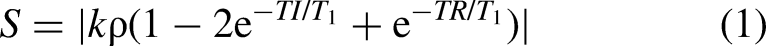Abstract
The magnetosome, a membrane-bound iron biomineral formed within magnetotactic bacteria, is a unique model for a magnetic resonance imaging (MRI) reporter gene contrast agent. We are translating this technology to mammalian cells by expressing essential magnetosome genes mamI, mamL, mamB, and mamE into MDA-MB-435 human melanoma cells. Currently, these genes are individually expressed in the cell although the future goal is to express all four genes together. We examined the influence of these genes on cellular MRI relaxation rates by culturing cells in the presence and absence of iron-supplemented medium and scanning them at 3 Tesla using a gelatin phantom. Total cellular iron was measured by inductively-coupled plasma mass spectrometry and correlated with relaxation rates obtained from phantom experiments. Apart from mamE, magnetosome genes that are individually expressed in mammalian cells grown in iron supplement significantly affected cellular transverse relaxation rates compared to cells grown without iron supplement. Interestingly, mamI, mamL, mamB, and mamE (even though the latter had no effect on relaxation rate) significantly affected cellular iron content. This developing gene-based contrast agent will equip MRI with improvement to imaging sensitivity and the technology to track cellular activities long term.
Keywords
Introduction
Molecular imaging of biological processes 1 permits the monitoring of diseases from diagnosis through therapy. Molecular probes in the form of contrast agents have been developed as molecular imaging tools for modalities such as computed optical imaging, positron emission tomography, and magnetic resonance imaging (MRI). 2 Among these, MRI is ideal for multiparametric imaging and discerning anatomic detail at any tissue depth; 3 however, MRI lacks the sensitivity and specificity needed to effectively detect and track cellular and molecular activities. To address this, gene-based, magnetic resonance (MR) contrast agents are being developed for molecular imaging. 4 These endogenous contrast agents originate from the cell and remain throughout its life cycle, providing many advantages for long-term, repetitive imaging of disease progression.
However, compared to the production of gene contrast as applied to optical imaging, including fluorescent imaging and bioluminescent imaging, which is dependent on the transfection of a single gene, developing an effective gene reporter probe for MRI has many additional challenges and complexities. It was recognized about 20 years ago that the iron particle known as a magnetosome produced by magnetotactic bacteria (MTB) could be the foundation for such an MRI reporter gene. Magnetosome formation compartmentalizes iron biominerals in membrane-enclosed vesicles. 5 This not only provides genetically controlled iron biomineralization but also prevents iron cytotoxicity by sequestering the crystal within a lipid bilayer. Initially, it was thought to require a single gene (i.e., MagA, Mms6); however, the size of the change on the MRI signal of these single gene reporters has been disappointingly small compared to changes observed in MTB which involves all the associated genes. For example, the increase in the spin–spin relaxation time (R2) of the magA or mms6 reporter gene in mammalian cells is of the order of 15 s−16–11 whereas for MTB within mouse tumors it is more than 300 s−1. 12
A large body of work has been undertaken to show how the production of magnetosomes by MTB can be genetically engineered to produce iron particles of different sizes 13 which are enclosed within a membrane and can be coated with different proteins as biomarkers for specific molecular targets.14–16 The development of the MRI reporter gene can instead do all this through incorporation of the needed genes in the DNA material in addition to advancing molecular imaging and precision medicine to the next level.
Hence, we have started to investigate the potential of creating a reporter that incorporates four of the genes on the genomic island that are considered essential. If we could engineer a reporter that would result in a magnetosome concentration within mammalian cells identical to the concentration within the MTB, then we have determined that the detection limit for large animals and humans would be less than 3000 cells and for small animals (i.e., mouse) less than three cells. 4 Even if within the entire mammalian cell magnetosome numbers were similar to the number in the much smaller MTB, the cell detection limit would still be significant (300,000 cells for large subjects and 300 cells for the mouse). This would be a substantial improvement compared to conventional MR imaging.
In MTB, magnetosome formation is a stepwise, protein-directed process that begins with vesicle formation and culminates with iron biomineralization. 17 As the biology of magnetosome formation by MTB matured, it became apparent that as many as 30 genes are involved. 16 Introducing such a large number of genes is not practical given the genetic engineering difficulties and the potential for transfection rejection in the mammalian cell. Fortunately, studies of deletion mutants revealed several essential genes involved in the various stages of magnetosome synthesis.5,16–19 To improve the localization and compartmentalization of a rudimentary magnetosome-like particle in mammalian cells, we are introducing select genes deemed essential for the initial stages of magnetosome formation. The targeted genes, mamI, mamL, mamB, and mamE, are clustered on the mamAB operon and highly conserved in species of MTB.5,18 The putative roles of mamI, mamL, and mamB involve magnetosome vesicle formation17,19 and may even provide docking site(s) for additional protein(s) 4 that facilitate iron biomineralization, like MamE.20,21 MamE also has a reported role in protein sorting and has an essential role in recruiting specific proteins onto the magnetosome membrane. 22 Additionally, MamI and MamB have been suggested to have nonessential iron-handling roles. The luminal loop of MamI has been shown to bind to magnetite. 23 Although studies in AMB-1 have not been conducted, it has been reported that MamB in MSR-1 dimerizes to transport iron and binds iron ions at the C-terminal domain. 19 To date, no studies have revealed any iron-handling abilities in MamL.
