Abstract
CDC2-like kinase 2 is a promising target for the therapy of knee osteoarthritis. A series of natural and commercial triterpenoid derivatives were evaluated for their inhibitory activity against CDC2-like kinase 2 using ADP-Glo assay by GloMax® Navigator Microplate Luminometer. Most of these triterpenoid derivatives exhibited the moderate inhibitory activity against CDC2-like kinase 2, but in particular, Methyl ganoderate A showed the highest inhibitory activity with an IC50 value of 0.69 μM against CDC2-like kinase 2. Methyl ganoderate A showed the inhibitory activity against CLK1, CLK2, CLK3, CLK4, DYRK1A, and DYRK2 with IC50 values of 8.79 μM, 0.69 μM, >20 μM, 10.75 μM, 9.06 μM, and >20 μM, indicating that it is a novel and selective CDC2-like kinase 2 inhibitor. Preliminary structure–activity relationships (SARs), molecular docking, and molecular dynamics simulation of Methyl ganoderate A against CDC2-like kinase 2 were studied. The investigation of mechanisms indicated that Methyl ganoderate A inhibited cartilage degradation and exerted anti-inflammatory effects in vitro.
Keywords
Introduction
Knee osteoarthritis as a typical degenerative disease causes disability in older adults. 1 Pain and disability of knee osteoarthritis severely affect the quality of life for patients. 2 The treatment of knee osteoarthritis imposes a heavy economic burden in the current society. CDC2-like kinase 2 (CLK2) as a member of the CDC2-like kinase family is the key protein kinase that phosphorylates splice factors. 3 CLK2 could affect chondrocyte function, bone differentiation, and inflammatory response, making CLK2 a promising target for the treatment of knee osteoarthritis. 4
Heterocyclic compounds display various activities, 5 and several heterocyclic derivatives as CLK2 inhibitors are designed, synthesized and evaluated. 6 Indazole derivative LBM22 (Figure 1) exhibited potent inhibition of CLK2 with an IC50 value of 3.9 nM and inhibited SR protein phosphorylation in a concentration-dependent manner. 7 2,4,5-Trisubstituted-7H-pyrrolo[2,3-d]pyrimidine derivative CC-671 displayed a dual inhibition profile targeting CLK2 and Monopolar spindle 1 (Mps1, also known as TTK). 8 The isoquinoline derivative cirtuvivint exhibited the broad-spectrum inhibitory activity against CLK1, CLK2, CLK3, and CLK4 with an Adenosine triphosphate (ATP)-competitive manner. 9 Indazole derivative SM04755 as a dual CLK2/DYRK1A inhibitor could suppress the Wnt pathway and reduced the inflammation response. 10 Indazole derivative Lorecivivint as a dual inhibitor against CLK2 and DYRK1A entered a phase 2b trial for the treatment knee osteoarthritis. 11 Indazole derivative LQ23 inhibited Wnt/CLK2 signaling pathway and exhibited antiosteoarthritis effects in vitro and in vivo. 12 These representative CLK2 inhibitors were shown in Figure 1, and most of these reported compounds demonstrated the low selectivity for CLK2. Due to the similarity and high homology of CDC2-like kinase family members, discovery of novel and selective CLK2 inhibitors is extremely challenging.
