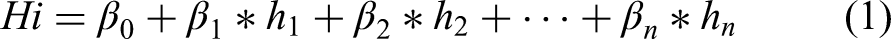Abstract
Introduction
Axillary nodal burden reflects the biological aggressiveness and prognostic behavior of breast cancer. This study aimed to develop a subregional habitat radiomics model based on multiparametric magnetic resonance imaging (MRI) and to evaluate its performance in predicting high axillary nodal burden in patients with breast cancer.
Methods
In this retrospective study, a total of 221 patients who underwent axillary lymph node dissection were categorized as having limited (0-2 metastatic nodes) or high (≥3 metastatic nodes) nodal burden based on pathological findings. Morphological MRI features were visually evaluated by experienced radiologists. A clinical model was established using univariate and multivariate logistic regression analyses. Conventional radiomics (C-radiomics) and habitat radiomics features were extracted from the whole tumor and its subregions, respectively, based on multiparametric MRI. The clinical, C-radiomics, and habitat radiomics models were then integrated into a comprehensive nomogram for quantitative prediction of axillary nodal burden.
Results
In predicting axillary nodal burden, the habitat radiomics model outperformed both the C-radiomics and clinical models, achieving areas under the curve (AUCs) of 0.791 (0.712-0.870) and 0.798 (0.686-0.911) in the training and validation cohorts, respectively. The C-radiomics model achieved AUCs of 0.733 (0.631-0.836) and 0.738 (0.612-0.865), while the clinical model achieved AUCs of 0.753 (0.663-0.843) and 0.733 (0.596-0.870). The combined nomogram demonstrated the highest diagnostic performance, with AUCs of 0.895 (0.839-0.951) and 0.885 (0.802-0.969) in the training and validation cohorts, respectively.
Conclusions
The integrated nomogram combining clinical, C-radiomics, and habitat radiomics models demonstrated strong predictive efficacy for preoperative assessment of axillary nodal burden in breast cancer. Future multicenter prospective studies are warranted to validate these results and refine the model's clinical applicability.
Introduction
Axillary lymph node (ALN) status is a key indicator of tumor aggressiveness and an essential prognostic determinant in breast cancer. It directly influences surgical planning and guides adjuvant treatment decisions, including the need for chemotherapy or radiotherapy.1,2 Axillary lymph node dissection (ALND) remains the gold standard for evaluating ALN status and removing metastatic lymph nodes (LN), particularly in patients with substantial nodal involvement.3–5 However, extensive ALND is associated with complications such as arm lymphedema and shoulder dysfunction, which can profoundly affect a patient's quality of life.6–8 Consequently, minimizing unnecessary ALND procedures has become a critical clinical objective.
The American College of Surgeons Oncology Group Z0011 (ACOSOG Z0011) trial demonstrated no significant difference in local recurrence or 10-year overall survival between patients with fewer than three axillary lymph node metastases (ALNMs) who underwent ALND and those who did not. 9 In contrast, patients with three or more ALNMs, indicating a high axillary nodal burden, had a higher risk of local recurrence and are generally considered candidates for ALND or neoadjuvant chemotherapy.10–12 Therefore, accurate preoperative assessment of ALNMs and reliable identification of patients with high axillary nodal burden are essential for optimizing individualized treatment strategies. 13
The use of preoperative imaging modalities such as mammography, ultrasound, and computed tomography is increasingly important for evaluating axillary nodal burden in patients with breast cancer and is now widely adopted.14–17 In contrast to single-modality imaging, multiparametric magnetic resonance imaging (MRI) offers a comprehensive and multidimensional framework for axillary evaluation. Over the past decade, conventional radiomics (C-radiomics) has emerged as a powerful tool for extracting high-throughput quantitative features from medical images, transforming complex visual information into measurable data that reflect tumor biology. This technique has been successfully applied in breast cancer staging and prognostic evaluation.18–20 However, C-radiomics typically treats the tumor as a homogeneous entity, thereby overlooking intratumoral heterogeneity, which may reflect distinct biological behaviors and influence metastatic potential.21,22
Unlike C-radiomics, habitat imaging involves segmenting the entire lesion into multiple regions of interest (ROIs). These “habitats” are subregions generated by clustering voxels based on intensity or entropy patterns within the segmented ROI. The concept is that each subregion, comprising voxels with similar imaging signatures, may reflect shared tumor microenvironmental characteristics. Habitat radiomics, therefore, extends beyond global tumor analysis, enabling a more refined exploration of intratumoral heterogeneity and its biological correlates. Through this voxel-level stratification, habitat-based analysis can provide insights into the tumor's spatial heterogeneity and its interactions with the surrounding tissue environment.23–26 Habitat radiomics has been shown to characterize the heterogeneity of breast cancer and correlate with response to neoadjuvant chemotherapy, 23 lymphovascular invasion (LVI),24,25 and patient prognosis 26 in patients with breast cancer.
