Abstract
BACKGROUND:
Hepatocellular carcinoma (HCC) is one of the most common and aggressive cancer worldwide and chronic infection of hepatitis B virus (HBV) serve as one of leading causes of HCC.
OBJECTIVE
: This study aimed to identify the novel long noncoding RNAs (lncRNAs) biomarkers for HBV-associated HCC.
METHODS:
The lncRNA and mRNA expression profiles of HCC patients with HBV infection were downloaded from The Cancer Genome Atlas. The differentially expressed lncRNAs (DElncRNAs) and mRNAs (DEmRNAs) between HCC and adjacent tissues were identified. The optimal diagnostic and prognostic lncRNA biomarkers for HCC were identified by using feature selection procedure and classification model. Functional annotation of DEmRNAs co-expressed with these lncRNAs biomarkers were performed. Receiver operating characteristic (ROC) curve and survival analysis of these lncRNAs biomarkers were performed. qRT-PCR validation was performed.
RESULTS:
A total of 82 DElncRNAs and 805 DEmRNAs between HBV-associated HCC and normal tissues were identified. CAPN10-AS1, LINC01093, RP5-890E16.2, FENDRR and C17orf82 were selected as optimal diagnostic and prognostic lncRNA biomarkers for HBV-associated HCC that were co-expressed with 105, 86, 70, 30 and 1 DEmRNAs, respectively. Based on the DEmRNAs co-expressed with these five lncRNAs biomarkers, Jak-STAT signaling pathway and retinol metabolism were two significantly enriched pathways. The result in qRT-PCR validation were consistent with our analysis based on TCGA, generally.
CONCLUSIONS:
This study identified five potential lncRNAs biomarkers for HBV-associated HCC with great diagnostic and prognostic value and provided clues for their functions in HBV-associated HCC.
Introduction
Hepatocellular carcinoma (HCC) is one of the most common and aggressive malignant tumors worldwide that is the third leading cause of cancer-related deaths all over the world [1]. Chronic infection of hepatitis B virus (HBV) serve as one of leading causes of HCC worldwide. The etiology and pathogenesis of HCC remain elusive. Due to lack of specific clinical manifestations the majority of patients with HCC were diagnosed at the late stage and over 4/5 of these patients are left poor prognosis [2]. Hence, it is urgent to develop novel diagnostic and prognostic biomarkers and effective therapeutic targets.
Long noncoding RNAs (lncRNAs) are a class of non-coding RNAs with more than 200 nucleotides which was broadly distributed in human genome [3]. LncRNAs have been indicated to involve with various biological processes and modulate gene expression with cis or trans effects [4]. Accumulated evidence demonstrated that lncRNAs involve with various cancer including HCC, colorectal cancer and breast cancer [5, 6, 7]. Several lncRNAs such as MEG3, MVIH, HULC, HOTAIR and MTIDP were reported to be associated with HCC [5, 6].
In this present study, the lncRNA and mRNA expression profile of a large number of patients with HBV-associated HCC we obtained from TCGA and the differentially expressed lncRNAs (DElncRNAs) and mRNAs (DEmRNAs) between HCC and normal tissues were identified. The optimal diagnostic and prognostic lncRNA biomarkers for HBV-associated HCC were identified by using feature selection procedure and classification model. The functions of these lncRNA biomarkers in HBV-associated HCC were further explored by functional annotation of their co-expressed mRNAs.
