Abstract
Background:
Recently, many studies have demonstrated that long non-coding RNAs (lncRNAs) are abnormally expressed in hepatocellular carcinoma (HCC) and may serve as a potential molecular biomarker to evaluate the prognosis of hepatocellular carcinoma. Therefore, we accomplished a meta-analysis built on current studies to assess the prognostic value of lncRNAs in hepatocellular carcinoma.
Methods:
The PubMed database was carefully searched to collect all eligible studies until February 20, 2019. The pooled hazard ratios (HRs) and 95% confidence intervals (CIs) of the overall survival, relapse-free survival, and progression-free survival were calculated to evaluate the prognostic significance of lncRNAs expression in hepatocellular carcinoma using Stata12.0 software. Heterogeneity, sensitivity analysis, and publication bias were also evaluated.
Results:
The results showed that the expression level of lncRNAs was significantly correlated with clinical outcomes. Abnormally expressed lncRNAs predicted poor overall survival (HR=2.19, 95% CI: 1.99-2.42, P<0.001; I2=44.7%, P=0.005), relapse-free survival (HR=2.68, 95% CI: 1.74-4.14, P<0.001; I2=0.0%, P=0.763) and progression-free survival of hepatocellular carcinoma patients (HR=2.44, 95% CI: 1.53-3.89, P<0.001; I2=0.0%, P=0.336). Statistical significance was also noted in subgroup meta-analyses that were stratified by follow-up time, cutoff value, and quality score. Moreover, the pooled results indicated that lncRNAs expression was significantly associated with tumor size (HR=1.48, 95% CI: 1.24-1.79), tumor number (HR=1.34, 95% CI: 1.08-1.66), and tumor node metastasis stage (HR=2.10, 95% CI: 1.48-2.99), but not liver cirrhosis and tumor differentiation (P>0.05).
Conclusions:
This meta-analysis indicates that lncRNAs are strongly associated with prognosis in hepatocellular carcinoma and may serve as a promising indicator for prognostic evaluation of patients with hepatocellular carcinoma. But larger clinical studies are needed to verify its feasibility.
Introduction
The global burden of cancer is increasing. Liver cancer in males is the fifth most common cancer diagnosed worldwide but the second most common cause of cancer death; in females it is the seventh most commonly diagnosed cancer and the sixth leading cause of cancer death. 1 Metabolic or nutritional disorders often result in hepatocellular carcinoma (HCC), the most common form of liver cancer in the world. 2 HCC represents the major histologic subtype of primary hepatocellular carcinoma, accounting for 90% of the global burden of liver carcinoma. 3 Surgical resection and liver transplantation are the main treatments for early HCC. 3 The prognosis of HCC depends on early detection, and the goal is to detect HCC early in order to apply curative therapies such as liver transplantation, surgical resection or ablation, and to improve the prognosis. 2 However, only a small proportion of patients (5%–10%) meet the criteria for resection, and patients undergoing hepatectomy may continue to have underlying inflammatory processes that require ongoing monitoring and treatment. 4 The overall 5-year survival rate for HCC is very low at 20%, which is associated not only with a delay in diagnosis but also with the late stage of disease. 3 HCC is most commonly diagnosed as advanced, primarily because of the slow growth of the tumor and the patient’s asymptomatic state early in the disease. 2 Sorafenib is a small-molecule oral multi-target tyrosine kinase inhibitor (TKI), which is the gold standard for first-line treatment authorized by U.S. Food and Drug Administration (FDA) for advanced liver cancer, which is difficult to operate on. 5 However, the prognosis of HCC patients treated with sorafenib remains poor due to its side effects and the patients’ increased resistance to the drug.5,6 Studies have shown that the MAPK pathway and BRAF signaling play a vital role in regulating the proliferation and survival of HCC cells: TKI inhibitors exert their activity on HCC cells that inhibit BRAF signaling, and sorafenib can block the MAPK pathway-mediated signal transduction. 7 For advanced cancers with poor prognoses, such as HCC, these two signaling pathways can be used to investigate their mechanisms to enhance the success rate of treatment. Owing to the lack of biomarkers for early diagnosis and prognosis, as well as the limitations of drug resistance, it is urgent to seek new biomarkers for the diagnosis and prognosis of HCC. Recent literature has reported that non-coding RNAs (ncRNAs) can modulate drug resistance in HCC, providing us with insights into the role of ncRNAs as hypothetical biomarkers and/or therapeutic targets for HCC in the future. 7
Long non-coding RNA (lncRNA), a kind of ncRNA with a length of more than 200 nucleotides, is a functional RNA molecule that cannot be translated into proteins and plays a significant role in numerous biological processes. It has been reported that lncRNAs are abnormally expressed in many cancers, which are closely linked to the occurrence, metastasis, and staging of tumors, 8 and may be used as non-invasive biomarkers and therapeutic targets. For instance, Qu et al. 9 identified that lncARSR is up-regulated in primary renal T-IC and is associated with poor prognosis in clear cell RCC (ccRCC). The regulatory mechanisms of lncRNAs in cancers are generally as follows: they interact with transcription factors to interfere with transcription; adsorb miRNAs as sponges; interact directly with proteins to regulate protein activity or with mRNAs to inhibit translation. 10 lncRNAs have been reported to change gene expression, which contributes to the pathophysiology of liver disease. 11 For instance, Wang et al. 12 found that lncRNA NEF may participate in the regulation of gastric cancer cell proliferation by regulating the expression of Runx1.