Currently, limited evidence exists regarding the application of habitat radiomics for preoperative prediction of axillary LN burden in breast cancer. Therefore, this study aimed to develop a multiparametric MRI–based habitat radiomics model to predict high axillary LN burden (ALNMs ≥ 3) in breast cancer. To the best of our knowledge, this represents the first investigation to explore this specific application.
Methods
Patients and Baseline Characteristics
This retrospective, single-center study included female patients diagnosed with breast cancer who underwent multiparametric MRI and subsequent breast surgery at the Central Hospital affiliated with Shandong First Medical University between June 2018 and December 2023. These patients were consecutively enrolled in the study between June 2018 and December 2023. The inclusion criteria were: (i) clearly visible primary breast lesions on MRI manifesting as masses, (ii) histopathologic confirmation of breast cancer after surgery, (iii) breast surgery including ALND, and (iv) no preoperative chemoradiotherapy before MRI. The exclusion criteria were: (i) patients with multiple systemic diseases or bilateral breast abnormalities, (ii) poor-quality MRI images showing motion artifacts, chemical shift artifacts, or inadequate fat suppression, and (iii) an interval longer than 2 weeks between MRI and surgery. A patient selection flowchart is shown in Figure 1. The dataset was randomly divided into training and validation cohorts in a 2:1 ratio. The training cohort was used to construct and optimize the predictive model, and the validation cohort was used for independent performance evaluation and generalizability testing.

Patient Enrollment Flowchart.
Ethical Approval Statement
The study protocol was approved by the Institutional Review Board of the Central Hospital affiliated with Shandong First Medical University (Approval No. 20250120011; Date: February 8, 2025).
All patient data were fully de-identified prior to analysis, ensuring that no personal identifiers, medical record numbers, or traceable imaging information were retained. Given the retrospective nature of this study and the anonymization of patient data, the requirement for written informed consent was waived by the Institutional Review Board.
MRI Protocol and Image Preprocessing
Breast MRI was performed using a 3.0-T superconducting magnetic resonance (MR) scanner (Discovery 750 W; GE Healthcare, Milwaukee, WI). Patients were positioned prone, with the breasts naturally dependent and the head facing forward. The imaging protocol included the following sequences: fast spin -echo (FSE) T1-weighted imaging (T1WI) with a slice thickness of 4 mm, field of view (FOV) of 320 × 320 mm, repetition time (TR) of 551 ms, and echo time (TE) of 15 ms; and FSE T2-weighted imaging (T2WI) with identical slice thickness and FOV, TR of 6828 ms, and TE of 81 ms. Diffusion-weighted imaging (DWI) was acquired using a single-shot echo-planar imaging sequence with b-values of 0 and 1000 s/mm2, TR of 3923 ms, and TE of 76.3 ms. Dynamic contrast-enhanced (DCE) imaging was performed using a breath-hold three-dimensional (3D) volume-interpolated gradient-echo technique with TR of 4.4 ms, TE of 2.1 ms, slice thickness of 1.5 mm, and FOV of 350 × 350 mm.
Gadolinium–diethylenetriaminepentaacetic acid was administered intravenously at 0.1 mmol/kg using an automated power injector at 2.0 mL/s through the dorsal hand vein, followed by an equal volume of normal saline. Eight sequential post-contrast phases were obtained at 55-s intervals, yielding a total scan duration of approximately 8-10 min.
MRI Preprocessing
All MRI scans were first subjected to N4 bias field correction to compensate for low-frequency intensity inhomogeneities inherent to MRI acquisition, which can significantly affect gray-level–based radiomics features. Next, intensity normalization was applied to harmonize signal intensity distributions across patients and imaging sessions, thereby reducing variability caused by subtle differences in acquisition parameters. Finally, all images were resampled to a uniform voxel size to standardize spatial resolution, ensuring that radiomics feature extraction is robust and comparable across the dataset. 27
MR Morphological Features Analysis
Two radiologists, with 8 and 15 years of experience in breast MRI, independently conducted a double-blind review to evaluate the MR morphological features of breast cancer. The assessed imaging features included tumor size (maximum lesion diameter), location, background parenchymal enhancement, fibroglandular tissue composition, time–signal intensity curve, peritumoral edema, subcutaneous edema, necrosis, internal enhancement pattern, and MRI-detected ALN status (MR-ALN).
