Abstract
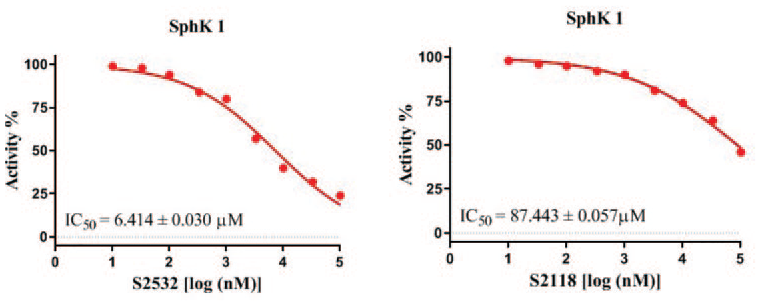
Sphingosine kinases (SphKs) are a class of lipid kinases, that have received extensive attention as important rate-limiting enzyme in tumor. Inhibition of the activity of SphK1 can lead to an anticancer effect. Herein, we describe the discovery process and biological characteristics of a new SphK1 inhibitor, ascorbyl palmitate, discovered through computer-aided drug design. Biochemical experiments show that ascorbyl palmitate has a strong inhibitory effect on SphK1, with an IC50 value of 6.4 μM. The MTT experiment showed that ascorbyl palmitate had anti-cancer effects toward the U87, A549, 22RV1, and A375 cell lines. Among them, ascorbyl palmitate has prominent inhibitory activity against the 22RV1 cell line, with an IC50 value of 41.57 μM. To explore the structure–activity relationship, four ascorbyl palmitate derivatives were synthesized and tested for kinase activity. The outstanding effect of ascorbyl palmitate toward SphK1 and its known non-toxicity suggest that ascorbyl palmitate may be a lead compound for the development of effective SphK1 anti-cancer inhibitors.

Introduction
Sphingosine kinases (SphKs), as lipid kinases, have two different subtypes (SphK1/SphK2). They are closely related to the occurrence and development of cancers. 1 They participate in the progression of cancers in two different ways: one is the role of SphKs as an oncogene, 2 and the other is the role of their metabolite sphingosine-1-phosphate (S1P). 3 They are involved in the regulation of a variety of signaling pathways through the production of S1P, which mediates a variety of biological functions such as calcium mobilization, mitogenesis, apoptosis, cell motility, and angiogenesis. Reducing the production of S1P by inhibiting SphKs can inhibit the occurrence and development of some diseases. 4 As far as SphK1 is concerned, it has been shown to be highly expressed in a variety of cancer cells such as ovarian, cervical, colon, stomach, lung, and brain cancer.5,6 Overexpressed SphK1 can not only stimulate cell growth but also cause malignant transformations of normal cells. 7 In contrast, SphK2 is much more complicated, and its effect on cells mainly depends on the degree of overexpression of SphK2. 8 Also, due to the uncertainty of the structure and function of SphK2, SphK1 inhibitors are still more frequently used. 9 In recent years, SphKs inhibitors have been studied; some of which have entered clinical trials. 10 For example, ABC294640 has an inhibitory effect on the growth of prostate cancer and colorectal cancer, is also effective for other solid tumors, and has completed a phase I clinical trial (NCT01488513). 11 There are some excellent SphK1 inhibitors, such as PF-543, Com23, Com83, SKI-II, Com54, SK1-I, and SKI-178 (Figure 1).12,13

Examples of SphK1 inhibitors.