RNA sequencing and microarray gene profiling data from the Gene Expression Omnibus (GEO) showed that thousands of lncRNAs were differentially expressed in HCC tissues compared with normal tissues. 13 Some of these lncRNAs are significantly related to the prognosis of HCC patients and may play a critical role in the development, progression, and recurrence of HCC by regulating cell invasiveness. For instance, Cui et al. 13 confirmed that knockdown of SNHG7 and PVT1 impaired the invasion of HCC cells. Because HCC cells can evade immune surveillance and promote tumor growth and development by activating different immune checkpoint pathways, immune checkpoint inhibitors (ICIs), such as Nivolumab and pembrolizumab, have been approved for the second-line treatment of HCC. 14 Due to its low response rate, the combined application of TKIs, different ICIs, and other drugs will improve the anti-cancer effect. Transcription or post-transcriptional regulation of lncRNA expression can significantly influence the progression of the disease, and this process typically involves miRNAs acting as sponges. For instance, lncRNA MIAT promotes the proliferation and invasion of HCC cells through sponge miR-214. 15 lncRNA ANRIL may serve as a miR-122-5p sponge to regulate the function of miR-122-5p in HCC. 16
Numerous studies on lncRNAs have shown that lncRNAs may have potential as biomarkers for the diagnosis and prognosis of HCC. Previous studies have shown that knockdown of GHET1 inhibited the proliferation of HCC cells, and lncRNA GHET1, as a growth regulator, maybe a novel prognostic biomarker and therapeutic target for HCC. 17 Therefore, we conducted a meta-analysis to quantitatively evaluate the clinical feasibility of lncRNAs as new biomarker candidates and their useful insights into the prognosis of HCC.
Materials and methods
Data sources and search strategy
We collected all the relevant literature about the relationship of lncRNAs and the prognosis of hepatocellular carcinoma from the PubMed database. The detailed search strategy is as follows: (“long non-coding RNA”, “lncRNA”, “lncRNAs”, “Long noncoding RNAs”, “Long non-coding RNAs”) AND (“hepatocellular carcinoma”, “hepatocellular carcinomas”, “hepatocellular cancer”, “hepatocellular cancers”, “liver cell carcinoma”, “liver cell carcinomas”, “liver cancer”, “liver cancers”, “hepatic cancer”, “hepatic cancers”, “liver neoplasms”, “liver neoplasm”, “hepatic neoplasms”, “hepatic neoplasm”, “HCC”). All included studies were English articles.
Inclusion and exclusion criteria
Studies were incorporated for the following criteria: (a) The clinical related study; (b) research objects being human; (c) quantitative reverse transcription-polymerase chain reaction (qRT-PCR), reverse transcription-polymerase chain reaction (RT-PCR), or quantitative polymerase chain reaction (qPCR) was used to detect lncRNA expression; (d) patients undergoing liver resection; and (e) we could obtain sufficient data including overall survival (OS), the hazard ratio (HR), and 95% confidence interval (95% CI) directly from the full text. Studies were excluded for the following criteria: (a) being irrelevant to HCC; (b) not being a study on lncRNA; (c) letters, reviews, or meta-analysis; (d) short of full text; and (e) no prognosis.
Data extraction and quality assessment
According to inclusion and exclusion criteria, the characteristics of these collected articles were as follows: author, year of publication, the nation of the population enrolled, the type of lncRNAs and expression, HR and 95% CI (OS/PFS/relapse-free survival (RFS)), cutoff value, follow-up and quality score (Table 1). If the data were unavailable, we did our best to contact study authors to request the missing data. The information was extracted and recorded using a standardized form. We assessed the quality of each included study by using the Newcastle-Ottawa Scale (NOS) to obtain the quality score.
Characteristics of studies included in the meta-analysis.
