Abstract
Objective:
The purpose of this systematic review and meta-analysis was to evaluate the efficacy of fecal Fusobacterium nucleatum (Fn) for detecting colorectal cancer. It is the first systematic review and meta-analysis to focus exclusively on fecal Fn for colorectal cancer.
Materials and methods:
Comprehensive searches of several databases before January 2018 were conducted. Fecal Fn for detecting colorectal cancer was evaluated via pooled sensitivity, specificity, positive likelihood ratio, negative likelihood ratio, diagnostic odds ratio, and area under the curve. Heterogeneity was explored using meta-regression and subgroup analyses. The publication bias and the overall quality of evidence were also analyzed.
Results:
Our analysis included six papers containing seven case-control studies in the systematic review and meta-analysis. Overall sensitivity and specificity were 0.68 (95% confidence interval (CI) 0.64, 0.72) and 0.78 (95% CI 0.75, 0.81), respectively. The positive likelihood ratio and negative likelihood ratio in detecting colorectal cancer were 2.87 (95% CI 1.62, 5.10) and 0.40 (95% CI 0.30, 0.54) respectively. The diagnostic odds ratio (OR) was 8.75 (95% CI 4.86, 15.78) and the area under the curve was 0.80. A subgroup analysis showed that in Asia, the colorectal cancer sample size ⩾50 had higher specificity (specificity 0.85, 95% CI 0.80, 0.88). No publication bias existed. The GRADEpro showed a moderate level of the available evidence.
Conclusions:
Compared to other examinations, the fecal Fn test seems a good choice for detecting colorectal cancer. It also has better diagnostic performance in Asians. However, more clinical trials with large sample sizes and strict randomization are needed to further verify the evidence.
Introduction
Colorectal cancer (CRC) represents the third most commonly diagnosed malignancy and is the fourth leading cause of cancer death in the world 1 . According to the International Agency for Research on Cancer report, the global burden of CRC is expected to increase by 60% to more than 2.2 million new cases and 1.1 million cancer deaths by 2030 2 . Despite massive efforts in surgery, chemotherapy, radiotherapy, and targeted therapy, these therapies suffer from novel problems including acquired resistance, adverse effects, and high costs. As such, accurate approaches for the early diagnosis of CRC are required for intensive monitoring to optimize the likelihood of timely and successful treatment.
In general, endoscopy is the gold standard for the initial diagnosis of CRC, which should be confirmed by histological examination of biopsy specimens 3 . However, the U.S. Preventive Services Task Force found adequate evidence of harm associated with endoscopy 4 . The invasive nature, complications, time, and high cost represent the main barriers 4 . Thus, there is a need for inexpensive and non-invasive methods with improved sensitivity and specificity for the detection of colorectal lesions.
Intestinal microbiota represent a complex ecosystem, including firmicutes, bacteroidetes, actinobacteria, and proteobacteria 5 . Its development is the result of interactions between physiological process in the host and micro-organisms 6 . Alterations in the intestinal microbiota, named dysbiosis, are often linked to disease development, especially CRC5,7,8. Recently, specific bacterial populations have been identified to play a role in CRC initiation and progression, including Streptococcus bovis, Fusobacterium nucleatum (Fn), Enterococcus faecalis, Escherichia coli, and Bacteroides fragilis. Among them, Fn, an adherent, invasive, and proinflammatory bacterium, has been verified as one of the most important factors for CRC9,10. Since the detection of molecular biomarkers in fecal samples for CRC may be a more promising alternative than blood biomarkers or histological examination to be implemented in present clinical settings, an increasing number of diagnostic trials are being carried out to explore the efficacy of the fecal Fn test in CRC detection. However, its diagnostic performance, determined by polymerase chain reaction (PCR), although easy and less costly, varied across these studies (Table 1), resulting in its diagnostic accuracy being unclear. Therefore we conducted a systematic review and meta-analysis to assess the efficacy of fecal Fn levels for the detection of CRC between CRC patients and individuals in the control group according to the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines 11 .
Characteristics of eligible studies included in the meta-analysis.
