Abstract
Background
Recently, microRNA-221 has been found to be abnormally expressed in hepatocellular carcinoma; however, its clinical value has not been summarised. This meta-analysis aimed to assess the prognostic significance of miR-221 in hepatocellular carcinoma.
Material and Methods
PubMed, Science Direct, Web of Science, Scopus, Ovid MEDLINE, EMbase, Google Scholar, the Cochrane Library, CNKI, CBM, VIP and Wanfang databases were searched for eligible articles. The endpoints included overall survival, progression-free survival, recurrence-free survival, metastasis-free survival, disease-free survival. Hazard ratios with 95% confidence intervals were used to explore the relationship between miR-221 expression and clinical survival results of liver cancer patients. Subgroup analysis and sensitivity analysis were performed. Begg’s test and Egger’s test were conducted to evaluate publication bias.
Results
A total of nine studies including 607 patients were recruited for this meta-analysis. The pooled hazard ratios displayed that high miR-221 expression was remarkably associated with poorer overall survival (hazard ratio = 1.91, 95% confidence interval: 1.53–2.38, p < 0.01) and unfavourable progression-free survival/recurrence-free survival/metastasis-free survival/disease-free survival (hazard ratio = 2.02, 95% confidence interval: 1.58–2.57, p < 0.01). The results of Begg’s test and Egger’s test did not exhibit obvious publication bias.
Conclusions
High expression of miR-221 can predict poor outcome of hepatocellular carcinoma. miR-221 can be used as a promising prognostic biomarker of hepatocellular carcinoma.
Introduction
Hepatocellular carcinoma (HCC) is a common malignant tumour of the digestive system, the incidence rate of which ranks sixth among tumours in the world.1,2 However, China is one of the countries with high incidence and mortality of liver cancer in the world. 3 The occurrence of HCC is a multiple-step, multiple-stage process and evolution. Despite rapid development of treatment methods for liver cancer (surgical resection, transarterial chemoembolisation, radiofrequency ablation, targeted therapy and immunotherapy), the overall survival (OS) regarding liver cancer is still poor.4,5 There is an urgent need to study molecular mechanisms in depth and search for markers for early diagnosis or prognosis of liver cancer.
Previous studies have confirmed that miRNAs are abnormally expressed in tumours and participate in several stages of tumour pathophysiology, including cell proliferation, invasion and metastasis.6–8 It is believed that the biological functions of miRNAs vary depending on tissue sources and their target genes. And some miRNAs can be considered to be biomarkers for the diagnosis and prognosis of diseases.
MicroRNA-221 (miR-221) is a member of the miRNA family. It is located on the Xp11.3 region of the human chromosome and distributes in clusters. miR-221 was abnormally expressed in leukemia, 9 oral squamous cell carcinoma, 10 bladder cancer, 11 liver cancer, 12 breast cancer, 13 glioma 14 and so on. In Liu’s study, 10 up-regulating the expression of miR-221 could effectively reduce the expression of matrix metalloproteinases (MMPs) and superoxide dismutase, suggesting miR-221 could inhibit cell invasion and might act as anti-oncogene in oral squamous cell carcinoma. Contrary to the effect of miR-221 in oral squamous cell carcinoma, miR-221 acted as a proto-oncogene in liver cancer. Compared with liver cirrhosis patients or healthy people, the expression level of miR-221 was higher in patients with HCC. 15 The significantly abnormal expression of miR-221 might act as a potential prognostic biomarker for HCC. Our study evaluated the prognostic significance of miR-221 for liver cancer through meta-analysis, in order to provide references for its future research as a biomarker and scientific basis for clinical practice.
