Abstract
The interactions between radicicol and four proteins (catalase, trypsin, pepsin, and human serum protein) are investigated by spectroscopic techniques and molecular docking. A static quenching process is confirmed. The binding constant value between radicicol and human serum protein is the largest among the four proteins. Results reveal changes in the micro-environment of the protein by the addition of radicicol. It is found that radicicol shows an inhibitory effect on the activity of proteins (catalase, trypsin, and pepsin). Molecular docking results are consistent with the thermodynamic experimental results. This work provides clues to the elucidation of the mechanisms of the interactions between radicicol and proteins.
The characteristics of the binding between radicicol and four proteins are investigated.

Introduction
Radicicol is a natural antifungal macrocyclic compound obtained from the metabolites of cucumber anthracnose using macroporous resin extraction and silica gel column chromatography separation technology; 1 its structure is shown in Figure 1. Early research found that radicicol is a highly effective and specific small molecule heat shock protein 90 (Hsp90) inhibitor.2–4 Radicicol can change the conformation of Hsp, thereby inhibiting the specific binding of Hsp90 to the target protein. 5 Radicicol, as a recognized Hsp90 functional inhibitor, has been widely used in biological research and has been used in clinical trials as an antitumor drug. 5 Rohner et al. 6 treated the offspring of Mexican dark-environment cavefish and surface waters with radicicol and detected that the mRNA levels of two Hsp90 inhibitory marker genes, BCL2-associated athanogene-3 (BAG3) and heat shock protein beta-1 (HSPB1), were upregulated, proving that Hsp90 function is inhibited. Radicicol has attracted wide attention from researchers and is an interesting candidate for clinical research.