We used the human melanoma tumor cell line MDA-MB-435 to evaluate the effect(s) of magnetosome proteins MamI, MamL, MamB, and MamE on cellular MR relaxation rates and total cellular iron content. While each magnetosome protein increases elemental iron content in response to iron-supplemented cell culture, only MamI, MamL, and MamB increase MR relaxation rates. Truncation of the extramembranous MamL C-terminal peptide does not alter the novel iron-handling properties of MamL, whereas the MR silent nature of MamE provides a new context for interpreting its localization and functionality in a foreign cell environment.
Materials and Methods
Molecular Cloning
Magnetosome genes mamI and mamL were amplified by PCR from the genomic DNA of Magnetospirillum magneticum strain AMB-1 (ATCC 700264) using custom primers. Briefly, the mamI and mamL amplicons were purified using a PCR clean-up kit (Invitrogen, Life Technologies, Burlington, Canada); digested with appropriate restriction enzymes; and purified once more prior to insertion in the molecular cloning vectors pEGFP-C1 (Clontech) and ptdTomato-C1 (Clontech), respectively. After propagation in Escherichia coli strain XL10GOLD, vector-insert plasmid constructs were purified and used for mammalian cell transfection.
For cloning of Tomato-MamLtrunc, the last 15 amino acids from the C-terminus of MamL were removed by PCR site-directed mutagenesis. Briefly, primers were designed that included a stop codon 45 nucleotides upstream from the end of mamL. These primers were then used in PCR amplification of the truncated mamL gene, which was then inserted into the ptdTomato-C1 vector with restriction enzymes EcoRI and BglII. The vector-insert construct was propagated in E. coli strain XL10GOLD; purified; and then used for mammalian cell transfection.
For cloning of Tomato-MamB, the mamB gene was amplified by PCR from AMB-1 genomic DNA using custom primers. The amplicon was purified with a PCR clean-up kit; digested with restriction enzymes; purified once more; and inserted into the ptdTomato-C1 vector (Clontech). The vector-insert construct was propagated in E. coli strain XL10GOLD; purified; and then used for mammalian cell transfection.
For cloning of EGFP-MamE, the mamE gene was amplified by PCR from AMB-1 genomic DNA using custom primers. The amplicon was purified with a PCR clean-up kit; digested with restriction enzymes; purified once more; and inserted into the pEGFP-C1 vector (Clontech). The vector-insert construct was propagated in E. coli strain XL10GOLD; purified; and then used for mammalian cell transfection.
Cell Culture
MDA-MB-435 cells (ATCC HTB-129; derived from an adult female and characterized as a melanoma cell line) are a model of aggressive tumorigenesis. 8 Cells were cultured in 100-mm polystyrene-coated cell culture dishes (CELLSTAR, VWR International, Mississauga, Canada) with Dulbecco's Modified Eagle Medium containing 1 g/L glucose (Gibco, Life Technologies, Burlington, Canada), 10% fetal bovine serum (Gibco), 4 U/mL penicillin, and 4 µg/mL streptomycin at 37 °C with 5% CO2.
Cell lines expressing fluorescent fusion proteins, enhanced green fluorescent protein (EGFP)-MamI and EGFP-MamE, or tdTomato (Tomato)-MamL, Tomato-MamLtrunc, or Tomato-MamB, were each maintained in medium containing 500 µg/mL Geneticin (G418; Gibco).
Cell Harvest and Phantom Preparation
A spherical MR phantom was constructed using two 9 cm plastic hemispherical molds. 7 One hemisphere was filled with 4% gelatin (porcine type A)/phosphate-buffered saline pH 7.4 (PBS; 137 mM NaCl, 2.7 mM KCl, 8 mM Na2HPO4, 2 mM KH2PO4) while the other half contains sample wells embedded in 4% gelatin/PBS (Figure 1).

MRI phantom. (A) Representative locator images show an MRI phantom containing cells stably expressing Tomato-MamB. The 3-mm slice (thin lines denoted with diamond symbols, left panel) was positioned perpendicular to the wells and within the volume occupied by cells. An axial cross section of the phantom (right panel) shows the layout of Ultem wells (white circles) and reference peg (black circle). (B) A representative image of the phantom in part A is displayed at TE = 60 ms. Wells 1 and 2 contain cells grown in iron-supplemented medium whereas wells 3 and 4 contain cells grown in nonsupplemented medium. The scale bar represents signal intensity (arbitrary units).
