Abstract
Background:
ADAMTS5 has different roles in multiple types of cancers and participates in various molecular mechanisms. However, the prognostic value of ADAMTS5 in patients with hepatocellular carcinoma (HCC) still remains unclear. We carried the study to evaluate the prognostic value and identified underlying molecular mechanisms in HCC.
Methods:
Firstly, the association of ADAMTS5 expression and clinicopathological parameters was evaluated by in GSE14520. Next, ADAMTS5 expression in HCC was performed using GSE14520, GSE36376, GSE76427 and The Cancer Genome Atlas (TCGA) profile. Furthermore, Kaplan-Meier analysis, Univariate and Multivariate Cox regression analysis, subgroup analysis was performed to evaluate the prognostic value of ADAMTS5 in HCC. Finally, GO enrichment analysis, gene set enrichment analysis (GSEA) and weighted gene co-expression network analysis (WGCNA) were performed to revealed underlying molecular mechanisms.
Result:
The expression of ADAMTS5 was positively correlated with the development of HCC. Next, high ADAMTS5 expression was significantly associated with poorer survival (all P < 0.05) and the impact of ADAMTS5 on all overall survival (OS), disease-free survival (DFS), relapse-free survival (RFS), disease specific survival (DSS) and progression free interval (PFI) was specific for HCC among other 29 cancer types. Subgroup analysis showed that ADAMTS5 overexpression was significantly associated with poorer OS in patients with HCC. Finally, ADAMTS5 might participate in the status conversion from metabolic-dominant to extracellular matrix-dominant, and the activation of ECM-related biological process might contribute to high higher mortality risk for patients with HCC.
Conclusion:
ADAMTS5 may play an important role in the progression of HCC, and may be considered as a novel and effective biomarker for predicting prognosis for patients with HCC.
Introduction
HCC, accounting for 75-85% cases of liver cancer, is considered as the sixth common occurring cancer and the fourth most common cause of cancer-related death worldwide, 1 especially in East Asia. 2 HCC is characterized by high invasiveness, high metastasis potential, high rate of recurrence and low survival rate. In the past 2 decades, death rates associated with HCC still continue to increase. Additionally, multiple predisposing parameters are known to contribute to the development of HCC, the most common of those being chronic viral hepatitis (B and C), alcohol abuse, and exposure to aflatoxins, 3 and it is predicted that the incidence of HCC will go to the peak in 2030. 4 Hence, effective prognostic biomarkers and novel therapeutic targets are urgently required.
ADAMTS5 belongs to the ADAMTS family, which participated in normal biological as well as pathological processes, such as cancers. 5 Recently, increasing evidence indicated the potential role of ADAMTS5 in multiple types of cancers, overexpression of ADAMTS5 promote the development of human glioblastomas, 6 and downregulation or hypermethylation of ADAMTS5 is reported to inhibite many kinds of cancers, including human colorectal cancer, 7 gastric cancer, 8 melanoma. 9 To date, few study reported the association between HCC and ADAMTS5, Chongyi Li et al 10 have reported low expression of ADAMTS5 protein associates with progression and poor prognosis of HCC. However, multiple online websites showed high ADAMTS5 expression was an independent risk factor for HCC development, thus the exact role of ADAMTS5 in HCC is unclear and contradictory. In the study, we performed a systemic analysis to reevaluate prognostic value and revealed molecular mechanisms in HCC.
Materials and Methods
Three gene expression profiles(GSE14520, GSE36376, GSE76427) were downloaded from GEO database (http://www.ncbi.nlm.nih.gov/geo/), the clinical information of GSE14520 was also identified form GEO database, the detailed clinical information was shown in Table 1. GSE14520 contained 219 HCC tumor samples and 241 nontumor samples, GSE36376 contained 240 HCC tumor samples and 193 adjacent non-tumor samples, GSE76427 contained 52 HCC tumor samples and 145 adjacent non-tumor samples. The gene expression profile contained 415 candidates (365 patients and 50 normal candidates) was downloaded from TCGA (https://cancergenome.nih.gov/).
Correlation of ADAMTS5 Expression With Clinicopathological Parameters.

ADAMTS5 mRNA Expression Analysis
ADAMTS5 mRNA was compared between tumor and non-tumor samples by performing R Limma package. Furthermore, ADAMTS5 mRNA was investigated by analyzing GSE14520 and TCGA among different TNM stages, Tumor Size, CLIP stage, T.
The samples were divided into high- and low- ADAMTS5 expression groups according to median ADAMTS5 mRNA level, Kaplan-Meier analysis for OS and DFS was performed to compare the survival rate by analyzing GSE14520 and TCGA dataset. Univariate and multivariate logistic-regression analysis were performed to investigate whether ADAMTS5 could be independent of other clinical parameters. Besides, the association between ADAMTS5 expression and OS, DFS, DRF, PFS, DSS, progression free survival(PFS) was analyzed by data mining in Kaplan-Meier plotter (http://kmplot.com/analysis/), GEPIA(http://gepia.cancer-pku.cn/index.html) and Oncolnc (http://www.oncolnc.org/). Additionally, the Kaplan Meier plotter (http://kmplot.com/analysis/) was used for subgroup survival analysis for OS in HCC patients. Furthermore, the association between ADAMTS5 expression and OS, DFS was studied across 29 cancer types to evaluate the differences related to cancer type by using GEPIA. Finally, the association between ADAMTS5 mRNA and OS, DSS, PFI was studied across 29 cancer types to evaluate the differences related to cancer type by using TCGA dataset.