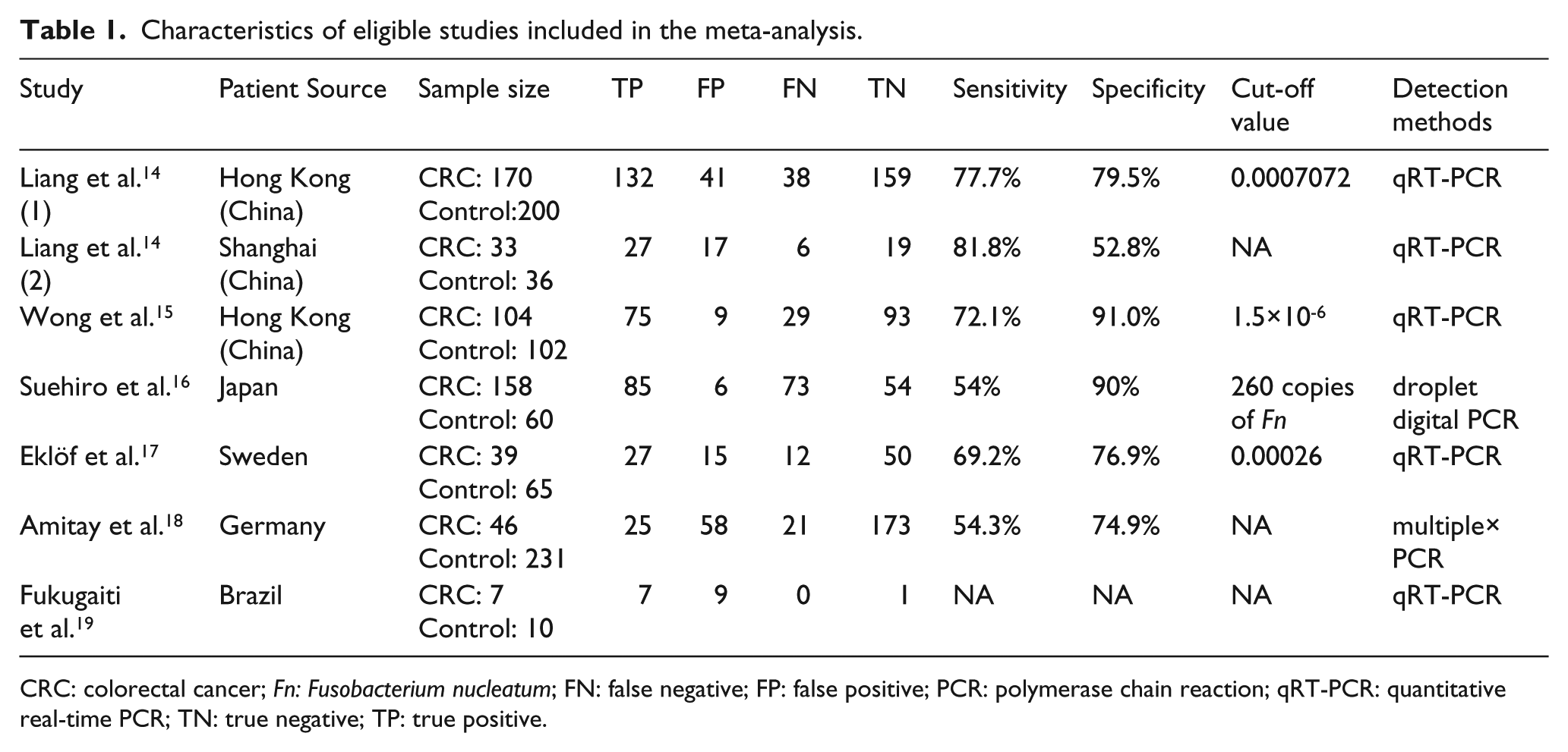
CRC: colorectal cancer; Fn: Fusobacterium nucleatum; FN: false negative; FP: false positive; PCR: polymerase chain reaction; qRT-PCR: quantitative real-time PCR; TN: true negative; TP: true positive.
Materials and methods
Inclusion criteria
The aim of the study selection was to evaluate the performance of Fn using human fecal samples. Studies included in our review met the following criteria: (i) samples came from feces; (ii) there were at least two independent cohorts including CRC patients and controls in each study; (iii) each study provided sufficient data for constructing the diagnostic four-fold (2×2) contingency table; (iv) all the papers were in English or Chinese; and (v) if data or data subsets were used in more than one article, the article with the most detail or the most recent article was chosen.
Search strategies
The search was performed from 1 January 2017. According to inclusion criteria, we comprehensively searched for potentially eligible literature published from 1 January 1990 to 1 January 2018 through PubMed, Embase, Web of Science, the Cochrane Library, Chinese Wan Fang, and the China National Knowledge Infrastructure (CNKI) database. Search terms included CRC, fecal, and Fn. Reference lists of the reports selected in the original search were also examined. The search strategy is summarized in Supplementary Table 1.
Exclusion criteria
We excluded the following: (i) duplicate publications; (ii) studies of case reports, letters, or reviews; (iii) studies without usable data; (iv) studies with cell lines or animals; and (v) samples not from feces.
Date extraction
A team of two reviewers (Huang Q and Peng YH) screened all titles, abstracts and full texts independently and in duplicate. A third reviewer (Xie FW) acted to resolve any disagreements. We extracted the following details from the eligible studies: the first author, date of publication, country of origin, ethnicity, number of patients, detection methods for fecal Fn, and outcome data (true positives (TP), false positives (FP), false negatives (FN), and true negatives (TN)). If the statistical data were described in the study, we extracted them directly. Otherwise, they were calculated from available numerical data in the articles. Three independent researchers calculated data independently to reduce variability. We also sent emails to the corresponding authors of eligible articles for original data needed for the meta-analysis.