The absence of MR-ALN was defined by one major criterion or more than two minor criteria. The major criterion was loss of the central fatty hilum on both T1- and T2-weighted images. The minor criteria included: (1) short-axis diameter >1 cm, (2) eccentric cortical thickening, (3) peripheral displacement of the fatty hilum, (4) irregular cortical contour, (5) matted lymph nodes, and (6) decreased signal intensity compared with other lymph nodes on T2-weighted images. 28
For quantitative parameters, the mean of the two radiologists’ measurements was calculated to reduce interobserver variability. For qualitative parameters, any discrepancies were resolved through consensus discussion.
Tumor and Subregion Segmentation
Figure 2 illustrates the study workflow, which began with whole-lesion identification followed by subregional ROI generation. The primary radiologist who performed the ROI delineation had more than 8 years of experience in chest imaging, and refinement was provided by a second radiologist with 15 years of experience. ROI segmentation was carried out using the open-source software ITK-SNAP version 4.0 (http://www.itksnap.org/pmwiki/pmwiki.php). After segmentation, ROI registration was completed in 3D Slicer version 4.6, labeled across five imaging sequences: T1WI, T2WI, DWI, apparent diffusion coefficient (ADC), and DCE imaging.

Schematic Overview of the Study Design. The Radiologist Delineated the Entire Lesion Area, Followed by k-Means Clustering to Segment Subregions. Radiomics Features Were Extracted from Both the Entire Lesion and Each Subregion, then Subjected to Dimensionality Reduction and Selection. C-Radiomics and Habitat Radiomics Models were Subsequently Constructed, and These Were Combined with the Clinical Model to Develop a Comprehensive Nomogram for Quantitative Prediction of Axillary Nodal Burden in Breast Cancer.
Manual ROI delineation was first performed on the DCE images to ensure precise lesion contouring, after which the same ROIs were propagated to other MRI sequences using the 3D Slicer general registration module (Elastix). Elastix offers multiple preset registration algorithms; for this breast-specific application, the 3D DCE-MRI (breast) preset was chosen because it is optimized for the anatomical complexity of breast tissue. To maintain spatial fidelity, registration was performed separately for each sequence according to its unique FOV and slice-thickness characteristics. Registration quality was independently assessed by two radiologists using a 4-point scale (1 = severe misregistration, 4 = excellent alignment). Images with a quality score ≤ 2 were manually corrected, and 95% (209 of 221) of the registered images achieved a score ≥ 3, ensuring accurate alignment across sequences.
Lesion subregional heterogeneity, reflecting the tumor's ecological diversity, was quantified using a clustering-based approach. Radiomic feature maps derived from first-order intensity and texture metrics (including GLCM entropy, GLRLM short-run emphasis, and GLSZM zone variance) were normalized and used as the input for k-means clustering. The optimal number of subregions was determined using the Calinski–Harabasz (CH) index, which measures the ratio of between-cluster dispersion to within-cluster cohesion (variance ratio criterion). A higher CH value indicates better-defined and more compact clusters. To ensure clustering stability, CH values exceeding one standard deviation from the mean were excluded as potential outliers. Cluster configurations ranging from three to seven were evaluated, and the configuration yielding the highest CH value was selected as the optimal clustering solution. The lower limit (three) was chosen to ensure adequate representation of intratumoral heterogeneity, while the upper limit (seven) was set to prevent over fragmentation and unstable small clusters, as supported by previous radiomic subregion studies.24–26
C-Radiomics and Habitat Radiomics Generation
C-radiomics and habitat radiomics features were generated through standardized extraction using the PyRadiomics package, with the entire workflow adhering to the guidelines of the Image Biomarker Standardization Initiative to ensure consistency and reproducibility of radiomic analyses 29 ; specifically, for habitat radiomics, multimodal MRI sequences including T1WI, T2WI, DWI, ADC maps, and DCE-MRI were used as the data sources, and 111 quantitative radiomic features were extracted from each pre-identified tumor subregion—these features encompassed 19 histogram-based features (capturing intensity distribution characteristics), 17 shape-based features (describing lesion morphological properties), and 75 texture-based features (quantifying spatial intensity relationships); meanwhile, to facilitate comparative analysis between the two radiomics approaches, the same set of 111 features was also extracted from the manually delineated entire lesion ROI to generate C-radiomics features, ensuring that the feature composition and extraction parameters were consistent across both C-radiomics and habitat radiomics.