The structure of radicicol.
Interactions between small molecules and proteins can affect their pharmacodynamics and pharmacokinetic properties7,8 and may also affect the structure of proteins.9,10 Therefore, studies on the interactions between small molecules and proteins at the molecular level are of great significance for understanding the role of these molecules in the human body. Commonly used methods for such studies are UV-Vis absorption spectroscopy, fluorescence spectroscopy, infrared spectroscopy, and circular dichroism (CD).10–15 In recent years, mass spectrometry, 16 nuclear magnetic resonance, 17 isothermal titration calorimetry, 13 and other technologies have gradually been utilized for the study of proteins. In our previous work, we studied the interactions between radicicol and the FTO protein and proved that radicicol is an effective inhibitor. 18 Human serum proteins are used as transport and storage proteins in organisms and can be used as transport intermediates for many compounds, 19 providing a basis for the study of some small molecules entering the body. Trypsin plays a very important role in the digestion and decomposition of food, and its level can be used to judge pancreatic function and pathological changes. 20 Pepsin is a type of hydrolase which functions as a digestion protein and has specificity for certain amino acid sequences. 21 Catalase is an antioxidant and a potential target for some diseases. 22 The study of the interaction between radicicol and proteins has theoretical significance for the design and development of new drugs. 23
Results and discussion
Fluorescence quenching spectra
The fluorescence spectra of the four proteins in the absence and presence of increasing radicicol concentrations are shown in Figure 2. The concentrations and volumes of radicicol used with the four proteins are identical. It can be seen from the figure that the excitation wavelength is 280 nm and that the maximum emission peaks of the four proteins are all around 340 nm. After adding the same volume of radicicol, the fluorescence of the four proteins exhibited quenching effects, and the fluorescence intensities of the proteins gradually decreased.
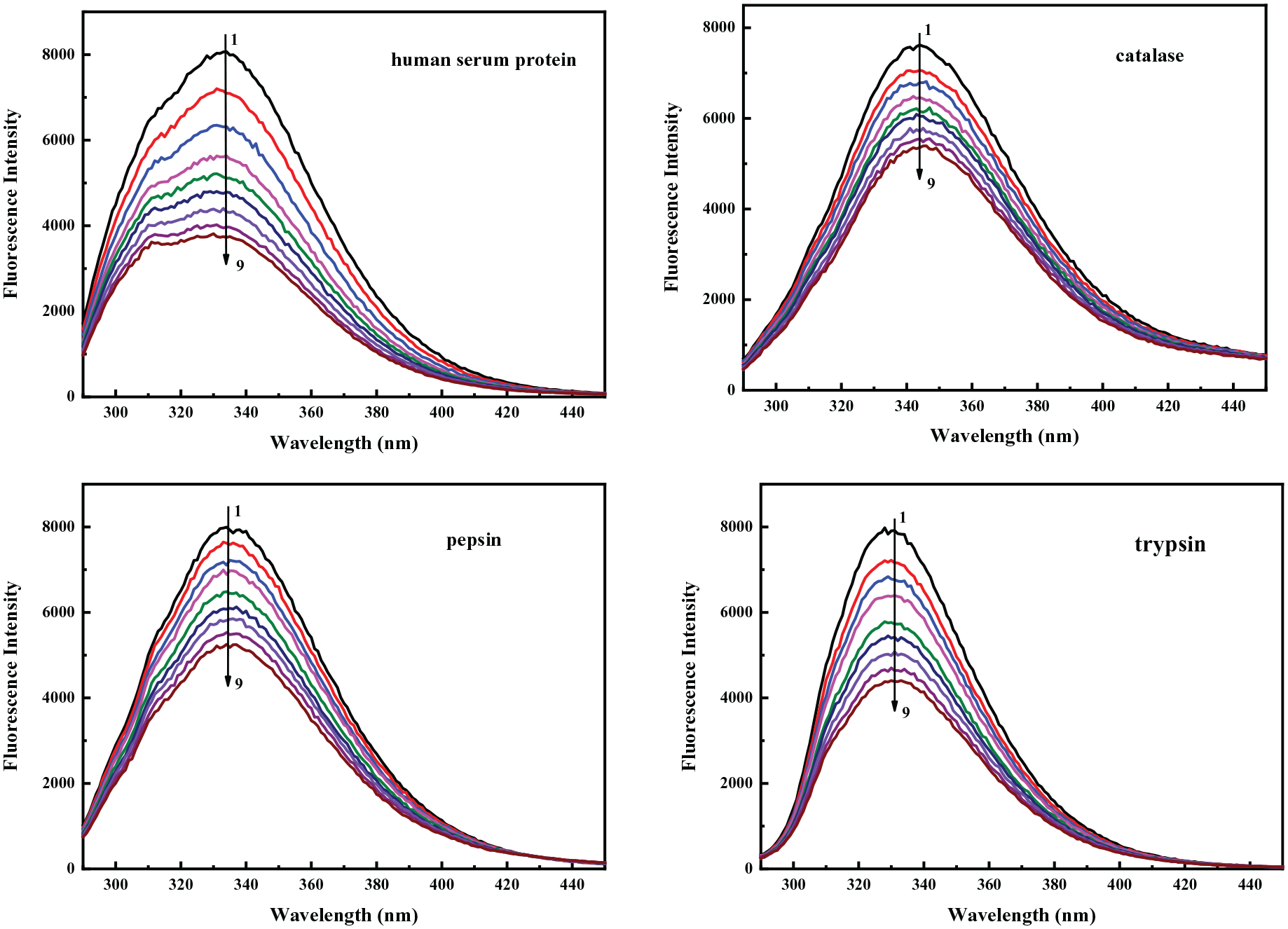
Effects of radicicol on fluorescence spectra of the four proteins (T = 301 K, pH = 7.4, λex = 280 nm, from 1 to 9 means the concentration of radicicol is 0, 1.0, 2.0, 3.0, 4.0, 5.0, 6.0, 7.0, 8.0 × 10−6 mol L−1, CHsa = 3.2 × 10−6 mol L−1, CPep = 3 × 10−7 mol L−1, CCat = 4 × 10−6 mol L−1, CTry = 3.75 × 10−6 mol L−1).
The type of fluorescence quenching is usually divided into static quenching and dynamic quenching. 24 To study the quenching types of radicicol on the four proteins, we used the Stern–Volmer equation (1) to calculate the Kq values at three different temperatures to determine the quenching mechanism 25