The samples were divided into high- and low- ADAMTS5 expression group according to median ADAMTS5 mRNA level. DEGs between high- and low- ADAMTS5 expression group from GSE14520 were identified using the Limma version 3.36.2 R package, with corrected P-value < 0.05 and absolute log fold change (FC) > 0.5 being considered as the cutoff criterion. GO biological process enrichment analysis was performed for up-DEGs and down-DEGs. Besides, co-expression analysis also was performed to identified significant genes associated with ADAMTS5 by mining TCGA dataset, with P-value < 0.001 and spearman correlation coefficient r > 0.6. Finally, GSEA (http://software.broadinstitute.org/gsea/index.jsp) was conducted to investigate pathways and groups of genes that may be associated with ADAMTS5, NOM P-value < 0.05 was considered significantly enriched.
To furtherly find the gene modules associated with ADAMTS5, The samples were divided into high- and low- ADAMTS5 expression group according to median ADAMTS5 mRNA level, high and low ADAMTS5 group were considered as external sample traits. Gene co-expression network was constructed to explore the modules highly associated with sample traits, 9 was selected as the soft thresholding power to produce a weighted network. The enrolled genes were clustered into 6 modules except the gray module, the modules with high correlation coeficient were considered candidates relevant to traits, which were picked out to perform further pathway enrichment analysis using an online-based web tool “Metascape” (http://metascape.org/).
Result
The Association of ADAMTS5 Expression With Clinicopathological Parameters
As showed in Table 1, we found the ADAMTS5 mRNA has a high association with liver cirrhosis (P value = 0.001), main tumor size (P value = 0.0059), multinodular (P value = 0.005), ALT (P = 0.005), early recurrence (P value = 0.047), CLIP stage (P value = 0.001), TNM stage (P value = 0.001). However, there was no significant correlation between age (P value = 0.066), gender (P value = 0.111), AFP (P value = 0.251), HBV viral status (P value = 0.086). Logistic-regression analysis were performed to assess association between ADAMTS5 mRNA and clinicopathological parameters, which revealed the similar result, the ADAMTS5 mRNA has a higher association with liver cirrhosis (P value = 0.003), main tumor Size (P value = 0.005), multinodular (P value = 0.005), ALT (P value = 0.005), early recurrence (P value = 0.049), CLIP stage (P value = 0.003), TNM stage (P value = 0.0009), and there was no significant correlation between age (P value = 0.051), gender (P value = 0.11), AFP (P value = 0.25), HBV viral status (P value = 0.13)(Table 2).
Logistic Regression of ADAMTS5 Expression Association With Clinical Features.

The Expression of ADAMTS5 Was Positively Correlated With the Development of HCC
As showed in Figure 1, ADAMTS5 mRNA was significant higher in HCC tumor samples than normal samples in TCGA (Figure 1AC), GSE14520 (Figure 1D), GSE36376 (Figure 1E) and GSE46427 (Figure 1F), and ADAMTS5 mRNA was significant higher in HCC tumor than adjacent peritumoral tissues (Figure 1B). Next, we analyzed the ADAMTS5 mRNA in HCC specimens from different tumor TNM stages, T, main tumor size, CLIP stage in the TCGA and GSE14520 database. In GSE14520, ADAMTS5 mRNA was significant higher in main tumor size >5 cm than main tumor size <=5cm (Figure 1G), as the CLIP stage (Figure 1H) and TNM stage (Figure 1I) increased, the ADAMTS5 mRNA increased with statistical significance. In TCGA dataset, as the T (Figure 2J) and TNM stage (Figure 2K) elevated, the ADAMTS5 mRNA increased with statistical significance. GEPIA also indicated that ADAMTS5 mRNA had an elevated trend with TNM stage increased (Figure 1L), while the P value = 0.0977.