Computer-aided drug design has been recognized as an important tool for the discovery of drugs. 14 Structure-based virtual screening is a common technique used by most pharmaceutical companies as well as certain academic groups in the early stages of drug development. Analysis of various SphK1 protein crystal structures also provides convenience for the virtual screening of SphK1 inhibitors. The crystal structure of the SphK1 protein was first reported in 2013. 15 It was proposed that there is a polar head and a hydrophobic tail in the SphK1 protein cavity, and the key amino acids Asp81, Leu268, and Asp178 play a key role in the polar part. Therefore, in this study, we used the SphK1 inhibitors mentioned above (Figure 1) to construct some pharmacophore models, and the pharmacophore models are then evaluated. The compound library was initially screened using the optimal pharmacophore model, and then the selected compounds were molecularly docked with SphK1 (3vzb). Finally, in order to obtain potential SphK1 inhibitors, the compounds with the best docking scores were tested for SphK1 kinase activity. Figure 2 shows a flowchart of the virtual screening.

The flowchart of the virtual screening.
Results and discussion
Establishment and evaluation of the pharmacophore model
In this study, all operations related to computer-aided drug design were performed with BIOVIA Discovery Studio 2020 (DS2020). First is the construction and validation of pharmacophore models. Based on the principle of the diversity of active molecular structures of a training set, seven SphK1 antagonists with high activity reported in the literature were selected to construct the pharmacophore models. The training set is shown in Figure 1. The construction and evaluation of the pharmacophore model refer to earlier reports.16,17 The Supporting Information provides the detailed process of the pharmacophore model construction. In this study, the hydrophobic (H) characteristics, hydrogen bond acceptor (HBA), hydrogen bond donor (HBD), and aromatic ring center (R) were selected as the characteristic pharmacophore elements. Using the Common Feature Pharmacophore Generation module of the DS2020 software, 10 models were constructed based on the active conformations of the training set molecules. Figure 3(a) shows the scoring situation of 10 models. From the scoring ranking and pharmacodynamic feature elements of Figure 3(a), we find that models 1–3 have more pharmacophore characteristics and higher rankings. In order to further evaluate the effect of these models, 15 active compounds and 15 inactive compounds (Supplemental Figure S1) were used as a testing set to evaluate these 10 models again. As shown in the heat map Figure 3(b), the pharmacophore model 3 had a high matching value with the 15 active compounds, and a low matching value with the 15 inactive compounds. 18 Figure 3(c) shows the composite diagram between model 3 and the SphK1 inhibitor PF-543. It can be found from Figure 3(c) that the polar head of PF-543 has HBA characteristics. The rest of the pharmacophore features are hydrophobic characteristics. This model coincides with the “J-shaped” hydrophobic cavity of SphK1. Therefore, model 3 was selected for preliminary screening of the compound library.

(a) The scores of 10 models (H stands for hydrophobic characteristics, R stands for aromatic ring center, D stands for hydrogen bond donor, and A stands for hydrogen bond acceptor). (b) The heat map of 10 models (the red square represents the higher the match between the compounds and the models). (c) The composite diagram between model 3 and PF-543 (the green ball represents the hydrogen bond acceptor (HBA), and the blue ball represents the hydrophobic characteristic).
Virtual screening of databases
Based on the results of the pharmacophore evaluation, we chose model 3 as the optimum model to screen the Selleck database. Next, the Selleck database containing 99,040 compounds was filtered to rule out compounds that do not maintain drug-likeness and the remaining 51,020 compounds were optimized by the Prepare Ligand Procedure module in the DS2020 software. And these 51,020 compounds were virtually screened by the optimal pharmacophore model 3. According to the ranking of fit values, the top 4324 compounds were selected for molecular docking. The 100 compounds with the highest docking scores were then analyzed for interactions with SphK1. According to the interactions of the compounds with the key amino acid residues Asp81 and Asp178, 12 compounds were selected as potential SphK1 inhibitors. Figure 4 shows the structures of these 12 compounds.
