Abstract
Comparative molecular docking studies on creatine and guanidinoacetic acid, as well as their phosphorylated analogues, creatine phosphate, and phosphorylated guanidinoacetic acid, are investigated. Docking and density functional theory studies are carried out for muscle creatine kinase. The changes in the geometries of the ligands before and after binding to the enzyme are investigated to explain the better binding of guanidinoacetic acid and phosphorylated guanidinoacetic acid compared to creatine and creatine phosphate.

Introduction
Creatine kinase (CK) is a crucial enzyme of cellular bioenergetics that regulates energy fluxes by the transfer of phosphates between different phosphorylated substrates and adenine nucleotides. 1 Phosphocreatine (CrP) seems to be the primary CK substrate with the highest affinity for CK reported in both in vivo and in vitro studies.2,3 However, many other guanidino compounds can act as substrate analogues for the enzyme, which can be particularly important in cases of CrP deficiency. Kan et al. 4 reported that phosphorylated guanidinoacetic acid (GAAP) can partly compensate for the lack of CrP, with GAAP used for high-energy phosphoryl transfer through reaction with CK in the skeletal muscle of mice. However, limited data are available concerning the comparison between GAAP and CrP in terms of CK kinetics. So far, there are no studies evaluating the docking environment of CK as well as differences between specific substrates (such as creatine and guanidinoacetic acid) and their phosphorylated analogues before and after docking. Molecular docking techniques are well-documented computational tools that suggest the most acceptable mechanism for intermolecular interactions of ligands with biomolecules. 5 Therefore, molecular docking studies on the above compounds are expected to provide valuable information about the mechanism of binding to CK and are highly justified.
Results and discussion
Density functional theory analysis
A molecule of phosphocreatine (CrP) forms a seven-membered ring (Figure 1(a)) through intramolecular hydrogen bonding between the H5-atom of the iminium group and the oxygen atom from the carboxylate group (length 1.70 Å). The angles between bonds from the guanidine groups C2–N3–C4 (121.1°), N3–C4–N6 (121.1°), and C4–N6–H5 (113.8°) are close to the values obtained from X-ray analysis of guanidinium chloride 6 and guanidinoacetic acid 7 (121.5°, 120.8°, and 114°, respectively). Besides, the angle between the N3–C2–C1 bonds (109.6°) is almost tetrahedral, which indicates that the formation of H-bonds does not disturb the geometry of the CrP molecule and contributes to its stability. However, steric hindrance between the methyl group on N3 and the phosphate group restricts free rotation about the C4–N5 bond compared to the molecule of N-phosphoguanidinoacetic acid (GAAP). The values of the rotation barriers depend on the dihedral angles between the N3–C4–N5–P bonds and are presented in Figure S1. As can be seen from Figure S1, the amount of energy needed for rotation around the C4–N5 bond in CrP is extremely high, even larger than the dissociation energy of a covalent bond. This result indicates that rotation in the range of 30°–70° will cause breaking of the C4–N5 bond, causing a restriction of free rotation. The existence of a methyl group on N3 in CrP and its rotation in the stated range leads to overlapping of the Van der Waals volumes of the CH3 group and the NH2 group, as shown in Figure S2. The existence of the methyl group and steric hindrance along with overlapping of the Van der Waals volumes leads to a higher amount of energy being required for rotation in comparison with GAAP in which there is no methyl group in this position.

Optimized structures of molecules of (a) CrP and (b) GAAP.
The molecule of GAAP forms six- and seven-membered rings through two intramolecular H-bonds (Figure 1(b)). The seven-membered ring is similar that of the CrP, but the H-bond in GAAP is shorter (1.66 Å) compared to that of CrP, and has large deviations of the bond angles from the preferred tetrahedral geometry. The second H-bond (length 1.58 Å), between H3 and O3 from the phosphate group, leads to the formation of a six-membered ring (–O3–H3–N3–C4–N5–P–), which results in the almost planar geometry of this part of the molecule. From Table 1, it can be seen that the values of some of the dihedral angles are close to zero, indicating a high ring strain. Considering these results, the formation of this H-bond creates a rigid structure with restricted ability for free rotation and with relatively unfavorable geometry in the six-membered ring.