Materials and methods
Protocol and registration
This meta-analysis was registered with the International Platform of Registered Systematic Review and Meta-Analysis Protocols (INPLASY, https://inplasy.com), and the registration number was INPLASY2021130014. Besides, the study was performed according to the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines. 16
Search strategy and literature inclusion criteria
The search strategy used (‘hepatocellular cancer’ or ‘hepatocellular carcinoma’ or ‘hepatic neoplasm’ or ‘hepatic cancer’ or ‘liver cancer’ or ‘cancer of the liver’ or ‘liver neoplasms’ or ‘liver neoplasm’) and (‘microRNA-221’ or ‘miRNA-221’ or ‘miR-221’ or ‘has-miR-221’). The search strings were used to search PubMed, Science Direct, Web of Science, Scopus, Ovid MEDLINE, Embase, Google Scholar, the Cochrane Library, EMbase, CNKI, CBM, VIP and Wanfang databases. The deadline was 10 October 2020. The corresponding reference documents and conference paper abstract were searched manually. Screening titles, abstracts and full texts were read to distinguish eligible studies. The study was performed according to the PRISMA guidelines.
Article inclusion criteria: (a) research object: history of HCC; (b) research type: only randomised clinical trials were included, and similar articles published by the same author were selected recently; (c) outcome indicators: OS, progression-free survival (PFS), recurrence-free survival (RFS), metastasis-free survival (MFS), disease-free survival (DFS) or Kaplan–Meier curve can be obtained from the original article or contact the original author. If the reported information lacked detailed information, or the data had already been reported (same institution, repeated period of patient recruitment), the study would be excluded. In addition, reviews, editorials, abstracts, letters, case reports, and expert opinions were eliminated from meta-analysis.
Data screening and quality evaluation
The two researchers independently screened all articles and read the title, abstract and body text to exclude all irrelevant and repetitive articles that did not meet the inclusion and exclusion criteria. Relevant information was extracted, including the first author’s name, year of publication, country, study design, tumour type, detection method, OS, PFS, RFS, MFS, DFS, hazard ratio (HR) and corresponding 95% confidence interval (CI). The quality of each included study was evaluated according to the Newcastle–Ottawa Quality Assessment Scale (NOS). 17 NOS scores of 7–9 indicated high quality. However, NOS scores of 0–3 and 4–6 suggested low and moderate quality, respectively.
Statistical analysis
Review Manager 5.0 (Cochrane Collaboration, Oxford, UK) and STATA 15.0 software (STATA, University of Texas Station, USA) were applied for data analyses, using HRs and 95% CI to analyse the pooled data. A fixed effects model was used when I2 was <50% or p < 0.1. And we also conducted subgroup analyses. Sensitivity analysis was used to test stability of meta-analysis. We finally estimated publication bias by Begg's test and Egger's test. It was considered to be statistically significant when p < 0.05.
Results
Study characteristics and risk of bias within studies
Through the above-mentioned search strategy, we conducted a literature search and initially retrieved a total of 852 articles. After deleting 395 duplicate publications, 457 articles remained. We further excluded 439 articles through screening the titles and abstracts. After full-text review, we removed nine articles. Ultimately, nine studies including 607 patients were recruited for this meta-analysis.18–27 The literature selection process is displayed in Figure 1.