where F0 and F are the fluorescence intensities of the four proteins in the presence and absence of radicicol. Kq is the bimolecular quenching constant, τ0 is the lifetime of the fluorescence in the absence of quencher (τ0 = 10−8 s), 26 [Q] is the concentration of radicicol, and Ksv is the Stern–Volmer quenching constant. Therefore, the above formula can be used to obtain Ksv by linear regression of a plot of F0/F and [Q].
The F0/F-[Q] curves obtained based on the fluorescence emission spectra of the interactions between radicicol and the four proteins at three temperatures (293, 301, and 309 K) are shown in Figure 3. The quenching constants Ksv at different temperatures are evaluated and are given in Table 1. Since the fluorescence lifetime of the biopolymer is 10−8 s, the quenching rate constant Kq can be calculated based on Ksv = Kqτ0. The reported maximum scattering collision quenching constant is 2 × 1010 L mol−1 s−1. 27 The data in Table 1 show that the value of Kq is much larger than the reported maximum scattering collision quenching constant. Therefore, the rate constant of the protein quenching initiated by radicicol is greater than the value of Kq for the quenching mechanism and proves that radicicol is static for the four protein macromolecules. From Table 1, the decrease in Kq with increasing temperature also indicates a static quenching mechanism.

Stern–Volmer curves of the interactions between radicicol and the four proteins at three temperatures (293, 301, and 309 K). The concentrations of radicicol are 0, 1.0, 2.0, 3.0, 4.0, 5.0, 6.0, 7.0, 8.0 × 10−6 mol L−1.
Quenching constants (Ksv), quenching rate constants (Kq), and correlation coefficients (R2) of the interactions between radicicol and the four proteins.

Binding constants and number of binding sites
The binding constant (KA) and the number of binding sites (n) in the static quenching reaction can be obtained according to the double logarithmic equation (2)28,29
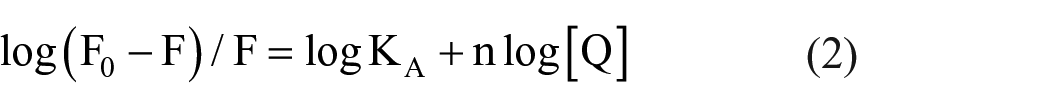
Figure 4 shows the fitting curve for lg [Q] versus lg [(F0 − F)/F] at different temperatures. The binding constant (KA) and the number of binding sites (n) can be obtained by extrapolating the slope and intercept of the curve and are listed in Table 2.

Logarithmic curve of fluorescence quenching of the interaction between radicicol and four proteins at three temperatures.
The binding constant (KA) and the number of binding sites (n) of the interaction between radicicol and the four proteins at three temperatures.

It can be seen from the plots in Figure 4 that a linear fit was obtained between lg[(F0 − F)/F] and lg[Q]. The KA data in Table 2 show that the order of the binding ability is Hsa > Pep > Cat > Try. The values of KA are in the range of 104–106 L mol−1, which are in agreement with the common affinities of drugs for proteins. 30 The number of binding sites is about 1. The data in Table 2 indicate that the binding constant values decrease with increasing temperature due to a reduction of the stability of the radicicol–protein complex and a change of the surrounding protein caused by the high temperature, which leads to a weakening of the binding ability of radicicol. It was further confirmed that radicicol quenched the fluorescence of the four proteins as static quenching.
Number of thermodynamic parameters and binding force type
To determine the type of force between radicicol and the four proteins, the thermodynamic parameters ΔHθ (enthalpy change), ΔSθ (entropy change), and ΔGθ (Gibbs free energy) can be calculated according to the van’t Hoff equation (3) and the thermodynamic equation (4) for comparison 31


The thermodynamic curves of the interaction between radicicol and the four proteins are shown in Figure 5, and the thermodynamic parameters obtained by calculation are listed in Table 3.

van’t Hoff curves of the interaction between radicicol and the four proteins at three different temperatures (293, 301, and 309 K).
Thermodynamic parameters of the interactions between radicicol and the four proteins at different temperatures.