Dihedral angles before and after docking of CrP and GAAP.

Crp: creatine phosphate; GAAP: phosphorylated guanidinoacetic acid.
HOMO–LUMO orbitals play important roles in the stabilizing interactions between ligands and receptor proteins. Hence, the orbital energies of both the HOMO and the LUMOs were calculated to estimate the chemical reactivity of the selected compounds using density functional theory (DFT). HOMO–LUMO plots were generated to analyze the atomic contribution of these orbitals. The HOMO and LUMO plots show the positive electron density in red color and the negative electron density in blue (Figure 2). The HOMO and LUMO energy gap (ΔE = ELUMO – EHOMO) explains the eventual charge transfer interactions taking place within the molecule. As can be seen from Figure 2, a lower energy gap was exhibited by GAAP and CrP compared to non-phosphorylated analogues. The higher values of HOMO–LUMO gap indicate the higher kinetic stability of the molecules. On the phosphorylation of creatine and guanidinoacetic acid, the values of ΔE are lowered, indicating less kinetic stability of the phosphorylated analogues. Considering the chemical hardness, GAA and Cre molecules are harder than GAAP and CrP.

Spatial distribution of the HOMO and LUMO molecular orbitals and the corresponding ΔE values of the investigated compounds. Ppositive electron density is shown in red, while negative electron density is shown in blue.
The molecular electrostatic potential (MEP) surfaces of the investigated ligands are shown in Figure 3. The color code is used to better visualize the density of the electrons, where blue indicates a minimal concentration of electrons and red indicates a high concentration of electrons. The regions with negative potential values were concentrated on