Quality assessment
We used the Quality Assessment of Diagnostic Accuracy Studies (QUADAS-2) tool to assess the methodological quality of the included studies 12 . The QUADAS-2 scale contains four domains: patient selection, index test, reference standard, and flow and timing. We assessed all domains for potential risk of bias, and the first three domains for concerns regarding applicability. Each question was assigned a “yes,” “no,” or “unclear” response when every eligible study was evaluated.
Data analysis
We extracted all accuracy data from each study (TP, FP, TN, and FN) to obtain pooled sensitivity, specificity, positive likelihood ratio (PLR), negative likelihood ratio (NLR), diagnostic odds ratio (DOR), and their 95% confidence interval (CI). A summary receiver operating characteristics (SROC) curve was constructed for recruited studies and area under ROC curve (AUC) was calculated to estimate the overall accuracy. A preferred test had an AUC close to 1, while a poor test had an AUC close to 0.5.
Heterogeneity was assessed using a bivariate boxplot, a Chi-square test, and an inconsistency index (I2). If I2 was greater than 50%, significant heterogeneity would be considered to exist among the studies 13 . Then we would use the random-effects model and perform subgroup analyses by different ethnicities, detection methods, CRC sample size, or patient number if data were available. We also performed a likelihood ratio scatter gram to evaluate the exclusion and confirmation capacities of the index test.
Finally, we used Egger’s publication bias plot to assess the publication bias in our meta-analysis. Quality of evidence was assessed across important outcomes using the GRADE approach to support management recommendations by the GRADEpro software (version 3.6).
The statistical analysis was conducted using Meta-Disc Version 1.4 software (XI Cochrane Colloquium, Barcelona, Spain) and STATA vision 15.0 (Stata Corporation, College Station, TX, USA). Meta-disc software was used to perform data analysis, meta-regression analysis, and heterogeneity testing. STATA software was used to determine if publication bias was present, as well as to calculate the relative risks of fecal Fn for the detection of CRC. A P value <0.05 was considered to be statistically significant.