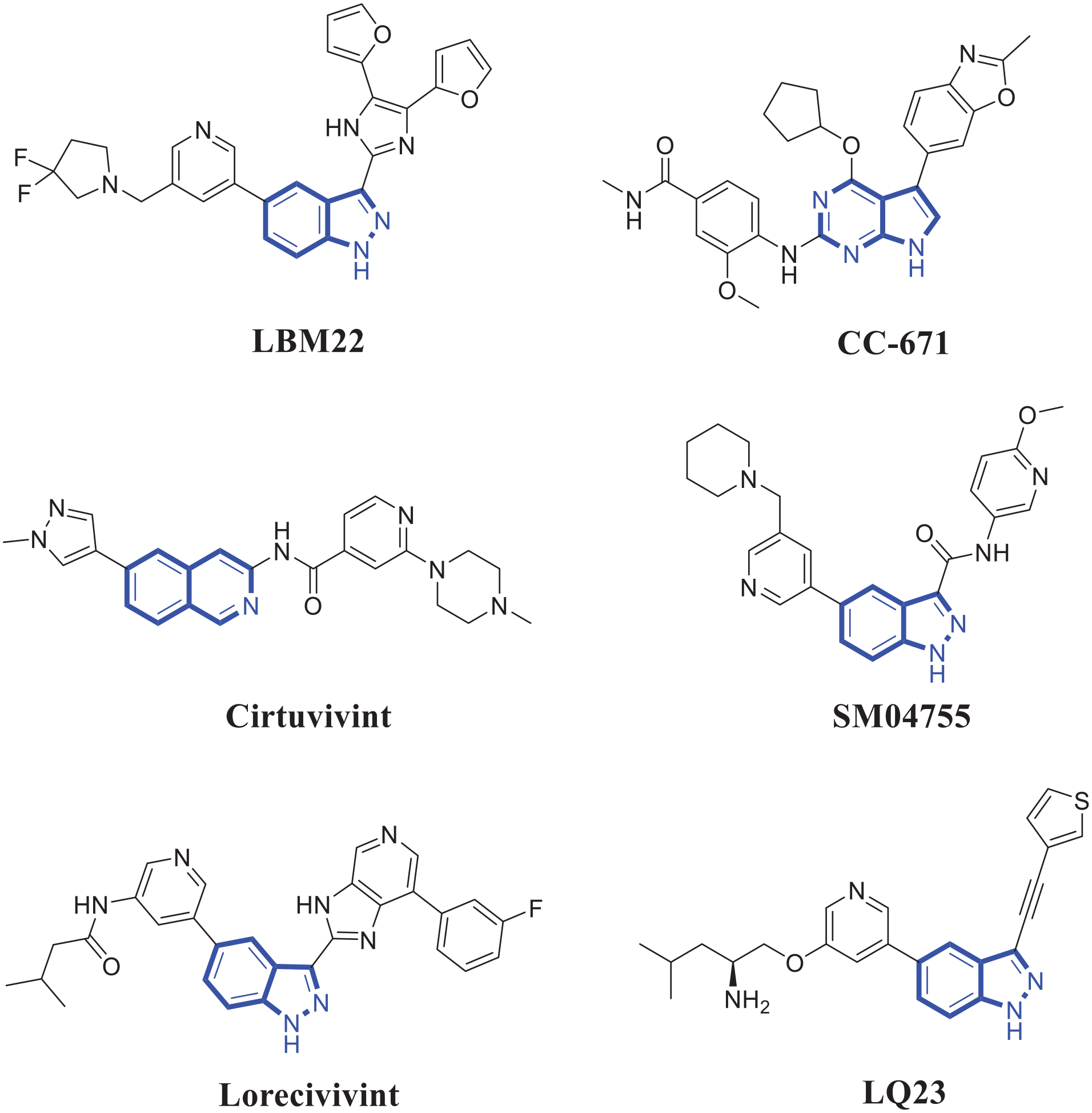
The representative CLK2 inhibitors.
In this work, a series of triterpenoid derivatives were evaluated for their inhibitory activity against CLK2. Among these triterpenoid derivatives, Methyl ganoderate A (MGA) was identified as a novel and selective CLK2 inhibitor. In addition, MGA suppressed cartilage degradation and exhibited anti-inflammatory effects in vitro. To the best of our knowledge, there are no reports of triterpenoid derivatives as novel and selective CLK2 inhibitors for the treatment of knee osteoarthritis so far.