oxygen and phosphorus, whereas regions with positive potentials were concentrated on the hydrogen atoms of NH2, NH, CH2, and CH3 groups. A region with the highest positive potential is located over the hydrogens from the guanidino group, and this site of the molecule has an essential role in hydrogen-bonding interactions with the negative sites of CK. However, from Figure 3, it is evident that introduction of a phosphate group leads to a more pronounced negative potential, causing stronger H-bond donor ability in the case of GAAP and CrP compared to GAA and Cre.

Molecular electrostatic potential (MEP) surfaces: (a) GAA, (b) Cr, (c) GAAP, and (d) CrP.
Molecular docking
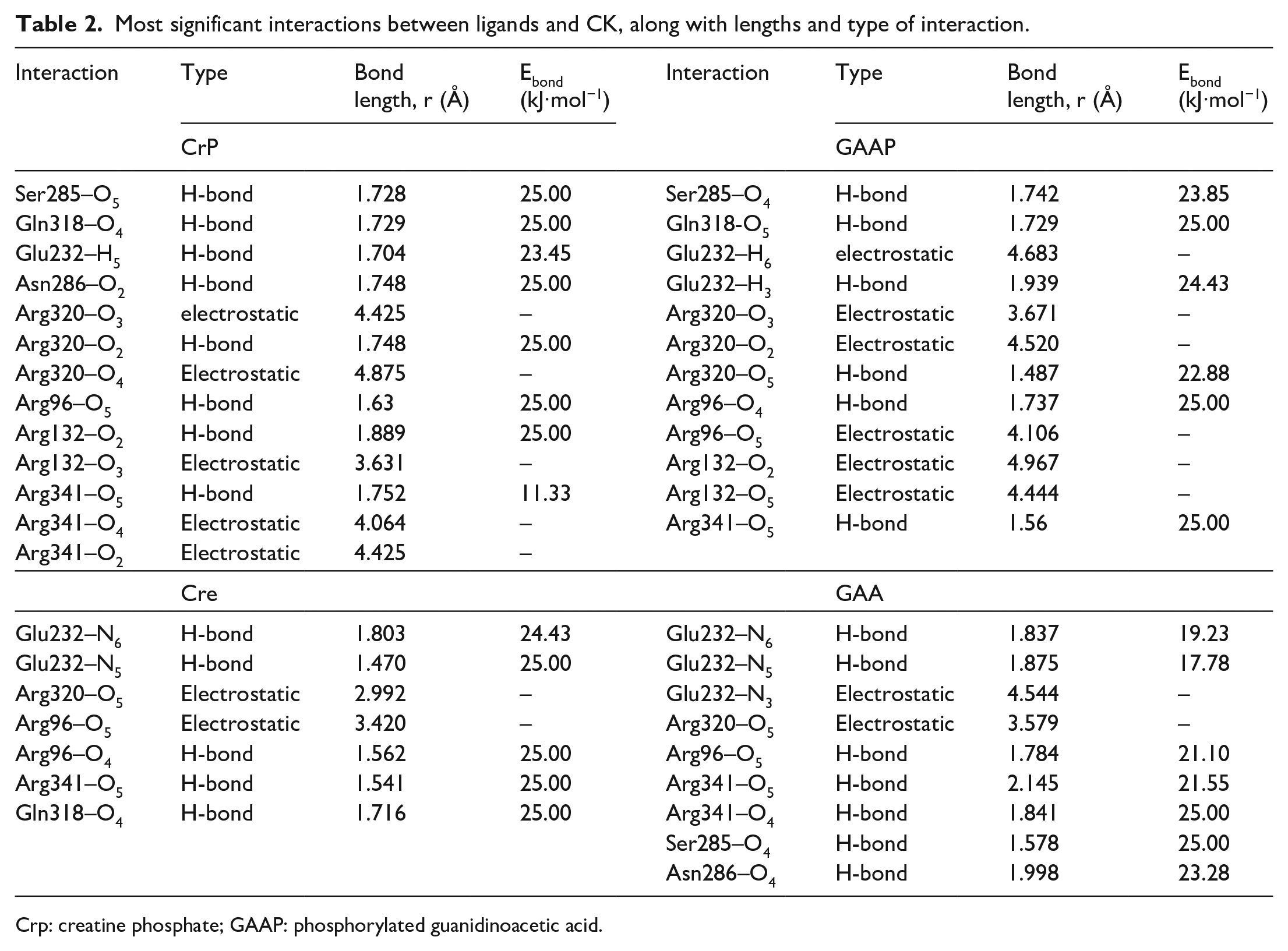
Docking results for the enzyme CK with GAAP, CrP, GAA and Cre are shown in Figure 4, and the most significant interactions are tabulated in Table 2. As can be seen from Figure 4, the enzyme CK has strong interactions with the oxygen atoms of the carboxylate groups of GAAP and CrP, through hydrogen bonding with arginine side chains (Arg 96, Arg 320, Arg 341), glutamine (Gln 318), and serine (Ser 285), which are in a good agreement with the results of Bong et al., 8 Lim et al., 9 and Jourden et al. 10 These H-bond interactions decrease the electron density on the carboxylate anion, which result in breaking of intramolecular H-bonds between the oxygen atom of carboxylate group and a hydrogen atom from N6 and allows for rotation around C2–N3 for 130° in the case of both molecules.

Docking results between the enyzme creatine kinase and different ligands: (a) GAA, (b) Cre, (c) GAAP, and (d) CrP.
Most significant interactions between ligands and CK, along with lengths and type of interaction.
Crp: creatine phosphate; GAAP: phosphorylated guanidinoacetic acid.