Results
Literature screening
As shown in Figure 1, the literature search process is summarized in a flow diagram according to PRISMA. We retrieved 656 original studies from electronic bibliographic databases published between 1990 and 2018. After 285 duplications were excluded, we screened the titles and abstracts of 371 publications, whereby another 353 papers that did not meet the inclusion criteria were excluded. We downloaded the full text of the remaining 18 publications for future screening, of which 2 studies with reported overlapping patients and 10 studies with incomplete data were excluded. Finally, 6 articles containing 7 case-control studies met the inclusion criteria and were completely analyzed14-19.
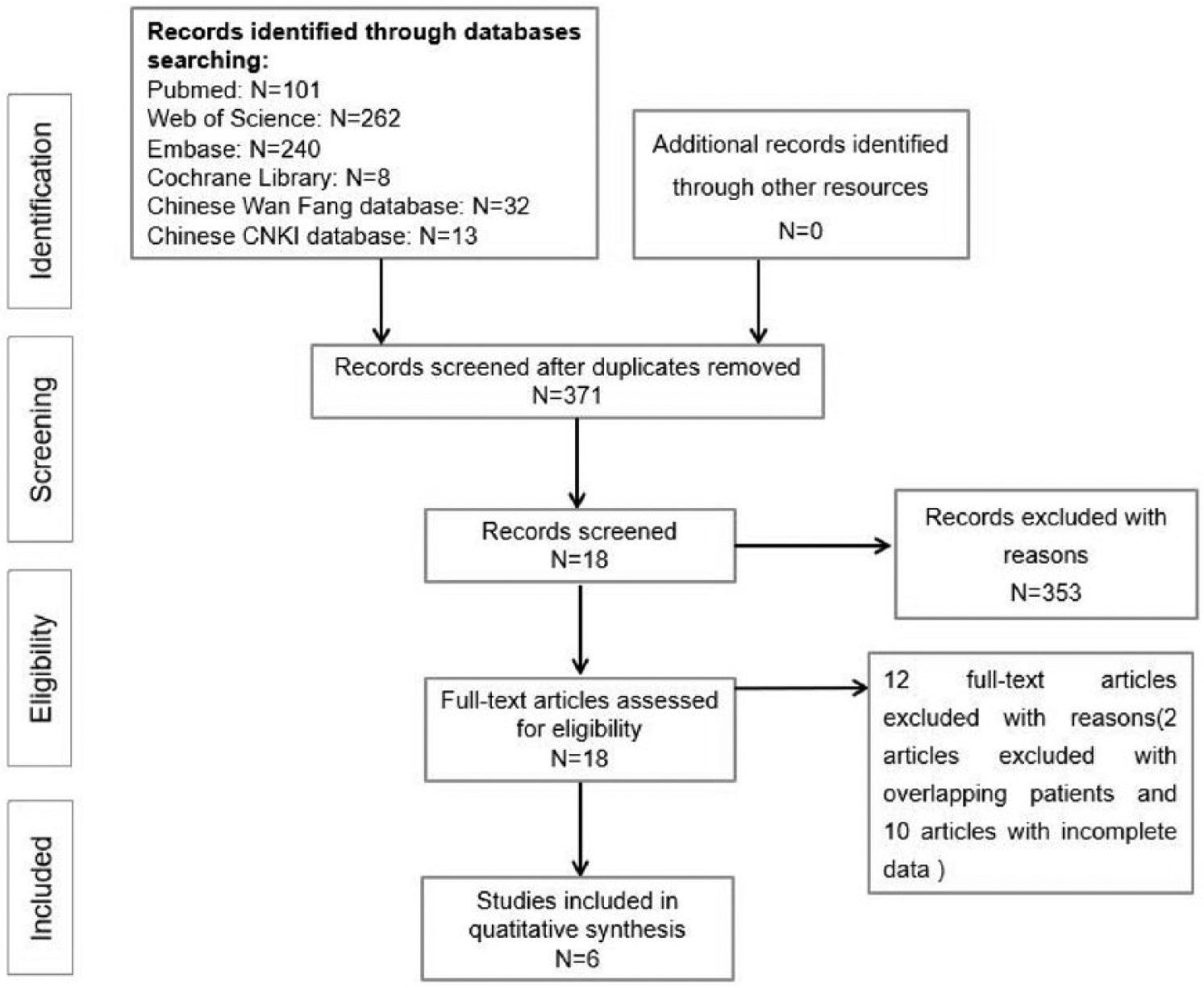
Flow chart of the systematic review and meta-analysis process.
Characteristics of eligible studies
The clinical details of the included studies are presented in Table 1. Regarding the six included articles, one was conducted in both Hong Kong and in Shanghai, the others were in Hong Kong, Japan, Sweden, Germany, and Brazil. All of these studies had been published in the last 3 years (2015–2017). Because the patients in Liang’s study 14 were from two regions and were calculated separately, we have regarded them as two experiments. Fecal Fn was evaluated by quantitative real-time PCR (qRT-PCR), droplet digital PCR, or multiplex PCR. All studies included in the meta-analysis used the histopathological examination as the gold standard. Figure 2 shows an overview of the methodological quality results. In general, the overall quality of the eligible studies was high.
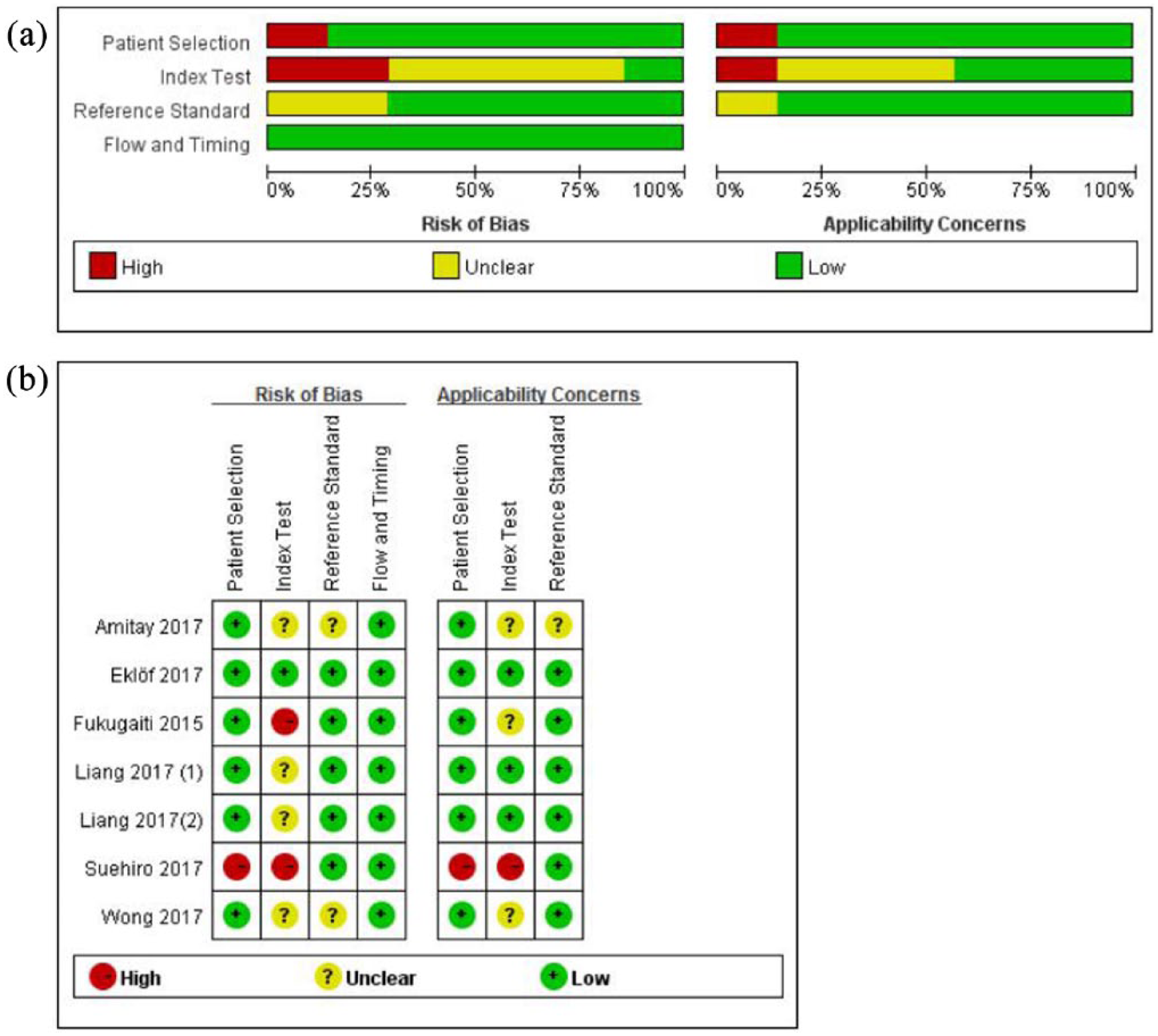
An overview of the methodological quality results. (a) Risk of bias and applicability concerns summary. (b) Risk of bias and applicability concerns graph.