Results and discussion
A series of triterpenoid derivatives exhibited the inhibitory activity against CLK2
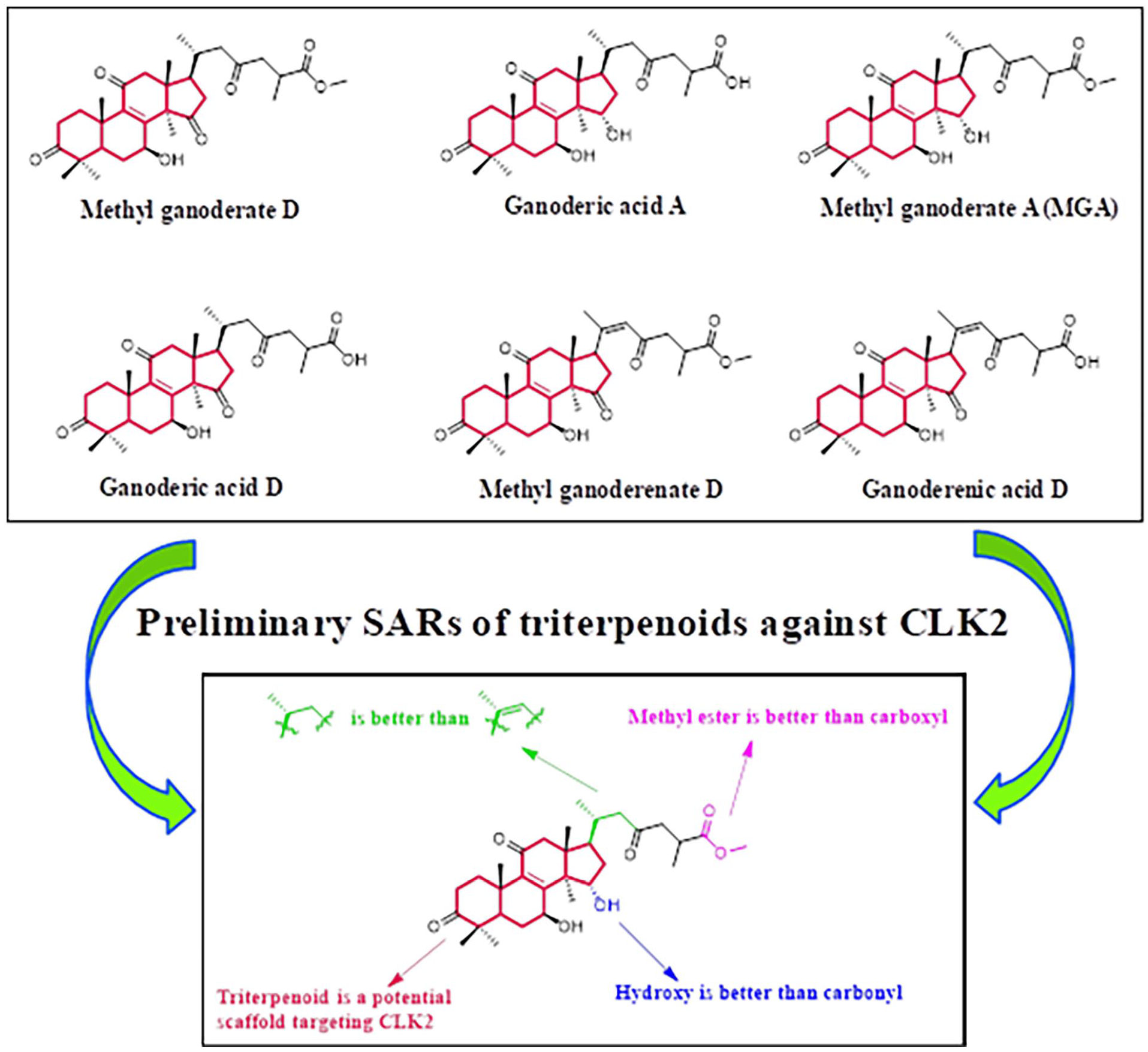
Ganoderic acid A as a natural triterpenoid extracted from Ganoderma was reported that it could improve knee osteoarthritis by regulating the ratio of RANKL/OPG. 13 Therefore, we hypothesized that Ganoderic acid A derivatives may have the potential inhibitory activity against CLK2 for the treatment of knee osteoarthritis. In this work, six natural triterpenoid derivatives (Methyl ganoderate D, Ganoderic acid A, MGA, Ganoderic acid D, Methyl ganoderenate D, and Ganoderenic acid D) were evaluated for their inhibitory activity against CLK2. Their chemical structures and inhibitory rates against CLK2 are shown in Figure 2. When the concentration is 10 μM, the inhibitory rates of Methyl ganoderate D, Ganoderic acid A, MGA, Ganoderic acid D, Methyl ganoderenate D and Ganoderenic acid D are 38.3%, 45.7%, 96.7%, 32.3%, 29.0%, and 22.7%, respectively. When the concentration is 20 μM, the inhibitory rates of Methyl ganoderate D, Ganoderic acid A, MGA, Ganoderic acid D, Methyl ganoderenate D, and Ganoderenic acid D are 52.0%, 63.3%, 97.7%, 43.0%, 40.3%, and 33.3%, respectively. All results indicated that these natural triterpenoid derivatives might inhibit CLK2 in a concentration-dependent manner.

