Abstract
Extracellular vesicles (EVs), which include exosomes, microvesicles, and apoptotic bodies, are nanosized structures that are secreted by various cells and act as important mediators in intercellular communication. Recent studies have shown that exosomes carrying bioactive molecules are generated from multivesicular bodies and are present in various body fluids. mRNAs and microRNAs (miRNAs) are encapsulated in exosomes and have been found to be involved in multiple pathophysiological processes. Here, we provide a review of tumor-associated exosomal mRNAs and miRNAs and their roles in metastasis and drug resistance. In particular, we emphasize their clinical application potential as diagnostic and prognostic biomarkers of cancer and in cancer therapy.
Introduction
Extracellular vesicles (EVs) are crucial factors in mediating crosstalk between tumor cells and their microenvironment. EVs, known as nanosized vesicles, are secreted by nearly all types of cells, including cancer cells,1-4 and act as transporters of proteins, nucleic acids, and lipids in a variety of physiological processes (innate and/or acquired immunity, immune modulation, blood coagulation, autophagy, and tissue regeneration) or as a consequence of pathological developments (tumor initiation, invasion, angiogenesis, metastasis, chemoresistance, recurrence, and immunosuppression).3,5-7 The diversity of phenotypic states and functions is due to the high heterogeneity of EVs. Although much of the literature broadly classifies EVs into three major categories—namely, exosomes, microvesicles (MVs) and apoptotic bodies—the nomenclature is still a controversial topic.
To better define the subtypes of EVs and understand the mechanisms of EV biogenesis and functionality, the need for highly purified EVs has become particularly important. Ultracentrifugation has been considered the most common approach for EV isolation, but other diverse isolation methods are still being investigated to improve the purity and specificity of separation, including size exclusion chromatography, ultrafiltration, polymer-based precipitation, microfluidics, density gradient isolation using sucrose, and antibody-based affinity capture.8-12
Exosomes are small extracellular vesicles carrying 5-nucleotidase activity, range from 30 to 100 nm in diameter, and play a vital role in intercellular communication.2,13 Tumor-derived exosomes transfer functional proteins and nucleic acids, and can support cancer progression and metastasis through continual trafficking to recipient cells. The proteins in exosomes include transcription factors (Rab GTPases), heat shock proteins (HSP70, HSP90), transport proteins (ESCRT complex), adhesion molecules (integrins, cadherins), tetraspanins (CD9, CD63, CD81, and CD82), enzymes, receptors, and epithelial cell adhesion molecules (EpCAM).14,15 Moreover, the nucleic acids carried by exosomes include dsDNA (c-Myc), mRNAs, microRNAs (miRNAs), and other noncoding RNAs (lncRNAs, tRNAs, rRNAs, snoRNAs, snRNA, circRNAs, and piRNAs), all of which are protected by the exosome lipid bilayer.16,17 Exosomal RNA profiles can influence protein expression patterns and change the pathophysiological activities and functions of parental cells. Currently, to highlight the importance of exosomes, attractive databases for exosomal RNA target identification and function have been developed. For example, both the ExoCarta database (http://exocarta.org/, containing 4563 proteins, 1639 mRNAs, and 764 miRNAs)18,19 and exoRBase (http://www.exoRBase.org/, containing 58,330 circRNAs, 15,501 lncRNAs and 18,333 mRNAs) 20 are widely used to examine the features of RNA expression profiles.
Notably, in recent years, increasing attention has been given to functional exosomes isolated from cancer cells and stromal cells. In this review, we focus on tumor-associated exosomal mRNAs and miRNAs by presenting recent advances in the biogenesis of exosomes and their capacities in cancer diagnosis. Furthermore, we emphasize the roles of exosomal mRNAs and miRNAs in metastasis and assess the potential use of exosomal RNAs as prognostic biomarkers. Finally, we discuss connections between exosomal RNAs and drug resistance, and highlight the prospective use of these RNAs as therapeutic agents in cancer. The scientific literature was comprehensively searched using the electronic PubMed database for all relevant studies from inception till 2019.
Biogenesis, secretion, and uptake of exosomes
To better understand the biogenesis of exosomes, novel methods must be developed to obtain a more thorough description of their structures. Insights into exosome morphology have been obtained by applying several different techniques. Using conventional transmission electron microscopy (TEM), a unique cup-shaped morphology of exosomes can be observed.21,22 Cryo-electron microscopy (cryo-EM), which avoids the influence of structure shrinking, was utilized to observe a close-to-native state and showed exosomes as round structures. 23 Because most investigations of exosomes have focused on the purification methods rather than on structural observation, knowledge regarding the exosome structure and surface molecules is still lacking. A new ultra-sensitive low force atomic force microscopy (AFM) approach, similar to field emission scanning electron microscopy (FESEM), was used to examine the substructure of individual human saliva exosomes, which cannot be resolved with TEM, and the nanoscale structures of exosomes under varying forces were well accounted. 24 Compared to the round vesicle structures of exosomes without substructure details observed under TEM, three-dimensional structures of single isolated sub-100 nm exosome and their reversible elastic nanomechanical properties can be revealed by AFM. This study is regarded as an important development and supplies critical data for developing engineered exosome drug-delivery platforms possessing inherent biocompatibility and native nanoarchitecture.
Unlike microvesicles, which directly bud from the plasma membrane, exosomes are generated from multivesicular bodies (MVBs, known as mature endosomes) through accumulation of intraluminal vesicles (ILVs) in the lumen of endosomes. Recently, several distinct mechanisms have been identified in the biogenesis of exosomes, which is considered to be complex. It seems that exosomes can be mainly formed through both endosomal sorting complexes required for transport (ESCRT)-dependent and ESCRT-independent mechanisms.25,26 Although some cytosolic proteins modified by ubiquitylation have been found to be segregated in ILVs and in exosomes, the underlying mechanisms are still unclear. 27 Multiprotein subcomplexes are required in the ESCRT-dependent mechanism for exosome biogenesis and act in a stepwise manner, including ESCRT-0, ESCRT-I, ESCRT-II, and ESCRT-III.28,29 In general, ubiquitinated cargoes are recognized by ESCRT-0 and ESCRT-I subunits and are clustered and recruited on microdomains of the limiting membrane of MVBs; then, budding and fission of microdomains are mediated by ESCRT-II and ESCRT-III subunits. 30 In addition, other essential players, such as ALG-2-interacting protein X (ALIX), vacuolar protein sorting-associated protein (VPS4), and ATPase, are involved in the ESCRT-dependent pathway. However, MVBs can still be formed even though all components of ESCRT complexes were depleted, 31 indicating that ESCRT-independent pathway is another critical manner in exosome biogenesis and sorting. This notion is supported by some lipids, such as sphingosine 1-phosphate and sphingolipid ceramide are identified to be involved in the regulation of the budding or release of exosomes under ESCRT-independent pathway.32,33 Additionally, endosomal sorting through the ESCRT-independent pathway can also be regulated by the tetraspanin family (CD9, CD63, CD81, and CD82).34,35
In a cancer type- or stage-specific manner, various miRNAs can be differentially sorted into MVBs depending on binding to specific motifs. 36 Moreover, this process can be performed by different regulators, including ESCRT-II subcomplex, tetraspanin-enriched microdomains, RNA-induced silencing complex (RISC), the protein Argonaute 2 (AGO2), major vault protein, and Y-box-binding protein 1 (YBX1).37-42 Because the main interest in exosomal nucleic acids has been focused on noncoding RNAs rather than on coding RNAs, the mechanism by which mRNA sorting into exosomes is regulated is still poorly understood. Specific mRNAs, including mostly fragmented mRNAs with enriched UTRs and some full-length mRNAs, were reported to be transferred into exosomes by binding of multifunctional protein YB-1 and methyltransferase NSUN2 to specific motifs of mRNAs (ACCAGCCU, CAGUGAGC, and UAAUCCCA).43,44 Further research is essential to understand the process by which mRNAs are sorted into exosomes.
