-122 and ssc-miR-129b, and the target genes in inflammatory process. The ssc-miR-152 and ssc-miR-17-5p have more abundant expression in PBMCs compared to other differentially expressed miRNAs. Previous studies reported that MiR-152 can impair innate response by targeting calcium/calmodulin-dependent protein kinase II (CaMKIIα).38,39 CaMKIIα, a major downstream effector of calcium (Ca2+), could promote TLR-triggered production of pro-inflammatory cytokines and type I IFN. 40 MiR-17-5p is a well-known oncogenic miRNA, which regulates tumorigenesis, proliferation, invasiveness and cell-cycle progression by targeting various tumour suppressors.41–43 A recent study by Ji et al. found that miR-17-5p expression significantly decreased in PBMCs isolated from mice which were injected with 50 mg/kg LPS for 4 h. 44 These data are consistent with our results. In addition, Ji et al. found that overexpression of miR-17-5p inhibited LPS-induced IL-1β and TNF-α expressions in macrophages (RAW264.7 cells) by targeting TLR4. 44 According to our miRNA sequencing data, ssc-miR-129b were poorly expressed (<10 TPM per miRNA) in porcine PBMCs, while the four miRNAs (ssc-miR-122, ssc-miR-152 and ssc-miR-17-5p) were modestly expressed (10–10,000 TPM per miRNA). Therefore, further studies must be carried out in porcine PBMCs to identify the potential functions and target genes of ssc-miR-122, ssc-miR-152 and ssc-miR-17-5p in inflammatory process. However, ssc-miR-129b existed at low levels and might have little impact on immunity or inflammation in porcine PBMCs.
In addition, we performed function analysis of the potential targets of the novel differentially expressed miRNAs. Target gene and KEGG pathway predictions indicated a significant enrichment in GO functional categories related to diseases, immunity and inflammation, including the pathways in cancer, endocytosis, MAPK signalling pathway, HTLV-I infection, the chemokine signalling pathway, the TNF signalling pathway, the TLR signalling pathway and the T-cell receptor signalling pathway. For example, we predicted that IL12RB1 was the potential target of the novel miRNA 17_42478. IL12RB1 is a Mendelian susceptibility to mycobacterial disease (MSMD) gene, which is essential for mycobacterial disease resistance and T-cell differentiation. 45 In addition, we predicted TRAF3 was the potential target of the novel miRNA 5_14697. In TLR and RIG-I-like receptor (RLR) signalling pathways, TRAF3 is a highly versatile regulator that positively controls type I IFN production but negatively regulates MAPK activation and alternative NF-κB signalling. 46 TRAF3 is also known to function as a resident nuclear protein, and to impact B-cell survival and activation. 47 CXCL2 (macrophage inflammatory protein-2 (MIP-2)), a critical chemokine for neutrophils, controls the early stage of neutrophil recruitment during inflammation. 48 TLR4-expressing tissue macrophages release CXCL2 following stimulation with LPS. 49 CEBPB is a transcription factor activated by LPS or IL-17, and itself regulates numerous genes involved in inflammation, including IL-6 and IL-23R.50,51 We predicted CXCL2 and CEBPB were the potential targets of the novel miRNA 2_6974 and 6_17355, respectively.
Here, we have provided the changes of miRNA expression in LPS-stimulated PBMCs of pigs, which will serve as a reference point for future functional studies or challenge experiments directed to uncover the role of miRNAs in immunity and disease pathogenesis.
Supplemental Material
INI901560 Supplemental material1 - Supplemental material for Analysis of microRNA expression profiles in porcine PBMCs after LPS stimulation
Supplemental material, INI901560 Supplemental material1 for Analysis of microRNA expression profiles in porcine PBMCs after LPS stimulation by Jing Zhang, Xin Xu, Xingfa Huang, Huiling Zhu, Hongbo Chen, Wenjun Wang and Yulan Liu in Innate Immunity
Supplemental Material
INI901560 Supplemental material2 - Supplemental material for Analysis of microRNA expression profiles in porcine PBMCs after LPS stimulation
Supplemental material, INI901560 Supplemental material2 for Analysis of microRNA expression profiles in porcine PBMCs after LPS stimulation by Jing Zhang, Xin Xu, Xingfa Huang, Huiling Zhu, Hongbo Chen, Wenjun Wang and Yulan Liu in Innate Immunity
Supplemental Material
INI901560 Supplemental material3 - Supplemental material for Analysis of microRNA expression profiles in porcine PBMCs after LPS stimulation
Supplemental material, INI901560 Supplemental material3 for Analysis of microRNA expression profiles in porcine PBMCs after LPS stimulation by Jing Zhang, Xin Xu, Xingfa Huang, Huiling Zhu, Hongbo Chen, Wenjun Wang and Yulan Liu in Innate Immunity
Supplemental Material
INI901560 Supplemental material4 - Supplemental material for Analysis of microRNA expression profiles in porcine PBMCs after LPS stimulation
Supplemental material, INI901560 Supplemental material4 for Analysis of microRNA expression profiles in porcine PBMCs after LPS stimulation by Jing Zhang, Xin Xu, Xingfa Huang, Huiling Zhu, Hongbo Chen, Wenjun Wang and Yulan Liu in Innate Immunity
Supplemental Material
INI901560 Supplemental material5 - Supplemental material for Analysis of microRNA expression profiles in porcine PBMCs after LPS stimulation
Supplemental material, INI901560 Supplemental material5 for Analysis of microRNA expression profiles in porcine PBMCs after LPS stimulation by Jing Zhang, Xin Xu, Xingfa Huang, Huiling Zhu, Hongbo Chen, Wenjun Wang and Yulan Liu in Innate Immunity
Supplemental Material
INI901560 Supplemental material6 - Supplemental material for Analysis of microRNA expression profiles in porcine PBMCs after LPS stimulation
Supplemental material, INI901560 Supplemental material6 for Analysis of microRNA expression profiles in porcine PBMCs after LPS stimulation by Jing Zhang, Xin Xu, Xingfa Huang, Huiling Zhu, Hongbo Chen, Wenjun Wang and Yulan Liu in Innate Immunity
Footnotes
Acknowledgements
This study was supported by the National Natural Science Foundation of China (31702203), the Open Project of Hubei Key Laboratory of Animal Nutrition and Feed Science (grant number 201908), the Project of the Hubei Provincial Department of Education (T201508) and the projects of Wuhan Science and Technology Bureau (2018020401011304).
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