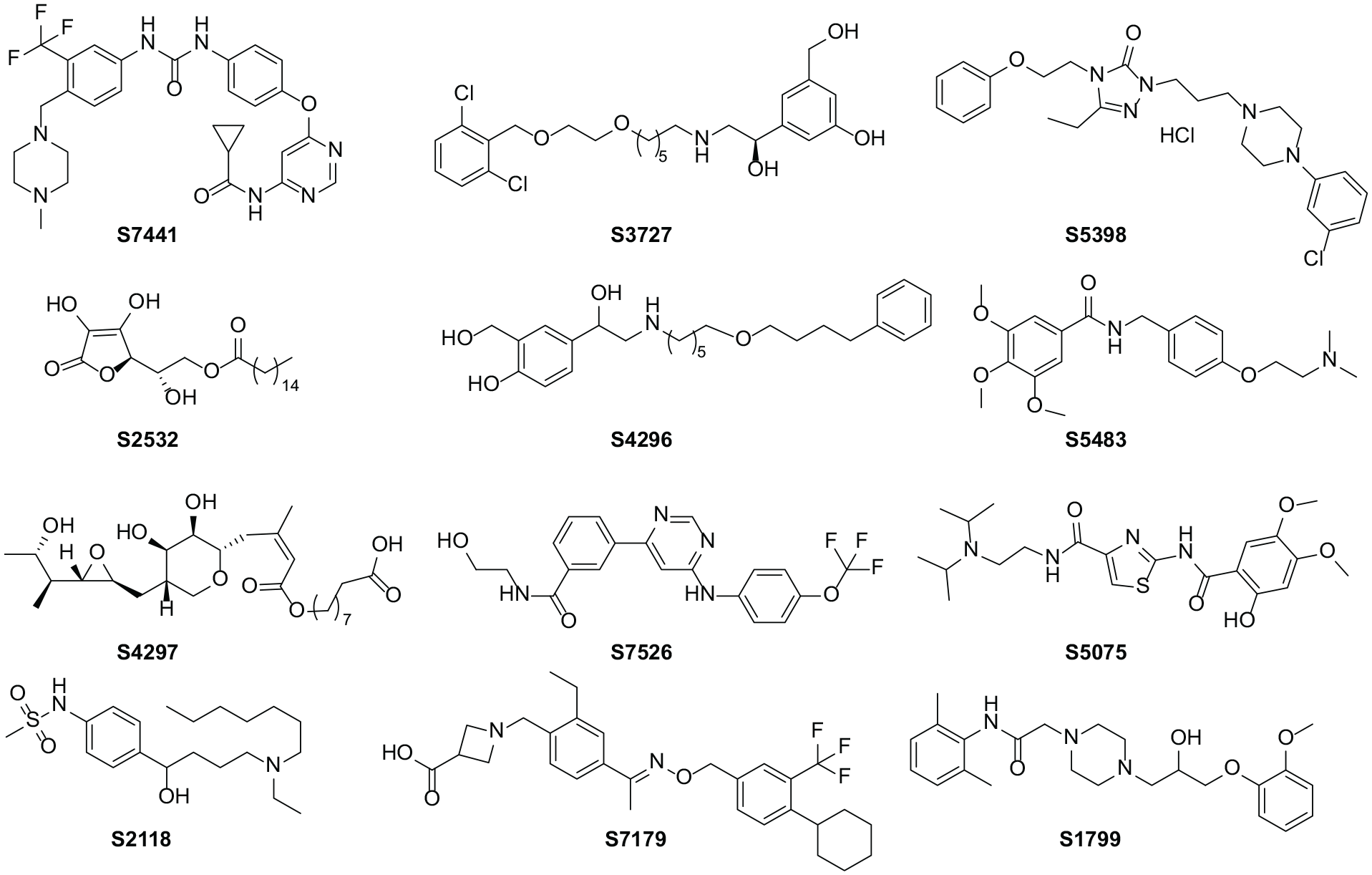
The structures of the 12 compounds selected for study of the inhibition of SphK1 (the number of the compound corresponds to its catalog number in the Selleck database).
Kinase inhibitory activities
These 12 compounds were tested for their inhibition rate against SphK1 at a concentration of 10 μM. Their inhibition rates on SphK1 at a concentration of 10 μM are shown in Table 1. From Table 1, it is seen that at a concentration of 10 μM, most of the compounds had no inhibitory effect on SphK1. However, the two compounds numbered
The inhibition rates of the 12 selected compounds against SphK1 at 10 μM.

