Abstract
Anti-gastric cancer activity of 18 synthesized compounds was studied using theoretical and computational method. The studied compounds were docked against mesenchymal–epithelial transition factor (PDB ID:
Keywords

Introduction
Cancer, a malignant neoplasia is mostly originated as a narrowed infection, which over time spread all over the human body.1,2 Many ailments in human body are signs for the existence of cancer such as lengthened cough, weight loss, rare blood loss, and bumps. 3 More so, gastric cancer, a type of cancer is a nuisance which ensues in human gastrointestinal system. 4 This type of cancer has been rated to be fifth among many other diseases that cause death globally.5,6 More so, men are much more affected than women.7,8
Moreover, mesenchymal–epithelial transition factor is one of the major and important measures for metastasis of several carcinomas. 9 According to Christine et al., 10 mesenchymal–epithelial transition factor plays a serious role at the concluding phases of carcinoma development, and several types of cancer, such as breast cancer, gastric cancer, lung cancer, and melanoma, have revealed its upturned manifestation relative to average tissue.11–14
The part taken by heterocycles in finding and development of innovative drugs is vital. Triazole and pyrimidine are two major compounds found in 1,2,3-triazole-pyrimidine-urea, and it has been discovered by several researcher to have anti-inflammatory, anticancer, antimicrobial, diuretics, analgesics, and antioxidant properties.15–18