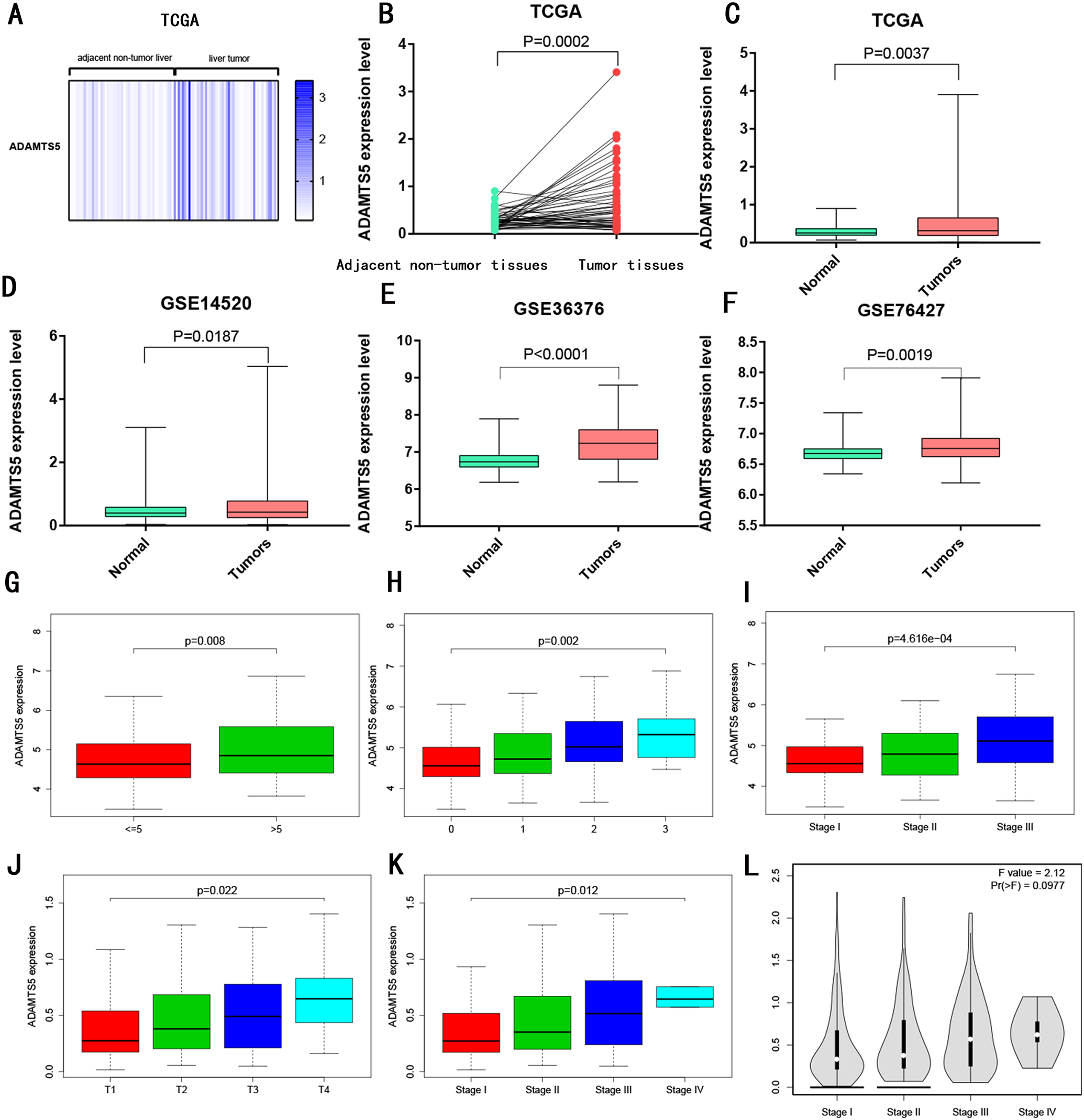
Expression level of ADAMTS5. MRNA expression level of ADAMTS5 between normal samples and HCC tumor samples in (A, C) TCGA; (D) GSE14520; (E) GSE36376; (F) GSE76427. MRNA expression level of ADAMTS5 between adjacent peritumoral tissues and HCC tumor tissues in (B) TCGA. MRNA expression level of ADAMTS5 based on different pathological parameters in (G, H, I) GSE14520; (J, K) TCGA and (L) GEPIA. (G) Main tumor size; (H) CLIP stage; (I, K, L) TNM stage.

Kaplan-Meier survival analysis based on (A) GSE14520; (B-D) TCGA; (E) oncolnc; (F-G) GEPIA; (H-K) Kaplan-Meier plotter. (A, B, G, I) DFS; (C, J) DSS; (D) PFI; (E, F, H)OS; (K) PFS.
As showed in Figure 3, HCC patients with high ADAMTS5 expression were associated with poorer OS(log-rank p = 0.003) in GSE14520(Figure 3A) and poorer OS(log-rank P value < 0.001) in TCGA(Figure 3B). With the increasing ADAMTS5 mRNA, patients have a worse OS in GSE14520 (Figure 3C) and worse OS(log-rank P < 0.001) in TCGA (Figure 3D). Univariate Cox regression analysis and multivariate Cox regression analysis also displayed ADAMTS5 was a powerful and independent factor in GSE14520 (Figure 3E, 3G) and TCGA dataset (Figure 3F, 3H). Besides, HCC patients with high ADAMTS5 mRNA expression were associated with worse DFS (log-rank P value = 0.006) in GSE14520 (Figure 2A), and worse DFS(log-rank P value = 0.023) in TCGA (Figure 2B). Additionally, high ADAMTS5 mRNA significantly contributed to worse DSS(log-rank P value = 0.025, Figure 2C) and PFI(log-rank P value = 0.006, Figure 2D) in TCGA dataset. Next, we validated the prognosis value in online website, Oncolnc website showed that HCC patients with high ADAMTS5 mRNA expression were associated with worse OS(log-rank P value = 6.5e-5, Figure 2E); GEPIA website indicated HCC patients with high ADAMTS5 mRNA expression were associated with worse OS(log-rank P value = 9.8e-5, Figure 2F) and DFS(log-rank P value = 0.0069, Figure 2G); Kaplan Meier plotter indicated high ADAMTS5 mRNA expression significantly contributed to worse OS(log-rank P value = 1e-6, Figure 2H), relapse-free survival (RFS, log-rank P value = 0.0044, Figure 2I), DSS(log-rank P value = 0.00029, Figure 2J) and progression free survival (PFS, log-rank P value = 0.00014, Figure 2K).