Triterpenoid derivatives exhibited the inhibitory activity against CLK2. (a) Chemical structures of natural triterpenoids. (b) Inhibitory rates against CLK2.
Preliminary structure–activity relationships of triterpenoid derivatives against CLK2
Based on the above results, all triterpenoid derivatives exhibited moderate to good inhibitory activity against CLK2. These results indicated that triterpenoid is a potential scaffold targeting CLK2. Compared with the inhibitory rate of Ganoderic acid A, MGA showed a better inhibitory activity against CLK2, demonstrating that methyl ester group is better than carboxyl group. The inhibitory rates between Methyl ganoderate D and Ganoderic acid D also illustrated the importance of methyl ester group. When the carbon–carbon double bond on the side chain of Methyl ganoderenate D was replaced with a carbon–carbon single bond on the side chain of Methyl ganoderate D, the inhibitory activity was obviously increased. The results indicated that carbon–carbon single bond on the side chain of triterpenoid derivative is better than carbon–carbon double bond. Replacement of the hydroxy group of Ganoderic acid A with a carbonyl group of Ganoderic acid D led to a decrease of the activity, suggesting that hydroxy group is better than carbonyl group. The preliminary structure–activity relationships (SARs) of these triterpenoid derivatives against CLK2 were summarized in Figure 3.

Preliminary structure–activity relationships of triterpenoids against CLK2.
Discovery of MGA as a novel and selective CLK2 inhibitor
Among all triterpenoid derivatives, MGA displayed the best inhibitory activity against CLK2. Therefore, MGA was selected to evaluate for its selectivity against different kinases (CLK1, CLK2, CLK3, CLK4, DYRK1A, and DYRK2). Based on the results in Figure 4, MGA exerted different inhibitory activities against CLK1, CLK2, CLK3, CLK4, DYRK1A, and DYRK2 with IC50 values of 8.79 μM, 0.69 μM, >20 μM, 10.75 μM, 9.06 μM, and >20 μM, respectively. These results showed that MGA is a novel and selective CLK2 inhibitor.