Following the formation of MVBs, exosome secretion occurs when MVBs fuse with the plasma membrane and release their cargos to the extracellular environment. Furthermore, several Rab GTPases (Rab5, Rab7, Rab9, Rab11/35, Rab22a, and Rab27) and their respective effectors (synaptotagmin-like protein 4, exophilin 5) have been reported to be required for regulation of the secretory pathways of exosomes.45-50 Thus, decreasing production of exosomes by inhibiting GTPases seems to be a novel method to suppress tumor growth and metastasis. 51 Once released into the extracellular environment, exosomes are taken up by recipient cells through the following mechanisms: membrane fusion, pinocytosis, phagocytosis, endocytosis, and receptor-mediated endocytosis. 52 In a Dicer-dependent manner, the transcriptome of target cells can be altered by exosomal miRNAs to promote tumor progression, resulting in efficient and rapid mRNA silencing (Figure 1). For example, a specific cell-independent capacity in breast cancer cell-derived exosomes, which process premiRNAs into mature miRNAs, was shown to be associated with RISC loading complex proteins, Dicer, TRBP, and AGO2. 53 Exosomes also play an important role in degradation of unwanted RNAs. There is now evidence that competition between human homologue of mRNA transport mutant 4 (hMTR4) and Aly/REF export factor (ALYREF) for associating with arsenic resistance protein 2 (ARS2) is required for exosome-mediated degradation and export of nuclear mRNAs. 54 However, how export-defective mRNAs are recognized by exosomes requires further investigation.
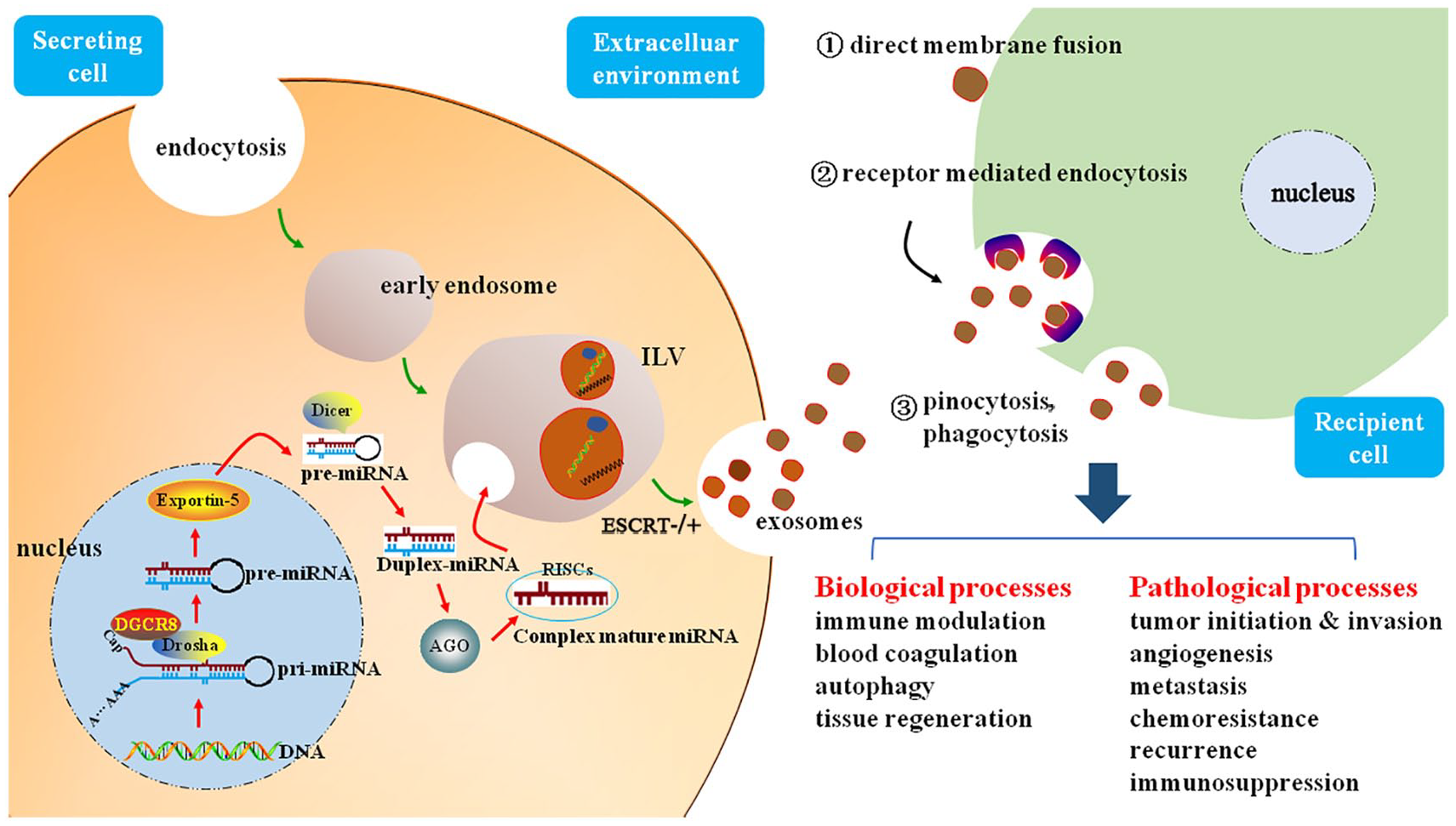
Schematic illustration of exosomal miRNA biogenesis and their communication with recipient cells. Canonically, miRNA genes in the nucleus are transcribed to generate pri-miRNAs, then these pri-miRNAs are transmitted into pre-miRNAs by the catalytic effect of DGCR8 and Drosha complex. Hairpin pre-miRNAs are then exported out of the nucleus by exportin-5. By the digestion of Dicer complex, pre-miRNAs in the cytoplasm are processed into duplex miRNAs, which are converted into single-stranded ones. Mature miRNAs are sorted into exosomes in miRISC-related pathway, single-stranded miRNAs bind to AGO proteins to form RISC, and then the exosomes loaded with miRNAs are released to the extracellular environment. Through the pathways of direct membrane fusion, receptor mediated endocytosis, pinocytosis, and phagocytosis, exosomes are taken up by recipient cells to delivery exosomal miRNAs, participating in various biological and pathological processes.