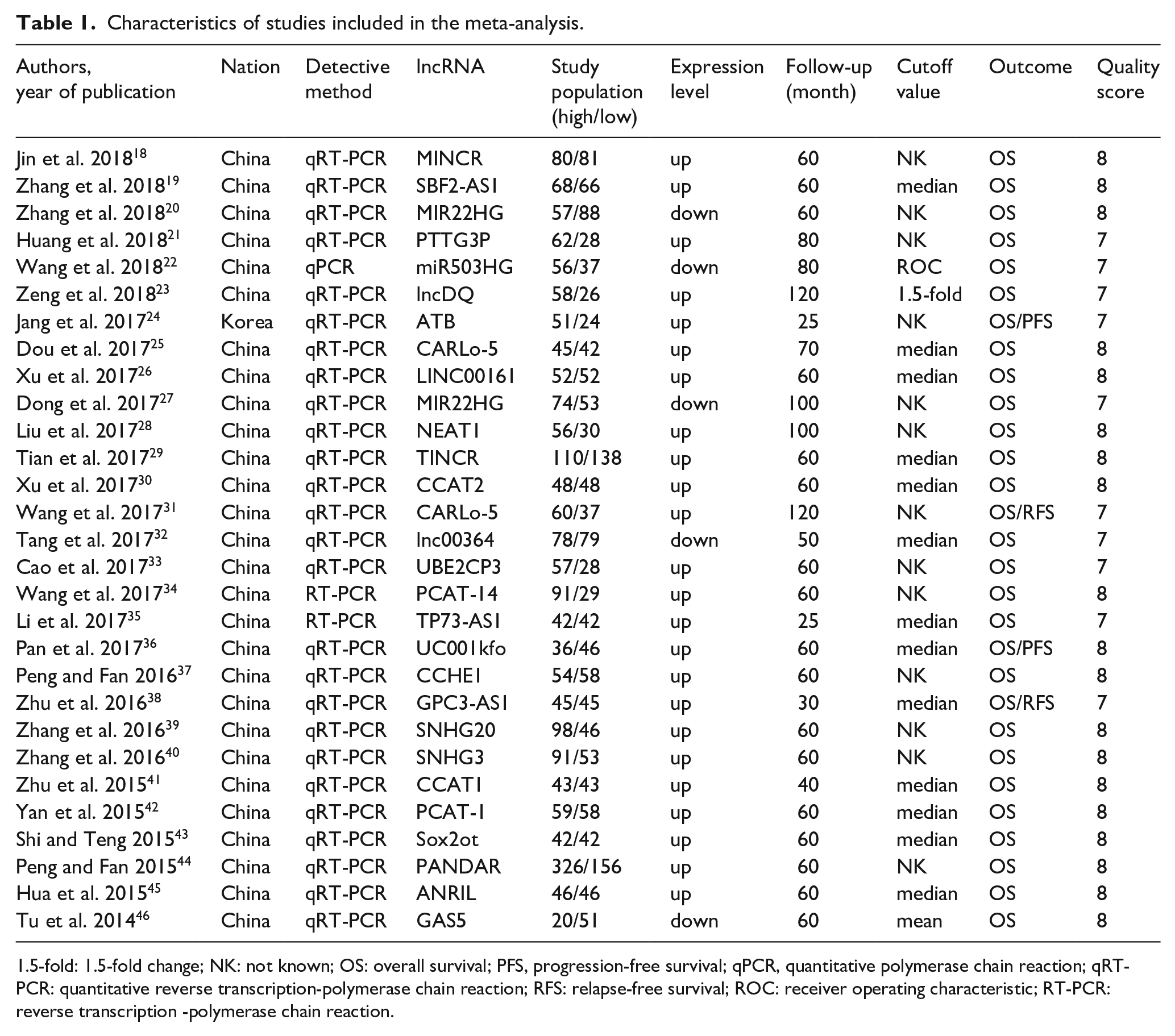
1.5-fold: 1.5-fold change; NK: not known; OS: overall survival; PFS, progression-free survival; qPCR, quantitative polymerase chain reaction; qRT-PCR: quantitative reverse transcription-polymerase chain reaction; RFS: relapse-free survival; ROC: receiver operating characteristic; RT-PCR: reverse transcription -polymerase chain reaction.
Statistical analysis
Stata12.0 software was used for the meta-analysis. Outcome variables were represented by HR and 95%CI. The results were shown in forest plots and P < 0.05 was considered statistically significant. The heterogeneity of the studies was indicated by I2 statistics, which was acceptable when I2 < 50%. When the heterogeneity test result was P > 0.10, the homogeneity could be considered, and the fixed effect model could be used to calculate the combined statistics. On the contrary, subgroup analysis should be carried out to analyze the causes of heterogeneity. Sensitivity analysis was also performed to evaluate the stability of the results. Publication bias was expressed by Begg’s funnel plot and Egger’s publication bias plot.
Results
Study inclusion and characteristics
A total of 362 articles were initially retrieved from the PubMed database and 1 of Chinese literature was excluded. After the titles and abstracts were scanned, 48 reviews, 16 meta-analysis, 7 animal experiments, and 127 irrelevant articles were removed. According to the inclusion and exclusion criteria, the full texts of the remaining 163 articles were read for further evaluation, and then 20 articles without the full text, 14 without original data, 34 without the prognosis, and 66 articles without sufficient data were excluded. Finally, 29 studies18-40, 41-46 were included in the meta-analysis (Figure 1).