Materials and methods
Integrated profiles in TCGA
The Cancer Genome Atlas (TCGA) has produced huge amounts of cancer genomics data, which provide unprecedented opportunities to reveal molecular mechanisms of cancer. The clinical data of patients with HCC was downloaded from TCGA (
Identification DElncRNAs and DEmRNAs between HCC and normal tissues
First, we filtered difficultly detected mRNAs (mRNAs with read count value
Then, the DElncRNAs and DEmRNAs between HCC and normal tissues were calculated via R-bioconductor package DESeq2. Benjamini and Hochberg method was used to conduct multiple comparisons to obtain the false discovery rate (FDR). The threshold for the expression of both DElncRNAs and DEmRNAs was FDR
Identification of optimal diagnostic lncRNA biomarkers for HCC
LASSO algorithm were fit using the glmnet package (
To further identify the optimal lncRNA with diagnostic value for HCC, feature selection procedures were conducted as follows: 1). By using the random forest analysis, the importance value of each DElncRNA ranked according to the mean decrease in accuracy; 2). The optimal number of features was found by subsequently adding one DElncRNA at a time in a topdown forward-wrapper approach; 3). By using support vector machine (SVM) at each increment, the accuracy was assessed and the optimal diagnostic lncRNA biomarkers for HCC were identified. Hierarchical clustering analysis of these diagnostic lncRNA biomarkers for HCC were conducted by using R package “pheatmap”. Based on the obtained optimal diagnostic lncRNA biomarkers for HCC, we used e1071 package (
Hierarchical clustering analysis of DElncRNAs and the top 100 DEmRNAs between HCC and normal tissues. A) DElncRNAs; B) The top 100 DEmRNAs. Row and column represented DElncRNAs/DEmRNAs and tissue samples, respectively. The color scale represented the expression levels.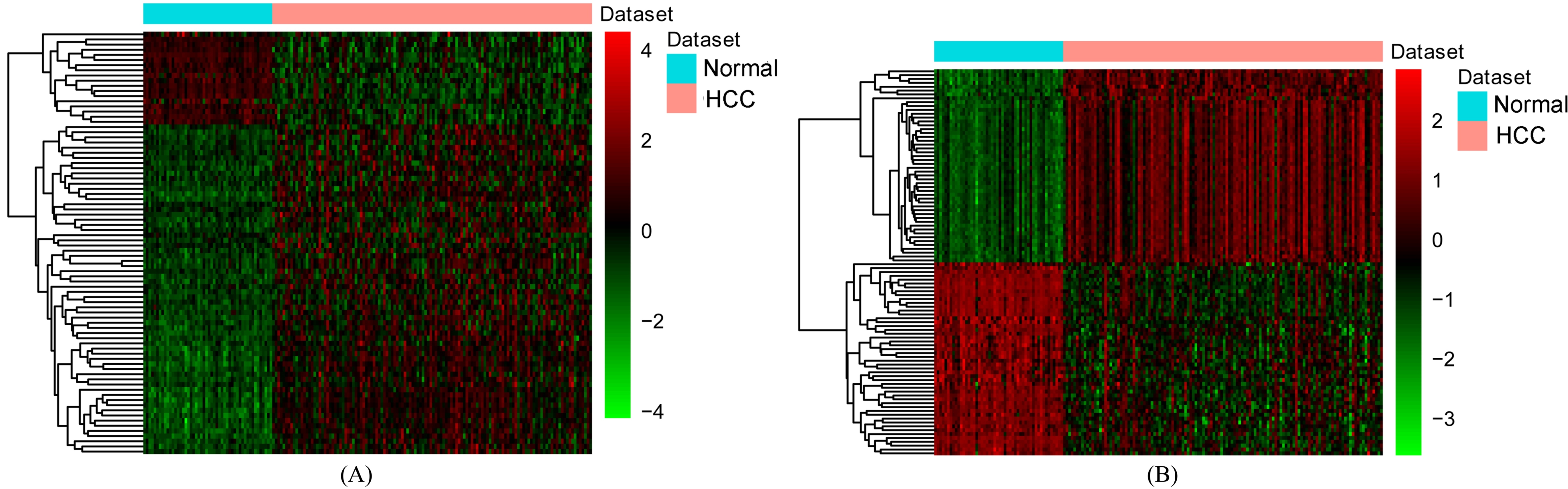
By using the survival (
DElncRNA-DEmRNA co-expression analysis
WGCNA (Weighted correlation network Analysis) method was employed to construct the DElncRNA-DEmRNA co-expression network as follows: 1) The pairwise Pearson correlation coefficients between these optimal lncRNAs biomarkers and DEmRNAs were calculated; 2) The threshold for DElncRNA-DEmRNA co-expression pairs was
Functional annotation
To uncover the biological functions and detect the potential pathways of the optimal DElncRNAs with diagnostic value for HCC, functional annotation, including Gene Ontology (GO) classification and pathway enrichment analysis of DEmRNAs co-expressed with these optimal DElncRNAs with diagnostic value for HCC was conducted by using online software GeneCodis. Statistical significance was defined as false discovery rate (FDR)
Confirmation by qRT-PCR
To verify the expression of DEmRNAs and DElncRNAs between HCC and normal controls, we collected 4 pairs of matched HBV-associated HCC cancer and adjacent normal tissues. Based on manufacturer’s protocol, total RNA of samples was isolated with the Trizol reagent (Invitrogen, Carlsbad, CA, USA). The reverse transcription of mRNA and lncRNA was performed by using the FastQuant RT Kit (Tiangen, China). The qRT-PCR reactions were performed with SuperReal PreMix Plus (Invitrogen, Carlsbad, CA, USA) in ABI 7500 Real-time PCR Detection System. Relative gene expression was analyzed with
Results
DEmRNAs and DElncRNAs between HCC and normal tissues
The clinical data of 377 patients with HCC were obtained from TCGA. The 124 HCC patients with chronic HBV infection were enrolled in this study. The media age of these 124 patients was 60 years old. Male and female of them were account for 66.1% and 33.9%, respectively. The ethnicity of them was Hispanic or Latino (93.5%) and Not Hispanic or Latino (2.1%). The tumor stage of them was stage I (28.2%), stage II (27.4%), stage III A–B (40.3%) and stage IV A–B (1.6%), respectively. The mRNA and lncRNA expression profiles of 124 HBV-associated HCC tumor tissues and 50 adjacent normal tissues were obtained. After filtering the difficultly detected mRNAs and lncRNAs, a total of 15260 mRNAs and 3195 lncRNAs were retained for analysis.