NT: no activity detected.
Values are percent inhibitions of SphK1 at 10 μM, averages of two separate experiments, standard deviations were ±5%.
The SphKs assays were carried out as described previously.
19
From Table 1, it can be seen that the compounds numbered
The inhibitory effects of compounds numbered

Cytotoxic activity
From the structure of compound numbered
Anti-proliferative effects of
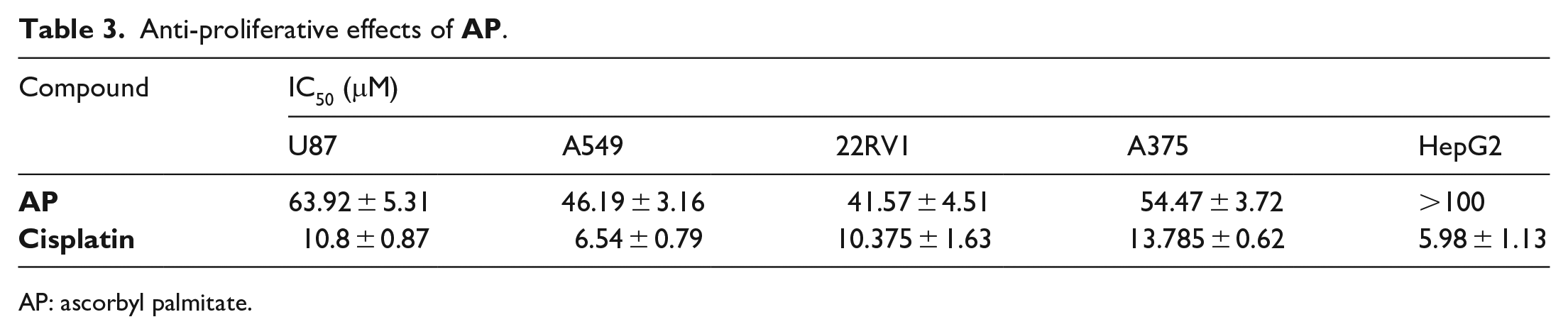

AP: ascorbyl palmitate.
Structure derivatization of AP
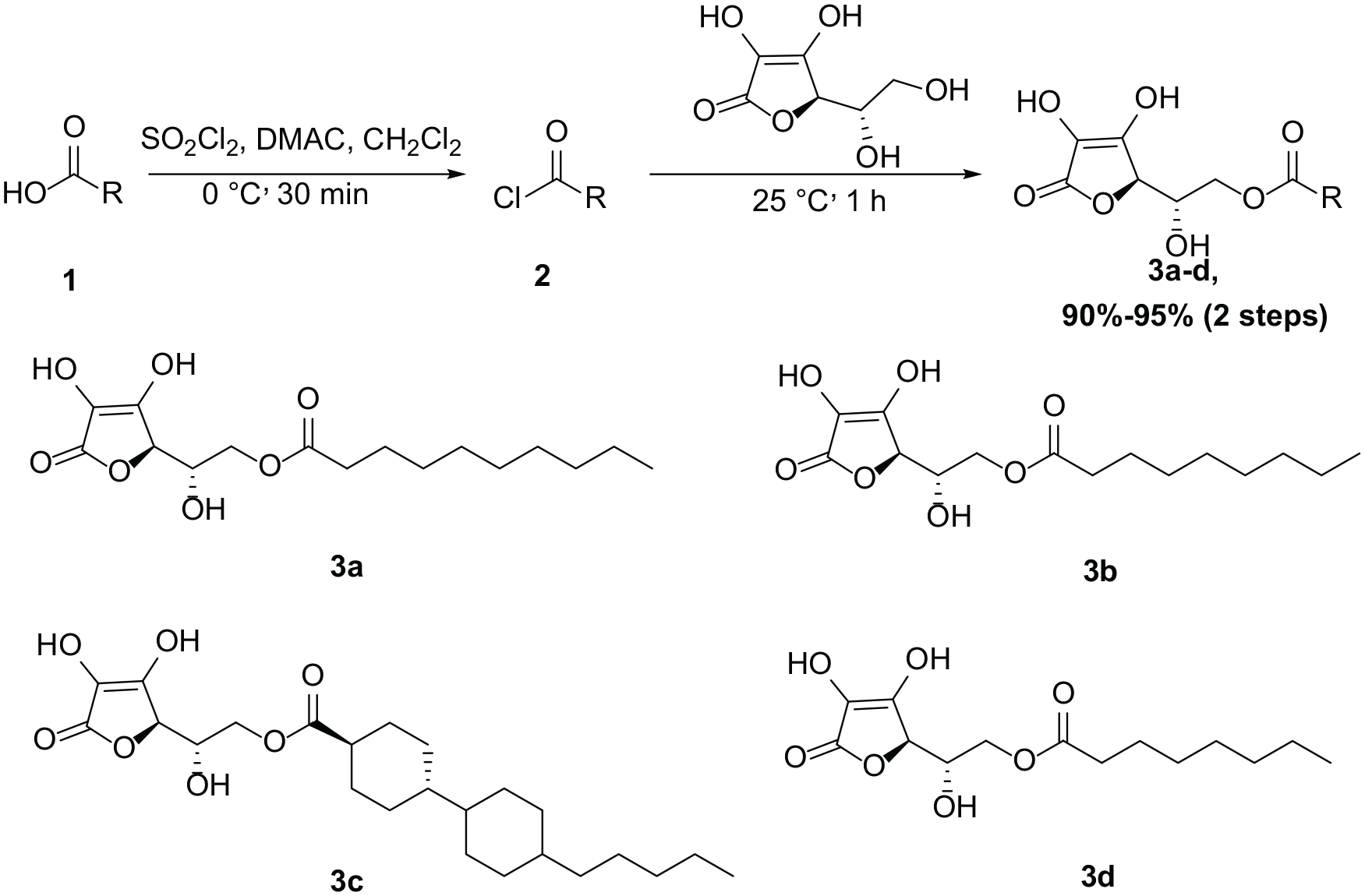
Synthetic route toward
Kinase activity of the AP derivatives 3a–d
In order to further study the structure–activity relationship between the
Inhibition studies of

VC: vitamin C; NT: no activity detected.