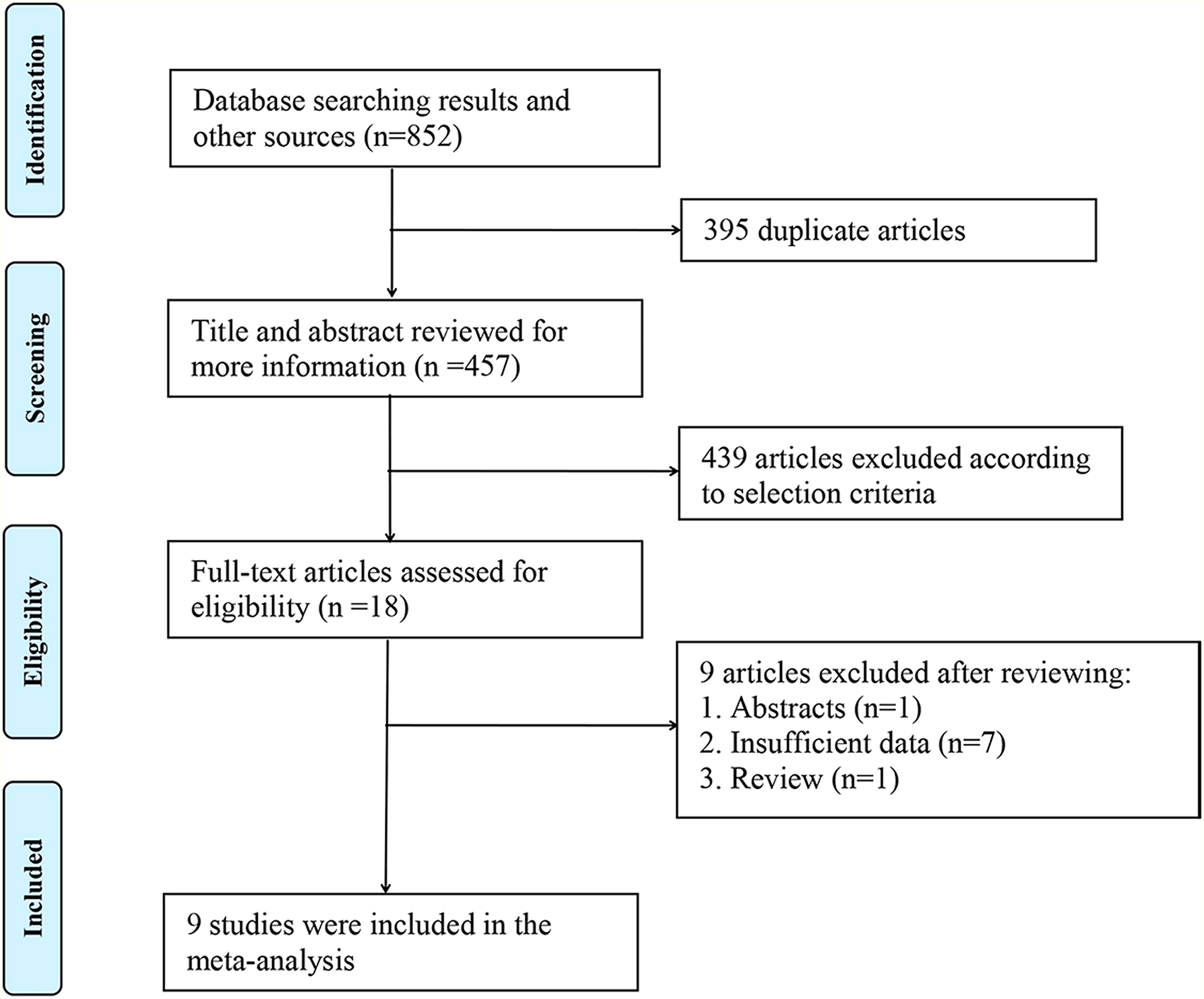
Flow diagram of the literature search.
The nine included studies were all retrospective studies. There were five studies from China, three in Europe and one in Korea. However, three studies detected miR-221 in frozen tissue (FT), five in formalin-fixed, paraffin-embedded tissues (FFPET) and one in serum. The average score of NOS of the included studies was 7.7. More details are displayed in Table 1.
Characteristics of nine included studies.