In this study, the formation of a radicicol–protein complex was found to be spontaneous as was evident from the negative ΔG values. The negative values of ΔG and the ΔS values for the interactions of Pep (or Hsa) with radicicol indicated that hydrogen bonding and weak van der Waals interactions are the predominant forces that contribute to the Gibbs free energy change of the interactions. ΔG < 0, ΔH < 0, and ΔS > 0 indicate that the reaction between radicicol and Cat (or Try) is an entropy-driven self-heating reaction, and that the type of forces may be electrostatic and hydrophobic.
The effect of radicicol on the conformations of the four proteins
Synchronous fluorescence and three-dimensional (3D) fluorescence are commonly used to study the influence of small molecules on protein conformations. 32 It was reported that the change in the maximum emission position corresponds to the change in polarity around the chromophore molecules. 33 When the wavelength interval (Δλ) between the excitation and emission wavelengths is stable at 15 or 60 nm, the synchronous fluorescence gives the characteristic information of tyrosine residues or tryptophan residues. The synchronous fluorescence map of the four proteins and radicicol is shown in Figure 6. It can be seen that the fluorescence intensity of tryptophan residues is much greater than that of tyrosine residues. With the continuous addition of radicicol, the fluorescence intensity of the tryptophan residues and tyrosine residues is reduced to varying degrees. When the maximum emission wavelength of Cat and Pep at Δλ = 15 nm remains unchanged and the maximum emission wavelength at Δλ = 60 nm is slightly red-shifted, this is because the addition of radicicol changes the micro-environment of the tryptophan residues around the two proteins. Try and Pep remain unchanged, indicating that the micro-environment of these two amino acid residues is not changed.

Synchronous fluorescence spectra of the interactions between the four proteins and radicicol, (301 K, Δλ = 15, or 60 nm), the concentrations of the small molecules from 1 to 9 is 0, 1.0, 2.0, 3.0, 4.0, 5.0 6.0, 7.0, 8.0 × 10−6 M.
The 3D fluorescence spectra are shown in Figure 7. The intensities of Peak 1 and Peak 2 decrease significantly in the presence and absence of radicicol. The uniformity further proves that radicicol can change the micro-environments and conformations of the four proteins.

The three-dimensional fluorescence spectra of the four proteins and the three-dimensional fluorescence spectra after the addition of radicicol at 301 K.
CD measurements
CD can be used to measure the secondary structure of proteins in the far ultraviolet region (190–260 nm). At these wavelengths, the chromophore is the peptide bond, and the signal arises when it is located in a regular, folded environment. 34 The effect of radicicol on the CD spectra of the four proteins is shown in Figure 8. Hsa and Cat have two negative peaks at 208 and 220 nm, which are typical α-helixes. Both Pep and Try have a strong negative peak, which is a typical β-sheet. 35 When radicicol binds to proteins, the negative band intensities decreased. This clearly indicated changes in the protein secondary structure, with some apparent loss of helical stability. The shapes of the CD spectra of the proteins in the absence and presence of radicicol were observed to be similar, which indicated that the structures of the proteins were predominantly α-helical and β-sheets even after binding with radicicol. The CD results are expressed in terms of the mean residue ellipticity (MRE) in degree cm2 dmol−1 according to equation (5) 36