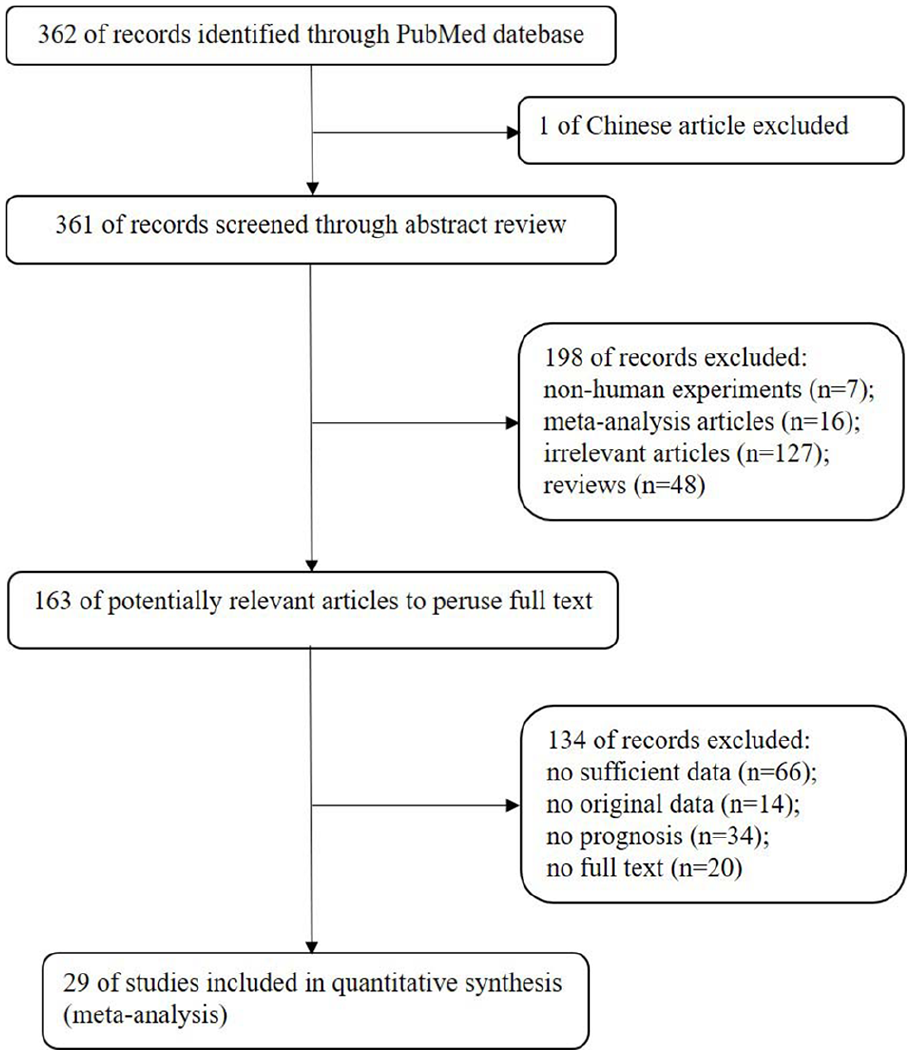
Flowchart of the strategy used for the selection of reports used in our analysis.
A total of 3577 patients from 29 studies were incorporated into the present meta-analysis, according to the inclusion and exclusion criteria. A total of 27 lncRNAs were studied in these articles, of which 24 were up-regulated and 5 down-regulated. In addition, there were two lncRNAs duplicated in the literature. All of the 3577 patients, who were treated with routine hepatectomy, were divided into two groups (high and low expression of lncRNA) according to the qRT-PCR, RT-PCR, or qPCR measurement results. The cutoff value for calculating lncRNAs expression was the mean, median, or other. No patient received chemotherapy or radiotherapy before surgery. All the articles included provided OS results, with two providing PFS and two providing RFS. Additionally, NOS score ranged from 0 to 9, and all included studies got NOS score of 7 or more, illustrating their high methodological quality.
Meta-analysis between lncRNA expression level and HCC survival
A total of 29 studies reported the OS of HCC based on different expressions of lncRNA. We extracted HRs and their associated 95% CIs of OS from the included studies, and HR >1 implies a poor prognosis. According to the results of the forest plot, these data are homogeneous (I2=44.7%, P=0.005), so the fixed-effect model was used to calculate the combined effect size (HR=2.19, 95% CI: 1.99-2.42, P<0.001) (Figure 2(a)), suggesting a significant association between lncRNAs expression level and OS in HCC patients. The prognostic significance of lncRNAs in RFS and PFS was evaluated in two studies with 187 patients and in two studies with 157 patients, respectively. LncRNAs was up-regulated in all four articles evaluated by RFS and PFS results. The results showed that the expression of lncRNAs was significantly associated with RFS (HR=2.68, 95% CI: 1.74-4.14, P<0.001; I2=0.0%, P=0.763)(Figure 2(b)) and progression-free survival (PFS) of HCC patients (HR=2.44, 95% CI: 1.53-3.89, P<0.001; I2=0.0%, P=0.336)(Figure 2(c)).