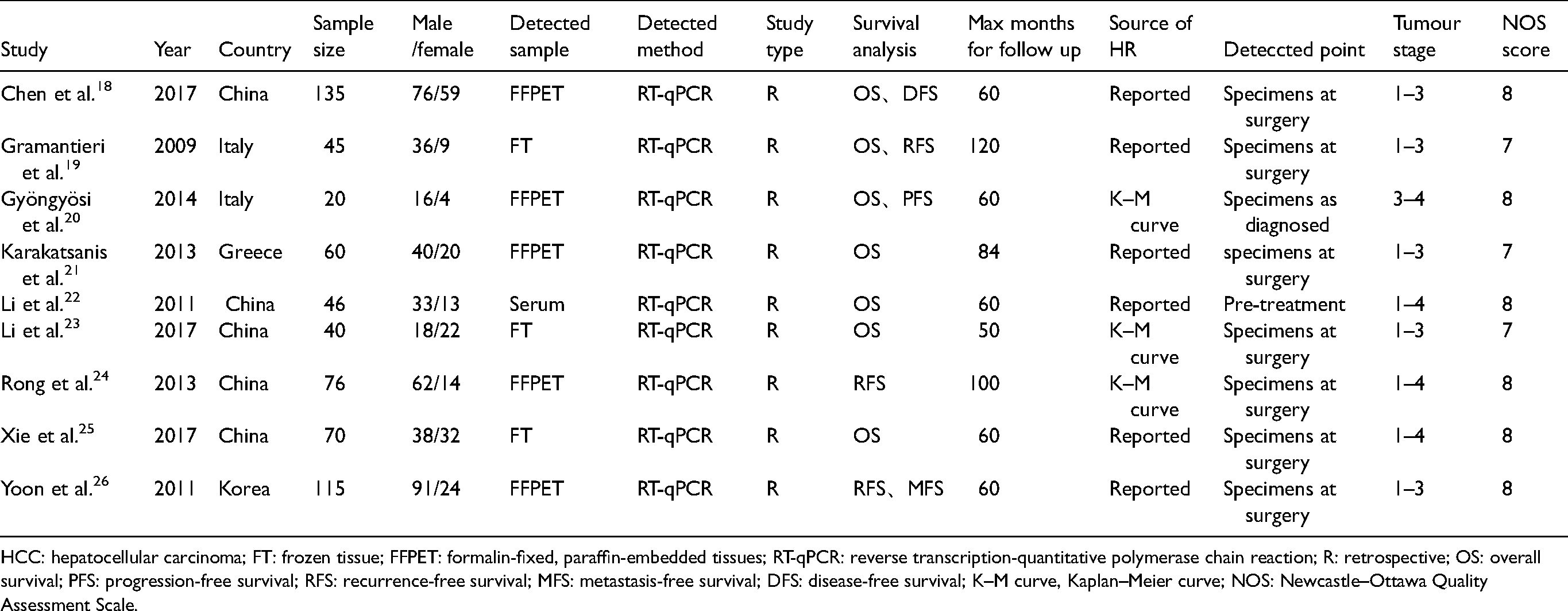
HCC: hepatocellular carcinoma; FT: frozen tissue; FFPET: formalin-fixed, paraffin-embedded tissues; RT-qPCR: reverse transcription-quantitative polymerase chain reaction; R: retrospective; OS: overall survival; PFS: progression-free survival; RFS: recurrence-free survival; MFS: metastasis-free survival; DFS: disease-free survival; K–M curve, Kaplan–Meier curve; NOS: Newcastle–Ottawa Quality Assessment Scale.
High miR-221 expression and OS
Seven articles consisted of 416 patients explored the relationship between miR-221 expression and prognosis using OS. The I2 = 0% and p = 0.75; so, we used a fixed effect model. For high expression of miR-221, the pooled HR for OS was 1.91 (1.53–2.38), suggesting high miR-221 was related to markedly poorer prognosis for liver cancer patients. The forest plot was shown in Figure 2.