Identification of optimal lncRNA biomarkers for HCC. A) The importance value of each DElncRNA ranked according to the mean decrease in accuracy by using the random forest analysis; B) The variance rate of classification performance when increasing numbers of the predictive DElncRNAs; C) Hierarchical clustering analysis of HCC-specific five lncRNAs biomarkers (CAPN10-AS1, LINC01093, RP5-890E16.2, FENDRR and C17orf82) between HCC and normal tissues; Box-plot displayed the expression levels of CAPN10-AS1 (Fig. 2D), LINC01093 (Fig. 2E), RP5-890E16.2 (Fig. 2F), FENDRR (Fig. 2G) and C17orf82 (Fig. 2H) between HCC and normal tissues. The X-axis represented normal and case (HCC) groups. The Y-axis represented gene expression levels.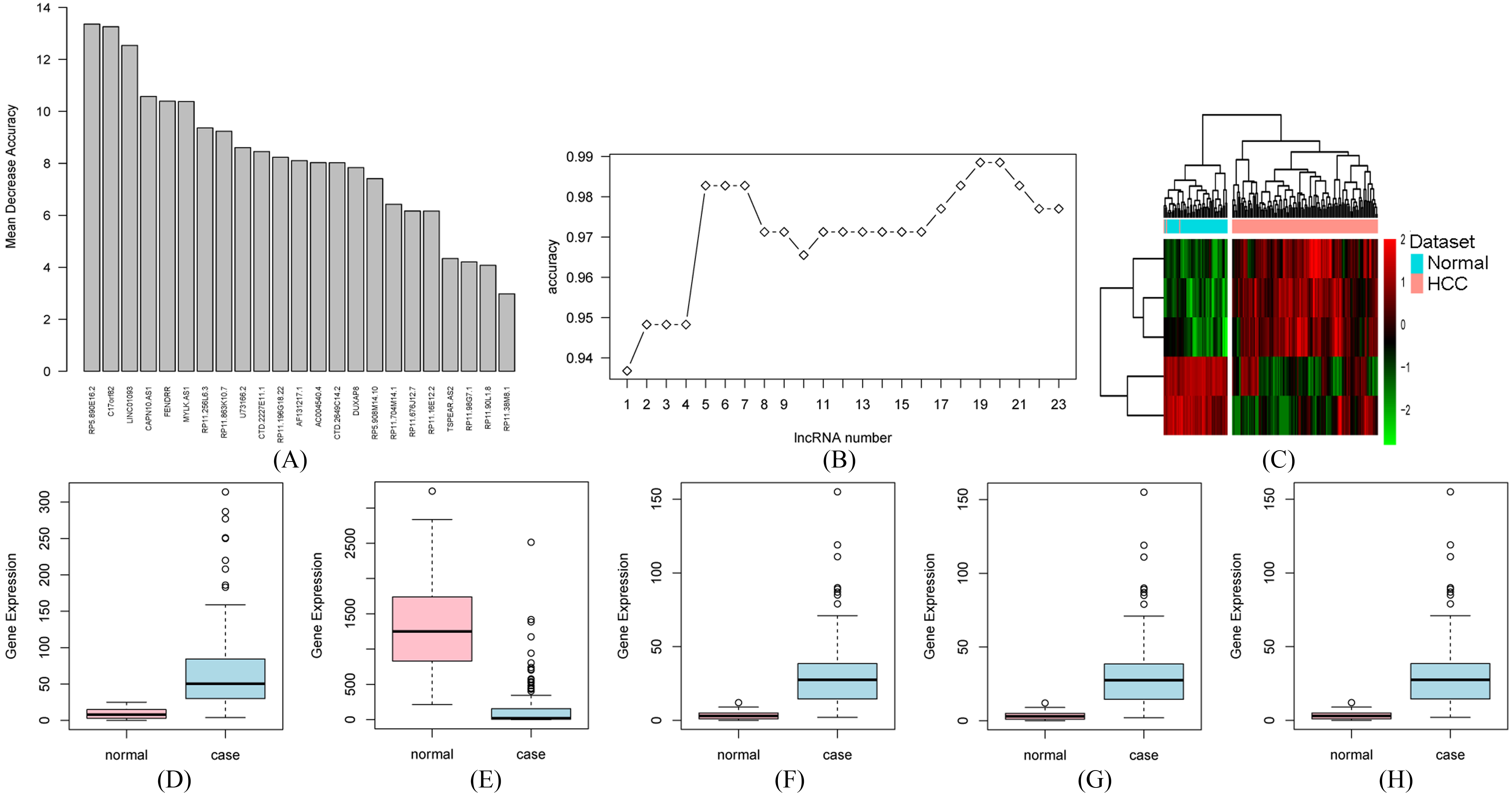
A total of 82 DElncRNAs (18 down-regulated and 64 up-regulated lncRNAs, Supplemental Table S1) and 805 DEmRNAs (159 down-regulated and 646 up-regulated mRNAs, Supplemental Table S2) between HCC and normal tissues were identified with FDR
DElncRNAs between HCC and normal tissues after reduced dimensions of data
Five optimal diagnostic lncRNA biomarkers for HCC