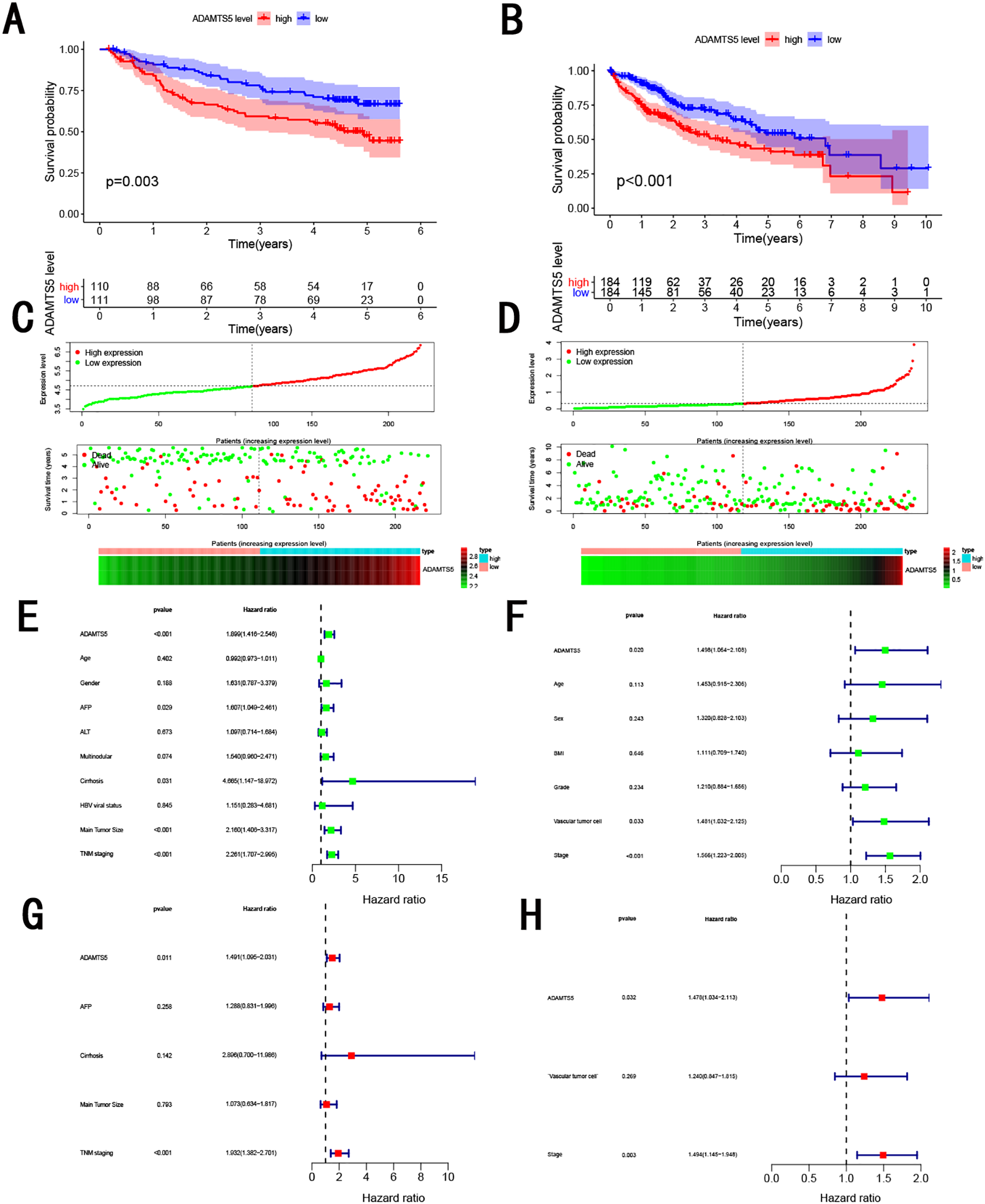
Survival analysis for OS in (A, C, E, G) GSE14520 and (B, D, F, H) TCGA. (A, B) Kaplan-Meier curves of OS; (C, D) distribution of patient survival durations, gene expression levels; (E, F) prognostic value detection of the ADAMTS5 via univariate survival-related analysis; (G, H) prognostic value detection of the ADAMTS5 via multivariate survival-related analysis.
Furthermore, we perform the survival analysis stratified by clinical variables(main tumor size, CLIP stage, TNM stage) in GSE14520 and by clinical covariates(T, TNM stage) in TCGA dataset. Stratification analysis indicated that high ADAMTS5 expressing HCC patients showed significantly worse OS rate than low ADAMTS5 expressing HCC patients in all the subgroups (Figure 4). Meantime, we make a stratification analysis by using online website Kaplan Meier plotter, high ADAMTS5 expressing HCC patients belonging to male, female, vascular invasion- Micro, vascular invasion- None, hepatitis virus infection-positive, hepatitis virus infection-negative, stage I-II, stage III-IV, stage I, stage II and stage III groups showed significantly worse OS rate than those with low ADAMTS5 expressing HCC patients (Figure 5).

Kaplan–Meier survival analysis of OS based classifier in subgroups of patients with pathological factors in GSE14520 and TCGA.