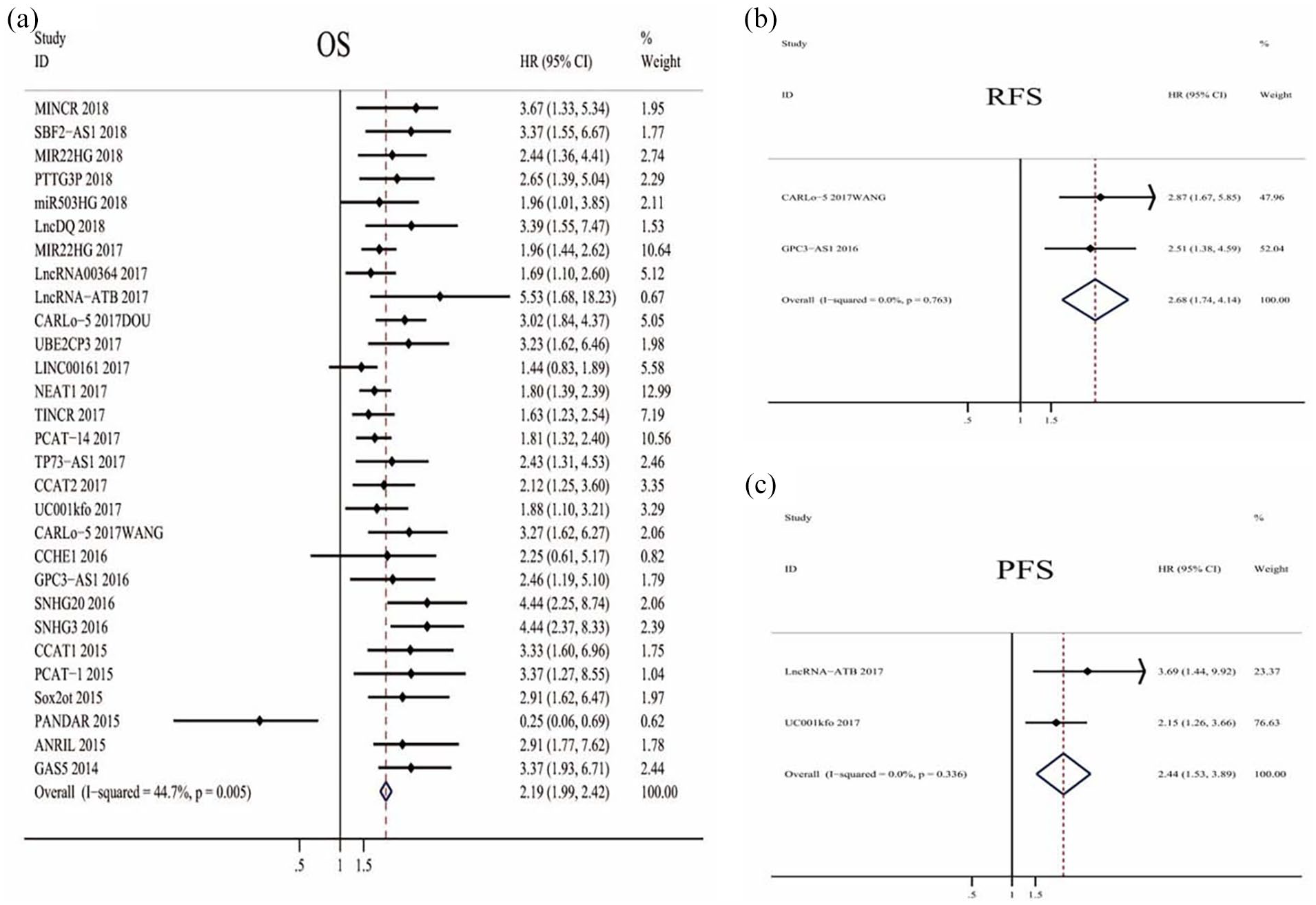
Meta-analysis of the pooled HRs of OS (a), RFS (b), and PFS (c) for HCC patients with increased lncRNAs transcription level depending on the treatments.
Subgroup analyses
Subgroup analyses were performed based on follow-up time, cutoff value, and quality score (all data are shown in Table S1). The follow-up time was divided into two categories, with 60 months as a threshold. In this analysis, the expression of lncRNAs was a strong prognostic marker for OS both in longer follow-up time (HR=2.18, 95% CI: 1.97-2.42) and shorter follow-up time (HR=2.28, 95% CI: 1.72-3.03). Stratified analysis of the cutoff value indicated a significant association between abnormal lncRNAs expression and OS in the studies using the median as cutoff value (HR=2.18, 95% CI: 1.97-2.42). Additionally, when the quality score was considered, there was a significant relevance between lncRNAs and OS in the two groups divided into 7 score (HR= 2.26, 95% CI: 1.90-2.70) or 8 (HR=2.16, 95% CI: 1.93-2.43) score.
Association of lncRNAs expression with clinicopathological features of HCC
The pooled HRs and 95% CIs of the clinicopathological characteristics in the included studies are shown in Table S2. The pooled results indicated that lncRNA expressions were significantly associated with tumor size (HR=1.48, 95% CI: 1.24-1.79, random-effect), tumor number (HR= 1.34, 95% CI: 1.08-1.66, fixed-effect), and tumor node metastasis (TNM) stage (HR=2.10, 95% CI: 1.48-2.99, random-effect). However, no significant relationship was observed between lncRNA expressions and liver cirrhosis (P>0.05), and tumor differentiation (P>0.05). Because of insufficient data, we could not evaluate the correlation between lncRNA expressions and other clinicopathological characteristics.