To prepare cell samples, MDA-MB-435 cells expressing MamI, MamL, MamLtrunc, MamB, or MamE were cultured in the presence or absence of medium supplemented with 250 µM ferric nitrate as previously described. 7 For each sample, approximately 150 million cells were grown and harvested using 0.05% trypsin ethylenediamine tetraacetic acid (EDTA, Gibco). Harvested cells were collected by centrifugation at 400×g for 5 min at 15 °C and washed three times in PBS to remove culture media. The final cell slurry was loaded into a custom-made Ultem well (inner diameter 4 mm, height 10 mm; Lawson Imaging Prototype Lab), using centrifugation to fill the well with cells (approximately 100 million cells). The remaining cells were analyzed for elemental iron and zinc content, as described below. Each cell phantom contained four sample wells.
Magnetic Resonance Imaging Sequences
Cell phantoms were scanned at 3 Tesla (T) on a Biograph mMR (Siemens AG, Erlangen, Germany) using previously described sequences to acquire longitudinal and transverse relaxation rates (R1 = 1/T1; R2 = 1/T2; R2* = 1/T2*). 7 All image acquisitions used for relaxation measurement had a slice thickness of 3 mm and field of view of 120 × 120 mm2 (Figure 1).
To measure R1 longitudinal relaxation rates, an inversion recovery (IR) spin echo sequence was used. 7 The matrix size was 128 × 128 and voxel size was 1 × 1 × 3 mm3. Repetition time (TR) was 4,000 ms; excitation flip angle was 90°; and inversion times (TI) were 22, 200, 500, 1,000, 2,000, and 3,900 ms.
For acquisition of R2 transverse relaxation rates, a single-echo (SE) spin echo sequence was used with matrix size of 192 × 192 and voxel size 0.6 × 0.6 × 3.0 mm3. Echo times (TE) were 13, 20, 25, 30, 40, 60, 80, 100, 150, and 200 ms; TR was 2,200 ms; and excitation flip angle was 90°.
For acquisition of R2* transverse relaxation rates, a multi-echo gradient echo (GRE) sequence was used with matrix size of 192 × 192 and voxel size 0.6 × 0.6 × 3.0 mm3. Echo time values were 6.12, 14.64, 23.16, 31.68, 40.2, 50, 60, 70, and 79.9 ms; TR was 2,000 ms; and flip angle was 60°.
MR Data Analysis
Longitudinal relaxation rates were determined using MATLAB 7.9.0 (R2010b). A 9-voxel circular region of interest (ROI) was selected, corresponding to 22 mm3 and encompassing the area containing cells without including the wall of the Ultem well. The average signal intensity within the ROI was determined at each TI and then entered in a MATLAB script to determine R1 values, fitting the curve using a standard IR equation for magnitude signals:
Protein Assay
Cells remaining after preparation of the MRI cell phantom were mixed with 850 μL radioimmunoprecipitation assay buffer and 150 μL Complete Mini protease inhibitor cocktail (Roche Diagnostic Systems, Laval, Canada). Cells lysis was completed by sonication using three 12-s bursts of a Sonic Dismembrator (model 500, Thermo Fischer Scientific, Ottawa, Canada) at an amplitude of 30%. Total amount of protein was quantified using the bicinchoninic acid assay. 25
Sample Preparation for Inductively-Coupled Plasma Mass Spectrometry
Sample protein was adjusted to 2 mg/mL for elemental iron and zinc analysis. Each sample represents the cell population mounted in a single well of the MR phantom. Samples were stored at −20 °C until analyzed by inductively-coupled plasma mass spectrometry (Biotron Analytical Services, Western University). Data are reported as total cellular iron or zinc content normalized to total amount of protein. As zinc values change minimally compared to iron values, interactions (or lack of) with zinc are not discussed further in the text but values are reported in Supplementary Table S1.
Statistical Analysis
All statistical tests were performed using GraphPad Prism version 8.0.1, with p values < 0.05 considered the criterion for statistical significance. Unpaired t-tests were used to identify significant differences between pairs of iron-supplemented and nonsupplemented samples. Spearman's correlation coefficients (r) were calculated to identify significant correlations between relaxation rates and cellular iron (Fe) content, with statistically significant r values ≤ 0.30 considered weak correlations, 0.31–0.69 moderate correlations, and ≥0.70 strong correlations, as described in other publications.