IC50 values of MGA against different kinases.
Molecular docking of MGA targeting CLK2
In order to explore the binding mode of CLK2 and MGA, CLK2 protein was used to perform docking studies (PDB code: 6FYK). As the docking results in Figure 5(a), MGA was located in the active pocket of CLK2. In addition, MGA (sapphirine structure, Figure 5(b)) was docked into a similar pocket as the original ligand (light red structure, Figure 5(b)). As shown in Figure 5(c), MGA formed three H-bonds with Asp327, Leu169, and Lys292 of CLK2.
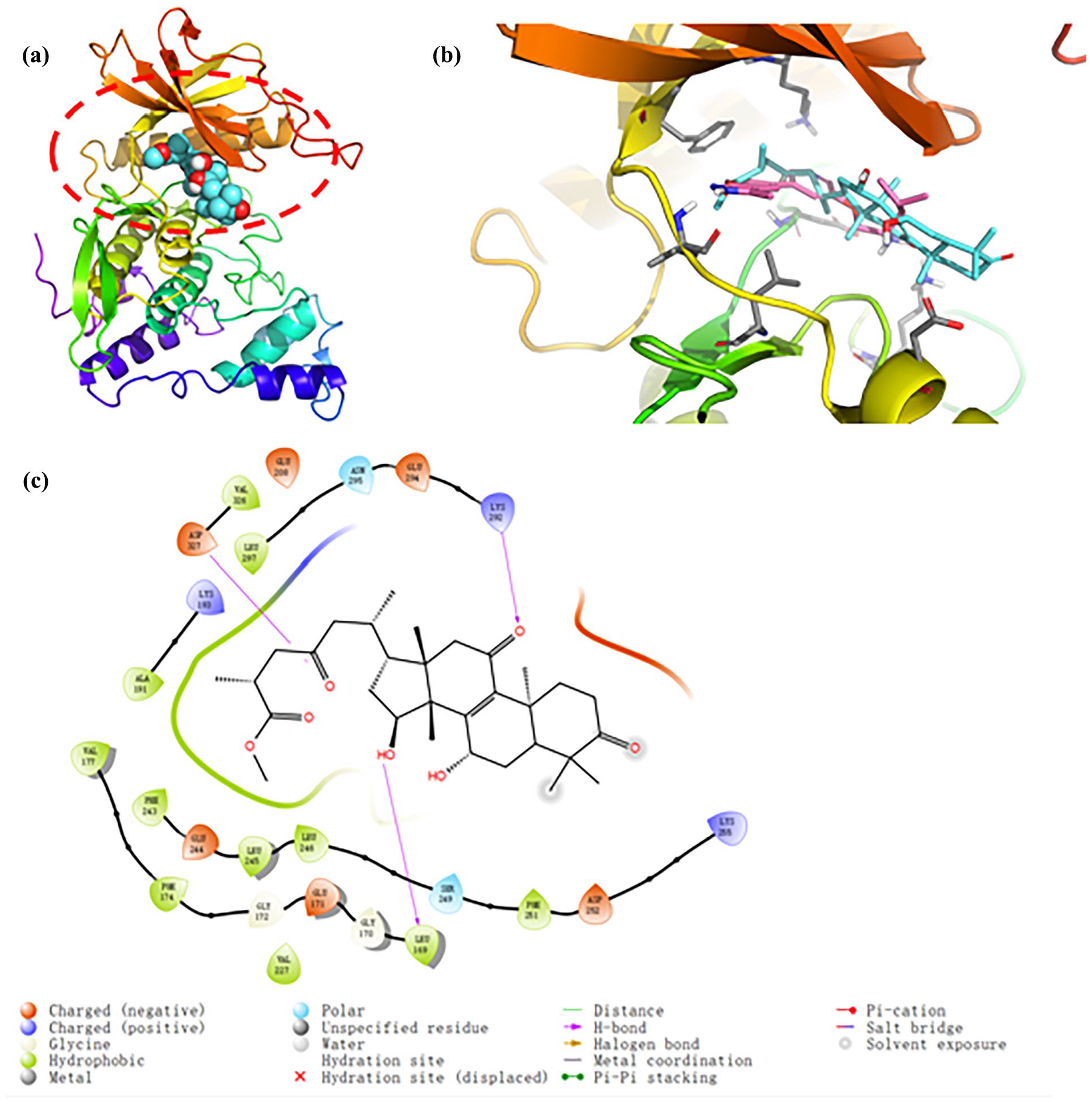
Binding mode of CLK2 and MGA. (a) 3D binding mode of protein surface and MGA, (b) a similar pocket between MGA and the original ligand, and (c) hydrogen bonds and different binding effects.
Molecular dynamics simulation of MGA
Molecular dynamics simulation of MGA targeting CLK2 protein was performed in this work, and results of root mean square deviation (RMSD), radius of gyration, polar surface area, solvent accessible surface area (SASA), root mean square fluctuation (RMSF), and molecular surface area were shown in Figure 6. At least on the time scale of 100 ns, values of RMSD, radius of gyration, polar surface area, SASA, and molecular surface area for MGA do not display any distinctive trends and fluctuate around small constants. During the Molecular dynamics (MD) simulation, RMSF of residues is crucial for evaluating the structural stability of protein. From the results in Figure 6, most residues exhibit low RMSF values, indicating their high stability throughout the simulation. However, some residues have significantly higher RMSF values. This suggests they are in highly flexible regions or underwent major conformational changes during the simulation.