Publication bias and sensitivity analysis
For the meta-analysis of the association between lncRNA expression levels and OS, we used a funnel plot to observe whether there was bias in the results of a certain system evaluation or meta-analysis. Hence, Begg’s test (P=0.023) (Figure 3(a)) and Egger’s linear regression test (P=0.009) (Figure 3(b)) showed no significant publication bias in the study. Sensitivity analysis is a method to re-evaluate the combined effect size of the remaining studies except those whose effect size deviates significantly. We concluded that the direction and magnitude of OS estimation summary results were not significantly affected, indicating that none of the studies led to our results (Figure 4), which further confirmed the stability of our results.
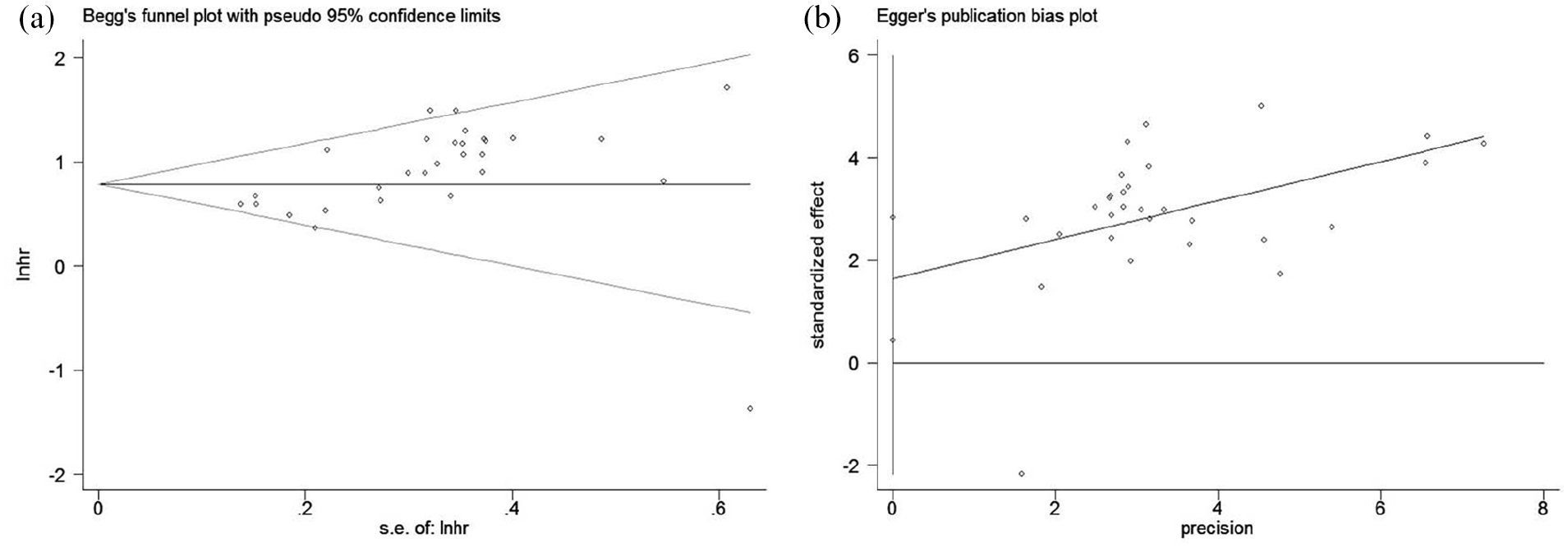
Publication bias analysis of the association between lncRNAs expression and OS of HCC were assessed by Begg’s funnel plot (a) and Egger’s publication bias plot (b).