Results
The human melanoma MDA-MB-435 cell line has been used to track changes in MR measures of both cells7,24 and the tumors they form 8 using gene-based contrast from MTB iron-handling protein. Here, we expand this data set by characterizing MRI measures in cells expressing essential magnetosome proteins MamI, MamL, MamB, or MamE. While removal of anyone of these conserved proteins from MTB precludes the formation of a bacterial magnetosome, their precise role in bacteria or mammalian cells is ill-defined. To better characterize the iron-handling nature of these magnetosome proteins, we expressed them in mammalian cells and cultured these in the presence (+ Fe) and absence of iron supplementation, measuring MR relaxation rates under each condition.
Transverse Relaxation Rates in Mammalian Cells Expressing Essential Magnetosome Proteins
In all expression systems, transverse relaxation times displayed a monoexponential signal decay, as shown for MamB-expressing cells (Figure 2A; Supplementary Figure S1). Shortening of both T2* and T2 was evident not only in iron-supplemented cells expressing MamB but also in those expressing MamI or MamL (Supplementary Figure S2). However, unlike MamB, MamI, and MamL, MamE expression did not shorten transverse relaxation times in iron-supplemented mammalian cells (Figure 2B).

Essential magnetosome proteins display monoexponential decay in mammalian expression systems. Cells were cultured in the presence (+ Fe) and absence of iron supplementation; harvested; mounted in a gelatin cell phantom; and scanned at 3T. (A) Representative data from MamB-expressing cells indicate a monoexponential signal decay of transverse relaxation times. In addition, the expression of MamB resulted in shortening of both T2* and T2 when cells were grown in the presence of an iron supplement (circle and square symbols) compared to nonsupplemented medium (upright and inverted triangle symbols). (B) Conversely, regardless of iron supplementation, MamE-expression alone does not significantly shorten the monoexponential decay of T2* or T2. Data are the mean signal intensity ± SD from all 21 voxels in the ROI.
Compared to nonsupplemented cells and the parental control (Supplementary Table S1), iron-supplemented cells expressing fluorescent fusion proteins of MamI, MamL, and MamB significantly increased R2* (Figure 3A) and R2 (Figure 3B) transverse relaxation rates. For MamI- and MamL-expressing cells, there was also a significant increase in the iron-specific R2′ component (R2*–R2; Figure 3C) secondary to iron supplementation.

Transverse relaxation rates of mammalian cells expressing essential magnetosome proteins. Human MDA-MB-435 cells expressing fluorescent magnetosome fusion proteins were cultured in the absence (solid bars, no Fe) or presence (hatched bars, +Fe) of iron-supplemented medium prior to mounting in a gelatin cell phantom for 3T MRI. Bar graphs show transverse relaxation rates of the parental control relative to those cells stably expressing MamI, MamL, MamLtrunc, MamB, or MamE. Compared to controls, iron-supplemented cultures of all magnetosome protein expression systems increased total transverse relaxation rates (R2*, A) and the irreversible component (R2, B) with the exception of MamE-expressing cells. An increase in magnetic field inhomogeneities, as characterized by R2′ (R2*–R2, C) was only evident in iron-supplemented MamI and MamL expression systems. Sample number is indicated either within each bar (A and B) or above them (C). Data are the mean ± SEM (Supplementary Table S1; n = 4–12; * p < 0.05, ** p < 0.01, *** p < 0.001).
The potential role of a MamL C-terminal peptide in modulating MRI relaxation rates was considered. When grown in the presence of an iron supplement, cells expressing MamLtrunc exhibited R2* comparable to full-length MamL: 39.84 ± 5.20 s−1 versus 37.07 ± 5.62 s−1, respectively (Figure 3A; Supplementary Table S1). The R2 and R2′ components (Figure 3B and 3C) were likewise similar in magnitude and relationship to the total transverse relaxation rate (R2*) despite the removal of 15 residues from the MamL C-terminal.
Relative to MamL, the structural model of MamB proposes a larger cytoplasmic-facing domain (Supplementary Figure S3). In MamB-expressing mammalian cells, the influence of iron supplementation is largely accounted for by an increase in R2 (Figure 3B). However, for MamE, the largest of the essential membrane-associated magnetosome proteins examined here, the structural model predicts a large extramembrane domain facing the (magnetosome) vesicle lumen 26 (Supplementary Figure S4). Mammalian cells expressing MamE did not display any difference in R2* or R2 between iron-supplemented and nonsupplemented samples: 16.52 ± 1.02 s−1 versus 14.00 ± 1.22 s−1 and 10.31 ± 1.23 s−1 versus 10.22 ± 1.37 s−1, respectively (Figure 3A and 3B; Supplementary Table S1). Indeed, this cell type produced values for all transverse relaxation rates that were no different than the parental control.
In the absence of iron supplementation, MDA-MB-435 cells expressing MamI, MamL, MamB, or MamE exhibit R2 values that are comparable to those of the parental cell type.