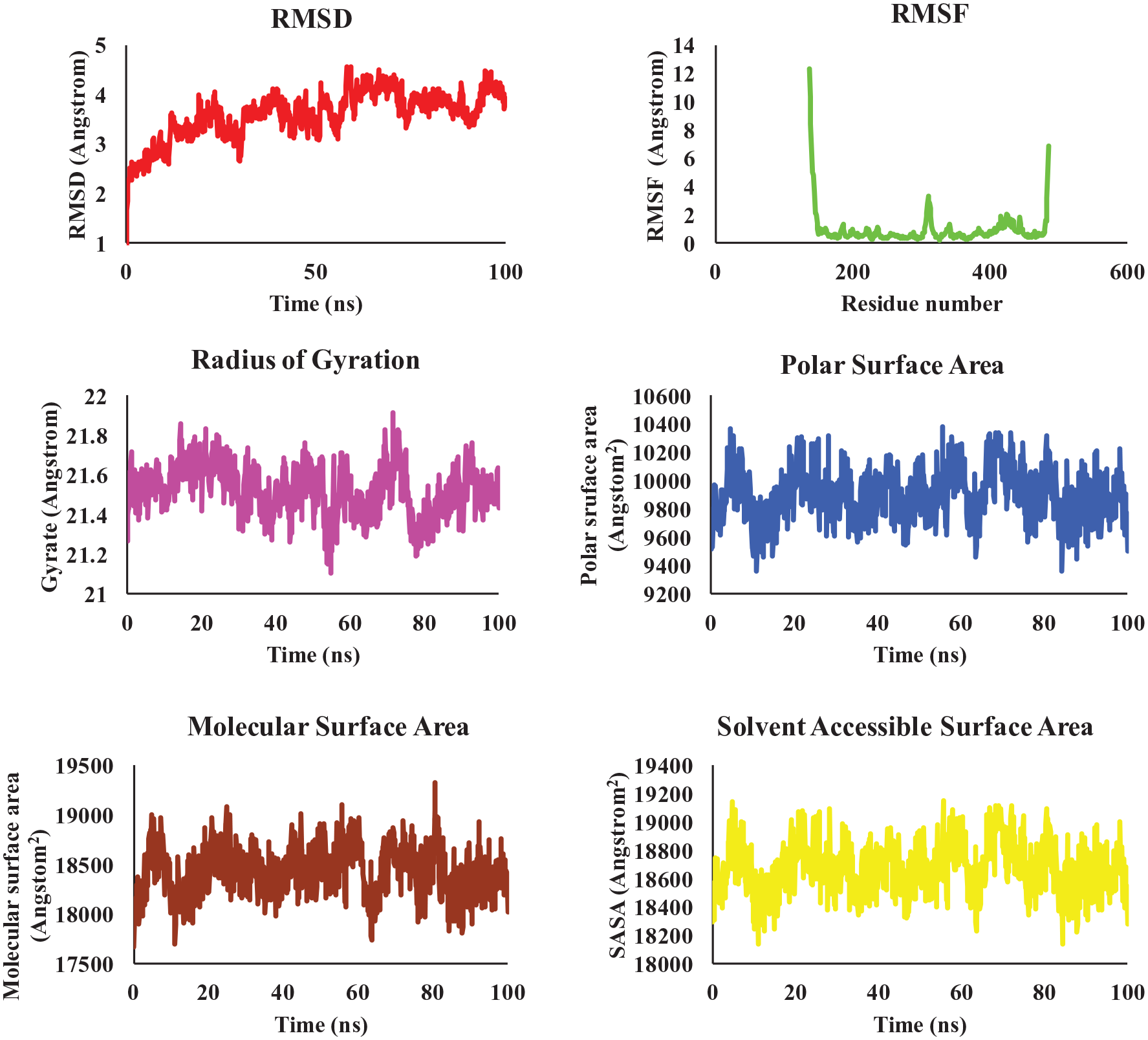
Dynamics of MGA binding to CLK2.
Molecular properties of MGA
Over the past decade, studies on molecular properties of kinase inhibitors have played important roles in drug discovery. 14 Molecular properties analysis of MGA, Lorecivivint, Cirtuvivint, SM04755, and LQ23 as CLK2 inhibitors was performed by a free online service ADMETlab2.0. The optimal values of molecular weight, number of hydrogen bond acceptors (nHA), number of hydrogen bond donors (nHD), number of rotatable bonds (nRot), number of rings (nRing), number of atoms in the biggest ring (MaxRing), number of heteroatoms (nHet), formal charge (fChar), number of rigid bonds (nRigs), topological polar surface area (TPSA), logP, and logD are 100~600, 0~12, 0~7, 0~11, 0~6, 0~18, 1~15, −4~4, 0~30, 0~140, 0~3, and 1~3, respectively. As shown in Table 1, MGA exhibited the acceptable molecular properties. However, some parameters of Lorecivivint, Cirtuvivint, SM04755, and LQ23 are not optimal (red color). These results indicated that MGA is a promising CLK2 inhibitor for further drug development.
Molecular properties analysis of MGA, Lorecivivint, Cirtuvivint, SM04755, and LQ23 as CLK2 inhibitors.