In addition, although direct visualization of the exosomal mRNA fate is lacking, it has been shown that mRNAs residing in EVs can be efficiently translated in recipient cells to regulate pathways involved in metastasis and drug resistance.55,56 Typically, USER codes of EV-resident mRNAs can be recognized and combined with regulatory RNAs (such as miRNAs, snRNAs, or siRNAs) or RNA binding proteins (RBPs), contributing to stability and translation. 57 Nevertheless, how exosomal RNAs are targeted to the cell surface is still unclear, and more studies are needed to comprehend this process.
Exosomal mRNAs and miRNAs in cancer diagnosis
Currently, cancer is still the second major cause of mortality in the United States. 58 Although many promising advances have been achieved over the last decade, early diagnosis, which is necessary to develop treatment strategies and enhance the survival rate, remains difficult. In this respect, reliable biomarkers with high sensitivity and specificity for early detection and accurate monitoring of cancer should be developed. Generally, diagnosis of most cancer types still relies on traditional imaging examinations (e.g. magnetic resonance imaging, computed tomography, or a positron emission tomography scan), biopsy strategies or analysis of serum biomarkers. 59 Notably, liquid biopsy has been proposed as a novel diagnostic approach, with the allure of providing a minimally invasive, convenient, quick, and economical procedure relative to tissue biopsies, and can be used for early diagnosis, monitoring of therapeutic response, forecasting recurrence, obtaining prognostic information, and even for population screening. 60 The main liquid biopsy approaches involve circulating tumor cells (CTCs), circulating tumor DNA (ctDNA), and exosomes. 61 Exosomes have been reported to be present in various body fluids, including blood, urine, saliva, cerebrospinal fluid, ascites, and bile.62-66 In addition, more exosomes are generated and released from cancer cells. 67 Compared with normal human blood, which contains approximately 2000 trillion exosomes, a conservative estimate indicated that approximately 4000 trillion exosomes are present in the blood of cancer patients. 68 Overall, these data support the notion that the high number of exosomes present in circulation offers a rich source of tumor material, compared with the limited number of detectable CTCs and ctDNA. Recently, circulating exosomes containing glypican-1 (GPC1) were demonstrated to be specifically enriched in pancreatic cancer patients, and were highly outperformed over serum-free GPC1 or serum carbohydrate antigen 199 (CA 19-9) for detection of early-stage pancreatic cancer. 68 These results argued that biomarkers developed from circulating exosomes are superior to traditional serological markers, and can be served as a potential non-invasive diagnostic method for pancreatic cancer. In addition, exosomes are regarded as ideal diagnostic biomarker candidates in many cancer types, including pancreatic cancer, melanoma, breast cancer, prostate cancer, lung cancer, gastric cancer, colorectal cancer (CRC), hepatocellular carcinoma (HCC), and glioblastoma (GBM).69,70
Similar to proteins, exosomal nucleic acids are receiving increased attention as a new diagnostic tool with the following advantages: (a) abundant resources, (b) high stability, and (c) real-time information. First, exosomes are abundant in circulation, as mentioned above, and exosomes are the main source of circulating nucleic acids. Tumor-derived exosomes contain an abundance of nucleic acids, exhibiting up to 100-fold enrichment relative to parental cells.3,71 Thus, analysis of enriched exosomal nucleic acids can provide direct information about a cancer. Second, nucleic acids are protected by the lipid bilayer membrane of exosomes, thereby avoiding degradation by RNases. 72 Moreover, it is reported that exosomal RNAs are stable even under long-term freezing conditions and after freeze-thaw cycles. 73 Finally, and most importantly, exosomes are secreted by cancer cells, and exosomal RNAs present a strong resemblance to parental cells; thus, real-time information can be provided by detecting tumor-derived exosomes. Interestingly, detecting tumor-derived exosomal RNA seems beneficial to overcome the challenge of limited ctDNA. It was reported that the sensitivity of epidermal growth factor receptor (EGFR) mutation detection could be improved by combining exoRNA and ctDNA detection in plasma, especially in the subgroup of M0/M1, a non-small-cell lung cancer with low levels of ctDNA. 74 In recent years, numerous studies have suggested that both exosomal mRNAs and miRNAs can be used as cancer biomarkers (Table 1). Especially, tumor-derived exosomal miRNAs have been extensively studied as diagnostic biomarkers and can be used to distinguish specific molecular subtypes of cancer. 75 Based on the literature, circulating exosomal miR-21, which is also associated with cancer metastasis and chemoresistance, has been proposed to be a universal diagnostic and prognostic biomarker to identify various cancers, including prostate cancer, lung cancer, colorectal cancer, laryngeal and esophageal squamous cell carcinoma, and hepatocellular carcinoma.76-79 Interestingly, compared with exosomal GPC1 and plasma CA 19-9, upregulation of miR-10b, miR-21, miR-30c, and miR-181a and downregulation of let7a detected in exosomes from plasma shows superiority as a diagnostic biomarker in pancreatic cancer. 80 This study also indicated the utility of multiple reaction monitoring (MRM), which is a rapid, sensitive, novel liquid chromatography-tandem mass spectrometry (LC-MS/MS)-based method that can be used for efficient high-throughput analysis of clinical samples. 81 Additionally, exosome-associated mRNAs are also considered promising diagnostic markers of cancer. For instance, high levels of human telomerase reverse transcriptase (exo-hTERT) mRNA in serum were found in diverse cancers and showed potential as a diagnostic “pan-cancer” marker. 82 Exosomal Wiskott-Aldrich syndrome protein family member 2 (WASF2) and ADP-ribosylation factor 6 (ARF6) mRNAs were significantly increased in serum from pancreatic cancer patients and provided excellent accuracy (AUC > 0.9) for distinguishing early pancreatic cancer patients from control subjects. 83
Exosomal mRNAs and miRNAs as cancer diagnostic and prognostic biomarkers.