where

The conformational changes observed from CD spectroscopic analysis are shown in Table 4. The CD spectra suggest that the secondary structures of the proteins were changed due to the addition of radicicol.
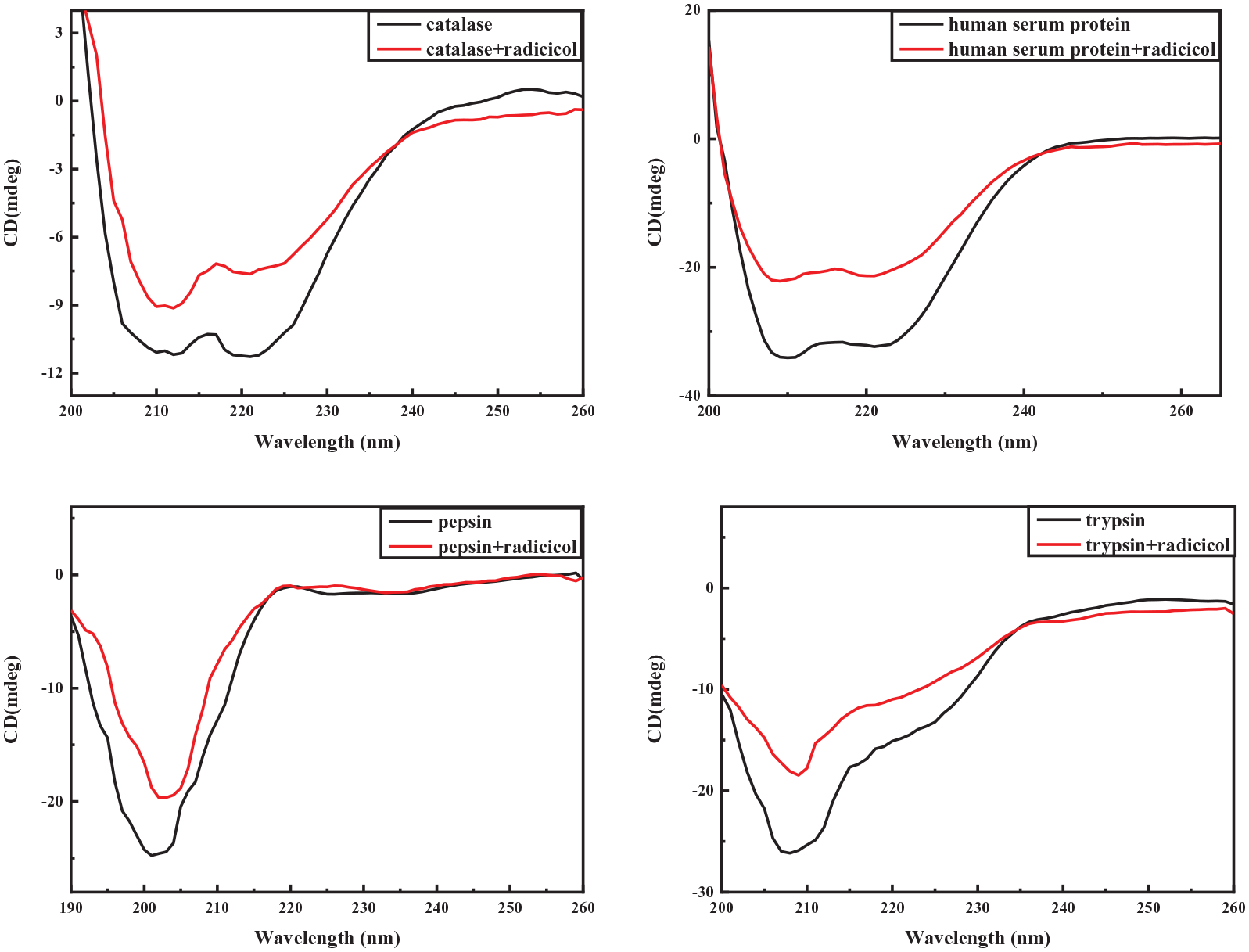
Circular dichroism chromatograms of radicicol and the four proteins (T = 25 °C, Cprotein = 40 µM, Cradicicol = 20 µM).
Secondary structures of the protein changes on addition of radicicol.

The effect of radicicol on protein activity
To reveal whether radicicol can affect the activity of proteins, the effect of radicicol on protein activity in vitro was investigated. The absorbance change was measured at 275 nm (OD275) to study the inhibition rate of pepsin activity. 37 The absorbance change was measured at 253 nm (OD253) to study the trypsin activity inhibition rate. 38 The absorbance change was measured at 240 nm (OD240) to study the inhibition rate of catalase activity. 39 The inhibition rate of protein activity can be calculated with the formula (7) 40

As showed in Figure 9, the changes in enzyme activity are induced by the addition of radicicol of different concentrations. With the rise of concentration of radicicol, the inhibition rate increases and the activity of the protein is decreased. It can be seen from the figure that there are obvious differences in the inhibitory effects of radicicol on several proteins, and the inhibitory effect on pepsin is the best. These results imply that radicicol can inhibit protein activity when it enters the human body.