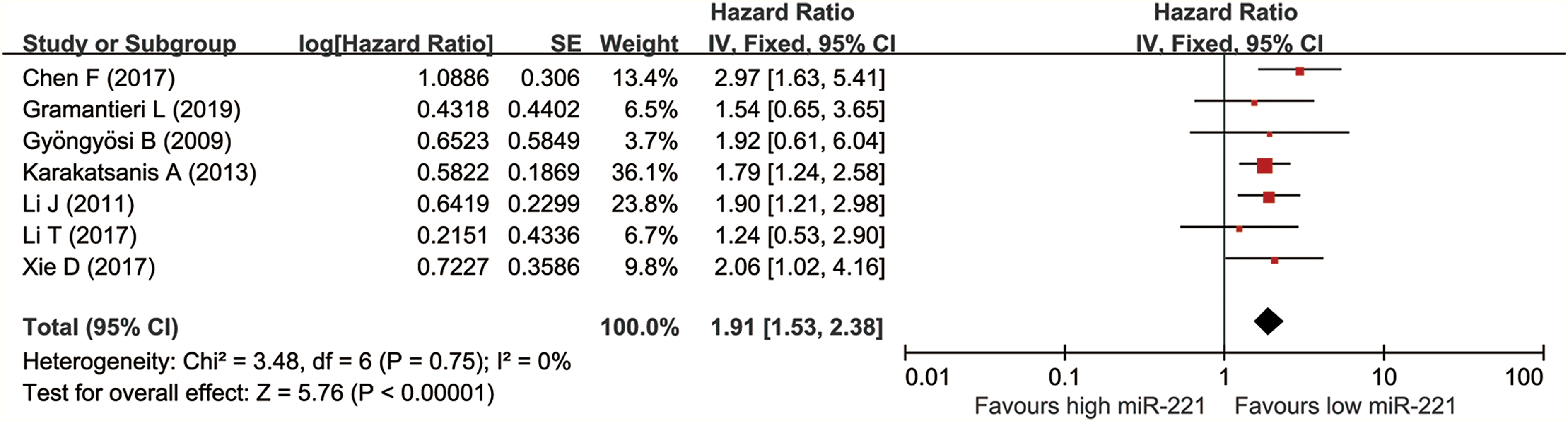
Forest plot of the relationship between high miR-221 expression and overall survival (OS).
High microRNA-221 expression and DFS/MFS/RFS/PFS
Five studies involving 391 patients documented the relationship between miR-221 expression and prognosis using DFS/PFS/RFS/MFS. The I2 = 41% and p = 0.13, suggesting no obvious heterogeneity was observed in these studies. Thus, a fixed effect model was adopted. Pooled HR for DFS/MFS/RFS/PFS was 2.02 (1.58–2.57). The high miR-221 could predict unfavourable DFS/MFS/RFS/PFS for liver cancer patients. More details are displayed in Figure 3.
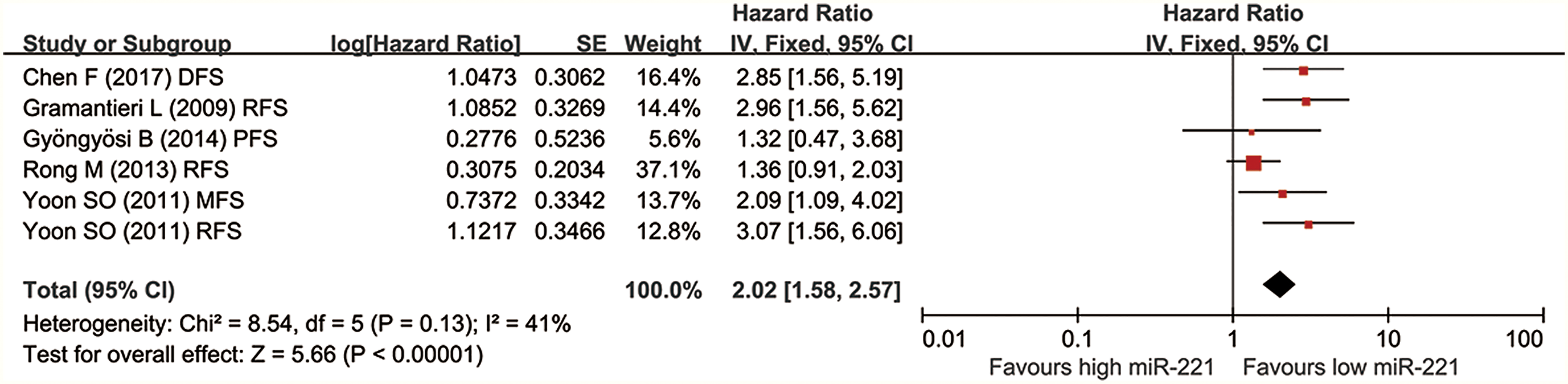
Forest plot of the relationship between high miR-221 expression and DFS/RFS/MFS/PFS.
