Abstract
BACKGROUND:
Early recurrence is the main obstacle for long-term survival of hepatocellular carcinoma (HCC) patients after curative resection.
OBJECTIVE:
We aimed to develop a long non-coding RNA (lncRNA) based signature to predict early recurrence.
METHODS:
Using bioinformatics analysis and quantitative reverse transcription PCR (RT-qPCR), we screened for lncRNA candidates that were abnormally expressed in HCC. The expression levels of candidate lncRNAs were analyzed in HCC tissues from 160 patients who underwent curative resection, and a risk model for the prediction of recurrence within 1 year (early recurrence) of HCC patients was constructed with linear support vector machine (SVM).
RESULTS:
An lncRNA-based classifier (Clnc), which contained nine differentially expressed lncRNAs including AF339810, AK026286, BC020899, HEIH, HULC, MALAT1, PVT1, uc003fpg, and ZFAS1 was constructed. In the test set, this classifier reliably predicted early recurrence (AUC, 0.675; sensitivity, 72.0%; specificity, 63.1%) with an odds ratio of 4.390 (95% CI, 2.120–9.090). Clnc showed higher accuracy than traditional clinical features, including tumor size, portal vein tumor thrombus (PVTT) in predicting early recurrence (AUC, 0.675 vs 0.523 vs 0.541), and had much higher sensitivity than Barcelona Clinical Liver Cancer (BCLC; 72.0% vs 50.0%), albeit their AUCs were comparable (0.675 vs 0.678). Moreover, combining Clnc with BCLC significantly increased the AUC, compared with Clnc or BCLC alone in predicting early recurrence (all
CONCLUSIONS:
Our lncRNA-based classifier Clnc can predict early recurrence of patients undergoing surgical resection of HCC.
Introduction
Hepatocellular carcinoma (HCC), is one of the most common types of cancer worldwide and is the second leading cause of cancer deaths in men [1]. Clinically, resection is the most widely used potentially curative treatment for HCC patients [2]. However, a high incidence of recurrence leads to the poor survival of HCC patients [3, 4]. It has been reported that the recurrence rate peaked at 1 year after HCC resection, and this recurrence pattern was mainly due to invasion or metastasis from the primary tumors [5]. Therefore, it is urgent to establish a classifier that can effectively stratify the HCC patients according to the risk of early recurrence and consequently enable optimal postoperative adjuvant therapies.
Although traditional staging systems, such as Barcelona Clinical Liver Cancer (BCLC) system, were useful for treatment guidance, they are inadequate for predicting survival who undergo hepatic resection [6]. Considerable efforts have been devoted to identifying molecular markers for predicting the recurrence of HCC patients, including immunohistochemical markers, protein-coding gene markers, microRNAs, and epigenetic biomarkers [7, 8, 9, 10]. However, due to the inconsistency of analytical platforms and complex etiologies of HCC, these molecular markers have not been widely introduced into clinical practice as expected initially, and most of them still need to be validated.
LncRNAs are RNAs longer than 200 nucleotides (nt) and lacking protein-coding abilities [11]. Abnormal expression of lncRNAs has been implicated in tumor development and progression, and accumulating studies have shown that the aberrantly expressed lncRNAs could potentially mark the spectrum of disease progression, and some of them have exerted diagnostic or prognostic values [12, 13, 14, 15]. However, whether incorporating multiple lncRNAs could improve the prediction of early recurrence for HCC remains unknown.
In this study, a screen for differentially expressed lncRNAs between HCCs and the adjacent normal liver tissues resulted in the identification of 15 upregulated and 5 downregulated lncRNAs. We further investigated the expression profiles of the 20 dysregulated lncRNAs in a training cohort of 160 HCC patients, and identified a nine-lncRNA classifier (Clnc) for early recurrence by using a linear support vector machine (SVM) model. Then the prognostic value of Clnc was validated in a test set of 161 patients. We also assessed the ability of the classifier to separate patients with respect to survival.