Dimensionality reduction was conducted using a three-step process.21,24,25 First, Pearson correlation coefficients were calculated, and features with coefficients ≤0.9 were retained. Second, a two-sample t-test was applied to compare positive and negative classifications, retaining features with P < .05. Third, the least absolute shrinkage and selection operator (LASSO) regression was applied to identify features with the strongest correlation to the outcome. The optimal λ parameter was determined through 10-fold cross-validation, and features with non-zero coefficients were retained.
A habitat radiomics index (Habitat index) was constructed by integrating selected subregional features weighted by their respective coefficients, while a C-radiomics index (Rad score) was derived using features from the entire lesion. The Habitat index can be formulated as follows:
Establishment of Multiple Diagnosis Models
To assess the risk of high axillary nodal burden in breast cancer, we developed and evaluated four diagnostic models: (1) the clinical model, (2) the C-radiomics model, (3) the habitat radiomics model, and (4) the integrated nomogram model. These models were constructed by combining clinical variables with radiomic features derived from multimodal MRI sequences. Univariate and multivariate logistic regression analyses were applied to identify key predictors of high axillary nodal burden (≥3 ALNMs). The models were then developed based on the significant clinical and radiomic features identified.
The clinical model was developed using clinical variables identified as significant predictors of high axillary nodal burden in univariate analysis. These variables were incorporated into the multivariate logistic regression model. The C-radiomics model was developed by extracting radiomic features from the entire tumor using conventional MRI sequences, with the Rad score computed as a combined metric. For the habitat radiomics model, features were extracted from subregions identified through habitat analysis, and the Habitat index was computed to represent intratumoral heterogeneity.
The nomogram model was constructed by integrating the variables identified in the multivariate clinical model with the Rad score from the C-radiomics model and the Habitat index from the habitat radiomics model. These integrated variables were then used to generate a comprehensive nomogram, which provides a quantitative prediction of high axillary nodal burden. The nomogram allows for individualized risk assessment by combining both clinical and radiomic features, providing a clinically useful tool for preoperative risk assessment.
Model Performance Evaluation
The performance of each model was evaluated using receiver operating characteristic (ROC) curve analysis, with the area under the curve (AUC), sensitivity, specificity, accuracy, positive predictive value (PPV), and negative predictive value (NPV) calculated to quantify diagnostic performance. The models were assessed for their ability to discriminate between patients with low and high axillary nodal burden, with the AUC serving as the primary indicator of overall model discrimination. To statistically compare model performance, DeLong's test was used to evaluate differences in AUC values among models. In addition, the Net reclassification improvement (NRI) and integrated discrimination improvement (IDI) metrics were calculated to further assess the incremental predictive value gained by the nomogram relative to the individual models. Finally, decision curve analysis (DCA) was performed to evaluate the clinical utility of the nomogram by quantifying the net benefit across a range of clinically relevant threshold probabilities.
Statistical Analysis
The reporting of this study conforms to the STROBE guidelines. 30 All statistical analyses were performed using R software (version 4.3.1; R Foundation for Statistical Computing, Vienna, Austria). The STROBE guidelines were followed for reporting. Patients who underwent axillary lymph node dissection (ALND) were categorized into limited nodal burden (0-2 ALNMs) or high nodal burden (≥3 ALNMs) groups based on pathological findings. Data normality was assessed using the Shapiro-Wilk test, and homogeneity of variances was tested with Levene's test. Continuous variables with normal distribution and equal variances were compared using the independent-samples t-test, while non-normally distributed variables were compared using the Mann-Whitney U test. Categorical variables were compared using the chi-square test, depending on the sample size and distribution. A two-tailed P-value < .05 was considered statistically significant.
Univariate logistic regression was performed to identify clinical and MRI features associated with the risk of high axillary nodal burden. Variables with P < .10 in univariate analysis were included in the multivariate logistic regression model to identify independent predictors. Multicollinearity among predictors was assessed using the variance inflation factor (VIF), with values < 5 indicating acceptable independence. Regression coefficients from the final multivariate model were used to construct the nomogram for individualized prediction.
Results
Baseline Patient Characteristics
A total of 221 patients with breast cancer who underwent ALND, and their preoperative MRI scans were included in this study. Among them, 52 (23.5%) were pathologically diagnosed with high axillary nodal burden. Patients were randomly assigned to the training cohort (n = 144) and validation cohort (n = 77). The median ages of the overall, training, and validation cohorts were 51 years (interquartile range [IQR], 45-58), 52 years (IQR, 45-58.2), and 50 years (IQR, 45-57), respectively. The distributions of clinical and imaging variables were comparable between the training and validation cohorts, with only one variable showing a statistically significant difference (Table 1). This high degree of cohort homogeneity ensured balanced feature distributions for subsequent model development and validation.