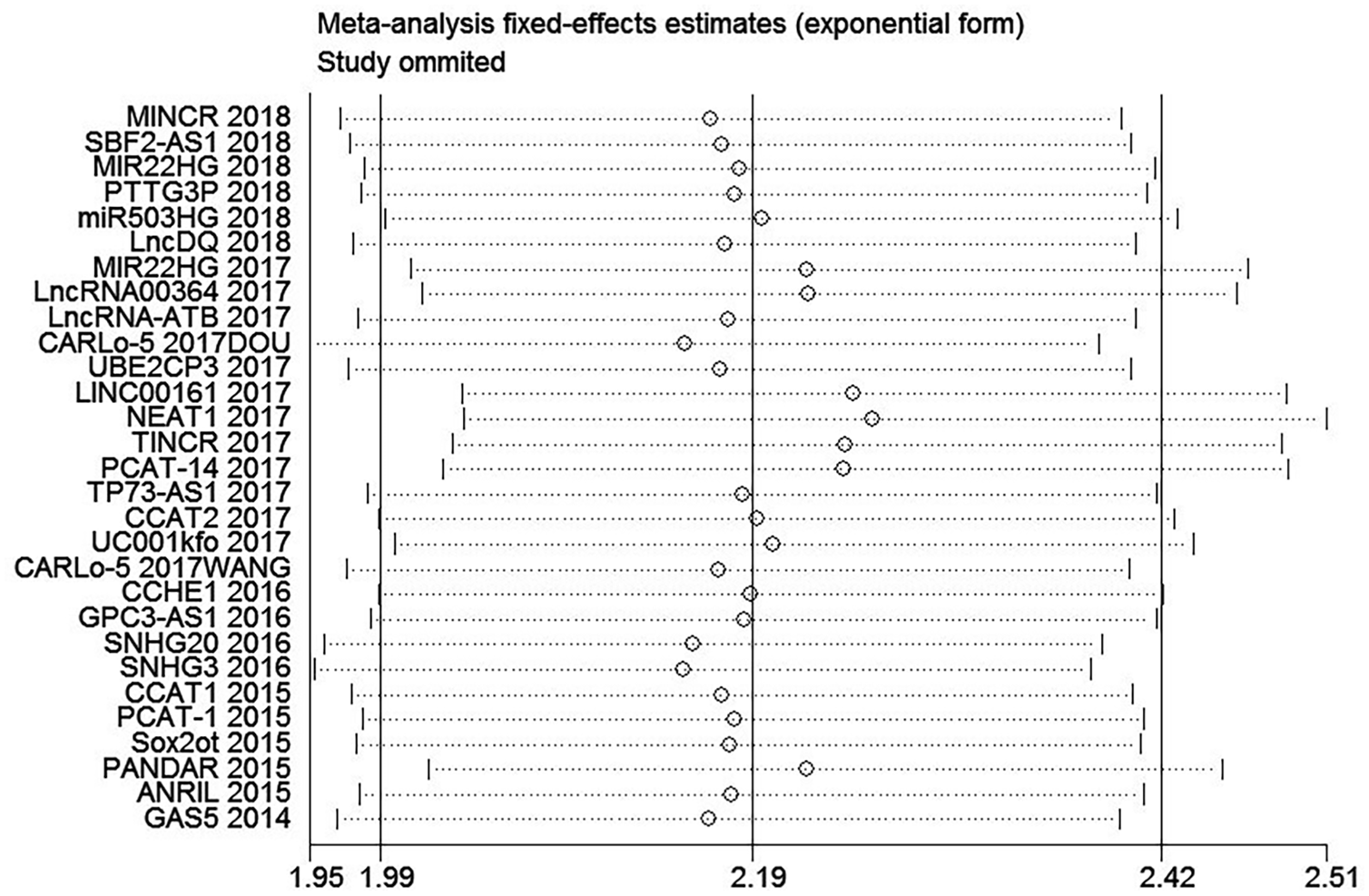
Sensitivity analysis of the association between lncRNAs expression and OS of HCC. Description: MINCR 2018: lncRNA + year; CARLo-5 2017DOU: lncRNA + year + author.
Discussion
Primary liver cancer, mainly HCC, is one of the most common malignant tumors in the world. 4 It is characterized by high morbidity, low survival rate, and poor prognosis after an operation. 3 Treatment options for this refractory disease are limited and most patients at the time of diagnosis are already too advanced to undergo surgical resection or liver transplantation. 47 Of course, when feasible, complete HCC resection is preferred 48 ; however, even with surgery, the prognosis is poor. Studies have demonstrated that among patients who underwent surgical HCC resection, the recurrence rate was about 70% in 5 years. 48 However, patients with advanced HCC are difficult to operate on and can only be treated with radiotherapy, chemotherapy, or medication. Drug therapy for HCC includes targeted drug therapy such as sorafenib, chemotherapy such as doxorubicin, and immunotherapy such as ICIs/monoclonal antibodies.5-6,14 However, as HCC patients gradually become resistant to drugs, recent studies have tended to evaluate several clinical and biological markers and their mechanisms to improve the choice of patients suitable for treatment with these drugs. Numerous studies found that several lncRNAs have been involved in drug resistance in HCC. lncRNA TUC338, ROR, and VLDLR, for example, have been found to be associated with HCC sorafenib resistance by transmitting intracellular signals through EVs.6,49 Also, the alteration of lncRNA HANR and lncARSR expression influenced the sensitivity of HCC to doxorubicin-mediated chemotherapy.6,49 A series of lncRNAs that have proved to be related to CSCs and EMT and can participate in various signaling pathways such as TGF-β, MAPK, and PI3K/AKT, including THOR, HULC, ANRIL, SNHG1, etc.6,49,50 Therefore, lncRNAs, as a new index for the diagnosis and prognosis of HCC, are crucial to the control of the occurrence, development, and recurrence of HCC.
In this meta-analysis, we attempted to collect all published papers to assess the prognostic potential of lncRNAs in HCC. We have determined that large amounts of lncRNAs are evaluated in the context of the HCC prognosis. Most of the included articles are only for one lncRNA to evaluate its relationship with the prognosis of HCC. Almost all published papers indicated that lncRNAs are statistically significant predictors of survival. However, few negative results have been reported in the literature, so publication bias is inevitable. It is well known that the published literature on prognosis and prognostic marker identification is characterized by poor methodological quality, significant publication bias, and broad heterogeneity in sample selection, such as follow-up and cutoff values.