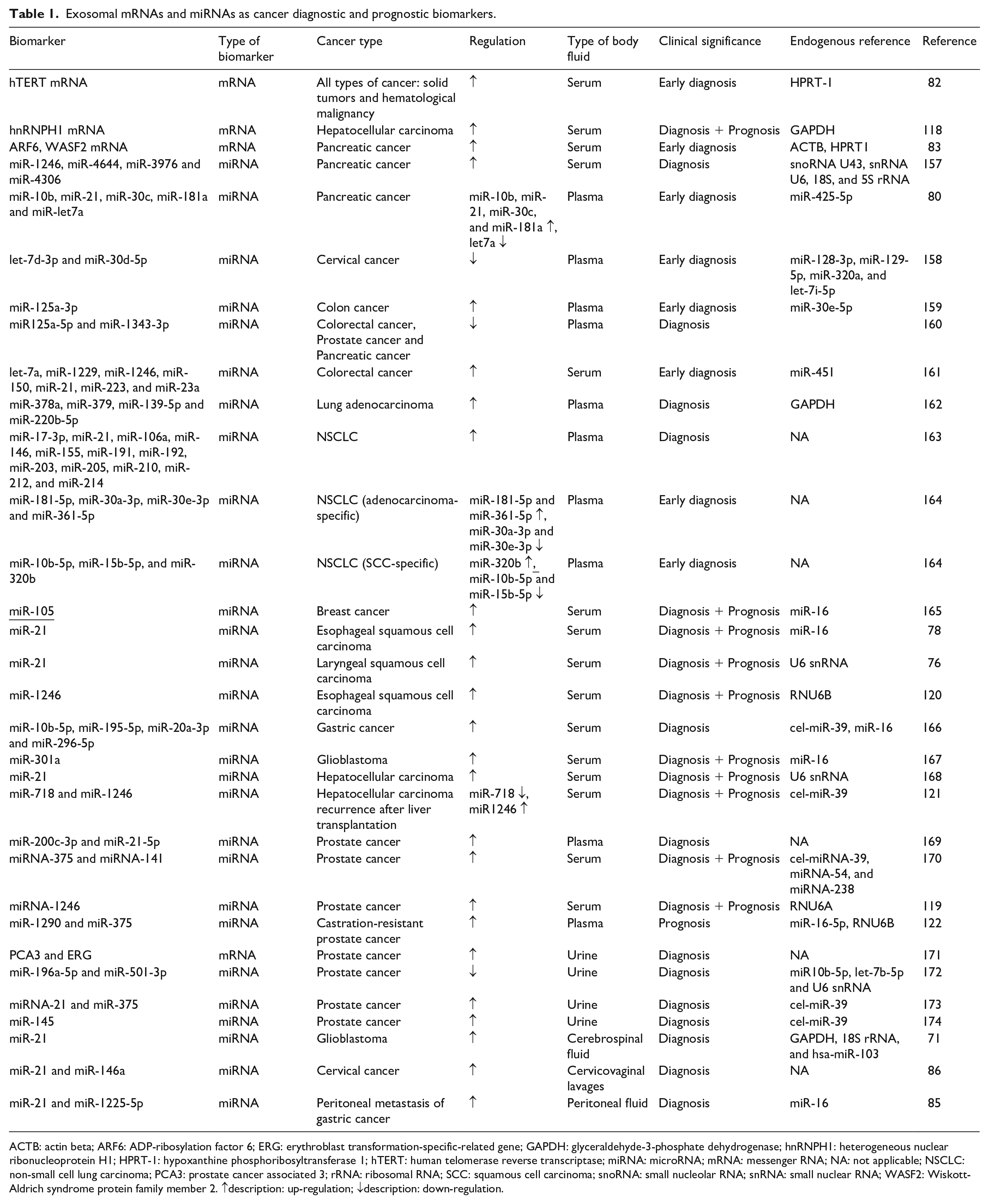
ACTB: actin beta; ARF6: ADP-ribosylation factor 6; ERG: erythroblast transformation-specific-related gene; GAPDH: glyceraldehyde-3-phosphate dehydrogenase; hnRNPH1: heterogeneous nuclear ribonucleoprotein H1; HPRT-1: hypoxanthine phosphoribosyltransferase 1; hTERT: human telomerase reverse transcriptase; miRNA: microRNA; mRNA: messenger RNA; NA: not applicable; NSCLC: non-small cell lung carcinoma; PCA3: prostate cancer associated 3; rRNA: ribosomal RNA; SCC: squamous cell carcinoma; snoRNA: small nucleolar RNA; snRNA: small nuclear RNA; WASF2: Wiskott-Aldrich syndrome protein family member 2. ↑description: up-regulation; ↓description: down-regulation.
Notably, although most studies have focused on circulating exosomes, detection of exosomal RNAs in other body fluids has become attractive, but more work is needed. For example, erythroblast transformation-specific-related gene (ERG) or prostate cancer associated 3 (PCA3) mRNA expression in urine exosomes highly discriminates high-grade (⩾ GS7) from low-grade (GS6) prostate cancer and benign disease, reducing unnecessary biopsies. 84 Evaluation of exosomal miR-21 and miR-1225-5p in peritoneum lavage fluid appears to be a novel approach for early diagnosis of peritoneal dissemination after curative resection of gastric cancer. 85 High levels of exo- miRNA-21 and miRNA-146a were detected in cervicovaginal lavages from cervical cancer patients, indicating their potential as biomarkers of cervical cancer. 86 Exosomal miR-21 harvested from cerebrospinal fluid can distinguish glioblastoma patients from healthy individuals, thus showing potential as a glioblastoma diagnostic biomarker. 71
Generally, detection of exosomal RNAs is based on real-time quantitative polymerase chain reaction (RT-qPCR); however, exosome purification and distinct internal controls remain the major limitations in detection approaches. Despite a variety of techniques have been employed to purify exosomes, the limitations of efficiency, specificity, and flexibility are still existed in methodologies. So far there is no gold standard for the purification of exosomes. Stable internal controls are essential for accurate miRNA quantification in qRT-PCR. With the lacking of a stably and abundantly expressed endogenous reference gene for circulating and exosomal miRNA normalization, the consensus over universally applicable internal controls has not been reached so far. Additionally, efficiently quantifying low levels of exosomal RNAs in early-stage cancer is a challenge. Recently, lipid-polymer hybrid nanoparticle-catalyzed hairpin DNA circuit (LPHN-CHDC), a novel and valuable nanoparticle-based biochip with signal amplification capability, was reported to selectively and sensitively identify low expression EV-associated mRNAs in serum from early pancreatic cancer patients. 87 Moreover, other novel and efficient detection techniques remain to be explored.
In sum, exosomal mRNAs and miRNAs from a body fluid sample, especially blood, can signal the presence of cancer without the use of invasive procedures. However, further work is still required before they can become reliable diagnostic tools for clinical application.