Comparison of Baseline Characteristics Between the Training and Validation Cohorts.

Assessment of the Clinical Model
Univariate and multivariate logistic regression analyses identified three independent factors significantly associated with high axillary nodal burden: tumor size (OR = 3.57, 95% CI = 1.43-8.91, P = .006), peritumoral edema (OR = 2.01, 95% CI = 0.82-4.96, P = .028), and MRI-detected axillary lymph node positivity (MRI-ALN; OR = 3.51, 95% CI = 1.27-9.76, P = .016) (Table 2). The clinical model achieved AUC values of 0.753 (95% CI = 0.663-0.843) in the training cohort and 0.733 (95% CI = 0.596-0.870) in the validation cohort, demonstrating good discriminatory ability and generalizability.
Univariate and Multivariate Logistic Regression Analyses for Selecting MRI Morphological and Clinical Characteristics in Model Development.

Generation of Lesion Subregions for Habitat Analysis
For habitat analysis, the original MR image data and complete lesion masks were integrated for subregional clustering. Image information from the segmented lesions was input into the k-means algorithm and processed on a slice-by-slice basis. The algorithm effectively partitioned intratumoral subregions and calculated CH index values, confirming the CH index's applicability across all MRI sequences (T1WI, T2WI, DWI, ADC, and DCE). Figure 3 illustrates the clustering workflow and the selection of the optimal cluster number. The maximum CH value occurred at five clusters in the training cohort, defining five distinct lesion subregions for subsequent radiomics feature extraction.

Determination of the Optimal Number of Clusters. The Calinski–Harabasz (CH) Index Peaked at Five Clusters in the Training Cohort, Corresponding to Five Distinct Tumor Subregions.
Assessment of the C-Radiomics and Habitat Radiomics Signature
Figure 4 illustrates the feature-reduction process and subsequent identification of the final radiomics signatures. A total of 555 (111 × 5) C-radiomics features were extracted from the entire tumor using multiparametric MRI sequences (T1WI, T2WI, DWI, ADC, and DCE). After Pearson correlation and t-test filtering, seven C-radiomics features were entered into LASSO regression (Figure 4a). Ten-fold cross-validation determined the optimal λ value (λ = 0.0016; Figure 4b), yielding four features with non-zero coefficients in the final model (Figure 4c).

Least Absolute Shrinkage and Selection Operator (LASSO) Regression and 10-Fold Cross-Validation for C-Radiomics (a–c) and Habitat Radiomics (d–f) During Model Training. A Total of 2775 (111 × 5 × 5) Habitat Radiomics Features Were Extracted From All Subregions Using Multiparametric MRI Sequences (T1WI, T2WI, DWI, ADC, and DCE). After Pearson Correlation and t-Test Screening, 15 Features Were Incorporated into LASSO Regression. Ten-Fold Cross-Validation Identified the Optimal λ (λ = 0.0066), Resulting in 10 Features With Non-Zero Coefficients.
Comparison of the Rad score and Habitat index between limited and high nodal burden groups revealed significantly lower scores in the limited group. This distinction is visualized in the waterfall plot (Figure 5). The C-radiomics model achieved AUCs of 0.733 (95% CI = 0.631-0.836) and 0.738 (95% CI = 0.612-0.865) in the training and validation cohorts, respectively. The habitat radiomics model demonstrated superior discriminative performance, with AUCs of 0.791 (95% CI = 0.712-0.870) and 0.798 (95% CI = 0.686-0.911) in the training and validation cohorts.