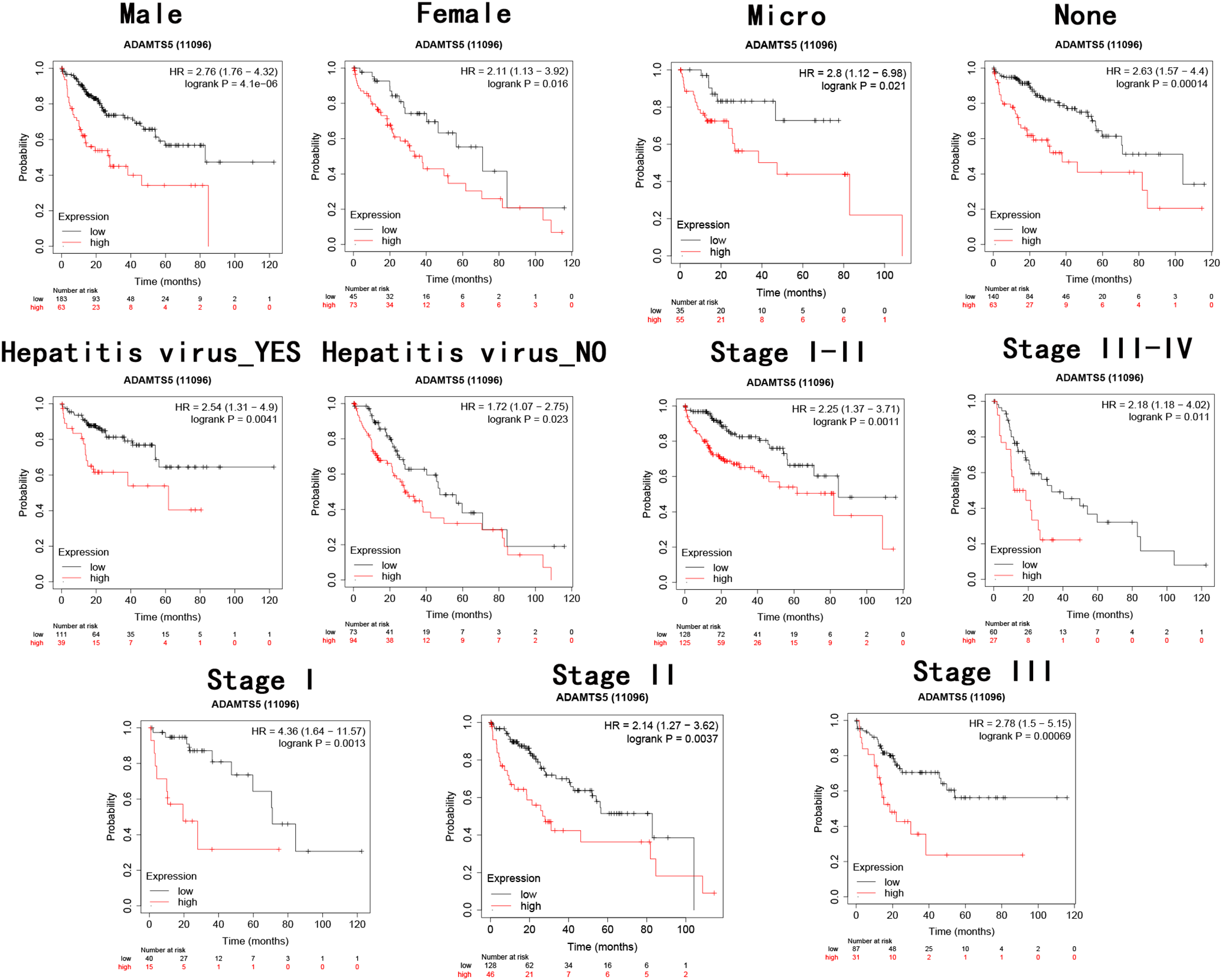
Kaplan–Meier survival analysis of OS based classifier in subgroups of patients with clinical characteristics online website Kaplan-Meier plotter.
The prognostic effect of ADAMTS5 in other 29 cancer types was investigated using GEPIA. As summarized in Table 3, HCC patients in high ADAMTS5 expression group were associated with worse OS(log-rank P value = 9.80E-05) and DFS(log-rank P value = 0.0069), the impact of ADAMTS5 on both OS and DFS was specific for HCC only. Next, we investigated the prognostic effect of ADAMTS5 in other 29 cancer types using TCGA dataset, HCC patients with high ADAMTS5 mRNA expression were associated with worse OS(log-rank P value = 0.000536), OS(Cox proportional hazards P-value = 2.47E-06), DSS(Cox proportional hazards P value = 0.001772), PFI(log-rank P value = 0.004402), the impact of ADAMTS5 on all OS, DSS and PFI was specific for HCC only (Table 4).
Survival Analysis Correlations for ADAMTS5 Across 29 Cancer Types From the Online Website (GEPIA).

Survival Analysis Correlations for ADAMTS5 Across 29 Cancer Types From the Cancer Genome Atlas (TCGA) Project.