Materials and methods
Patients and tissue samples
The patients enrolled in the study were diagnosed with primary HCC and underwent curative resection between 2001 and 2013 at the Sun Yat-sen University Cancer Center (Guangzhou, China). All the tissues were snapfrozen in liquid nitrogen or kept in RNAlater immediately after surgical resection and then stored at
Selection of candidate lncRNAs
Two strategies were used to select candidate lncRNAs. Firstly a bioinformatics analysis was conducted based on lncRNA expression profiles (GSE54238,
RNA extraction, reverse transcription (RT) and quantitative PCR (RT-qPCR)
Total RNA from HCC and non-cancerous tissues were extracted using TRIzol
Data process and supervised classifier construction
The gene expression levels were discretized before model construction. The point corresponding to the minimum value for (1-sensitivity)
The 160 patients in the training set were divided into two groups according to their recurrence state within 1-year post-operation. Then a linear SVM-based model with an exhaustive search method was used for the training cohort to construct a supervised classifier for predicting the postsurgical recurrence within 1 year. Each lncRNA combination was defined as one classifier. Leave-one-out (LOO) cross-validation was used to evaluate the performance of each classifier (Fig. 1). The optimal classifier was determined when the value of (1-sensitivity)
Study design. A schematic overview summarizes the construction and validation of the classifier. A total of 30 candidate lncRNAs were selected to analyze their expression levels in 20 HCC and non-tumor tissues. Among them, 20 lncRNAs were abnormally expressed in HCCs and then applied to establish the prognostic model. A total of 321 HCC patients from Sun Yat-sen University Cancer Center (Guangzhou, China) were collected and distributed into the training cohort (
Performance evaluation of Clnc classifier for early recurrence prediction. A. ROC curves for the Clnc classifier and different clinical characteristics including tumor size, tumor number, differentiation, PVTT, and BCLC stage in predicting recurrence within 1 year after hepatectomy in the training (Left, 
ROC curve analysis was carried out with MedCalc version 15.2.2, (MedCalc Software Ltd, Ostend, Belgium). The area under the curve (AUC) was used to measure prognostic or predictive accuracy. Survival analyses were performed using the GraphPad prism7.0 (GraphPad Software, San Diego, CA, USA). Survival curves are estimated for each group using the Kaplan-Meier method and the log rank test was used to compare the survival curves between groups. Regression analyses including Logistic regression and Cox Logistic regression were performed using the Statistical Package for the Social Sciences (SPSS) software (version 16, SPSS Inc. Chicago, IL). Logistic regression models were used to estimate the odds ratio (OR) and 95% confidence interval (CI), and to identify independent prognostic variables for early recurrence. The Cox regression models were applied to evaluate the prognostic variables for recurrence-free survival (RFS). Comparisons of quantitative data between two groups were analyzed by a paired
The predictive efficiency of Clnc and clinical features for early recurrence in the training and test sets
The predictive efficiency of Clnc and clinical features for early recurrence in the training and test sets
Abbreviations: AUC, area under the receiver operating characteristic curve; BCLC, Barcelona Clinic Liver Cancer stage; CI, confidence interval; N, number of patients; PVTT, portal vein tumor thrombus; SEN, sensitivity; SPE, specificity.