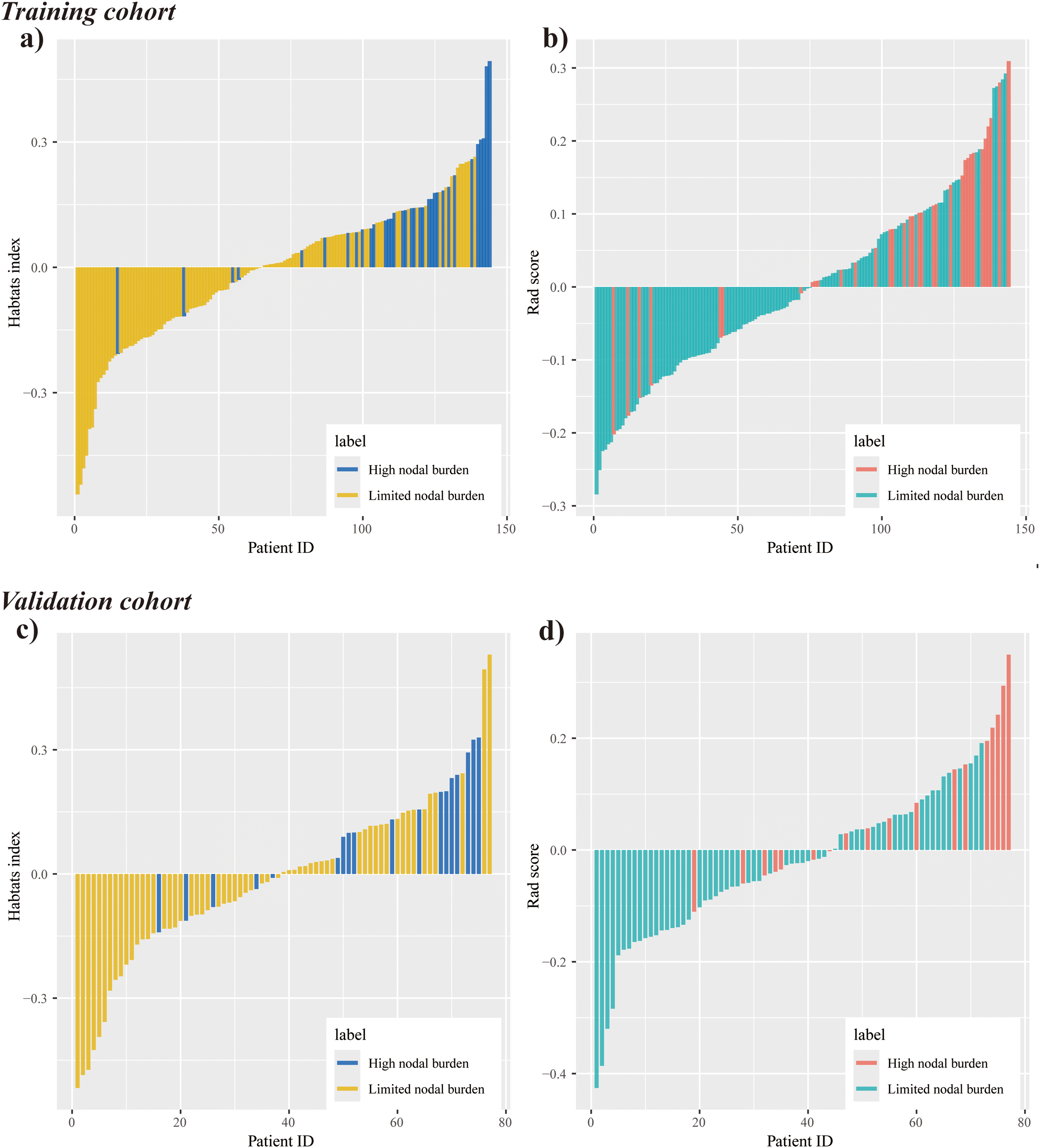
Waterfall Plots of the Habitat Index (a, c) and Rad Score (b, d) in the Training and Validation Cohorts, Comparing Patients with Limited versus High Axillary Nodal Burden. Both the Habitat Index and Rad Score were Significantly Lower in the Limited-Burden Group than in the High-Burden Group.
Nomogram Development
Significant predictors identified from the multivariate clinical model—namely tumor size, peritumoral edema, and MRI-ALN status—were combined with the Rad score and Habitat Index to construct a quantitative nomogram for estimating high axillary nodal burden. VIF values for tumor size (1.85), peritumoral edema (2.12), MRI-ALN (1.12), Rad score (2.47), and Habitat Index (1.98) indicated the absence of substantial multicollinearity among the selected variables. The final nomogram integrates these predictors to provide individualized estimates of the probability of high nodal burden (≥3 ALNMs), as illustrated in Figure 6. To further assess the robustness of the model, five-fold cross-validation was conducted. As presented in Supplementary Figure 1, the AUC values remained stable across folds (Folds 1-5 AUCs: 0.883, 0.948, 0.896, 0.799, and 0.939; mean AUC = 0.893 ± 0.053), supporting the generalizability of the nomogram across different data partitions.

Combined Nomogram for Preoperative Assessment of High Axillary Nodal Burden in Breast Cancer. The Nomogram was Constructed by Integrating The Rad Score and Habitat Index with Significant Clinical Predictors Identified through Multivariate Logistic Regression.