Diagnostic performance in meta-analysis
Heterogeneity in sensitivity and specificity were observed among these studies (I2=82.7% and 88.6%, respectively). Therefore, the random effects model was used. For fecal Fn, the sensitivity, specificity, PLR, NLR, diagnostic score, and DOR of these studies were performed by forest plots. As shown in Figure 3, the pooled sensitivity and specificity of fecal Fn are 0.68 (95% CI 0.64, 0.72) and 0.78 (95% CI 0.75, 0.81), respectively. Its PLR and NLR in diagnosis CRC were 2.87 (95% CI 1.62, 5.10) and 0.40 (95% CI 0.30, 0.54) separately. The DOR is 8.75 (95% CI 4.86, 15.78). As shown in the likelihood ratio scattergram (Supplement Figure 1), the summary PLR and NLR for fecal Fn was on the right under quadrant (LUQ), suggesting that the exclusion and confirmation of fecal Fn for detecting CRC were not bad. The SROC curve for the included studies is shown in Figure 3. The AUC was 0.80, indicating a good diagnostic accuracy of fecal Fn for CRC diagnosis. Furthermore, we used a Fagan diagram to evaluate the overall value of fecal Fn for the detection of CRC. As shown in Supplement Figure 2, the post-test probability (PLR: 41%, NLR: 8%) differed substantially from the pretest probability (20%).
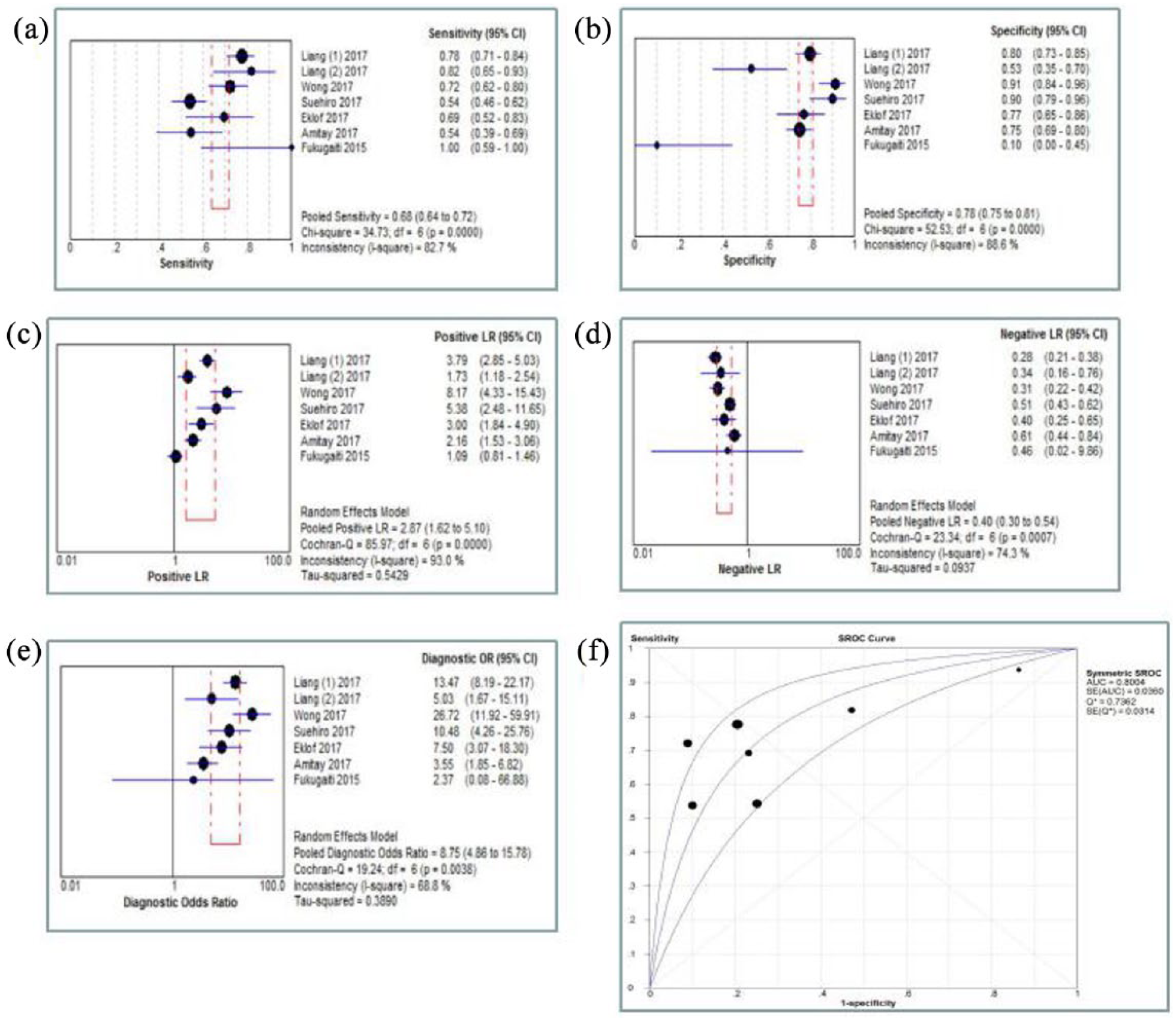
Meta-analysis of pooled sensitivity (a), specificity (b), PLR (c), NLR (d), DOR (e), and SROC curve (f).
Threshold effect and heterogeneity
It is known that heterogeneity can influence the accuracy of a diagnosis. The threshold effect, one source of heterogeneity, is partly due to differences of sensitivity and specificity. Spearman correlation coefficient of sensitivity and specificity is a good way to evaluate the threshold effect. 