Effect of radicicol on protein activity in vitro at 301 K (CPep = 20.0 μM, CCat = 0.5 mg mL−1, CTry = 10 mg mL−1).
Molecular docking
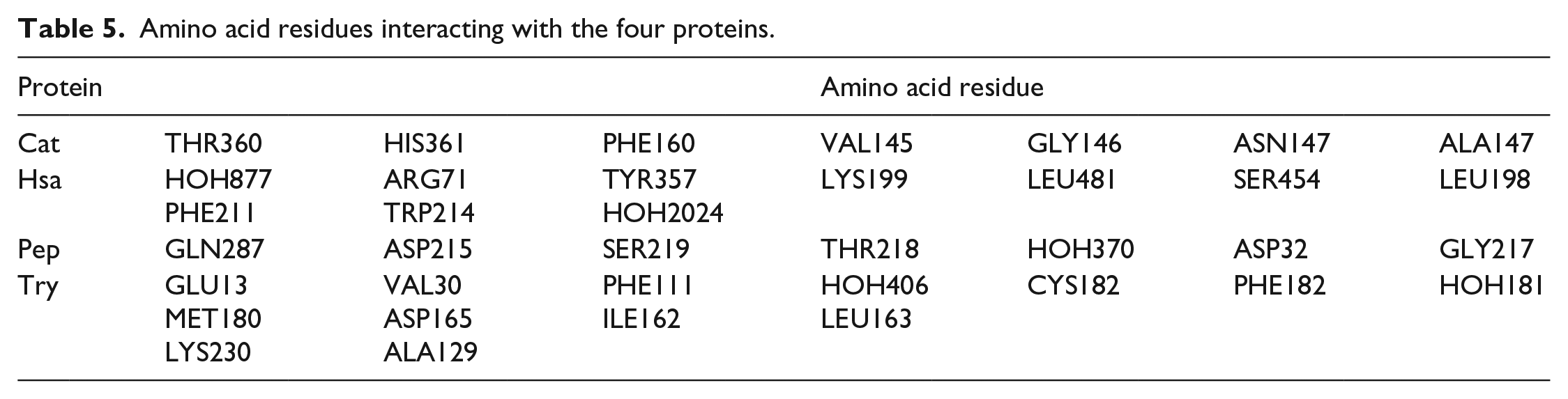
The docking results with the lowest energy obtained by molecular docking technology are shown in Figure 10. The amino acid residues that bind to the four proteins are listed in Table 5. The analysis has revealed that radicicol is embedded in the four proteins, and that the hydrophobic residues interacting with Hsa are PHE211, TRP214, LEU481, and LEU198. The hydrophobic residues that interact with Cat are PHE160, VAL145, GLY146, and ALA132. The hydrophobic residues that interact with Pep are GLY217, VAL30, and PHE111. The hydrophobic residues that interact with TRY are ILE162, LEU163, PHE181, and MET180. The existence of these hydrophobic residues proves the existence of hydrophobic forces in the interactions between radicicol and the four proteins. In addition, it can be seen from the table that there are hydrogen bonds in the residues of the four proteins that interact with radicicol, but the number of hydrogen bonds is different, which proves that hydrogen bonds also play a certain role in the interactions. This is consistent with the conclusions obtained from thermodynamic data analysis.
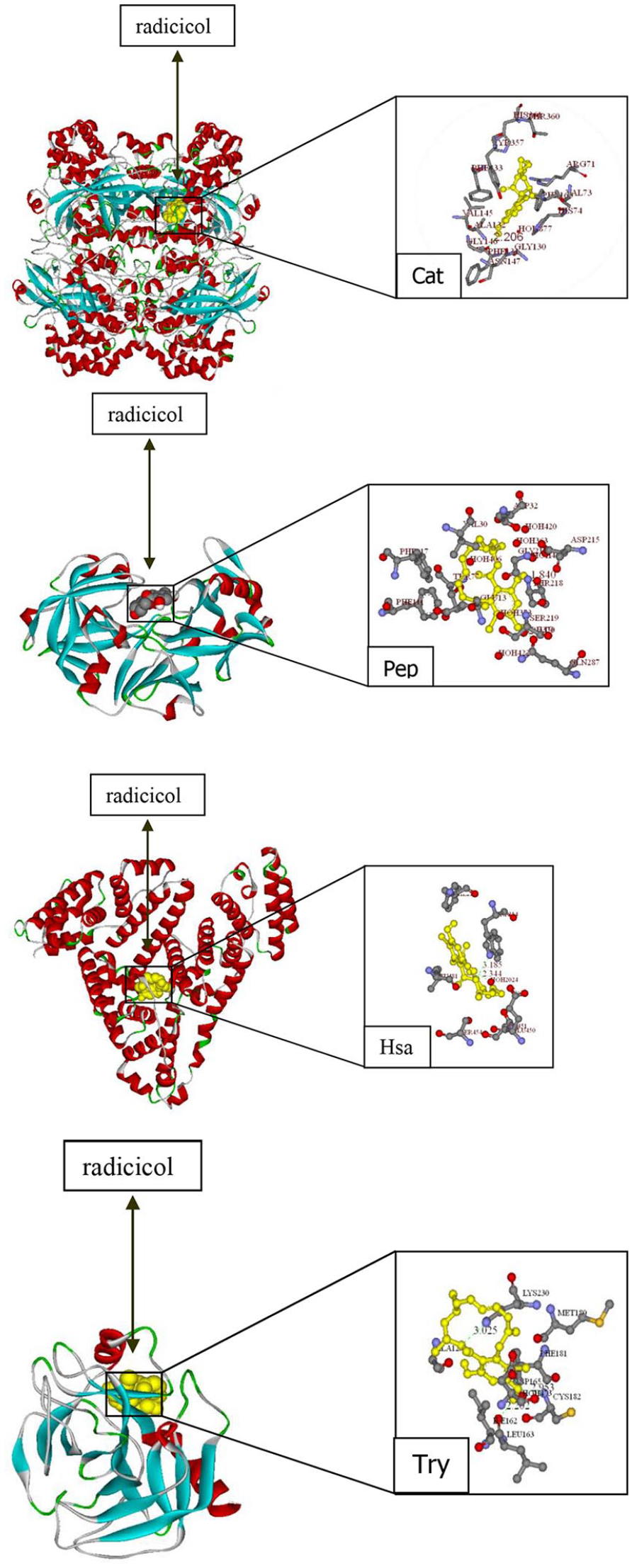
The docking results of radicicol and four proteins: the figures on the left show the ribbons model of the four proteins and the spherical models of radicicol. The figures on the right show the amino acid residues that interact with radicicol in the four proteins. The dotted lines and the numbers indicate the hydrogen bonds and bond lengths, respectively.
Amino acid residues interacting with the four proteins.
Conclusion
This work provides an approach for studying the interactions of four proteins with radicicol using different spectroscopic techniques under physiological conditions. The decreasing values of Kq with increase in temperature indicate the presence of a static quenching mechanism. Calculating the binding constant and the number of reactive binding sites at three temperatures shows that Hsa has the strongest binding capacity of the four proteins. According to the relevant laws of thermodynamics, the reaction interactions of radicicol and the four proteins can proceed spontaneously. The results