Longitudinal Relaxation Rates in Mammalian Cells Expressing Essential Magnetosome Proteins
Longitudinal R1 relaxation rates of nonsupplemented cells expressing MamI, MamL, MamLtrunc, MamB, or MamE were comparable to the parental cell line: between 0.9 and 1.0 s−1 (Figure 4; Supplementary Table S1). In the presence of iron supplementation, MamI-, MamL-, MamLtrunc-, and MamB-expressing cells displayed small but significant increases in R1 (Figure 4; Supplementary Table S1). As for transverse relaxation, removal of the MamL C-terminal peptide (MamLtrunc expression) had no effect on longitudinal relaxation of iron-supplemented cells compared to full-length MamL (1.12 ± 0.04 vs 1.18 ± 0.11, respectively). Finally, cells expressing MamE showed no substantial changes in R1 whether or not cultured in the presence of iron supplementation.

Longitudinal relaxation rates of mammalian cells expressing essential magnetosome proteins. Human MDA-MB-435 cells expressing fluorescent magnetosome fusion proteins were cultured in the absence (solid bars, no Fe) or presence (hatched bars, +Fe) of iron-supplemented medium prior to mounting in a gelatin cell phantom for 3T MRI. Bar graphs show longitudinal (R1) relaxation rates of the parental control, relative to those cells stably expressing MamI, MamL, MamLtrunc, MamB, or MamE. Apart from small increases in R1 values for iron-supplemented MamI, MamL, and MamB expression systems, the expression of essential magnetosome proteins exerted little influence on the longitudinal relaxation rates of mammalian cells. Sample numbers are indicated within each bar. Data are the mean ± SEM (n = 4-9; * p < 0.05, ** p < 0.01).
Effect of Magnetosome Proteins on Total Cellular Iron Content
Figure 5 demonstrates the influence of iron supplementation on the total cellular iron content of mammalian cells expressing magnetosome proteins. In the absence of iron supplementation, elemental iron analysis of cells expressing MamI, MamL, MamB, or MamE is no different than the parental cell line and less than 0.1 µg Fe/mg protein (Figure 5; Supplementary Table S1). However, in the presence of iron supplementation, all but the parental cell type displays increased cellular iron content, ranging between 0.9 and 1.5 µg Fe/mg protein. These data for iron-supplemented MamI-, MamL-, and MamB-expressing cells are consistent with increases in transverse relaxation rates. Likewise, iron-supplemented cells expressing MamLtrunc exhibit cellular iron content comparable to full-length MamL (0.93 ± 0.07 vs 1.11 ± 0.45 µg Fe/mg protein, Supplementary Table S1). Surprisingly however, iron-supplemented cells expressing MamE displayed similar increases in cellular iron content as all other mammalian cells herein expressing magnetosome proteins, even though transverse relaxation rates of MamE-expressing cells under the same conditions were not altered.

Total cellular iron content of cells stably expressing magnetosome proteins. Human MDA-MB-435 cells expressing fluorescent magnetosome fusion proteins were cultured in the absence (solid bars, no Fe) or presence (hatched bars, +Fe) of iron-supplemented medium prior to analysis by ICP-MS. The bar graph indicates total elemental iron content normalized to total cellular protein of the parental control compared to cells stably expressing MamI, MamL, MamLtrunc, MamB, or MamE. In all cells expressing magnetosome proteins, iron-supplemented cultures resulted in elevated cellular iron content. Sample numbers are indicated above each bar. Data are the mean ± SEM (n = 4-6; * p < 0.01).
Correlation Between MRI Relaxation Rates and Total Cellular Iron Content
By examining the correlation between relaxation rates and total cellular iron content of cells in standard culture (without additional iron supplement, -Fe), we show that R1, R2*, and R2 relaxation rates are not significantly correlated with cellular iron content, with p > 0.05 and low Spearman's correlation coefficients (r2 ≤ 0.04). Only R2′ displays a significant, moderate correlation to cellular iron content, regardless of culture condition.
When samples are cultured in iron-supplemented medium (+ Fe), a significantly strong correlation (r2 = 0.57) exists between R1 and cellular iron content. In addition, R2* displays a moderate to strong correlation (r2 = 0.42) to cellular iron content, unlike the lower but significant correlations with R2 and R2′ (both with r2 < 0.20). Moreover, as expected for these related transverse relaxation rates (R2* = R2 + R2′), the slope of the line is greatest for the correlation between R2* and cellular iron content.