MGA-inhibited cartilage degradation
Long-term treatment of knee osteoarthritis requires the inhibition of cartilage degradation. 15 Therefore, we evaluated the effects of MGA on the catabolism of chondrocytes and bone mesenchymal stem cells (BMSCs) under pathophysiological conditions. When chondrocytes and BMSCs were treated with Oncostatin M (10 ng/mL) and tumor necrosis factor (TNF)-α (20 ng/mL), the expression levels of ADAMTS5, MMP3, and MMP13 as matrix-degrading enzymes were sharply increased (model group, Figure 7(a) and (b)). After the treatment of MGA at 10 μM against chondrocytes and BMSCs under pathophysiological conditions, the expression levels of ADAMTS5, MMP3, and MMP13 were obviously decreased. Lorecivivint, a reported CLK2 inhibitor, was used as the control drug. All these results demonstrated that MGA as a novel and selective CLK2 inhibitor could suppress cartilage degradation.

MGA suppressed cartilage degradation in vitro. (a) Gene expressions of matrix-degrading enzymes (ADAMTS5, MMP3, and MMP13) in BMSCs. (b) Gene expressions of MMP3 and MMP13 in chondrocytes.
MGA exerted the anti-inflammatory activity in vitro
When chondrocytes were treated with Oncostatin M (10 ng/mL) and TNF-α (20 ng/mL), the expression levels of IL-6, TNF-α, and IL-β as inflammatory markers were obviously increased (model group, Figure 8). After the treatment of MGA or lorecivivint against chondrocytes, expression levels of IL-6, TNF-α, and IL-β were decreased. Thus, MGA exerted the anti-inflammatory activity in vitro.

Gene expressions of inflammatory markers (IL-6, TNF-α, and IL-β) in chondrocytes.
Conclusion
In conclusion, a series of natural triterpenoid derivatives were evaluated for their inhibitory activity against CLK2. Among them, MGA showed the highest inhibitory activity against CLK2. In addition, MGA showed the inhibitory activity against CLK1, CLK2, CLK3, CLK4, DYRK1A, and DYRK2 with IC50 values of 8.79 μM, 0.69 μM, >20 μM, 10.75 μM, 9.06 μM, and >20 μM, indicating that it is a novel and selective CLK2 inhibitor. Preliminary SARs, molecular docking, and molecular dynamics simulation of MGA against CLK2 were studied. Importantly, the investigation of mechanisms indicated that MGA inhibited cartilage degradation and exerted anti-inflammatory effects in vitro. To the best of our knowledge, there are no reports of MGA as a selective CLK2 inhibitor for the treatment of knee osteoarthritis so far. This work suggested that MGA is a potential candidate to treat knee osteoarthritis.
Experimental section
Source and purity of triterpenoid derivatives
Methyl ganoderate D, Ganoderic acid A, MGA, Ganoderic acid D, Methyl ganoderenate D, and Ganoderenic acid D as natural triterpenoid derivatives were purchased from Wuhan Tianzhi Biotechnology Co., Ltd. (Wuhan, China). CAS number of Methyl ganoderate D, Ganoderic acid A, MGA, Ganoderic acid D, Methyl ganoderenate D, and Ganoderenic acid D are 97210-12-3, 81907-62-2, 105742-78-7, 108340-60-9, 748136-03-0, and 100665-43-8, respectively. The purity of these natural triterpenoid derivatives is greater than 98%.
Inhibitory activity of kinases assay
ADP-Glo assay was used to evaluate the inhibitory activity against kinases. Magnesium chloride (10 mM), HEPES (50 mM), ethylene glycol tetra-acetic acid (1 mM), BriJ35 (0.01%), and dithiothreitol (2 mM) were added into wells of a 96-well plate. The concentrations of triterpenoid derivatives are 10 μM and 20 μM to perform the inhibitory rates against CLK1, CLK2, CLK3, CLK4, DYRK1A, and DYRK2. Dimethyl sulfoxide was used to prepare the solution of MGA at different concentrations (0.038 nM, 0.15 nM, 0.61 nM, 2.44 nM, 9.77 nM, 39.06 nM, 156.25 nM, 625 nM, 2500 nM, 10,000 nM, 20,000 nM, and 40,000 nM) for the kinase assays against CLK1, CLK2, CLK3, CLK4, DYRK1A, and DYRK2. Compound was added into the kinase buffer and incubated at 25 °C for 10 min. Then, ATP was added into the mixture and incubated at 25 °C for 60 min. Finally, ADP-Glo regent was added into the above system and incubated at 25 °C for 50 min. GloMax® Navigator Microplate Luminometer (Promega, Beijing, China) was used to read the plate. The inhibitory rates and IC50 values were calculated using GraphPad Prism 7.0. Above kinase assays were performed according to the reported study. 7
Molecular docking
The 3D structure of CLK2 was obtained from the Protein Data Bank and processed by AutoDock Tools, including the removal of water molecules and the addition of hydrogen atoms. The PDB code of CLK2 protein is 6FYK. During the molecular docking process, AutoDock was used to find the optimal binding site between MGA and CLK2. The calculation in molecular docking was performed for 100 independent docking times, and each calculation obtained a binding energy score. In order to improve the reliability of docking results, all docking studies were sorted by binding energy scores, and the best docking conformation was selected for the further analysis. PyMOL (https://pymol.org/) was used to visualize the molecular docking results, including hydrogen bonds and hydrophobic interactions.