show that the addition of radicicol can change the conformation of the four proteins. The CD spectra revealed that the presence of radicicol decreased the α-helix content of Hsa and Cat, meanwhile the presence of radicicol decreased the β-sheet content of Pep and Try. The spectrophotometric method directly proved that radicicol has an inhibitory effect on the proteins Pep, Cat, and Try. The results of molecular docking support the thermodynamic data results. The different numbers of residues in the docking results of the four proteins with radicicol also proved the different binding abilities.
Experimental
Materials and instruments
The four proteins and radicicol used in this experiment were purchased from Aladdin Reagent Company. Pepsin and catalase were stored in a refrigerator at 4 °C, Try and human serum protein were stored in a refrigerator at −20 °C. Human serum protein and catalase were dissolved in Tris-HCl buffer solution to prepare a stock solution, and gastric protein was dissolved in pH = 2 HCl-CH3COONa buffer solution. Trypsin was dissolved in PBS buffer to make a stock solution. Radicicol was dissolved in dimethyl sulfoxide (DMSO) to prepare a stock solution. Double distilled water was used, and the reagents used were all analytically pure. The stock solutions were all centrifuged and degassed after preparation. The protein stock solution was stored in a refrigerator at −80 °C, and the radicicol stock solution was stored in a refrigerator at −20 °C and then diluted according to the different experimental requirements.
An Agilent 8453 UV-Vis spectrophotometer (Agilent), a F-4600 fluorescence spectrophotometer (Japan Hitachi), a DZKW-4 constant temperature water bath (Beijing Zhongxing Weiye), a QL-866 vortex oscillator (Kirin, Haimen City medical instruments), an AUW120D analytical balance (SHIMADZU), and a Centrifuge 5424 small high-speed centrifuge (Ebende, Germany) were used.
Fluorescence spectra
The fluorescence spectra of the four proteins, both in the absence and presence of radicicol, were recorded over a wavelength range of 290–450 nm on excitation at 280 nm with 5 nm excitation and emission monochromator slit widths. Fluorescence quenching experiments were carried out at three different temperatures (293, 301, and 309 K) to evaluate the effect of the temperature on the interactions. 10