20 However, in our meta-analysis, the Spearman correlation coefficient of sensitivity and 1-specificity was 0.571 with a P-value of 0.180 (P>0.05), suggesting that a non-threshold effect existed among these studies. Then we performed the meta-regression based on the variables, including ethnicity (Asian/European/American population), detection methods (qRT-PCR vs. other PCR assays), patient number (n⩾150 vs. n<150), CRC sample size (n⩾50 vs. n<50) to explain this heterogeneity. Among the four factors, CRC sample size was identified as statistically significant (P=0.03), indicating that the CRC sample size was responsible for the relatively high heterogeneity (Supplement Table 2).
Subgroup analysis
The results of subgroup analysis of the different ethnicities (Asia vs. Europe & America) are shown in Supplement Table 3 and Supplement Figure 3. The sensitivity and specificity were 0.69 (95% CI 0.64, 0.73) and 0.82 (95% CI 0.77, 0.85) in Asia, which were higher than in Europe and America (sensitivity=0.64 (95% CI 0.53, 0.74), specificity=0.73 (95% CI 0.68, 0.78)). The AUC in Asia was 0.834, while the AUC in Europe or America was 0.736. When the CRC sample size<50, the sensitivity and specificity were 0.69 (95% CI 0.60, 0.77) and 0.71 (95% CI 0.66, 0.76). When the CRC sample size⩾50, the sensitivity and specificity were 0.68 (95% CI 0.63, 0.72) and 0.85 (95% CI 0.80, 0.88). Taken together, in Asia, the CRC sample size⩾50 had higher specificity (0.85, 95% CI 0.80, 0.88). However, similar sensitivity and specificity achieved between patient numbers ⩾150 and patient numbers <150.
Publication bias
To assess the publication bias for the diagnostic tests, we used Egger’s publication bias plot by STATA 15.0 software. The P-value obtained from the plot was 0.255, indicating that the presence of publication bias did not exist in this meta-analysis (Supplement Figure 4).