Construction of the classifier
A group of 30 candidate lncRNAs was selected either from lncRNA microarray data or literature, and their expression levels in 20 paired HCC and the adjacent normal liver tissues were analyzed by RT-qPCR. The dysregulation of 20 lncRNAs was confirmed, with 15 upregulated and 5 downregulated lncRNAs (Supplementary Fig. S2). Further analysis showed that 3 lncRNAs were significantly associated with early recurrence (
Based on the hypothesis that incorporating multiple lncRNAs could improve prediction performance, we incorporated all the 20 lncRNAs into a linear SVM model to build lncRNA-based classifiers for early recurrence. After an exhaustive search of all possible combinations and LOO cross-validation, a combination of nine lncRNAs, named as Clnc, was finally selected (Supplementary Fig. S3), which includes AF339810, AK026286, BC020899, HEIH, HULC, MALAT1, PVT1, uc003fpg, and ZFAS1(Supplementary Table 5). The discriminative value for Clnc as measured by the AUC was 0.753 (sensitivity, 0.726; specificity, 0.78) for predicting early recurrence after HCC resection, which was significantly higher than that of other traditional clinical features including tumor size, tumor number, portal vein tumor thrombus (PVTT), differentiation and BCLC stage (Fig. 2A, Left panel and Table 1). For the 160 patients, 61 were discriminated as high-risk for early recurrence and 99 as low-risk by Clnc. The recurrence probability in the high-risk group was remarkably higher than that in the low-risk group (60.7% (37/61) vs 14.1% (14/99) and the cumulative recurrence probability in the high-risk group was also significantly higher than that in the low-risk group (Fig. 2B, Left panel). Consistently, survival among high-risk patients was poorer than that of the low-risk patients (Supplementary Fig. S4; Left). To facilitate the usage of this classifier, we constructed a web-based application that allows the users to determine the risk for early recurrence of each patient according to the relative expression values of lncRNAs that constitute Clnc in HCC tissues (
Logistic regression analysis to evaluate factors that were associated with early recurrence for patients in the training and test sets
Logistic regression analysis to evaluate factors that were associated with early recurrence for patients in the training and test sets
Abbreviations: AFP, Alpha-fetoprotein; BCLC, Barcelona Clinic Liver Cancer stage; CI, confidence interval; HBV, hepatitis B virus; N, number of patients; NA, not available; OR, odds ratio; PVTT, portal vein tumor thrombus.