Nomogram Performance Evaluation
The proposed nomogram demonstrated excellent discriminative ability, with C-indices of 0.895 and 0.885 in the training and validation cohorts. Calibration curves showed close agreement between predicted and observed probabilities, and Hosmer–Lemeshow tests further confirmed good calibration (training cohort: P = .54; validation cohort: P = .905) (Figure 7). DCA indicated that the nomogram provided greater net clinical benefit than the individual clinical, C-radiomics, or habitat radiomics models (Figure 8). ROC analysis yielded AUCs of 0.895 (95% CI: 0.839-0.951) in the training cohort and 0.885 (95% CI: 0.802-0.969) in the validation cohort (Figure 9; Table 3). DeLong tests showed significantly higher AUCs for the nomogram compared with the habitat radiomics model (P = .039), C-radiomics model (P = .013), and clinical model (P = .019) in the training cohort, with similarly significant differences observed in the validation cohort (P = .042, 0.022, and 0.020, respectively). Supplementary Figure 2 provides confusion matrices for all models, demonstrating that despite lower PPV due to the limited number of high-burden cases (n = 34), the models maintained favorable sensitivity and NPV. Furthermore, positive NRI and IDI values in both cohorts (Supplementary Figures 3 and 4) confirmed the improved classification performance and incremental value of integrating clinical, C-radiomics, and habitat radiomics features in the nomogram.

Calibration Curves for the Combined Nomogram in the Training (a) and Validation (b) Cohorts. The Fitted Calibration Curves Closely Approximated the Ideal Diagonal, Indicating Good Agreement Between Predicted and Observed Probabilities.

Decision-Curve Analysis (DCA) Comparing the Clinical Utility of the Combined Nomogram, Clinical Model, C-Radiomics, and Habitat Radiomics in the Training (a) and Validation (b) Cohorts. The DCA Demonstrated a Higher Net Benefit for the Nomogram Than For Individual Models Across a Range of Threshold Probabilities.
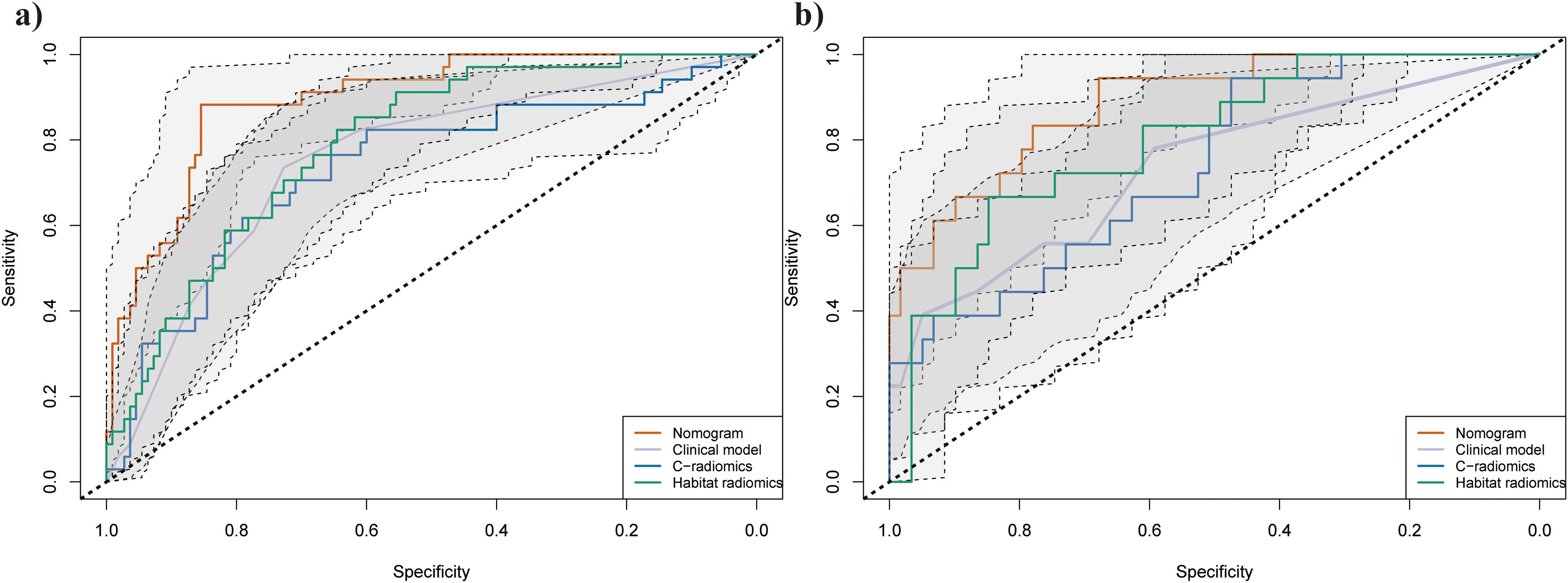
Receiver-Operating-Characteristic (ROC) Curves with 95% Confidence Intervals (CIs) Illustrating The Diagnostic Performance of the Combined Nomogram, Clinical Model, C-Radiomics, and Habitat Radiomics in the Training (a) and Validation (b) Cohorts. The Combined Nomogram Achieved the Highest Area-Under-the-Curve (AUC) Values, Indicating Superior Diagnostic Efficiency.