ADAMTS5 Expression Correlates With Extracellular Matrix (ECM) and Metabolic and Energy Regulation
HCC patients were divided into high- and low- ADAMTS5 expression group according to median ADAMTS5 expression level. We identified 89 up-upregulated DEGs and 140 down-regulated DEGs between high and low expression group (Figure 6A). Go enrichment analysis revealed that up-regulated genes were mainly involved in extracellular matrix organization, extracellular structure organization and extracellular matrix disassembly (Figure 6B), down-regulated genes were mainly involved in organic acid catabolic process, carboxylic acid catabolic process, small molecule catabolic process and alpha−amino acid metabolic process (Figure 6C). To further evaluate the function of ADAMTS5, we identified significantly co-expressed genes in HCC using TCGA dataset (P value < 0.0001, Figure 6D). A list of co-expressed genes including Spearman correlation coefficients 0.6 was shown in Table 5, Go enrichment analysis revealed that co-expressed genes were mainly involved in extracellular matrix organization, extracellular structure organization and cell−matrix adhesion (Figure 6E). In order to further explore the pathways regulated by ADAMTS5 in patients with HCC, we conducted GSEA between high and low ADAMTS5 expression group to identify the significant pathways (NOM P-value < 0.05), the genes in high ADAMTS5 expression group were mainly enriched in extracellular matrix (ECM), including KEGG_ECM_RECEPTOR_INTERACTION, KEGG_FOCAL_ADHESION, KEGG_GAP_JUNCTION, KEGG_PATHWAYS_IN_CANCER, KEGG_REGULATION_OF_ACTIN_CYTOSKELETON. As to ADAMTS5 low-expression group, the genes were enriched in metabolic pathways, such as KEGG_BUTANOATE_METABOLISM, KEGG_FATTY_ACID_METABOLISM, KEGG_GLYCINE_SERINE_AND_THREONINE_METABOLISM, KEGG_PRIMARY_BILE_ACID_BIOSYNTHESIS (Figure 6F).
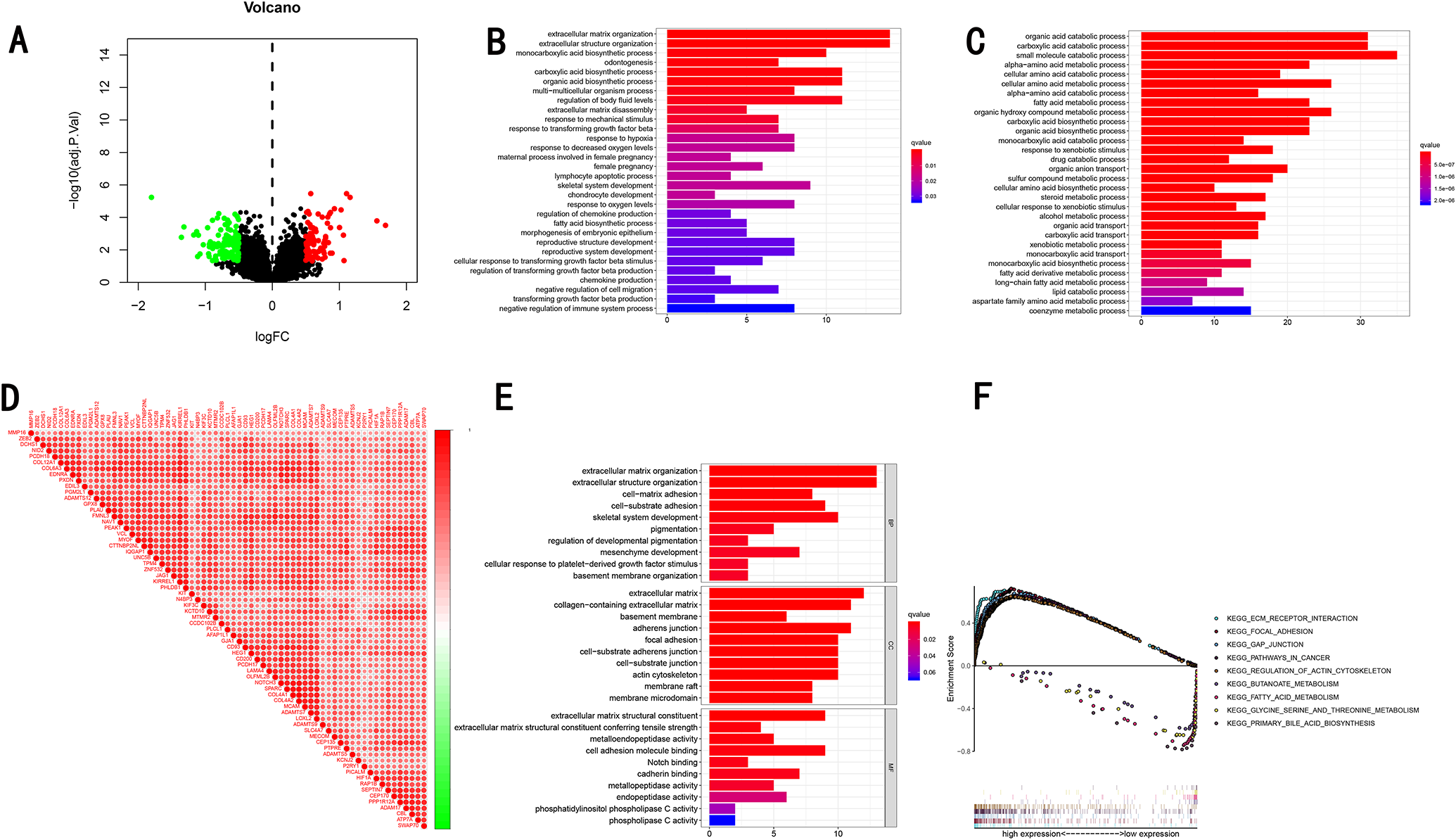
GO enrichment analysis based on DEGs and GSEA enrichment analysis revealed the biological processes and KEGG signaling pathways associated with ADAMTS5 in HCC. (A) Volcano plot of DEGs between high- and lowADAMTS5 expression group. The red points represent high expression genes, the green points represent low expression genes, the black points represent genes with no significant difference (corrected P value 0.5). (B) Biological processes associated with upregulated genes. (C) Biological processes associated with down-regulated genes. (D) Co-expressed genes associated with ADAMTS5 in HCC using TCGA (r > 0.600, p < 0.001). (E) Go enrichment analysis associated with co-expressed genes. (F) GSEA enrichment analysis revealed KEGG signaling pathways which were active in high expression or low expression group.