ROC and survival analysis of five HCC-specific lncRNA biomarkers. The ROC results of individual CAPN10-AS1 (Fig. 3A), FENDRR (Fig. 3B), LINC01093 (Fig. 3C), RP5-890E16.2 (Fig. 3D), and C17orf82 (Fig. 3E) and their combination based on support vector machine (SVM) model (Fig. 3F), decision tree model (Fig. 3G) and random forest (Fig. 3H). The x-axis shows 1-specificity and y-axis shows sensitivity. Figure 3I indicated that low sum-expression of these five lncRNAs were significantly associated with lower survival rate in patients with HBV-associated HCC (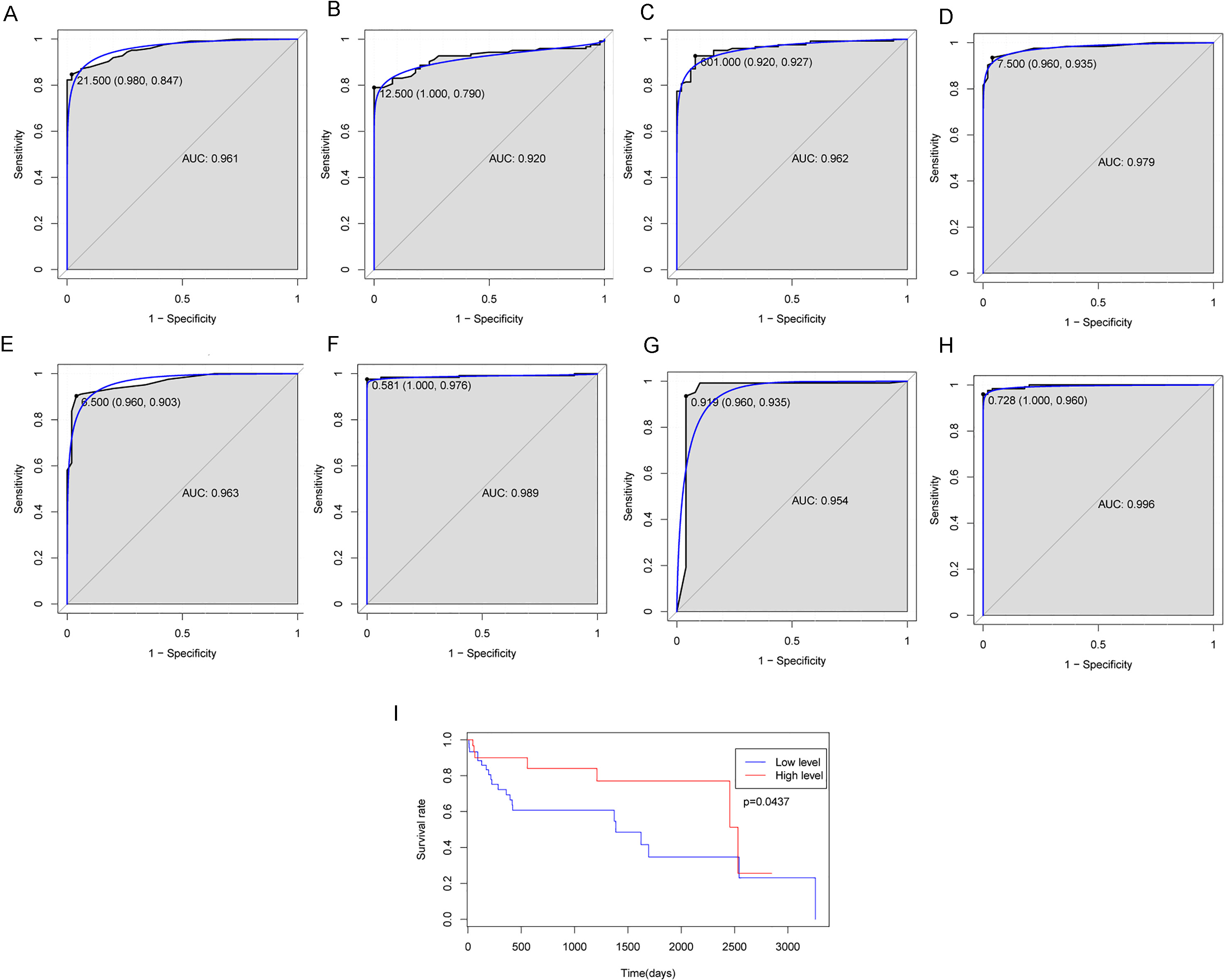
Based on reduced dimensions of the data, we obtained 23 DElncRNAs between HCC and normal tissues by using LASSO algorithm (Table 1). All these 23 DElncRNAs were ranked according to the mean decrease in accuracy by using the random forest analysis (Fig. 2A). Ten-fold cross-validation result indicated that the average accuracy rate of 5 DElncRNAs reached the highest point for the first time (Fig. 2B) and we defined these 5 DElncRNAs including three up-regulated DElncRNAs (RP5-890E16.2, CAPN10-AS1 and C17orf82) and two DElncRNAs (LINC01093 and FENDRR) as the optimal diagnostic lncRNA biomarkers for HCC (Table 2). Hierarchical clustering analysis of these 5 DElncRNAs between HCC and normal tissues were displayed in Fig. 2C. Box-plot displayed the expression levels of these 5 DElncRNAs between HCC and normal tissues (Fig. 2D–H).