Comparison of predictive power for early recurrence by the Clnc, BCLC stage, and their combinations. ROC curves in detecting HCC with early recurrence from training cohort (A) and test cohort (B) by the Clnc classifier, BCLC stage, and their combinations. The combination of Clnc and BCLC show the highest sensitivity in both training and test cohorts.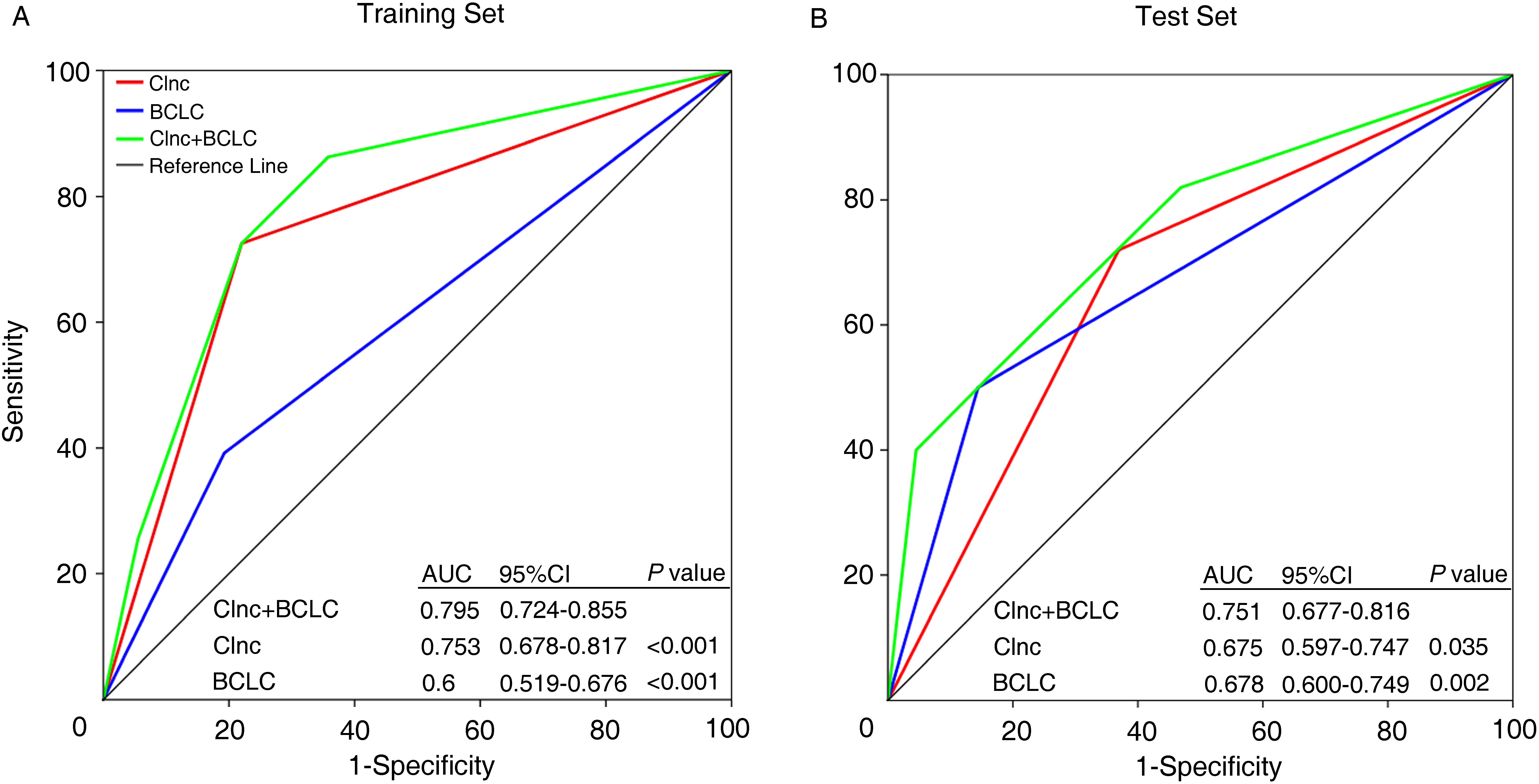
The predicted power of Clnc in the subgroups of patients in the combined training and test cohort
Abbreviations: AFP, Alpha-fetoprotein; BCLC, Barcelona Clinic Liver Cancer stage; CI, confidence interval; OR, odds ratio; Rec, recurrence.
The predictive efficiency of the Clnc was then evaluated in the test set. Clnc correctly predicted the early recurrence for 36 of 50 cases within the test set, and the odds ratio was 4.39 (95% CI 2.12–9.09),
BCLC staging system has been widely used as a prognostic factor, we then evaluated whether combining the Clnc and BCLC stage could increase the prediction efficiency. As shown, the combination of Clnc and BCLC (Clnc
Clnc is an independent prognostic factor for early recurrence and RFS in HCC patients
To explore the independence of Clnc from other clinical or pathological factors in predicting early recurrence, multivariable logistic regression analysis was performed using an entering variable selection method. Factors that were significantly associated with early recurrence in the univariate logistic regression analysis were included in the regression equation to adjust the coefficient. Clnc remained a powerful and independent factor to predict early recurrence in the training set (OR 9.408, 95% CI 4.298–20.592,
We then evaluated the prognostic value of Clnc in subgroups of different clinical stages. As shown, Clnc could identify patients with a high risk of early recurrence from low-risk patients in the subgroups divided according to AFP, tumor size, tumor number, differentiation, cirrhosis, and BCLC stage in the combined group including all the training and test cohorts (Table 3). These results suggested that Clnc had the prognostic potential for a subgroup of HCC.
Discussion
Early postsurgical recurrence remains a major cause of cancer-related mortality. Identifying patients at high risk of early relapse after resection could guide postoperative adjuvant therapies that confer the best survival and spare the low-risk patients from over chemotherapy. In this study, we developed a nine-lncRNA classifier, Clnc, and showed that Clnc could effectively discriminate patients at high risk of early recurrence from low-risk patients. Moreover, the Clnc classifier is an independent prognostic factor for the RFS of HCC patients.