Exosomal mRNAs and miRNAs in cancer metastasis and prognosis
Cancer metastasis—a dynamic and multistep process involving a complex molecular network—is the main cause of cancer-related death. Fully understanding the potential mechanisms underlying cancer metastasis is crucial to provide valuable information for new therapeutic strategies. Cancer metastasis is closely associated with coevolution of a tumor and its microenvironment. 88 It has been demonstrated that exosomes, which affect the local and distant environment by mediating intercellular communication, play an important role in cancer metastasis. According to Paget’s “seed and soil” metastasis theory, under the assistance of exosomes, cancer cells obtain migration and invasion capacity through epithelial-mesenchymal transition (EMT), and the microenvironment of secondary organs is modified to optimize conditions (termed the premetastatic niche) for migration.7,89-91 Then, active cancer cells colonize at favorable organ sites by inducing the formation of CTCs, and some cells develop into macroscopic metastases. Indeed, ovarian cancer cell-derived exosomes carrying matrix metallopeptidase 1 (MMP1) mRNA have emerged as important mediators of peritoneal dissemination and are closely related to poor prognosis. 92 However, the detailed molecular mechanisms by which exosomal RNAs regulate the metastasis process are still incompletely understood.
Emerging evidence suggests a significant role of tumor-derived exosomes in premetastatic niche formation, as well as remodeling of various cells in the tumor microenvironment.93-95 Moreover, metastatic behavior in moderately metastatic tumors can be significantly induced by exosomes derived from highly metastatic cells. 92 It has been confirmed that pancreatic cancer-derived exosomes play a key role in inducing establishment of the liver premetastatic niche, leading to increased liver metastatic burden. 96 Additionally, exosomal miR-122 derived from breast cancer cells has been demonstrated to promote metastasis by increasing nutrient availability in the premetastatic niche. 97 Diverse components are present in the architecture of the tumor microenvironment, including fibroblasts, endothelial cells, inflammatory and immune cells, extracellular matrix (ECM) constituents, adipose cells, signaling molecules, and soluble factors.98,99 In addition to cancer cells, the cells in the tumor microenvironment are collectively known as stromal cells. As mentioned above, stromal cells can be regulated by tumor-derived exosomal RNAs in preparation for metastasis. In this regard, exosomal miR-1247-3p secreted by highly metastatic HCC cells is capable of converting normal fibroblasts to cancer-associated fibroblasts (CAFs) in the lung premetastatic niche by activating the β1-integrin-NF-κB signaling pathway. 100 In addition, macrophages, the most abundant immune-related stromal cells, can be polarized into activated tumor-associated macrophages (TAMs) by hypoxic exosomal miR-301a-3p derived from pancreatic cancer cells, enhancing the metastatic potential of pancreatic cancer cells. 101 Meanwhile, exosomal miR-301a-3p detected in circulation is thought to be a novel prognostic biomarker. Similarly, TAMs have been shown to be stimulated by liposarcoma-derived exosomal miR-25-3p and miR-92a-3p to induce liposarcoma metastasis. 102 In contrast, GBM-derived exosomal miR-1, a tumor suppressor in various cancers, mitigates GBM invasion and progression by directly targeting annexin A2 (ANXA2) mRNA in exosomes derived from cells in the GBM microenvironment. 103 These findings provide evidence of complex communication networks in the tumor microenvironment and highlight the potential therapeutic value of exosomal miRNAs with distinct abilities.
Not only are cancer cell-derived exosomal RNAs essential in initiation and establishment of the premetastatic niche, but stromal cell-derived exosomal RNAs also play a crucial role. Recent evidence suggests that a reduction in exosomal miR-320a secreted by CAFs participates in promoting HCC cell migration and metastasis. 104 Moreover, exosomal miR-21-5p and miR-155-5p derived from TAMs have been identified as key factors promoting CRC metastasis through downregulation of brahma-related gene 1 (BRG1) expression in CRC cells. 105
Mesenchymal stem cells (MSCs) are multipotent stromal cells with a tissue reparative function, and they possess the capacity to differentiate into diverse cells and are present in several tissues, including bone marrow and adipose tissue. 106 MSCs are considered to be a promising cell therapeutic strategy for various diseases. More recently, it has been shown that MSCs can be recruited to tumor stroma and lead to cancer growth and progression by stimulating EMT in cancer cells, which is also promoted by the interaction between MSCs and cancer cells.7,94,107 Moreover, MSCs have been revealed to participate in the establishment of premetastatic niches. 108 Recent studies have demonstrated that cancer-associated MSCs can facilitate metastasis, 109 but the detailed mechanisms remain unclear. As researchers increasingly focus on the connection between MSC-derived exosomes and metastasis, this issue is becoming an attractive area. Many specific mRNAs (fibroblast growth factor 7 (FGF7), vascular endothelial growth factor A (VEGFA), and hepatocyte growth factor (HGF) mRNA) 110 and miRNAs (miR-143-3p, miR-10b-5p, miR486-5p, miR-22-3p, and miR-21-5p) 111 have been discovered to be enriched in exosomes secreted by bone marrow MSCs. Furthermore, high miR-1587 enrichment has been found to be present in exosomes derived from glioma associated-human mesenchymal stem cells (GA-hMSCs) and be conducive to acceleration of glioblastoma aggressiveness through downregulation of the tumor suppressive nuclear receptor co-repressor NCOR1 in glioma stem-like cells. 112 Additionally, upregulation of exosomal miR-214, miR-221, and miR-222 in gastric cancer tissue-derived mesenchymal stem cells (GC-MSCs) has been found to be closely related to venous invasion and lymph node metastasis, and proliferation and migration of gastric cancer cells can be facilitated by delivering GC-MSC-derived exosomal miR-221 targeting cancer cells. 113 These results highlight the significant effects of MSC-derived exosomal RNAs, which act as mediators in MSC-associated metastasis. However, detailed knowledge regarding how MSC-derived exosomal RNAs regulate cancer metastasis is still lacking and elusive.
Brain metastasis is the most serious of all metastasis patterns, resulting in an especially poor prognosis. It usual originates from various cancers, including breast cancer, lung cancer, and melanoma.114,115 Notably, brain metastasis involves specific and mysterious metastasis mechanisms that differ from conventional metastasis mechanisms, such as those underlying lung and liver metastasis. In addition, due to the protection of the blood-brain barrier (BBB), which possesses selective permeability, most therapeutic strategies for brain metastasis have failed. Thus, to alter the tragic reality, a complete understanding of the mechanism is crucial to address the current shortage of antimetastatic therapeutic approaches. Currently, accumulating evidence supports the notion that exosomal miRNAs are vital in the initiation and progression of brain metastasis. For example, exosomal miRNA-181c derived from brain metastatic cancer cells facilitates brain metastasis by destroying the BBB 116 via targeted binding to 3-phosphoinositide dependent protein kinase 1 (PDPK1) and downregulation of PDPK1 expression. In addition, astrocyte-derived exosomes containing phosphatase and tensin homologue (PTEN)-targeting miRNAs can be transferred to brain metastatic tumor cells, leading to PTEN loss. 117 Brain metastasis is promoted by PTEN loss, which results in reduced apoptosis and enhanced proliferation of brain metastatic tumor cells.