The AUC of CAPN10-AS1, FENDRR, LINC01093, RP5-890E16.2, and C17orf82 was 0.961, 0.920, 0.962, 0.979 and 0.963, respectively (Fig. 3A–E). The SVM, decision tree and random forest models were established based on these 5 DElncRNAs between HCC and normal tissues. The AUC of this SVM model was0.989 and the sensitivity and specificity of this SVM model was 97.6% and 100.0% (Fig. 3F). The AUC ofdecision tree model was 0.954 and the sensitivity and specificity of this decision tree model was 93.5% and 96.0% (Fig. 3G). The AUC of this random forestmodel was 0.989 and the sensitivity and specificity of random forest model was 96.0% and 100.0% (Fig. 3H). Taken together, the AUC of all these five lncRNAs and their combination was more than 0.9 which suggested that up-regulated RP5-890E16.2, up-regulated CAPN10-AS1, up-regulated C17orf82, down-regulated LINC01093, down-regulated FENDRR and their combination were associated with HBV-associated HCC and could predict the occurrence of HBV-associated HCC, respectively.
HCC-specific DElncRNA-DEmRNA co-expression network. The ellipses and rhombuses were represented the DEmRNAs and DElncRNAs, respectively. Red and blue color represented up- and down-regulation, respectively.