Like other studies, we used the same indicators (OS, PFS, RFS), the same method, the same system analysis software to evaluate the prognostic and got the same results. For instance, Zheng et al. 51 included 40 retrospective studies from 2011 to 2016 in their meta-analysis, and concluded that many lncRNAs are associated with OS and RFS of the HCC patients. But in our 29 studies, we also assessed the prognostic value of lncRNAs and their association with clinicopathological features. The results showed that the abnormal expression of lncRNAs was significantly correlated with tumor size, tumor number, and TNM stage, but not with liver cirrhosis and tumor differentiation. In addition, when we stratified the prognostic effects of lncRNAs in the subgroup analysis with follow-up time, cutoff value, and quality score, the results did not change. All studies evaluated the relationship between lncRNA expression and OS, and there was a statistically significant correlation between them. A number of these studies have also assessed the association between lncRNA expression and other types of survival, including RFS and PFS. The difference in lncRNA expression was statistically significant, which further proved the reliability of our results. Therefore, lncRNAs may be a novel way to predict poor prognosis and clinicopathological characteristics of HCC. Also, Chen et al. 52 confirmed that lncRNAs have high sensitivity and specificity in HCC patients and can be invoked as diagnostic factors.
In all lncRNAs that we evaluated, MIR22HG have been evaluated in two articles, as was CARLo-5. Down-regulation of MIR22HG expression is associated with poor prognosis of HCC, while the expression of CARLo-5 in HCC is up-regulated and associated with poor survival. This suggests that CARLo-5 has a carcinogenic effect in HCC, and MIR22HG may be a tumor suppressor for HCC. In the exploration of the molecular mechanism of MIR22HG, it was found that the role of MIR22HG as a tumor inhibitor in HCC was partly to regulate the level of miR-10a-5p by acting as a competitive endogenous RNA. 53 In addition, CARLo-5 inhibited the expression of miR-200b through EZH2 and significantly inhibited tumor growth by knocking down CARLo-5 in vivo, 25 while CARLo-5 over-expression significantly enhanced cell proliferation, migration, and invasion in vitro. 31
We strictly established inclusion and exclusion criteria. All the studies included were cohort studies, and all the selected patients were followed-up for prognosis after liver resection (no other treatment). Nonetheless, our analysis inevitably has some limitations. First, given that this report is based on search results from only a single database (PubMed), relevant papers from other databases may have been missing. Second, most studies tend to report positive rather than negative results, and we only included English papers. Therefore, relevant articles that may meet the inclusion criteria may be ignored, which may lead to selection bias. Third, there is no consensus on cutoff values for distinguishing high and low lncRNA expressions. Fourth, our research only analyzes from the perspective of univariate survival analysis, without further verification by multivariate analysis, so it may be affected by other confounding factors. Also, because we looked at several cohorts in this article, some key statistics could not be measured, and significant deviations may have affected control selection and caused bias. Finally, all eligible studies were conducted in Asia, which may affect the broad applicability of these results, and most of the papers are from China, with only one from another country. Therefore, we did not conduct a subgroup analysis of ethnicity, which might be the source of between-study heterogeneity. In addition, Chinese papers tend to use small samples, but the probability of reaching statistical significance is much higher than in other literature. Chinese literature differs from the rest of the world in that its meta-analyses focus on individual candidate genes rather than on genome-wide studies. Nevertheless, our research results still have some reference value and research significance.
Conclusions
In conclusion, our results show that many lncRNAs related to the clinicopathological characteristics and survival of HCC were identified in this meta-analysis. Almost all lncRNAs were statistically significant predictors of the HCC prognosis. However, the importance of these correlations is difficult to determine due to many uncontrollable factors. Therefore, large-scale studies with standardized testing, analysis, and reporting processes are required to further verify the prognostic value of these lncRNAs in HCC.
Supplemental Material
Supplementary_materials_8 – Supplemental material for The prognostic significance of long non-coding RNAs in hepatocellular carcinoma: An updated meta-analysis
Supplemental material, Supplementary_materials_8 for The prognostic significance of long non-coding RNAs in hepatocellular carcinoma: An updated meta-analysis by Jie Liu, Yue Wei, Tao Wang, Zhexiao Zhang, Hairong Huang, Chenfei Wang, Huawen Li, Qing Jia, Hongfu Wu and Tangbin Zou in The International Journal of Biological Markers
Footnotes
Author contributions statement
JL, YW, TW, ZXZ, HRH, CFW, HWL, QJ, HFW, TBZ contributed to the conception and design of the study, literature review and analysis, drafting and critical revision and editing, and approval of the final version. JL, YW, and TW contributed equally to this article.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the National Natural Science Foundation of China (81874260), the Natural Science Foundation of Guangdong Province (2019A1515011567) and the Young Innovative Talents Projects of Universities in Guangdong (2018KQNCX097, 4SG19003Gj).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