List of Genes Associated With ADAMTS5 Expression (r > 0.600, P < 0.001).

Extracellular Matrix (ECM) Related Process Were Associated With ADAMTS5
To furtherly investigate the potential mechanisms of ADAMTS5 in HCC, WGCNA was performed to cluster genes that highly correlated with the high ADAMTS5 group and low ADAMTS5 group based on the expression profile of GSE14520. We identified 6 modules (Figure 7A) in the study(excluding a gray module). As showed in Figure 7B, we found genes in green module was highly positively associated with high ADAMTS5 group(7x10-5) and genes in turquoise module was highly positively associated with low ADAMTS5 group(1x10-6). Go enrichment analysis was performed to explore the potential mechanisms of genes in green module and turquoise module using online-based web tool “Metascape” (http://metascape.org/), as showed in Figure 7C and Figure 7D, genes in the turquoise module were significantly enriched in metabolic process, such as monocarboxylic acid metabolic process, small molecule catabolic process, and carboxylic acid biosynthetic process. As showed in Figure 7E and Figure 7F, genes in the green module were significantly enriched in extracellular matrix (ECM), such as extracellular matrix organization, response to growth factor, and blood vessel development. Thus ADAMTS5 might participate in the status conversion from metabolic-dominant to extracellular matrix-dominant, and the activation of ECM-related biological process may might contribute to high higher mortality risk for patients with HCC.

WGCNA predicted biological pathways associated with ADAMTS5. (A) The cluster dendrogram of coexpression network modules. (B) The gene clusters obtained by WGCNA method. (C) Significantly enriched biological process of the co-expressed genes in turquoise module. (D) Significantly enriched biological process of the co-expressed genes in green module.
Discussion
ADAMTS5 has been reported to be associated multiple types of cancers with different role, 7,11 -13 which may functioned as anti-tumorigenic or pro-tumor role. However, the role of ADAMTS5 is still unclear in HCC, previous study has revealed that high ADAMTS5-expressed HCC had better OS than the patients without or with low expression of ADAMTS5, and the multivariate analysis indicated that ADAMTS5 expression was an independent prognostic factor for HCC patients. Our study demonstrated the opposite result, high ADAMTS5-expressed HCC had poorer OS than patients with low expression, ADAMTS5 expression is also associated with the development of HCC and is an independent prognostic factor for HCC patients. Moreover, the impact of ADAMTS5 expression on OS, DFS, RFS, DSS and PFI was specific for HCC among other 29 cancer types, which indicated the unique prognostic role of ADAMTS5 in HCC.
ECMs are highly specialized 3-D imensional macromolecular networks. which is composed of various proteins, such as collagen, proteoglycans, laminin, and fibronectin. 14 ECMs is dynamic and versatile to regulated cell function through direct or indirect means to regulate cell proliferation, migration, differentiation and invasiveness to maintain tissue homeostasis and functions, 14 -17 abnormal dynamics may lead to abnormal behaviors of cells in the local microenvironment through modulating the hallmarks of cancer, including dysregulated cellular metabolism, the evasion of immune destruction, sustaining proliferative signaling, evading growth suppressors, resisting cell death, enabling replicative immortality, inducing angiogenesis, activating invasion and metastasis. 18 Thus ECMs is critical for malignancy. During the normal condition, ECMs remodeling is tightly regulated through accumulation of inflammatory mediators, such as growth factors, cytokines and ECM-degrading enzymes, and ECM-degrading enzymes is considered as a crucial factor to control the ECMs remodeling activity, including matrix metalloproteinases (MMPs), plasminogen activators, a disintegrin and metalloproteinases (ADAMs), ADAMs with thrombospondin motifs (ADAMTS), 19 and abnormal ECMs remodeling is usually documented in previous studies related with various pathologies and is a crucial factor for initiation or progression of cancer. 15,20 Consistently, abnormal expression of many matrix-degrading enzymes are frequent in many types of human cancers. 21,22 Matrix-degrading enzymes can promote malignancy via various ways including degrade basement membranes, activate growth factors to enhance cell growth, invasion and neovascularization. 23,24 As an important member of matrix-degrading enzymes, ADAMTS family has been found to take part in various biological processes, including tissue morphogenesis, inflammation and angiogenesis. 25 As we know more about ADAMTS family, it surprised us that many of them(10 of 19) play an important role in the tumor pathological process, such as ADAMTS1, ADAMTS4, ADAMTS5, ADAMTS9 and so on. ADAMTS5 was first found in 1999 and was identified as a key enzyme in osteoarthritis by degrading aggrecan as an aggrecanase. 26 In recently, several studies have reported that aberrant ADAMTS5 expression in various cancers, including human glioblastomas, 6 human colorectal cancer, 7 human prostate cancer, 27 non-small cell lung cancer, 28 gastric cancer, 8 melanoma. 8 However, the functions of ADAMTS5 is not the same, they perform as an oncogene in human glioblastomas and non-small cell lung cancer. On the contrary, the ADAMTS5 functions as a tumor suppressor in breast cancer and human prostate cancer. Hence, ADAMTS5 may have different roles in different cancer tissues. To date, few study reported the association between HCC and ADAMTS5, Chongyi Li et al 10 indicated low ADAMTS5 expression is related with poor prognosis and functions as an anti-tumorigenic protein, and they reported that ADAMTS5 expression could downregulate VEGF level to inhibit tumor growth and angiogenesis in HCC cell line, lost expression of ADAMTS5 protein is associated with poor prognosis of HCC. However, multiple online websites present the opposite result that high ADAMTS5 expression is associated with poor survival in patients with HCC. Meantime, our result demonstrated that high ADAMTS5 was associated with poorer survival in HCC patients, and ADAMTS5 was an independent risk factor for survival in HCC. Our result had also suggested that the impact of ADAMTS5 on prognosis was specific for HCC. In addition, ADAMTS5 was associated with various types of survival evaluation index in HCC, including OS, RFS, DFS, DSS, PFI.