Significantly enriched GO terms and KEGG pathways of DEmRNAs co-expressed with DElncRNAs in HCC. The x-axis shows 
Confirmation by qRT-PCR. The expression of selected DElncRNAs and DEmRNAs in HBV-associated HCC tissues compared to adjacent normal tissues. The x-axis shows DElncRNAs and DEmRNAs and the y-axis shows log2 (Fold change). *indicated 
The overview of major finding in this present study.
Based on survival analysis, none of these five lncRNAs has prognostic value for HBV-associated HCC (
DElncRNA-DEmRNA co-expression analysis
A total of 292 DElncRNA-DEmRNA co-expression pairs consisted of 236 DEmRNAs and these five optimal lncRNA biomarkers for HCC were obtained. CAPN10-AS1, LINC01093, RP5-890E16.2, FENDRR and C17orf82 were co-expressed with 105, 86, 70, 30 and 1 DEmRNAs, respectively (Fig. 4).
Functional annotation
Based on the Functional annotation of DEmRNAs co-expressed with optimal DElncRNAs biomarkers for HCC, mitotic cell cycle (FDR
Confirmation by qRT-PCR
We performed the confirmation of four DEmRNAs (GHR, SOCS3, CYP4A11 and CENPF) and four DElncRNAs (CAPN10-AS1, LINC01093, RP5-890E16.2 and FENDRR) by qRT-PCR. CENPF and two DElncRNAs (CAPN10-AS1 and RP5-890E16.2) were up-regulated while two DElncRNAs (LINC01093 and FENDRR) and three DEmRNAs (GHR, SOCS3 and CYP4A11) were down-regulated in HBV-associated HCC tissues compared to adjacent normal tissues (Fig. 6). The qRT-PCR results were consistent with our bioinformatics analysis based on TCGA. The overview of major finding in this present study were displayed in Fig. 7.
Discussion
HCC is a common aggressive cancer worldwide and the mortality rate for HCC has increased at a greater rate compared to all other forms of cancer [9]. Exploring the accurate and specific biomarkers of HCC is essential. Accumulated evidence has indicated that lncRNAs play crucial roles in the progress of HCC.
In this present study, we obtained the DElncRNAs and DEmRNAs between HCC and normal tissues based on the lncRNA and mRNA expression profile of a large number of patients with HBV-associated HCC derived from TCGA. By using feature selection procedure and classification model five optimal diagnostic lncRNA biomarkers (CAPN10-AS1, LINC01093, RP5-890E16.2, FENDRR and C17orf82) for HBV-associated HCC were identified.
Down-regulated LINC01093 has been found in HCC tissues and cell lines in recent study [2]. Moreover, lower expression of LINC01093 was reported to be associated with poor prognosis of patients with HCC [2]. In this present study, LINC01093 was down-regulated in HCC tissues compared to normal tissues which provide evidence for previous study and emphasize its diagnostic value for HCC.
FENDRR (also known as FOXF1-AS1) is a long non-coding RNAs (lncRNA) that mediates dsDNA-RNA triplex formation to epigenetically regulate the expression of its target gene by recruiting polycomb repressive complexe 2 [10, 11]. Recently, down-regulated FENDRR has been found in various cancers including gastric cancer, lung adenocarcinoma and infantile hemangioma [10, 12, 13]. FENDRR could modulate the migration and invasion of gastric cancer cells by regulating the expression of fbronectin1 secreted matrix metalloproteinase (MMP) 2/(MMP) 9 [10]. Furthermore, low expression of FENDRR was closely associated with poor prognosis of patients with gastric cancer [10]. In this present study, FENDRR was found to be down-regulated in patients with HCC as well, which suggested its crucial role in process of HCC.
Moreover, the ROC results indicated the great diagnostic value of these five diagnostic lncRNA biomarkers (CAPN10-AS1, LINC01093, RP5-890E16.2,FENDRR and C17orf82) and their combination for HCC. Based on the survival analysis, combination of these five DElncRNAs owns great prognostic value for HCC as well. These finding provided new clues for developing novel diagnostic and prognostic biomarkers for HCC.