Currently, BCLC staging has been widely accepted as a powerful prognostic predictor and acted as the main reference to direct the treatment strategies in HCC. However, as a result of the heterogeneity of cancer at the molecular and genetic levels [19, 20], even patients with the same stage can have very diverse survival outcomes [21]. Recent studies have suggested a few lncRNAs with prognostic values in HCC, yet most of them only tested an individual lncRNA in a small size of patients, and the accuracy and robustness of these lncRNA markers need to be further validated. Here, we showed that Clnc was a significant predictive classifier for early recurrence and RFS in the validation cohort in multivariate analysis (Table 2 and Supplementary Table 7). Moreover, Clnc can identify high-risk patients in different subgroups divided according to AFP, tumor size, tumor number, differentiation, and even those who are in the early BCLC stage (Table 3), which suggests that Clnc may be used to refine the current prognostic factors. Furthermore, combining Clnc and BCLC together could significantly increase the accuracy of prognostic prediction.
Early recurrence represents a true metastasis and is associated with the dissemination of primary HCC tumor cells. Of the nine lncRNAs that make up the Clnc, five of them, including ZFAS1, HULC, HEIH, PVT1, and MALAT1, are well-known lncRNAs that were dysregulated in HCCs and have been implicated in the development and progression of HCCs. For instance, ZFAS1 functions as an oncogene in HCC progression by abrogating the tumor-suppressor ability of miR-150, and high levels of ZFAS1 in HCCs are correlated with poor recurrence-free survival [22]. Loss of function studies showed that knockdown of HULC, HEIH, or PVT1 suppresses migration and invasion of HCC cells [23, 24, 25], characteristics that contribute to the early dissemination of tumor cell, suggesting that overexpression of these lncRNAs may play a role in tumor metastasis. Consistently, retrospective cohort studies have revealed that patients with high expression levels of HULC, HEIH or PVT1 exhibited poor recurrence-free survival [23, 26, 27]. MALAT1 is upregulated in HCC and acts as a proto-oncogene by Enhancing mTOR-Mediated Translation of TCF7L2 [28]. Moreover, MALAT1 overexpression inhibits, while MALAT1 deficiency promotes migration and invasion of HCC cells, and higher MALAT1 expression in HCC patients was associated with worse RFS [29]. Taken together, our findings suggested that integrating multiple lncRNAs that were dysregulated in HCCs might develop an excellent prognostic classifier for HCCs.
Notably, the lncRNA genes we selected in this study concluded only those differentially expressed between HCC and the non-tumor liver tissues, considering that the aberrantly expressed lncRNAs in cancer could mark the spectrum of disease progression and these lncRNAs may have the potential to serve as biomarkers for early relapse. Given the heterogeneity of tumors, lncRNA genes showing similar expression levels between cancer and normal tissues may possibly predict the recurrence of HCC patients with high accuracy. Thus, a more sophisticated screening strategy to discover the prognosis related lncRNAs, ideally those differentially expressed in subgroups of HCC patients with different prognosis, may guarantee a more robust classifier. Besides, this study concludes only patients from one center, and the AUC for the classifier is moderate. Future efforts will focus on optimizing the algorithm to boost the AUC and multi-center external validation to assure the general applicability of our model.
In conclusion, we developed a lncRNA-based classifier that could identify the patients at risk for postsurgical recurrence of HCC, which might provide patients with a chance of optimal postoperative adjuvant therapies.
Authors’ contributions
Conception: Shuai He and Jin-E Yang.
Interpretation or analysis of data: Shuai He, Jin-Feng Li, Hao Tian, Ye Sang, Xiao-Jing Yang, and Gui-Xin Guo.
Preparation of the manuscript: Shuai He and Jin-E Yang.
Supervision: Jin-E Yang.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-210193.
sj-docx-1-cbm-10.3233_CBM-210193.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-210193.docx
Footnotes
Acknowledgments
This work was supported by the National Natural Science Foundation of China (81572400 and 81872259).