Overall quality of evidence by GRADE
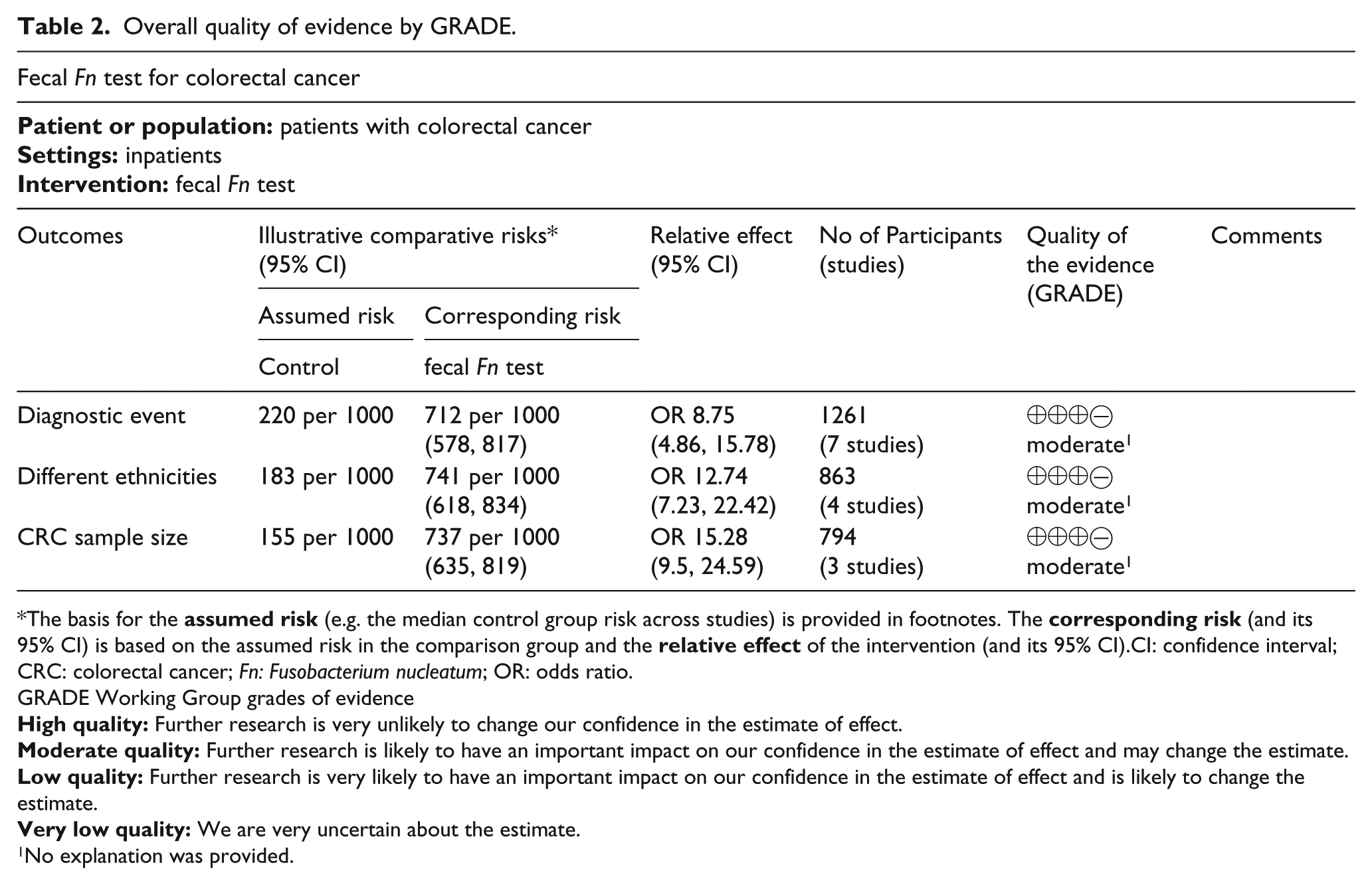
We graded the overall quality of available evidence by GRADE approach. The quality of evidence for all outcomes was moderate because of their inconsistency (Table 2).
Overall quality of evidence by GRADE.
The basis for the
GRADE Working Group grades of evidence
No explanation was provided.
Discussion and future directions
In this systematic review and meta-analysis, the fecal Fn test had pooled sensitivity of 68%, specificity of 78%, DOR of 8.75, and AUC of 0.80. Moreover, the test showed better discrimination ability for detecting CRC among Asians. To the best of our knowledge, ours is the first systematic review and meta-analysis to assess the diagnostic value of the fecal Fn test for screening CRC.
The pooled DOR was 8.75 (4.86–15.78), indicating the test’s relatively high discrimination ability. The pooled sensitivity and specificity was 68% and 78%, respectively, suggesting the test’s superior ability for ruling out patients without CRC. However, the PLR and NLR of the test was 2.87 (95% CI 1.62, 5.10) and 0.40 (95% CI 0.30, 0.54), respectively, which shows that patients with CRC have approximately 2.87 times higher possibility of testing positive compared with subjects without CRC, as well as a 40% chance of an individual having CRC if the test is negative. The performance of the fecal Fn test in the pooled PLR and NLR did not achieve the requirements of clinical practice, and remains to be modified for clinical confirmation and exclusion purposes. Anyway, the pooled results suggest that the fecal Fn test has good discrimination ability in clinical practice.