Subgroup analysis
Table 2 displayed the results of subgroup analysis based on region, cancer type and detected sample. High level of miR-221 was associated with unfavourable OS in both Asian (HR = 2.04, 95% CI: 1.51–2.76) and non-Asian patients (HR = 1.76, 95% CI: 1.28–2.44). The high miR-221 could predict poorer DFS/MFS/RFS/PFS in Asian (HR = 1.94, 95% CI: 1.48–2.55) and non-Asian patients (HR = 2.36, 95% CI: 1.37–4.06). As for the result of subgroup analysis of the detected sample, up-expressed miR-221 in FFPET indicated poor OS (HR = 2.04, 95% CI: 1.51–2.76) and DFS/MFS/RFS/PFS (HR = 1.89, 95% CI: 1.46–2.46). High miR-221 in FT indicated poor OS (HR = 1.64, 95% CI: 1.03–2.59) and DFS/MFS/RFS/PFS (HR = 2.96, 95% CI: 1.56–5.62). More details are shown in Supplemental Figure 1.
Subgroup analysis for OS and PFS/RFS/MFS/DFS in patients with high miR-221 expression.
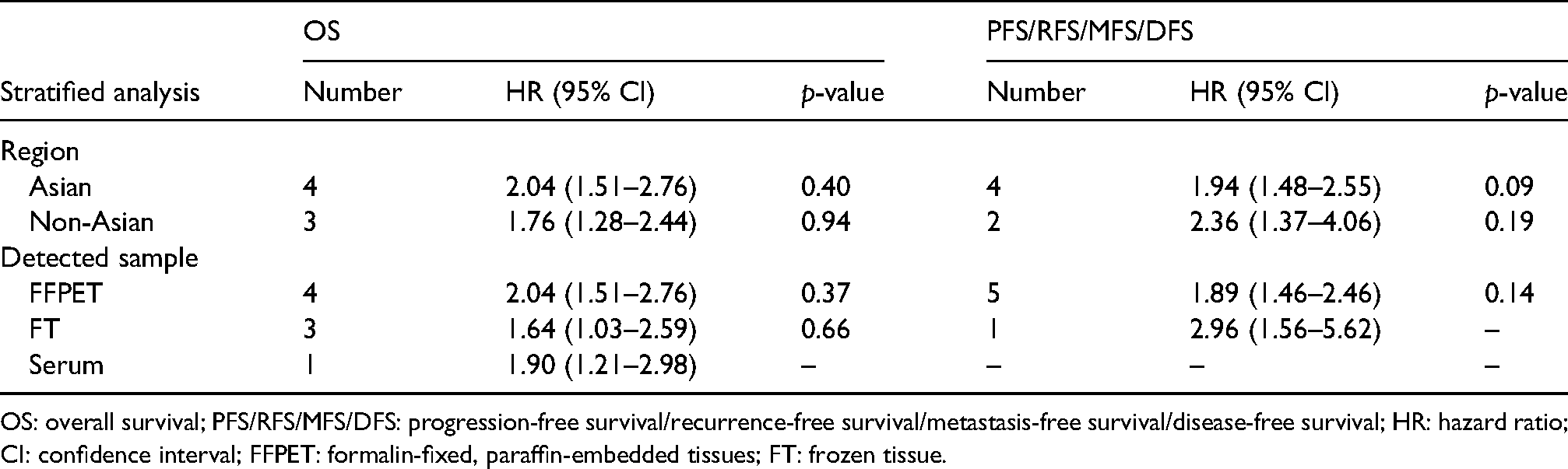

OS: overall survival; PFS/RFS/MFS/DFS: progression-free survival/recurrence-free survival/metastasis-free survival/disease-free survival; HR: hazard ratio; CI: confidence interval; FFPET: formalin-fixed, paraffin-embedded tissues; FT: frozen tissue.
Sensitivity analysis and publication bias
The results for OS and DFS/PFS/RFS/MFS were all stable, because no estimate exceeded the 95% CIs in sensitivity analysis (see Supplemental Figure 2). The Begg's funnel plots for OS and DFS/PFS/RFS/MFS were symmetrical overall (see Supplemental Figure 3). There was no proofed publication bias for the OS (Egger's test, p = 0.876) and DFS/PFS/RFS/MFS (Egger's tset, p = 0.373).