Thus, this work is aimed at calculating parameters with anti-gastric cancer properties as well as investigating non-bonding interactions (binding affinity, hydrophilic interactions, and hydrophobic interactions) between 1,2,3-triazole-pyrimidine hybrids derivatives (1,2,3-TPHDs) (Figure 1) and mesenchymal–epithelial transition factor (PDB ID:
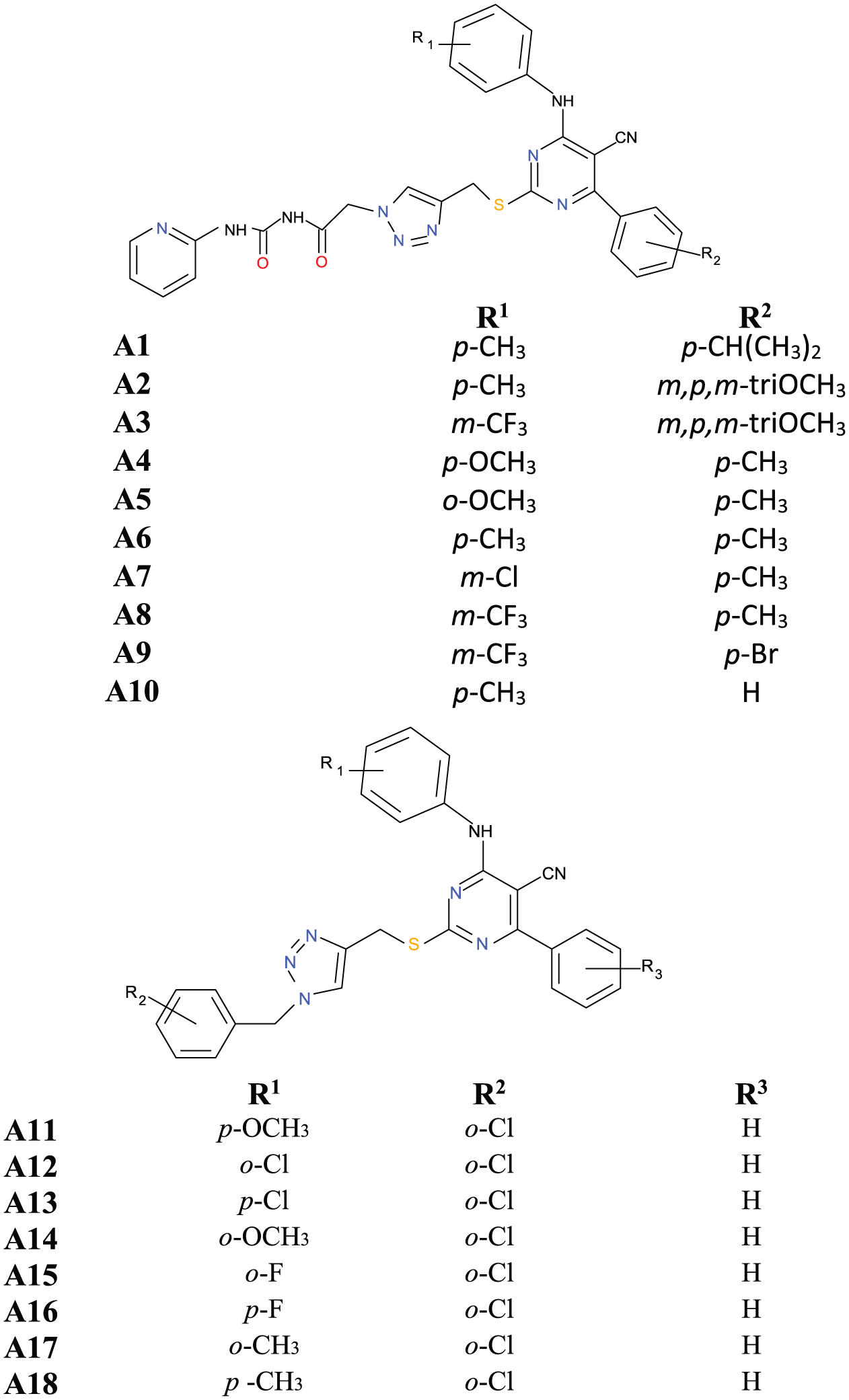
Methodology
Biological activities of 1,2,3-triazole-pyrimidine-urea hybrids derivatives was investigated using Spartan 14 software
22
for optimization and Discovery Studio client [ version v17.2.0.16349], AutoDock Tool [version 1.5.6 sep_17_14], and autodock_vina_1_1_2_win32 for molecular docking studies. The studied compounds were treated using Discovery Studio. In addition, AutoDock Tool helped in locating the active site of mesenchymal–epithelial transition factor (PDB ID:
Result and discussion
In this work, docking study was carried out on 18 molecular compounds in order to investigate the interactions between 1,2,3-TPHDs and mesenchymal–epithelial transition factor (PDB ID
Interactions between ligands and receptor (
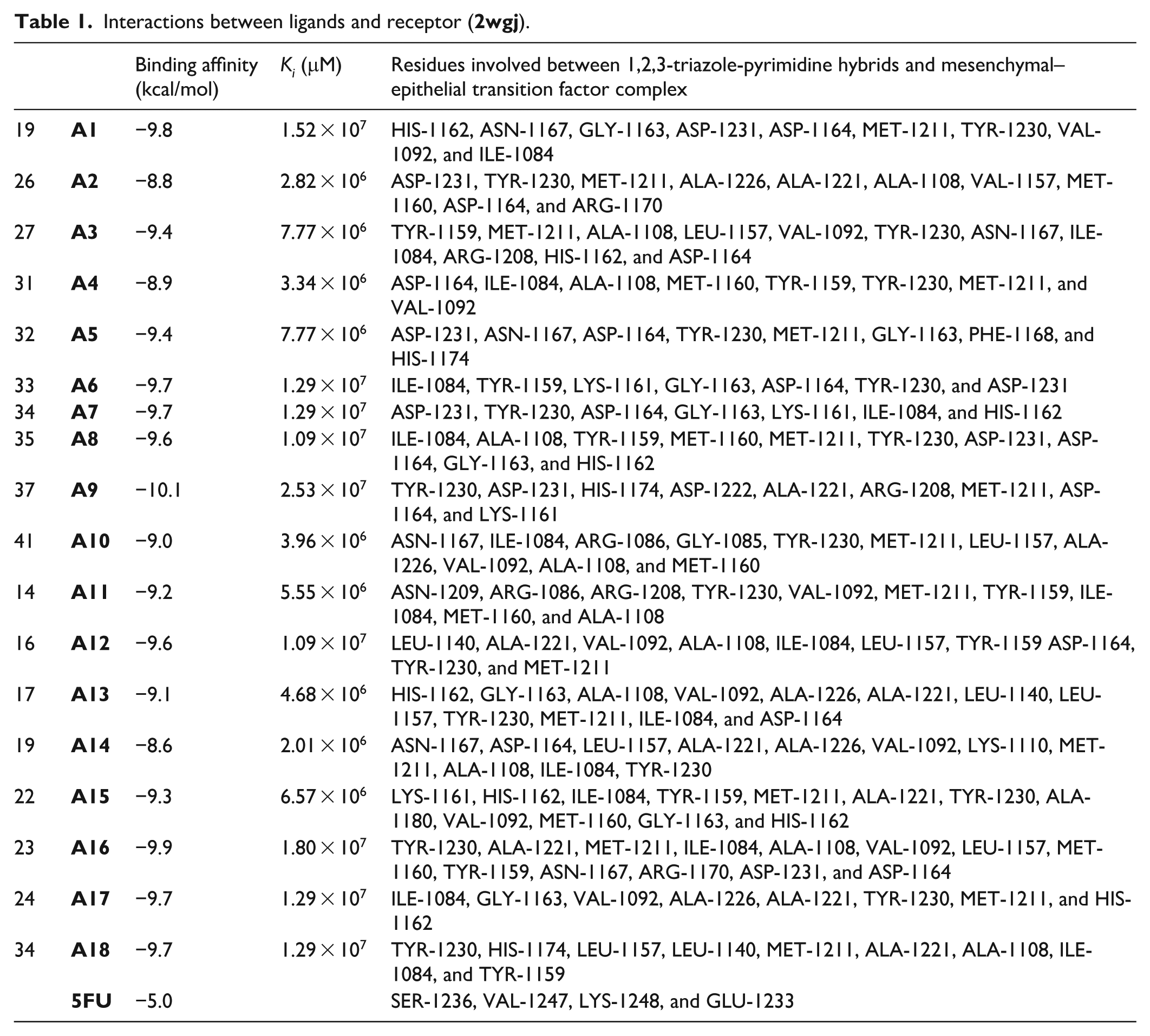
The residues involved between studied compounds and

Molecular binding interaction between
Conclusion
Bioactivity of 18 sets of 1,2,3-TPHDs were studied by optimization using density functional theory method via 6-31+G(d, p) as basis set. Also, the docking study was observed to reveal the binding interaction between the studied compounds and the receptor. Also, the studied compounds were compared to the standard used (
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.