In addition to our review, some articles confirmed that the fecal Fn test was promising in the detection of CRC from other aspects. For example, as for unity in different populations and races, Zeller et al. 21 used 21 species based on shotgun metagenomic sequencing, discovered in a French cohort, to classify patients with CRC in a German cohort. Higher accuracy (AUC 0.85) was reached, suggesting that metagenomic CRC prediction and detection are possible with high accuracy despite the differences between the study populations 21 . Yu and his colleagues 20 reached similar conclusions by comparing the fecal microbiome of 74 CRC patients and 54 controls from China, and 16 patients and 24 controls from Denmark. They identified four microbial gene markers that differentiated CRC and control microbiomes, including Fn 20 . Furthermore, these markers were enriched in early-stage (I-II) patient microbiomes, highlighting the potential of using fecal Fn for early detection of CRC. Considering the difficulty, the high cost, and the long process of sequencing analyses, the fecal candidates identified by metagenome sequencing were examined by PCR assays 20 . The result was that the PCR assays showed as good a correlation with Fn at the species level as with metagenome sequencing. Our review also verified the benefits of PCR in the detection of fecal Fn.
Furthermore, our included studies found that fecal Fn might complement the existing fecal immunochemical test (FIT)—another cheaper, less invasive stool-based test for CRC—in distinguishing healthy individuals from those with CRC. For example, Wong et al. 15 evaluated the diagnostic performance of Fn with respect to FIT, and validated the results in 181 subjects. They reported that Fn, when combined with FIT, showed superior sensitivity (92.3% vs. 73.1%, P<0.001) and AUC (0.95 vs. 0.86, P<0.001) than stand-alone FIT in detecting CRC. This combined test also increased the sensitivity (38.6% vs. 15.5%, P<0.001) and AUC (0.65 vs. 0.57, P=0.007) for detecting advanced adenoma. Liang et al. 14 reached a similar result through conducting a more comprehensive study. From the above, it is further concluded that fecal Fn can be an excellent marker for the non-invasive diagnosis of CRC.
We identified some weaknesses in our review. First, the heterogeneity test and meta-regression analysis showed significant heterogeneity across the included studies, and the CRC sample size may be the potential origin of the relatively high heterogeneity, which may also contribute to the overall quality of evidence. Second, the number of patients and studies included were relatively small. Finally, these studies lacked a unified criterion for Fn positive expression. Thus, larger-scale, multicenter, and higher-quality studies are required to confirm our findings in the future.
In conclusion, the fecal Fn test shows good discrimination ability for detecting colorectal cancer in our systematic review and meta-analysis. It also has better diagnostic performance in Asians. We certainly recognize that our suggested method is incomplete, mainly from the limited sample size, but it is indicated that the fecal Fn test represents at least a starting point toward non-invasive and accurate procedures in the detection of CRC. Larger-scale, multicenter, and higher-quality studies are required to confirm our findings in the future, allowing a better comparison of analytical results of Fn on clinical outcomes.
Supplemental Material
Supplementary_material – Supplemental material for Fecal fusobacterium nucleatum for detecting colorectal cancer: a systematic review and meta-analysis
Supplemental material, Supplementary_material for Fecal fusobacterium nucleatum for detecting colorectal cancer: a systematic review and meta-analysis by Qian Huang, Yonghai Peng and Fangwei Xie in The International Journal of Biological Markers
Supplemental Material
supplementfigure4 – Supplemental material for Fecal fusobacterium nucleatum for detecting colorectal cancer: a systematic review and meta-analysis
Supplemental material, supplementfigure4 for Fecal fusobacterium nucleatum for detecting colorectal cancer: a systematic review and meta-analysis by Qian Huang, Yonghai Peng and Fangwei Xie in The International Journal of Biological Markers
Supplemental Material
supplement_figure2 – Supplemental material for Fecal fusobacterium nucleatum for detecting colorectal cancer: a systematic review and meta-analysis
Supplemental material, supplement_figure2 for Fecal fusobacterium nucleatum for detecting colorectal cancer: a systematic review and meta-analysis by Qian Huang, Yonghai Peng and Fangwei Xie in The International Journal of Biological Markers
Supplemental Material
supplement_figure3 – Supplemental material for Fecal fusobacterium nucleatum for detecting colorectal cancer: a systematic review and meta-analysis
Supplemental material, supplement_figure3 for Fecal fusobacterium nucleatum for detecting colorectal cancer: a systematic review and meta-analysis by Qian Huang, Yonghai Peng and Fangwei Xie in The International Journal of Biological Markers
Supplemental Material
supplement_figure_1 – Supplemental material for Fecal fusobacterium nucleatum for detecting colorectal cancer: a systematic review and meta-analysis
Supplemental material, supplement_figure_1 for Fecal fusobacterium nucleatum for detecting colorectal cancer: a systematic review and meta-analysis by Qian Huang, Yonghai Peng and Fangwei Xie in The International Journal of Biological Markers
Footnotes
Acknowledgements
This work was supported through funds provided by two projects. The authors apologize to those colleagues whose contributions to this research field could not be cited due to space limitations, and thank all anonymous reviewers for their time, comments, and constructive criticism.
Declaration of conflicting interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The corresponding author Fangwei Xie is funded through the Natural Science Foundation of Fujian Province, P.R. China (Grant No. 2016J01579). The first author, Dr. Qian Huang, is funded through the Natural Science Foundation of Fujian Province, P.R. China (Grant No. 2017J01329).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