Discussion
HCC is a type of the most malignant solid tumours with poor prognosis. Alpha-fetoprotein is the most commonly used biomarker for HCC, but its sensitivity and specificity are flawed. 27 To investigate the mRNA level of miR-221 in HCC tissues, real-time quantitative reverse transcription polymerase chain reaction was performed in a previous study and results demonstrated that the miR-221 levels were higher in HCC tissues than in adjacent noncancerous tissues. 28 Rong et al. 24 believed that the overexpression of miR-221 was a key regulator and played a negligible role in the carcinogenesis of liver cancer. High miR-221 was found to be related to cell proliferation. They also observed the level of miR-221 was significantly associated with tumor, node, metastasis (TNM) staging, cell invasion and tumour recurrence. Consistent with the above results, study also reported that the expression level of miR-221 in HCC patients with T3, 4 or lymph node metastasis was higher than that in HCC patients with T1, 2, or without lymph node metastasis. 29 Therefore, there was close relationship between overexpression of miR-221 and poor prognosis of liver cancer patients. Our meta-analysis proved that miR-221 could be used as an effectively prognostic biomarker for HCC patients.
Our meta-analysis found that high miR-221 expression was significantly associated with unfavourable OS (HR = 1.91, 95% CI: 1.53–2.38, p < 0.01) and DFS/MFS/RFS/PFS (HR = 2.02, 95% CI: 1.58–2.57, p < 0.01). Subgroup analysis for OS displayed that high miR-221 expression was closely related to poor prognosis in both Asian (HR = 2.04, 95% CI: 1.51–2.76) and non-Asian patients (HR = 1.76, 95% CI: 1.28–2.44), suggesting that miR-221 overexpression can predict poor prognosis in both ethnic groups. Consistent with above results, high miR-221 expression was related to poorer DFS/MFS/RFS/PFS in Asian (HR = 1.94, 95% CI: 1.48–2.55) and non-Asian patients (HR = 2.36, 95% CI: 1.37–4.06). Besides, up-expressed miR-221 in FFPET of HCC patients indicated poor OS (HR = 2.04, 95% CI: 1.51–2.76) and DFS/MFS/RFS/PFS (HR = 1.89, 95% CI: 1.46–2.46). High miR-221 in FT indicated poor OS (HR = 1.64, 95% CI: 1.03–2.59) and DFS/MFS/RFS/PFS (HR = 2.96, 95% CI: 1.56–5.62). And higher miR-221 in serum can predict poor OS (HR = 1.90, 95% CI: 1.21–2.98). Therefore, miR-221 was a good prognostic biomarker for HCC patients, regardless of whether it was detected by FFPET, FT or serum. However, there was limited original research and sample size in the other subgroup. No obvious heterogeneity was observed in this meta-analysis. And no proofed publication bias was found for the OS and DFS/PFS/RFS/MFS. In consideration of the above results, we believed that miR-221 was a valuable prognostic indicator for HCC.