Molecular dynamics simulation
GROMACS 5.1.4 (http://www.gromacs.org/) software was used to simulate the molecular dynamics of the CLK2-MGA complex system under the constant pressure and temperature. AMBER FF99SB force field was used to optimize the structure of CLK2. PyMOL (https://pymol.org/) was used for the post-processing and analysis of simulated trajectories. All simulations were performed on a high-performance computing cluster with a total simulation time of 100 ns. Molecular dynamic studies were performed according to the reported study. 16
Cell culture
Rat chondrocytes were purchased from Shanghai Zhongqiao Xinzhou Biotechnology Co., Ltd. (Shanghai, China). Rat BMSCs were purchased from Shanghai Yiyan Biotechnology Co., Ltd. (Shanghai, China). Chondrocytes and BMSCs were cultured in Dulbecco’s modified Eagle medium containing 10% fetal bovine serum at 37 °C and 5% CO2. Dulbecco’s modified Eagle medium and fetal bovine serum were purchased from Thermo Fisher Scientific (MA, USA).
Gene expressions by quantitative real-time polymerase chain reaction
Chondrocytes and BMSCs were seeded in a six-well plate (Servicebio, Wuhan, China). Chondrocytes and BMSCs were treated with Oncostatin M (10 ng/mL) and TNF-α (20 ng/mL) for 72 h to construct the cell model of knee osteoarthritis. Total RNA was isolated from chondrocytes and BMSCs cells by a Trizol reagent (Servicebio, Wuhan, China) and reversed by an RT Master mixture (Servicebio, Wuhan, China). A Nano Spectrophotometer (Thermo Fisher Scientific, MA, USA) was used to analyze the purity and concentration of RNA. Gene expression in cDNAs was tested using a Q2000C quantitative real-time polymerase chain reaction (qPCR) system (Shanghai Mantong photoelectric Technology Co., Ltd., Shanghai, China). The reverse of rat ADAMTS5 is 5’-CTCGATACTTGCATGACTGT-3’, and the forward of rat ADAMTS5 is 5’-TCCTCTT-GGTGGCTGACTCT-3’. The reverse of rat MMP3 is 5’-GAGCAGCAACCAGGAATAGG-3,’ and the forward of rat MMP3 is 5’-TGGACCAGGGATTAATGGAG-3’. The reverse of rat MMP13 is 5’-AAGGCCTTCTCCACTTCAGA-3’, and the forward of rat MMP13 is 5’-CCCTCGAACACTCAAATGGT-3’. The reverse of rat IL-6 is 5’-AGAAGACCAGAGCAGATT-3,’ and the forward of rat IL-6 is 5’-TGTATGAACAGCGATGATG-3’. The reverse of rat TNF-α is 5’-CGCCACGAGCAGGAATGAGAAG-3,’ and the forward of rat TNF-α is 5’-AAAGGACACCATGAGCACGGAAAG-3’. The reverse of rat IL-β is 5’-AAAGAAGGTGCTTGGGTCCT-3,’ and the forward of rat IL-β is 5’-CAGGAAGGCAGTGTCACTCA-3.’
Statistical analysis
All experiments were repeated at least three times. GraphPad Prism 7.0 was used to analyze the statistical data (**P < 0.01;***P < 0.001).
Footnotes
Ethical considerations
This article does not contain any studies with human participants or animals performed by any of the authors.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Beijing Changping Hospital of Integrated Chinese and Western Medicine. This work was also supported by the Beijing University of Chinese Medicine Third Affiliated Hospital.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
The data are available from the corresponding author on reasonable request.