In brief, abundant evidence has confirmed the functions of exosomal RNAs in the cancer metastasis processes (Table 2); however, because most studies have focused on exosomal miRNAs, evidence showing how exosomal mRNAs regulate metastasis remains lacking. Furthermore, the majority of studies performed have utilized in vivo mouse xenograft models or in vitro cell experiments; thus, validation with human samples is still necessary.
Functional effects of exosomal mRNAs and miRNAs in cancer metastasis.
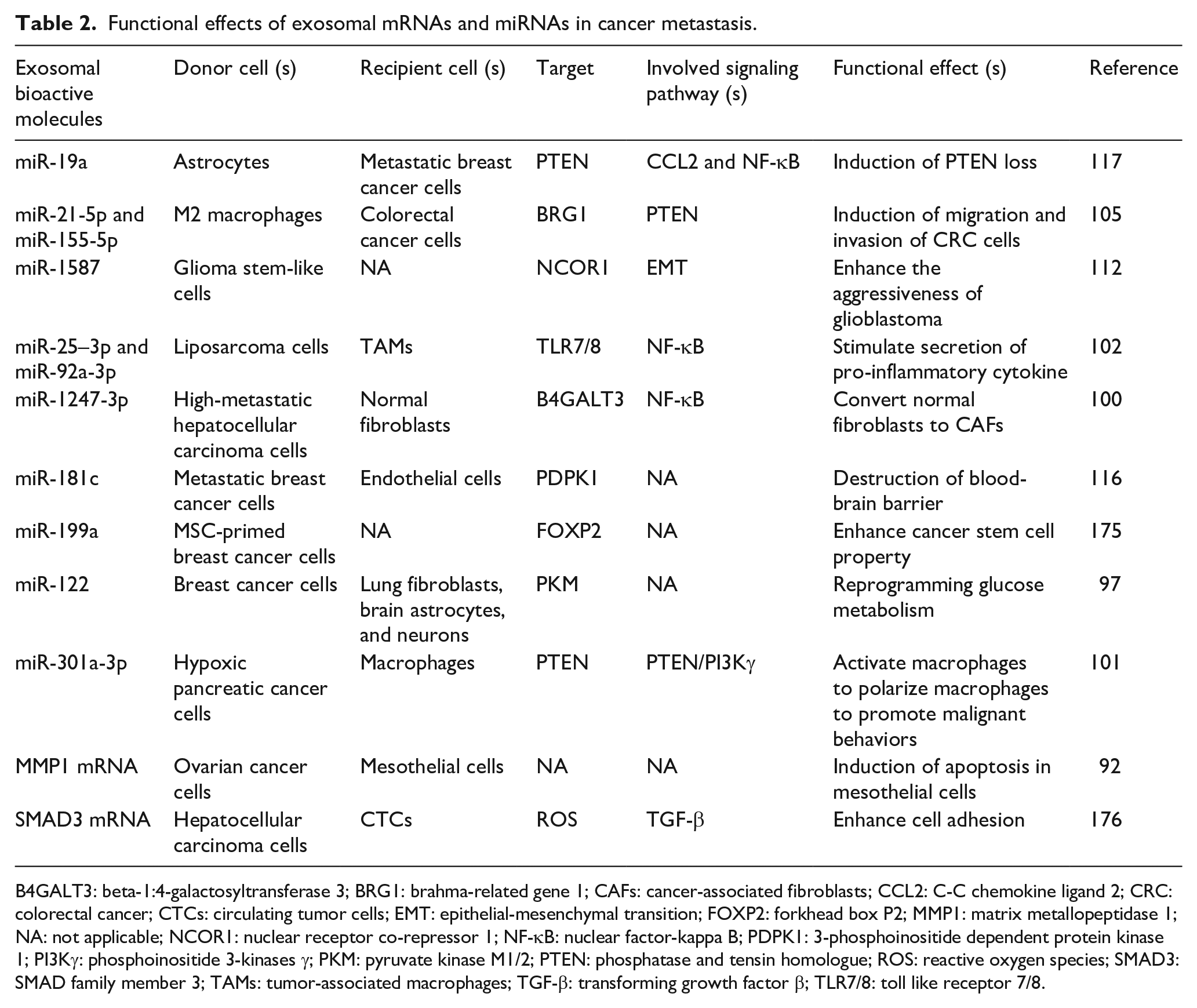
B4GALT3: beta-1:4-galactosyltransferase 3; BRG1: brahma-related gene 1; CAFs: cancer-associated fibroblasts; CCL2: C-C chemokine ligand 2; CRC: colorectal cancer; CTCs: circulating tumor cells; EMT: epithelial-mesenchymal transition; FOXP2: forkhead box P2; MMP1: matrix metallopeptidase 1; NA: not applicable; NCOR1: nuclear receptor co-repressor 1; NF-κB: nuclear factor-kappa B; PDPK1: 3-phosphoinositide dependent protein kinase 1; PI3Kγ: phosphoinositide 3-kinases γ; PKM: pyruvate kinase M1/2; PTEN: phosphatase and tensin homologue; ROS: reactive oxygen species; SMAD3: SMAD family member 3; TAMs: tumor-associated macrophages; TGF-β: transforming growth factor β; TLR7/8: toll like receptor 7/8.
Several metastasis-associated exosomal RNAs, particularly circulating exosomal RNAs, have emerged as extremely promising prognostic biomarkers to evaluate patient survival. For example, increased levels of serum exosomal heterogeneous nuclear ribonucleoprotein H1 (hnRNPH1) mRNA, which is associated with lymph node metastasis and portal vein tumor emboli, have prognostic potential for HCC in high HBV prevalence areas. 118 Moreover, upregulated exosomal miR-1246 in serum was capable of predicting recurrence, metastasis, and prognosis in multiple cancers, including HCC, aggressive prostate cancer, and esophageal squamous cell carcinoma.119-121 Finally, elevated levels of plasma exosomal miR-1290 and miR-375 have been confirmed to be promising prognostic biomarkers for castration-resistant prostate cancer patients. 122 In sum, these data support the idea that circulating exosomal mRNAs and miRNAs are extremely useful tools for prognosis assessment. Nevertheless, prospective validation of these promising molecules is required for their further clinical application.
Exosomal mRNAs and miRNAs in cancer drug resistance and therapies
As exosomal RNAs have been confirmed to participate in cancer progression and metastasis, they also have the potential to serve as therapeutic targets. Currently, emerging evidence suggests that exosomal RNAs participate in cancer drug resistance and that RNAs can be encapsulated in exosomes to serve as drug-delivery systems. A detailed discussion of recently reported examples is presented in the following section.