To meet the demand for comprehensive identification of miR-221, many researchers have verified over the years how miR-221 played a critical role in HCC. 30 It was reported that miR-221 was highly expressed in 71% of HCC patients, while CDKN1B/p27 and CDKN1C/p57 proteins were low expressed in 77% of cases. Furthermore, the expression level of miR-221 was significantly negatively correlated with CDKN1B/p27 and CDKN1C/p57. These results indicated that miR-221 had carcinogenic effects in liver cancer by targeting CDKN1B/p27 and CDKN1C/p57, hence promoting cell growth and proliferation. A study conducted by Gramantieri et al. 19 demonstrated that enhanced expression of miR-221 could lead to lower Bmf expression. Then, a luciferase reporter gene experiment confirmed that Bmf was one of the targets of miR-221. So, miR-221 inhibited liver cancer cell apoptosis by regulating the expression levels Bmf and caspase-3. In addition, miR-221 could also promote the proliferation of liver cancer through the PI3K/AKT/mTOR pathway by targeting DDIT4. 31 Garofalo et al. 32 showed that miR-221 induced resistance to tumour necrosis factor-related apoptosis-inducing ligand by targeting PTEN and TIMP3 and enhanced cell migration. Additionally, up-regulation of miR-221 could affect the proliferation, invasion, and migration of HCC through the JAK-STAT3 signalling pathway by targeting SOCS3. 33 Yuan et al. 34 found aryl hydrocarbon nuclear translocator (Arnt) messenger RNA was a new target of miR-221, which contributed to promoting proliferation of miR-221. Chen et al. 35 suggested that miR-221 modulated tumour cell proliferation by suppressing estrogen receptor-α (ERα), functioning as a tumour promoter. A previous study also reported that miR-221 suppression protected against endoplasmic reticulum stress-induced apoptosis. 36 miR-221 could promote epithelial–mesenchymal transition of HCC by targeting a new gene AdipoR1. 23 In addition, miR-221 regulated angiogenesis by Staphylococcal nuclease domain containing 1 (SND1) in HCC cells. 37 In HepG2 cells, miR-221 overexpression promoted cell growth, invasion and inhibition of apoptosis by activation of NF-κB and upexpression of bcl-2, VEGF and MMP-9. 38 These above results collectively suggested that miR-221 regulated tumour progression through various signalling pathways, reflecting the complexity of its biological mechanism.
In the meta-analysis, we proved miR-221 was a promising prognostic marker for HCC. Looking for biomarkers that can predict the prognosis of HCC will help the development of new molecular targeted drugs and provide choices of treatment options. Combining molecular targeted drugs with different mechanisms and multiple pathways will help improve the efficacy of targeted therapy. In the future, it is necessary to further identify new biomarkers and evaluate their clinical significance in patients with HCC, so as to improve clinical efficacy and achieve individualised treatment.
Although this study provided important information for understanding the clinical value and regulatory mechanisms of miR-221, some limitations of this study should be noted. Firstly, not all the included studies had a large sample size, which might impaired the statistical power of meta-analysis. Secondly, because some qualified studies did not directly provide survival data, the corresponding HR and 95% CI were extracted according to the survival curve, which may cause some micro-statistical errors. Thirdly, the cutoff value of each study was different, and the gold standard of the cutoff value should be verified to better assess the prognostic value of miR-221. In the future, more research should focus on the critical point and specific mechanism of miR-221 in HCC.
Conclusions
Based on all qualified evidence, our study showed that the upregulation of miR-221 had a statistically significant association with the poor OS and DFS/RFS/MFS/PFS of HCC. Through a comprehensive review of previous studies, we found that miR-221 was involved in the activation of various tumour signalling pathway by binding various targets. To sum up, miR-221 can be used as a promising prognostic marker for HCC.
Supplemental Material
sj-docx-1-jbm-10.1177_17246008211032689 - Supplemental material for The prognostic significance of microRNA-221 in hepatocellular carcinoma: An updated meta-analysis
Supplemental material, sj-docx-1-jbm-10.1177_17246008211032689 for The prognostic significance of microRNA-221 in hepatocellular carcinoma: An updated meta-analysis by Wenfeng Liu, Keshu Hu, Feng Zhang, Shenxin Lu, Rongxin Chen, Zhenggang Ren and Xin Yin in The International Journal of Biological Markers
Footnotes
Acknowledgements
This research was supported by the National Natural Science Foundation of China (No.: 81972889) and Excellent Backbone Foundation of Zhongshan hospital, Fudan University (No.: 2019ZSGG20).
Authors’ contributions
Wenfeng Liu was responsible for designing the work, data collecting and manuscript writing; Wenfeng Liu and Keshu Hu took part in data analysis and interpretation; Feng Zhang and Shenxin Lu helped collect the data; Rongxin Chen and Zhenggang Ren helped perform the analysis with constructive discussions; Xin Yin was responsible for the whole study and approved the final version.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Excellent Backbone Foundation of Zhongshan Hospital, Fudan University (grant nos. 2019ZSGG20 and 81972889).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