Values are percent inhibitions against SphK1 at 10 μM, averages of two separate experiments, standard deviations were ±5%.
Docking of AP and SphK1
We performed molecular docking analysis on the crystal structure of
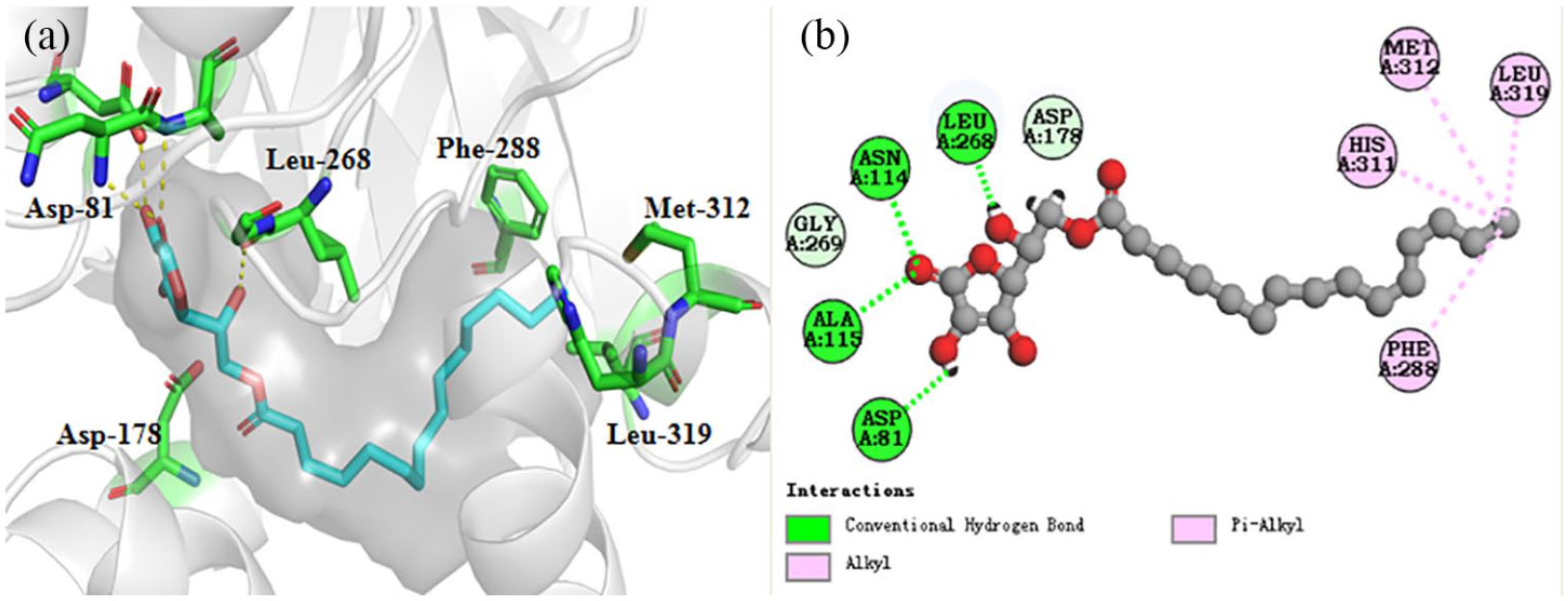
The schematic diagram of the three-dimensional (a) and two-dimensional (b) interactions between
Conclusion
In summary, we have reported the discovery of new SphK1 inhibitors by a computer-aided virtual screening approach and biological validations. It was determined by in vitro enzymatic activity that compounds numbered
Experimental
Materials
Vitamin C, thionyl chloride, N,N-dimethy-lacetamide, dichloromethane, sodium bicarbonate, fetal bovine serum, and other materials were obtained from commercial sources and were used without further purification.
Apparatus
High-resolution mass spectrometry (HRMS) was performed on an AB SCIEX X500R Accurate Mass Q-TOF using electrospray ionization (ESI). The NMR spectra were recorded on a Bruker AM-600 spectrometer (Billerica, MA, USA) with tetramethylsilane as the internal standard.
Synthesis
The synthesis of
(S)-2-((R)-3,4-dihydroxy-5-oxo-2,5-dihydrofuran-2-yl)-2-hydroxyethyl decanoate (3a )
White solid; 95% yield. 1H NMR (600 MHz, CD3OD): δ 4.73 (d, J = 2.01 Hz, 1H), 4.19-4.28 (m, 2H), 4.07-4.10 (m, 1H), 2.37 (t, J = 7.41 Hz, 2H), 1.57-1.67 (m, 2H), 1.24-1.38 (m, 12H), 0.89 (t, J = 6.46 Hz, 3H). 13C NMR (150 MHz, CD3OD): δ 173.76, 171.74, 152.69, 118.68, 75.81, 66.69, 64.41, 33.48, 31.62, 29.15, 28.99, 28.98, 28.77, 24.57, 22.30, 13.01. HRMS (ESI): m/z [M + H]+ calcd for C16H27O7: 331.1835; found: 331.1753.