Synchronous fluorescence spectroscopy
Synchronization was carried out at room temperature with Δλ = 15 or 60 nm. The scanning range is 240–340 nm; scanning speed is 1200 nm min−1. 10
3D fluorescence spectroscopy
The excitation wavelength and emission wavelength ranges were set to 210–290 nm and 310–380 nm, respectively; the scanning speed is 1200 nm min−1.
CD measurements
CD measurements were recorded in the far ultraviolet region at room temperature on a CD250 spectrometer. A 1.0-cm quartz cell was used. The molar ratio of protein and radicicol was kept constant, and the spectra of the four proteins were measured in the presence and absence of radicicol. The buffered spectrum is subtracted during scanning, and each sample is measured three times on average. The CD of Cat, has, and Try were measured at 200–260 nm, and the CD of Pep was measured at 190–260 nm.
Protein activity experiments
Radicicol was prepared as 2 mM stock solutions with DMSO. Cat was made up as a 0.5 mg mL−1 mother liquor with pH = 7 PBS solution, and hydrogen peroxide was used as the substrate. 39 Try was prepared as a 10 mg mL−1 stock solution, N-benzoyl-L-arginine ethyl ester (BAEE) was used as the substrate. 38 Pep is made into 20 µM stock solution with pH = 2 HCL-CH3COONa buffer, with hemoglobin as a substrate. 37 UV-Vis absorption spectroscopy was used to detect the absorbance changes of the three substrates and to study the changes in protein activity. 40
Molecular docking experiments
The radicicol structure was generated using ChemDraw, and the molecular docking simulation was performed with the software Autodock4.2, and the Lamarck genetic algorithm was applied to the docking process. Gaussian 09 was used for optimization. The density functional B3LYP was used to perform geometric optimization on the 6-31G (d,p) basis sets of C, H, O, N, and F atoms. The crystal structures of the four proteins are obtained from the Brook Haven Protein Data Bank (http://www.rcsb.org/pdb). The PDBID of human serum protein is 1H9Z, the PDBID of Cat is 4BLC, the PDBID of Pep is 5Pep, and the PDBID of Try is 2ZQ1. After the docking was completed, the optimal combination was found from 250 conformations.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was financially supported by the Natural Science Foundation of Henan Province (202300410478).