Thus, R1 and R2* are the most responsive to changes in cellular iron resulting from the magnetosome gene expression systems examined in this study. Similar correlations have been reported in iron overload disease. 27
Discussion
This study investigated the effect of four essential magnetosome proteins, MamI, MamL, MamB, and MamE, on the MR relaxation rates and elemental iron content of a human melanoma cell line. Although the similarity of relaxation properties of the systems studied here to those of ferritin-based systems 8 is intriguing, it does not necessarily indicate the presence of ferritin-based iron in our systems. An endoplasmic reticulum labile ferrous iron ion stain (BioTracker Far-red Labile Fe2+ Dye) was used to visualize iron ions in cells expressing magnetosome proteins (Supplementary Figures S5 and S6). Although we cannot confirm the form or state of iron that these magnetosome proteins interact with, these figures do not suggest an increase in ferrous iron ions when cells are grown in the presence of iron supplement. Thus, other forms of iron (such as free ferric ions, iron bound to proteins [ferritin], and/or iron crystals [magnetite]) are causing the increase in total cellular iron seen in Figure 5, and we can speculate that magnetosome proteins MamI, MamL, MamB, and MamE interact with these other forms of iron.
Using previously characterized fluorescent protein fusion protein constructs (Supplementary Figure S7), each stably expressed magnetosome protein exerted its influence in response to iron-supplemented cell culture. Under these conditions, all expression systems increased total cellular iron content more than 5- to 7-fold relative to the parental cell type (0.190 ± 0.007 vs 0.930 ± 0.073 to 1.390 ± 0.076). However, only mammalian cells expressing MamI, MamL, or MamB exhibited increases in MR relaxation rates, with moderate correlations between elemental iron and transverse and longitudinal relaxation rates. Although the MR-silent nature of MamE-expressing cells was surprising, these findings are consistent with distinct localization and proposed role(s) for this magnetosome protein. 6
Recent models of magnetosome assembly in MTB predict roles for MamI, MamL, and MamB in magnetosome vesicle formation and dual roles for MamB and MamE in iron biomineralization.20,28–30 In Magnetospirillum magneticum AMB-1, mamI and mamL have proposed function(s) related to invagination of the cell membrane for magnetosome vesicle formation 5 but no reported interactions with iron. However, in Magnetospirillum gryphiswaldense MSR-1, mamI may have a role in magnetite nucleation. 23 While there are few reports on bacterial mamL expression and/or interactions with iron, a dual role for mamB in iron biomineralization 24 and magnetosome membrane formation is proposed. 5 As a member of the cation diffusion facilitator (CDF) protein family, MamB may have a role in exporting divalent metal cations from the cytoplasm into intracellular compartments.28,31 The activities of MamE in MTB are also complex; involve proteolytic processing;21,29 and are required for iron biomineralization within the magnetosome vesicle.16,17 The need to better understand the role of essential magnetosome genes is key to manipulating the magnetosome and designing magnetosome-like structures in foreign cells, whether bacterial 32 or mammalian. 33
Influence of MamI, MamL, or MamB Expression on Cellular Relaxation Rates
Mammalian cells expressing MamI, MamL, or MamB share a number of features related to the increase in total cellular iron content in response to iron-supplemented culture. Small transmembrane proteins such as MamI and MamL were as effective as the putative CDF protein, MamB, at increasing total elemental iron in mammalian cells. Our findings that MamI increases cellular relaxation rates agree with reports in literature of the iron-handling activities of MamI. 23 Our study reports the first finding of the iron-handling ability of MamL. This iron-handling activity was not compromised by truncating the MamL C-terminal peptide, suggesting that membrane-spanning sequences regulate the sequestration of iron (Supplementary Figure S3). Our reports of the iron-handling ability of MamB from the AMB-1 species agree with reports of iron-handling function of MamB from the MSR-1 species 19 and suggest that the sequence similarities between the two MamB species could also indicate functional similarities. Appropriate increases in relaxation rates underlie the iron-handling activities of MamI-, MamL-, and MamB-expressing cells, with significant increases in iron-specific magnetic field inhomogeneities (R2′) related to MamI and MamL expression, accounting for approximately 50% of the correlation between total cellular iron and transverse relaxation rate.
The relation between R2 values and iron levels obtained herein is consistent with an earlier study 34 on iron deposits in brain tissues in-vitro. The correlation between R2 and iron levels (Figure 6) has a slope of 3.88 s−1/μg Fe/mg protein. Assuming that protein makes up approximately 20% of the cellular weight this slope can be converted from 3.88 s−1/μg Fe/mg of protein to 19.4 μg Fe/mg of total cellular weight. In Vymazal et al, 34 the corresponding slope when linearly extrapolated to 3 Tesla is approximately 31.9 s−1 mg Fe/g of tissue. In our study, the ratio of the slope of R2 versus iron content to that of R1 versus iron content is ≈ 13, which is similar to what was found in previous work 34 where the ratio of these slopes at 1.5T was approximately 10. Our finding that the slope of R2 versus iron content was very close to that for R2ʹ is consistent with a previous in-vivo brain study 35 which also found similar slopes for R2 and R2ʹ, using iron levels based on age and brain region. However, these slopes were approximately twice as large as in our study (and twice as large as in Vymazal et al 34 for R2).