(S)-2-((R)-3,4-dihydroxy-5-oxo-2,5-dihydrofuran-2-yl)-2-hydroxyethyl nonanoate (3b )
White solid; 90% yield. 1H NMR (600 MHz, CD3OD): δ 4.73 (d, J = 2.03 Hz, 1H), 4.16-4.27 (m, 2H), 4.06-4.12 (m, 1H), 2.37 (t, J = 7.4 Hz, 2H), 1.59-1.68 (m, 2H), 1.25-1.39 (m, 10H), 0.90 (t, J = 6.43 Hz, 3H). 13C NMR (150 MHz, CD3OD): δ 174.02, 171.73, 152.65, 118.69, 75.80, 66.68, 64.43, 37.08, 32.72 (2C), 32.05, 31.10, 26.24, 25.94 (2C). HRMS (ESI): m/z [M + H]+ calcd for C15H25O7: 317.1522; found: 317.1548.
(S)-2-((R)-3,4-dihydroxy-5-oxo-2,5-dihydrofuran-2-yl)-2-hydroxyethyl (1S,4r)-4’-pentyl-[1,1’-bi(cyclohexane)]-4-carboxylate (3c )
White solid; 94% yield. 1H NMR (600 MHz, CD3OD): δ 4.73 (d, J = 2.09 Hz, 1H), 4.24-4.27 (m, 1H), 4.17-4.21 (m, 1H), 4.07-4.10 (m, 1H), 3.67 (t, J = 4.54 Hz, 3H), 3.56 (t, J = 4.95 Hz, 3H), 2.37 (t, J = 7.43 Hz, 2H), 1.60-1.65 (m, 2H), 1.24-1.39 (m, 18H), 0.89 (t, J = 6.99 Hz, 3H). 13C NMR (150 MHz, CD3OD): δ 173.75, 171.73, 152.57, 118.69, 75.79, 72.12 (2C), 66.66, 64.40, 60.86 (2C), 33.47, 31.67, 29.33 (3C), 29.20, 29.06 (2C), 29.00, 28.78, 24.58, 22.33, 13.04. HRMS (ESI): m/z [M + H]+ calcd for C24H39O7: 438.2618; found: 439.2686.
(S)-2-((R)-3,4-dihydroxy-5-oxo-2,5-dihydrofuran-2-yl)-2-hydroxyethyl octanoate (3d )
White solid; 92% yield. 1H NMR (600 MHz, CD3OD): δ 4.74 (d, J = 2.02 Hz, 1H), 4.17-4.28 (m, 2H), 4.07-4.11 (m, 1H), 2.37 (t, J = 7.43 Hz, 2H), 1.57-1.67 (m, 2H), 1.24-1.39 (m, 8H), 0.90 (t, J = 6.42 Hz, 3H). 13C NMR (150 MHz, CD3OD): δ 173.85, 171.76, 152.61, 118.72, 75.83, 66.74, 64.42, 33.57, 31.46, 28.78, 28.70, 24.61, 22.29, 13.13. HRMS (ESI): m/z [M + H]+ calcd for C14H23O7: 303.1366; found: 303.1433. The 1H and 13C NMR spectra for compounds
The kinase test in vitro
The SphKs assays were carried out as described previously.
19
The assay was performed using Kinase-Glo Plus luminescence kinase assay kit (purchased from Promega, Fitchburg, USA). It measures kinase activity by quantitating the amount of ATP remaining in solution following a kinase reaction. Here, we used PF-543 (purchased from Selleckchem, Houston, TX, USA) as a positive control. The reaction system was 50 μL. Compounds
Supplemental Material
sj-pdf-1-chl-10.1177_17475198211001819 – Supplemental material for L-ascorbyl 6-palmitate as lead compound targeting SphK1: an in silico and in vitro investigation
Supplemental material, sj-pdf-1-chl-10.1177_17475198211001819 for L-ascorbyl 6-palmitate as lead compound targeting SphK1: an in silico and in vitro investigation by HaiJiao Chen, Xinmei Yang, Peng Sun, Ying Zhi, Qingqiang Yao and Bo Liu in Journal of Chemical Research
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the National Natural Science Foundation of China (grant no. 81903473) and the Academic Promotion Programme of Shandong First Medical University (no. 2019LJ003).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.