However, the most important difference between CrP and GAAP from an aspect of geometry and availability to bind with CK is a methyl group on N3 of the CrP molecule. As can be seen from Table 1, in the case of CrP, there are no significant changes in dihedral angles, except around the C2–N3 bond before and after binding with CK. However, the GAAP molecule has significant conformational changes after docking. Breaking the intramolecular H-bond between O3 and H3 in the GAAP molecule allows rotation around the N3–C4, C4–N5, and N5–P bonds (changes in the dihedral angles are given in Table 1). As a consequence of these geometry changes, a new H-bond between the O3 atom from phosphate group and H5 from the iminium group in GAAP (length 1.74 Å) is formed. These conformational changes lead to the formation of a new six-membered ring between H5–N6–C4–N5–P–O3, which is not planar and has a more stable conformation compared to GAAP before binding to CK. The formation of this new intramolecular H-bond in GAAP decreases the electron density on the oxygen atom from the phosphate group, resulting in a weaker interaction of this group with CK compared to a CrP molecule. Oxygen atoms from the
It is a well-known fact that magnesium ions play a crucial role in the formation of the transition state of the phosphoryl transfer reaction. Thus, in our case, the electrostatic interactions between magnesium ions and phosphate groups from CrP and GAAP were confirmed. Furthermore, the side chain of glutamic acid (Glu 232) had a significant role in binding CrP and GAAP molecules. A CrP molecule forms a H-bond between the H5 atom and the side-chain carboxyl group of Glu 232, while GAAP forms a H-bond through the H3 atom and a electrostatic interaction through H6 atom with the previously mentioned carboxyl group of Glu 232. These observations are in excellent agreement with the literature data, according to which the side-chain carboxyl group of Glu 232 plays a crucial role in orientating the substrate in the correct location for catalysis. 8 From these results, it can be concluded that the conformational geometry of docked GAAP changed more than the geometry of docked CrP, leading to relaxation in the molecular structure of GAAP. This relaxation is one of the main reasons for the more negative value of the binding energy of a GAAP molecule compared to CrP (Table 3).
Binding energies of selected ligands with the enzyme creatine kinase.

Crp: creatine phosphate; GAAP: phosphorylated guanidinoacetic acid.
A molecule of ADP, in the presence of GAAP and CrP, binds almost identically in both cases for CK. The NH2 group from adenine forms two H-bonds with an oxygen atom from the carbonyl group of serine (Ser 128) and glycine (Gly 294). Atom N1 from the adenine ring also forms one H-bond with the OH group of the serine (Ser 128) side chain. Histidine (His 296) plays the most significant role in ADP binding to CK, forming a π–π interaction with the adenine ring and a hydrogen bond with the 2′-OH group of the ribose ring. These results agree with the results of fluorescence quenching measurements on ADP docking on CK performed by Borders et al.,
11
and with X-ray crystallography measurements of docked ADP on CK by Lahiri et al.
12
and Bong et al.
8
Interactions between CK and oxygen atoms from phosphate groups occur via H-bonds and electrostatic interactions with positively charged side chains of arginine (Arg 130, Arg 132, Arg 292, Arg 320), as well as via H-bonds between the oxygen atom on the α-phosphate of ADP and the H-atom from the peptide bond between Val 325 and Gly 324. The crucial roles for optimal positioning of the molecule before the phosphoryl transfer reaction
10
have utilized
Stabilization of the carboxyl group of Cre, after transfer of the phosphate group, is less efficient compared to CrP. The Cre molecule forms five non-covalent interactions (three H-bonds and two electrostatic interactions), representing two less than CrP (five H-bonds and two electrostatic, Table 2). The difference in the efficiency of the stabilization of the carboxyl group is notably smaller in the case of GAA/GAAP. A molecule GAA has only one electrostatic interaction less with the side-chains of CK amino acids, compared to CrP. The total impact of all the interactions on the molecules of Cre and GAA and its phosphorylated forms is displayed via the electrostatic binding energies in Table 3, which show that the phosphorylated forms have stronger interactions with CK, and that the molecules of GAA and GAAP also are more strongly bond to the enzyme compared with Cre and CrP. Moreover, from the change of the atomic charges of the substrate before and after docking (Figure 5), it can be concluded that Cre and GAA have more significant changes in the electron structure after docking compared to CrP and GAAP. The existence of a phosphoryl group reduces the electron acceptor potential of the guanidino group, thereby decreasing the change in its electron structure.

Atomic charges before (left) and after (right) docking for (a) GAA, (b) Cre, (c) GAAP, and (d) CrP.