The GO enrichment analysis based on DEGs suggested that ADAMTS5 expression correlates with ECM and metabolic and energy regulation. The co-expressed genes associated with ADAMTS5 were mainly involved in extracellular matrix organization, extracellular structure organization and cell−matrix adhesion. Besides, the GSEA enrichment analysis revealed the genes in high ADAMTS5 expression group were mainly enriched in extracellular matrix (ECM), while the genes in low ADAMTS5 expression group were enriched in metabolic pathways. Furthermore, WGCNA method was performed to cluster gene module, and found genes in green module was highly positively associated with high ADAMTS5 group(7x10-5) and genes in turquoise module was highly positively associated with low ADAMTS5 group(1x10-6), GO enrichment analysis revealed genes in the turquoise module were significantly enriched in metabolic pathways, and extracellular matrix. These results implied that ADAMTS5 might participate in the status conversion from metabolic-dominant to extracellular matrix-dominant, and ADAMTS5 might affect the ECM biological process and consequently contributed to tumor progression. ECM could mediate microenvironment to regulate the metastasis, 29 it has reported that ADAMTS can degrade the ECM to promote invasion and metastasis, including ADAM-8, ADAM-12, ADAM-15, and ADAM-28 in non-small cell lung cancer, 30 -33 ADAMTS4 and ADAMTS5 in glioblastomas, 6 ADAM-9 in liver cancer. 34 Thus, there may be 2 reasons to provide explanation of the contradictory conclusions between the result of Chongyi Li’ study and our research, Firstly, tumor microenvironment is a complex macromolecular networks, ADAMTS5 could participate in tumor microenvironment to influence the various types of hallmarks of cancer, including dysregulated cellular metabolism, the evasion of immune destruction, sustaining proliferative signaling, evading growth suppressors, resisting cell death, enabling replicative immortality, inducing angiogenesis, activating invasion and metastasis, the previous study solely focus on HCC cells and sole cancer hallmark, 18 which may be different to explain the role of ADAMTS5 in tumor microenvironment. Secondly, as an important member of matrix-degrading enzymes, ADAMTS5 may promote HCC via many ways, including degrade basement membranes, activate growth factors to enhance cell growth, invasion and neovascularization. 23,24 It have not clearly determined the mechanisms might regulate the effects of ADAMTS5 on HCC, and further functional experiments are necessary to reveal the elusive mechanisms of ADAMTS5 in HCC. Considered of the controversial results, it is suggested meta-analysis to evaluate the association between ADAMTS5 and HCC. Finally, almost all the evidence comes from bulk tissue RNA-seq data, which indicated ADAMTS5 expression level could be influenced both by cancer cells and surrounding tumor microenvironment in our study, further search should be carried to the release the clear role of ADAMTS5 in HCC. Even though, we could cautiously postulate that ADAMTS5 is a novel biomarker predicting unfavorable prognosis for HCC.
Conclusion
To conclude, Our data suggested that ADAMTS5 is a prognosis-related gene for HCC, and the impact of ADAMTS5 on all survival was specific for HCC only. With a potential role in ECM and metabolic and energy regulation. ADAMTS5 is a promising therapeutic target for patients with HCC when taking subsite into consideration.
Footnotes
Acknowledgements
The authors thank the staff members of TCGA and GEO database.
Authors’ Note
Zhipeng Zhu: Conceptualization, Methodology, Software, Writing—Original Draft, Formal analysis, Supervision Jiuhua Xu: Validation Xiaofang Wu: Investigation Sihao Lin: Resources Lulu Li: Data Curation Weipeng Ye: Data Curation Zhengjie Huang: Funding acquisition, Writing—Review & Editing. GEO and TCGA databases is publicly available and open-access, and do not involve animal experiments and human specimens, ethical approval was not necessary.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from Project of Xiamen Scientific and Technological Plan (No. 3502Z20194005,3502Z20184020).