Drug resistance
Despite the advances achieved in cancer therapy, chemotherapy, endocrine therapy, and molecular targeted therapy are the basic drug therapy approaches for advanced carcinoma, and drug resistance remains the main challenge in cancer treatment and can ultimately lead to the death of cancer patients. To date, exosomal RNAs have been found to be associated with resistance to multiple drugs in various cancer types (Table 3) and can be used to monitor the real-time anticancer drug efficacy.123-126 Moreover, drug resistance can be induced and promoted not only by stromal cell-derived exosomal RNAs but also by drug-resistant cancer cells. 127 For instance, miR-365 and miR-21 derived from TAM-associated exosomes induces gemcitabine resistance in pancreatic ductal adenocarcinoma and cisplatin resistance in gastric cancer, respectively.128,129 Meanwhile, exosomal miR-21 isolated from cancer-associated adipocytes (CAAs) and fibroblasts (CAFs) confers resistance to paclitaxel in ovarian cancer cells by targeting apoptotic peptidase activating factor 1 (APAF1). 130 Furthermore, exosomal miR-222-3p derived from gemcitabine-resistant cells promotes the gemcitabine resistance of parental sensitive cells in non-small cell lung cancer (NSCLC) by targeting suppressor of cytokine signaling 3 (SOCS3), and serum exosomal miR-222-3p contributes to monitoring of the gemcitabine sensitivity in NSCLC patients. 131 This discovery suggests the probability that circulating exosomal RNAs can also be used as biomarkers for monitoring drug resistance in cancer. Similarly, plasma exosomal androgen receptor splice variant 7 (AR-V7) mRNA acts as a predictor of hormonal resistance in advanced prostate cancer patients. 132 Moreover, tumor-derived exosomal O(6)-methylguanine-DNA methyltransferase (MGMT) and alkylpurine-DNA-N-glycosylase (APNG) mRNA levels detected in serum using a microfluidic chip showed a capacity to predict the real-time drug response in GBM patients. 123 Blocking the secretion of exosomal RNAs may serve as a novel approach to reverse drug resistance. 126 Hence, a sufficient understanding of the mechanisms underlying exosomal RNA-based drug resistance is still needed for further precision cancer therapies.
Exosomal mRNA- and miRNA-associated drug resistances in cancer therapy and potential mechanisms.
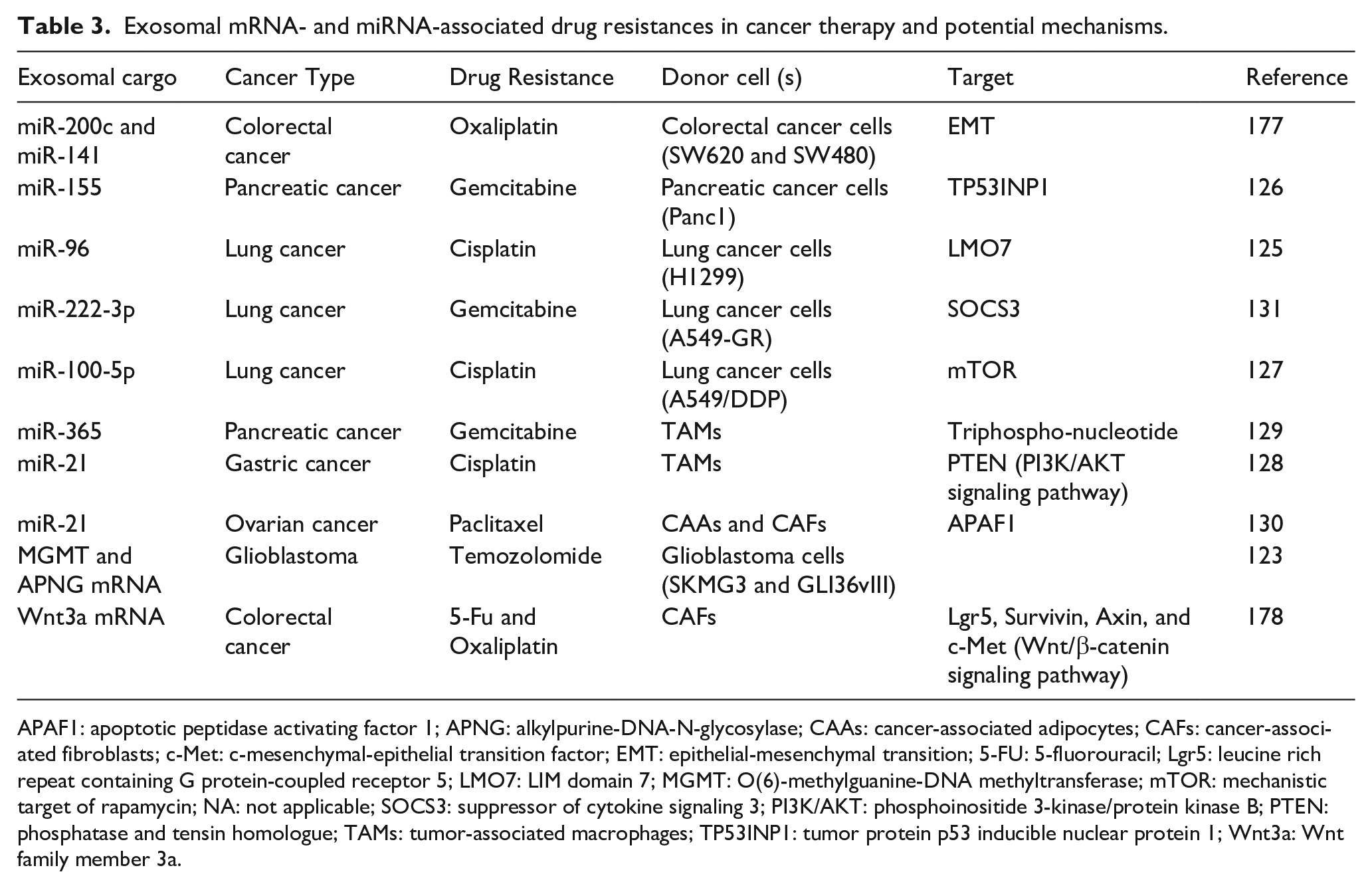
APAF1: apoptotic peptidase activating factor 1; APNG: alkylpurine-DNA-N-glycosylase; CAAs: cancer-associated adipocytes; CAFs: cancer-associated fibroblasts; c-Met: c-mesenchymal-epithelial transition factor; EMT: epithelial-mesenchymal transition; 5-FU: 5-fluorouracil; Lgr5: leucine rich repeat containing G protein-coupled receptor 5; LMO7: LIM domain 7; MGMT: O(6)-methylguanine-DNA methyltransferase; mTOR: mechanistic target of rapamycin; NA: not applicable; SOCS3: suppressor of cytokine signaling 3; PI3K/AKT: phosphoinositide 3-kinase/protein kinase B; PTEN: phosphatase and tensin homologue; TAMs: tumor-associated macrophages; TP53INP1: tumor protein p53 inducible nuclear protein 1; Wnt3a: Wnt family member 3a.