Correlation between relaxation rates and cellular iron content in mammalian cells expressing magnetosome proteins. Cells were grown in non-supplemented medium (Standard Fe, open symbols) or in 250 µM ferric nitrate/medium (Supplemented Fe, closed symbols); harvested; then mounted in a gelatin cell phantom for MRI while remaining cells were lysed for analysis by ICP-MS. Scatter plots relate MR relaxation rates (y-axis) to elemental iron content (x-axis) for the parental control (circles) and those cells stably expressing MamI (squares), MamL (upright triangles), MamLtrunc (inverted triangles), or MamB (diamonds). Under standard culture conditions (left-hand plots), nonlinear regression indicates no significant correlation between total cellular iron content and R1, R2*, or R2. Only R2´ was significantly correlated with cellular iron under both standard and supplemented (right-hand plot) culture conditions. Spearman's correlation coefficients indicate moderate correlations between iron content and R2 (+ Fe, r = 0.33), R2′ (+ Fe, r = 0.41), and R2′ (no Fe, r = 0.35). However, in iron-supplemented cultures, there were strong and moderate-to-strong correlations between iron and R1 (+ Fe, r = 0.75) and R2* (+ Fe, r = 0.65), respectively.
Furthermore, the slope of R2* (13.94 s−1/μg Fe/mg protein) is 3.6 times greater than the slope of R2 (3.88 s−1/μg Fe/mg protein), which indicates that dephasing appears to be intermediate between static dephasing regime (R2* ≫ R2) and mobile dephasing regime (R2* ≈ R2). This suggests the possibility that iron is aggregated or possibility in mineral form.
Structural models of MamI, MamL, and MamB36,37 position these transmembrane proteins in intracellular vesicles with extramembrane domains facing the cytoplasm (Supplementary Figure S3) where it is speculated to interact with accumulated iron. In this orientation, the interaction between water protons and the iron-rich intracellular milieu produces a distinct shortening of T2* and T2 relaxation times. Thus, in iron-supplemented mammalian cells expressing MamI, MamL, or MamB, MRI values are consistent with structural modeling based on protein sequence.
Influence of MamE Expression on Cellular Relaxation Rates
In iron-supplemented mammalian cells expressing MamE, total cellular iron content was 13-fold greater than the parental control, similar to values from iron-supplemented cells expressing MamI, MamL, or MamB. Despite this evidence of iron-handling activity, MamE expression did not alter cellular MR relaxation rates. As for R1, there was little correlation between elemental iron content and longitudinal relaxation rates. These findings raise the possibility that MamE-expressing cells interact with a form of iron that does not affect MR signal.
Structural modeling of MamE based on primary sequence (Supplementary Figure S4) predicts an intracellular protein with one N-terminal transmembrane domain (amino acids 22–42) and a large C-terminal domain facing the (magnetosome) vesicle lumen (amino acids 43–728).21,26 The fusion to EGFP should thus provide fluorescence on the cytosolic face and two heme-binding domains within the vesicle lumen. AlphaFold software predicts these iron binding sites are buried within the large luminal protein domain of MamE (Supplementary Figure S4) and are arranged as iron ions complexed in a porphyrin ring. This heme structure is speculated to cause the lack of MR relaxivity of MamE-expressing cells as iron ion (Fe2+ or Fe3+) in chelates have been reported to have low relaxation efficiency. 38 Other MTB proteins, which have no reported iron chelation sites, may interact with a different form of iron that alters the cellular MR signal. As well, the large protein structure of MamE will change the nuclear magnetic relaxation dispersion, which may reduce the relaxivity of the iron ions in the porphyrin rings at 3T. 39
Alternatively, the MR-silent nature of iron-supplemented MamE-expressing cells may be related to the active form of MamE when expressed in a mammalian system. In the AMB-1 strain, MamE is characterized as a trypsin-like protease in which autoproteolysis is required for biomineralization function. 21 Magnetotactic bacteria with a catalytically inactive MamE display a compromised phenotype in which biomineralization is altered and smaller magnetite crystals are produced.21,29 Hence, although immunodetection of EGFP-MamE fusion protein reveals full-length and lower molecular weight species of MamE in mammalian cell extracts, the correct processing of MamE remains uncertain and could compromise its full function (i.e., result in MR-silent activity).
Further studies on the form and function of MamE in foreign cell environments are required. Potentially, the interaction of MamE with iron is dependent upon the interaction of MamE with other essential magnetosome proteins such as MamI, MamL, and/or MamB. In MTB, MamE is a bifunctional protein 20 with a role in both protein localization and biomineralization. Studies examining the deletion of mamE report that localization of soluble proteins (i.e., MamC and MamF) and membrane-bound protein (MamI) were disrupted. 20 Since magnetosome formation is a stepwise process, the biomineralization function of MamE is potentially not activated unless MamE successfully interacts with other magnetosome proteins.