Except LINC01093 and FENDRR, three up-regulated DElncRNAs (CAPN10-AS1, C17orf82 and RP5-890E16.2) have never been reported in previous studies and their biological function remain unknown. Accumulated evidence has indicated that lncRNAs play key roles in regulating the expression levels of genes and proteins and participate in a variety of biochemical processes and diseases [14, 15, 16]. LncRNA-mRNA co-expression analysis was the most popular approach to identify potential target genes of lncRNAs and further explore the biological functions of lncRNAs in various disease [17, 18]. Based on these five HCC-specific lncRNA biomarkers, we constructed the DElncRNA-DEmRNA co-expression network and performed the functional annotation of DEmRNAs co-expressed with five lncRNAs.
Three DEmRNAs (GHR, SOCS2 and TSLP) co-expressed with LINC01093 and two DEmRNAs (LIFR and SOCS3) co-expressed with FENDRR were all DEmRNAs enriched in Jak-STAT signaling pathway. Previous studies have indicated that JAK/STAT signaling pathway was up-regulated in HCC tissues and could directly regulated the HCC tumorogenesis induced by HBV [19, 20]. Moreover, as a negative regulator of JAK/STAT signaling, SOCS3 was found to be silenced in most of HCC tumors [19, 21]. We speculated that both LINC01093 and FENDRR might involve with HCC tumorogenesis by regulating JAK/STAT signaling pathway.
CYP4A11, cytochrome P450, family 4, subfamily A, polypeptide 11, encodes a member of the cytochrome P450 superfamily of enzymes that is the major fatty acid omega-hydroxylase in human liver [22]. Recent study indicated that CYP4A11 was closely associated with metastasis risk and prognosis of HCC [23]. In this present study, CYP4A11 was a down-regulated gene that enriched in pathway of retinol metabolism. Retinoic acids (RAs), the metabolites of retinol, were reported to make contribution for prevention of liver fibrosis and carcinogenesis [24] which could inhibit the expression and activity of CYP4A11 [22]. Furthermore, higher pre-diagnostic serum retinol were closely associated with decreased risk of HCC with chronic HBV infection [25, 26]. In this present study, both LINC01093 and RP5-890E16.2 were co-expressed with CYP4A11. Moreover, five other DEmRNAs (CYP4A22, LRAT, CYP1A2, CYP2C19 and UGT2B7) enriched in pathway of retinol metabolism were co-expressed with LINC01093 as well. We make a hypothesis thatLINC01093 and RP5-890E16.2 might play roles in HCC by regulating the expression of CYP4A11 or retinol metabolism.
In this present study, CENPF, centromere protein F and FOXM1, forkhead box M1 were two up-regulated genes that co-expressed with CAPN10-AS1. Up-regulation of CENPF and FOXM1 in HCC was found in HCC in previous study as well [27, 28]. CENPF and FOXM1 served regulators of the malignancy of prostate cancer. Knockdown of CENPF and FOXM1 suppressed the proliferation of prostate cancer cells and tumor growth in cell-line-derived xenografts synergistically [27, 28]. A similar synergistic cooperation was speculated to exist in tumorogenesis of HCC and CAPN10-AS1 might involve with HCC by regulating it.
LGI3, leucine rich repeat LGI family member 3 was the only one co-expressed gene with C17orf82. Recent microarray study found the association between the expression of LGI3 and the prognosis of various cancers including brain, colorectal and lung cancer [29] which suggested that C17orf82-LGI3 co-expression might involve with HCC as well.
In conclusion, our study identified five DElncRNAs with great diagnostic and prognostic value for HBV-associated HCC which made contribution for developing the potential biomarkers for HBV-associated HCC. Function annotation of DEmRNAs co-expressed with these five lncRNA biomarkers provided clues for the functions of these five DElncRNAs in HCC which was benefit for exploring the mechanism of HBV-associated HCC. In our following research, whether these five lncRNAs were differentially expressed in bloods between patients with HBV-associated HCC and normal controls will be explored. Moreover, to detect whether these five lncRNAs have diagnostic value in early stage of HBV-associated HCC, further research with large sample size was needed to detect the expression of these five lncRNAs among individuals with different degree of HBV-associated HCC risk.
Footnotes
Acknowledgments
This study was supported by the Academician Workstation Fund of Zheng Shusheng and the Foundation of Zhejiang Health Department (2018KY900 and 2017KY 902). We thank Beijing Medintell Bioinformatic Technology Co., LTD for assistance in data analysis.
Conflict of interest
None declared.
Supplementary data
The supplementary files are available to download from