Conclusion
In this work, molecular docking of GAA, Cre, GAAP, and CrP toward muscle CK was performed, supported by DFT calculations, to better understand the binding affinity of these molecules. From the results of molecular docking, it is noted that optimal positioning of the molecule before the phosphoryl transfer reaction involves
Computational methods
The initial three-dimensional structure of muscle CK was retrieved from the protein data bank (PDB), its identifier being 1I0E. The raw CK structure was prepared using the Protein Preparation Wizard and the Glide module in the Schrödinger suite 2015-2 package. Desolvation was carried out by deleting the crystallized free water molecules beyond 5 Å. The enzyme structure integrity was checked and adjusted and missing residues and loops were added using Prime. The protonation, orientation, and tautomeric states of Asp, Glu, Arg, Lys, and His were adjusted to match a pH of 7.4 using the PROPKA option. The network of hydrogen atoms in the CK model was optimized at pH 7.4 using the optimized potential for liquid simulations (OPLS) force field (2005). The active sites of the modeled CK were investigated using SiteMap program implemented in Schrödinger Suite Package. The active pocket in CK was fixed with the help of the receptor generation program from the Glide module and used as such in the docking studies. This is a grid-based method with energetics that gives scores based on favorable interactions between a ligand and a protein. The binding sites were computed with a grid resolution of 0.3 Å. A Van der Waals scaling factor of 0.8 and partial charge cutoff of 0.15 were used in the docking process for all ligands.
The prepared ligands (Cre, GAA, GAAP, CrP, Mg2+, ADP, and ATP) were subjected to docking into the grid enclosed by active site of the enzyme, with flexible ligand sampling and using the extra precision mode. The grid size was set to 60 Å along all axes with a grid spacing of 0.5 Å, so the grid box allowed sufficient room for both translational and rotational freedom of the ligands. The docking parameters were as follows: population size 100, maximum number of iterations 2000, and the maximum number of steps 750. Interactions between the enzyme and the ligands were calculated based on the quality of the geometric contacts and their energies. Each ligand structure was geometrically optimized employing the empirically dispersion-corrected B3LYP exchange-correlation functional (B3LYP-D3) with the 6-31+G(d,p) basis set using Macro Model/Conformational Search and prepared for further docking using LigPrep with the force field OPLS-2005. All models were subjected to post-docking minimization and the best-docked structures for each ligand were determined based on the model energy score, which combines the energy grid score, the binding affinity, the internal strain energy, and the Coulomb–Van der Waals interaction energy scores. The models with the lowest energy were visualized using Maestro Pose Viewer (Schrödinger suite). The lowest binding energy conformer was chosen among 10 different conformers for each docking simulation and the one with the best score was used for further analysis after the post-docking minimization process had been finished. To estimate the relative binding affinity of the ligands and for comparison with experimental binding affinities, the free energies were computed via the Prime/Molecular Mechanics/Generalized Born Surface Area (MM/GBSA) approach. MM-GBSA considers contributions from OPLS molecular mechanics energies (EMM), polar solvation through the surface-generalized Born solvation model (GSGB), and a nonpolar solvation term composed of the nonpolar solvent-accessible surface area and Van der Waals interactions. The methodology adopts the surface-generalized Born model using a Gaussian surface for better representation of the solvent-accessible surface area. Residues in binding pockets of the protein were treated as flexible and ligand partial charges were assigned from initial charges. For a more reliable representation of the solvation effect as a dielectric medium, the generalized Born (GB) model was used.
The electronic effects of the bioactive molecules play an important role in the pharmacological effects. Subsequently, surfaces (molecular orbital, density, potential) and atomic electrostatic potential charges were monitored to compute the HOMO and LUMO energies. The HOMO energy represents the region of the small molecules which can donate electrons during binding, while the LUMO energy signifies the capacity of the molecule to accept electrons from the receptor. The difference in the HOMO and LUMO energies, known as the HOMO–LUMO gap energy, indicates the electronic excitation energy that is necessary to compute the molecular reactivity and stability of the compounds.
Supplemental Material
sj-pdf-1-chl-10.1177_1747519820978583 – Supplemental material for Molecular docking and density functional theory studies on creatine, guanidinoacetic acid, and their phosphorylated analogues binding to muscle creatine kinase
Supplemental material, sj-pdf-1-chl-10.1177_1747519820978583 for Molecular docking and density functional theory studies on creatine, guanidinoacetic acid, and their phosphorylated analogues binding to muscle creatine kinase by M Vraneš, S Ostojić, Č Podlipnik and A Tot in Journal of Chemical Research
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was financially supported by the Ministry of Education and Science of Serbia under project contract ON172012. The computational work was performed thanks to support received from Schrödinger Inc.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