Note that the development of this reporter gene would also benefit magnetic particle imaging (MPI).40,41 Potentially this could be an advance as important as bioluminescence has proven to be for optical imaging. Of course, there are some caveats. (1) Magnetosomes are normally large single magnetic domain nanoparticles 14 whereas MPI works best using SPION (super paramagnetic iron oxide nanoparticles) 40 which do not have a thermally stable permanent magnetic field. However, it has been shown that the linear chain appearance of magnetosomes in MTB can be alternately formed in a cluster if one gene is omitted, suggesting that the remnant magnetic moment may have been eliminated. 42 (2) Particle size is also a consideration, with an optimal core size for MPI around 30 nm in diameter; whereas, magnetosome particle size would have to be considerably less for zero remnant magnetic moment. However, it has been shown that MTB can be genetically programmed to produce different size magnetosomes. 13 Given the richness of the many genes associated with magnetosome formation and the many discoveries to come with respect to the role of each gene, this represents a cornucopia of bioengineering opportunities going forward once the first crude iron particle is formed from a reporter gene. One can speculate on the future of many different reporter gene signals in MPI, such as those now available in fluorescent optical imaging (i.e., GFP and Tomato), as the characteristics of the iron particle are sufficiently magnetically different for different reporter genes to give a different signal in MPI.
Conclusion
In this study, essential magnetosome genes mamI, mamL, mamB, and mamE were stably expressed in MDA-MB-435 human melanoma cells, demonstrating compatibility with mammalian cell expression systems. In this foreign environment, we studied the influence of these key magnetosome genes on the cellular iron content and MR relaxation rates. While all four gene expression systems increased total cellular iron content in response to iron-supplemented culture, only MamI, MamL, and MamB expression displayed a concomitant increase in transverse and longitudinal relaxation rates. It is interesting to note that the increases in spin–spin relaxation in cells expressing MamI, MamL, and MamB are similar to those observed for magA and mms6 gene expression: about 15 s−1. This suggests that more than one gene is needed to produce the sought after, large changes in spin–spin relaxation.
In the native bacteria (AMB-1), MamI, MamL, and MamB are essential to magnetosome membrane formation, while MamE is involved in protein recruitment to the magnetosome membrane and iron biomineralization. Although MamI, MamL, and MamB have no reported interactions with iron in AMB-1, MamI and MamB have reported iron-handling abilities in the MTB MSR-1. The evidence presented in the current eukaryotic cell study expands our understanding of the iron-handling nature of MamI, MamB, and MamE and introduces a previously unrecognized iron-handling function for MamL which, like MamI, depended on presumptive membrane-associated protein sequences. We anticipate that the MR-silent nature of MamE expression in iron-supplemented mammalian cells points to the importance of stepwise development of magnetosome particles, 5 assuming that MR measures in MamE-expressing cells depend on protein interaction(s) with other magnetosome components. In future studies, combined expression of these essential genes is expected to improve cellular MR relaxation rates and open paths to developing gene-based MR contrast technologies that can noninvasively monitor, identify, and track the expression of cellular and molecular biomarkers.
Supplemental Material
sj-docx-1-mix-10.1177_15353508241289765 - Supplemental material for Magnetic Resonance Imaging of Mammalian Cells Individually Expressing Membrane-Associated Magnetosome Proteins I, L, B, and E
Supplemental material, sj-docx-1-mix-10.1177_15353508241289765 for Magnetic Resonance Imaging of Mammalian Cells Individually Expressing Membrane-Associated Magnetosome Proteins I, L, B, and E by Qin Sun, PhD, Gabriel Varela-Mattatall, PhD, Prushoth Vivekanantha, PhD, Moeiz Ahmed, PhD, Salvan Hassan, PhD, Selinia Hong, PhD, Neil Gelman, PhD, R. Terry Thompson, PhD, Frank S. Prato, PhD and Donna E. Goldhawk, PhD in Molecular Imaging
Footnotes
Acknowledgements
The authors would like to thank Peggy Chou for her contributions to ICP-MS sample preparation and data analysis of the Tomato-MamB and EGFP-MamE expression systems.
Data Availability Statement
The authors confirm that the data supporting the findings of this study are available within the article and its Supplemental materials.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: this work was supplied in part by; a Canadian Foundation for Innovation no.11356 grant, a Natural Science and Engineering Research Council of Canada/Alliance ALLRP 576699 - 22 grant, and a Natural Science and Engineering Research Council of Canada/Discovery RGPIN-2020-09125 grant.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.