Cancer therapy
Clearly, exosomes are important mediators in cell-to-cell communication by transporting functional biomolecules. Exosomes have been demonstrated to influence the pharmacodynamics of nearby cells by transferring drugs intracellularly. 133 Thus, exosomes are regarded as a promising delivery system and present an attractive therapeutic strategy for cancer. Although exosome-based delivery platforms are still in the budding stage, they have garnered much attention due to their potentially superior low immunogenicity and high stability, specificity, efficiency, and biocompatibility. 134 Compared with traditional drugs, targeted delivery by exosomes can aid in conquering drug resistance and alleviating unwanted side effects.135,136 In recent years, emerging studies have suggested the utility of nucleic-acid-based therapeutics in cancer, which have entered clinical development; however, safer and more stable and efficient delivery systems are still needed for nucleic-acid-based anticancer drug development.137-139 Both exogenous and endogenous therapeutic molecules can be encapsulated in exosomes to serve as anticancer agents, and currently, exosome-based delivery is primarily centered around small molecule anticancer drugs, therapeutic proteins, and nucleic acids.140-142 Both miRNA and small interfering RNA (siRNA) for gene silencing and mRNA for gene expression have been contained in exosomes for nucleic acid delivery. Hence, exosomes are ideal vehicles for nucleic acid delivery, and handling of exosomal RNAs is presenting new options for customized, individualized anticancer therapies. For example, esophageal cancer-related gene 4 (ECRG4) mRNA can be delivered into recipient cells by exosomes, leading to suppressed cell proliferation and inhibited tumor growth, indicating the benefits of exosome delivery for cancer gene therapy. 143 Similarly, engineered exosomes have been used to deliver siRNA targeting oncogenic KRASG12D and were found to significantly suppress cancer growth and improve overall survival in mouse models of pancreatic cancer. 144 Moreover, tumor-suppressive miRNAs and anti-miRNAs can be delivered by exosomes to act as novel cancer therapy agents.145,146
Meanwhile, depending on their inherent ability to target cancer and penetrate natural barriers, especially the BBB, exosome-based drug delivery also brings a hopeful therapeutic strategy for brain cancer and metastasis. For instance, exosomes derived from brain endothelial cells were used to deliver the anticancer drugs doxorubicin and paclitaxel across the BBB and showed significant therapeutic efficacy in a zebrafish model of brain cancer by suppressing vascular endothelial growth factor (VEGF) RNA. 147 Currently, various evidence supporting RNA delivery by exosomes as an innovative anticancer therapeutic approach for brain treatment has been presented. For example, exosomal miRNA-142-3p from astrocytes has been found to suppress lung adenocarcinoma-associated brain metastases by consuming transient receptor potential ankyrin 1 (TRPA1). 148 In addition, genetically engineered exosomes enriched with therapeutic cytosine deaminase-uracil phosphoribosyltransferase (CD-UPRT) mRNA, which is a type of suicide gene mRNA, were shown to significantly decrease tumor growth in a subcutaneous xenograft glioblastoma tumor model after intratumoral injection. 149
Although exosome-based delivery systems have bright prospects for cancer therapy, there are still challenges in this field: (a) how to improve loading efficiency; (b) how to enhance delivery efficacy and targeted uptake by recipient cells; (c) how to reduce off-target effects; and (d) how to evaluate the appropriate dose and ensure quality control. To overcome these challenges, various engineering strategies have been applied for both presecretion cells and post-purification of exosomes. At present, anticancer cargos can be loaded into exosomes with high loading efficiency using a range of techniques, including electroporation, incubation, and sonication.135,150-152 Appropriate modification of surface structures is beneficial to improving the delivery ability and targeted therapeutic efficacy of exosomes. For surface modification, hydrophobic insertion, receptor-ligand binding, and multivalent electrostatic interactions are useful tools in established engineering technologies. 153 For example, exosomes incorporated with an aminoethylanisamide-polyethylene glycol (AA-PEG) vector have been discovered to target gather in lung cancer cells by binding to sigma receptor, enhancing their therapeutic efficacy. 154 Mouse immature dendritic cell (imDC)-derived exosomes modified by engineering the imDCs to express Lamp2b, an exosomal membrane protein, fused to the αγ integrin-specific iRGD peptide has been used to achieve better tumor targeting. 136 Nevertheless, in the aspects of reducing off-target effects, evaluating appropriate doses and ensuring quality control, there is still much confusion that needs to be resolved.
Conclusions and perspectives
Exosomes, an important subtype of EVs, are released by nearly all cell types and are involved in intercellular communication during various physiological and pathological processes. Currently, increasing evidence has confirmed that exosomes play a vital role in cancer, including in invasion, metastasis, chemoresistance, recurrence, and immunosuppression. Despite the various techniques developed for exosome isolation, identification and quantification, the lack of a uniform standard method is still a crucial obstacle in current studies of exosomes.155,156 Diverse proteins, nucleic acids and lipids are protected in exosomes and exist in various body fluids and can be used as novel and promising biomarkers of cancer. Detection of exosomal mRNAs and miRNAs can provide information for early diagnosis, monitoring of therapeutic response, forecasting recurrence, and determining prognosis in cancer patients. Additionally, understanding the mechanism of exosome biogenesis, secretion, and uptake, and how exosomal mRNAs and miRNAs participate in cancer metastasis and drug resistance can accelerate the development of novel approaches in cancer therapy. For example, cancer progression and metastasis can be inhibited by blocking metastasis-associated exosomal mRNAs and miRNAs, and a better anti-cancer drug response can be induced by disturbing drug-resistance-associated exosomal mRNAs and miRNAs. With the advantages of high biocompatibility and low immunogenicity, exosomes are an ideal delivery platform for mRNAs and miRNAs in cancer therapy. Nevertheless, the challenges of how to improve loading efficiency and targeted uptake and how to evaluate and quantify exosomes need to be addressed. Currently, most research is focused on exosomal miRNAs, and the data concerning exosomal mRNAs in cancer are still relatively limited; thus, more studies focused on exosomal mRNAs are needed in the future. In addition, although abundant research has revealed the bright prospects of exosomal mRNAs and miRNAs in cancer diagnosis, prognosis, and therapy, further work is required before clinical application can be realized.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (Grant No. 81572528, 81872427, 81672428), Outstanding Youth Fund of Heilongjiang Province (Grant No. JC2018024), and the Haiyan Fund Project of Harbin Medical University Cancer Hospital (Grant No. JJQN2019-02).