Assessing Diagnostic Efficacy Across Models for Discriminating Nodal Burden in Breast Cancer Cases.

Comparison with Machine Learning Classifiers
To further validate the superiority of the proposed nomogram, we conducted comparative experiments using three widely applied machine learning classifiers—support vector machine (SVM), random forest (RF), and extreme gradient boosting (XGBoost). All classifiers were developed following the same feature extraction and dimensionality reduction procedures as the nomogram to ensure methodological consistency. The diagnostic performance of each model is summarized in Supplementary Table 1. In both the training and validation cohorts, the AUCs achieved by SVM, RF, and XGBoost were consistently and markedly lower than those of the nomogram, further confirming the robustness and enhanced predictive ability of our proposed model.
Comparison Across MRI Modalities
A comparative analysis was conducted to evaluate the diagnostic efficacy of different MRI modalities for predicting high axillary nodal burden. As shown in Supplementary Table 2, single-sequence MRI models based on T1WI, T2WI, DWI, ADC, and DCE exhibited only moderate discriminative ability in both the training and validation cohorts, with AUCs ranging from 0.785 to 0.820 in the training cohort and 0.762 to 0.798 in the validation cohort. In contrast, the multiparametric MRI-derived nomogram demonstrated markedly superior diagnostic performance, achieving AUCs of 0.895 in the training cohort and 0.885 in the validation cohort.
Discussion
This study demonstrated the predictive potential of habitat radiomics derived from multiparametric MRI for identifying patients with high axillary nodal burden in breast cancer. Habitat radiomics, which quantifies subregional tumor heterogeneity, showed superior diagnostic performance compared with C-radiomics derived from the entire tumor and visually assessed MRI morphological features. Furthermore, habitat radiomics, C-radiomics, and MRI morphological variables, tumor size, MRI-ALN status, and peritumoral edema, were identified as independent predictors of high nodal burden and were integrated into a combined predictive model.
In multivariate logistic regression, high axillary nodal burden was independently associated with tumor size, peritumoral edema, and MRI-ALN status. Tumor size remains a well-established prognostic marker in breast cancer and correlates closely with ALNM.31,32 Larger tumors often reflect longer tumor dwell time and increased angiogenesis, whereas smaller, biologically aggressive tumors may metastasize early despite limited local growth. 33 Therefore, tumor size should be interpreted within the context of additional biological and imaging parameters.
Peritumoral edema was also strongly associated with high nodal burden. In breast cancer, peritumoral edema reflects multiple pathological processes, including angioectasia, interstitial fibrosis, necrosis, and LVI. Edema results from obstruction of lymphatic and blood vessels by tumor emboli, representing an early event in metastatic spread.34,35 This finding aligns with Song et al, 36 who identified peritumoral edema as an independent predictor of ALNM in early-stage triple-negative breast cancer.
MRI-ALN positivity emerged as the strongest predictor of nodal metastasis, consistent with prior studies demonstrating correlations between MRI-detected axillary node morphology and LVI.37–39 Similarly, Xue et al 28 confirmed the independent predictive role of MRI-ALN status in early-stage disease. Despite these advantages, visually assessed MRI features are inherently subjective, with substantial interobserver variability and overlapping manifestations across molecular subtypes. Consequently, reliance on MRI morphological assessment alone remains inadequate for accurately identifying high nodal burden.
Habitat radiomics, designed for quantitative assessment of intratumoral heterogeneity, is based on the premise that clustering subregions containing voxels with similar imaging characteristics may reveal shared biological properties.23–26 In this study, habitat radiomics demonstrated superior performance compared with C-radiomics and clinical models. These findings align with prior studies showing that habitat radiomics effectively captures the spatial heterogeneity of breast cancer, providing more objective diagnostic information than C-radiomics and visually assessed MRI morphological features. Previous investigations have linked habitat radiomics to therapeutic response, LVI, and prognosis in breast cancer.23–26
The nomogram integrating habitat radiomics, C-radiomics, and MRI morphological features exhibited the highest predictive accuracy among all models. DCA further confirmed its superior clinical utility, demonstrating higher net benefits than individual models across a wide range of threshold probabilities. This finding highlights the added value of incorporating habitat-derived information into predictive frameworks for assessing axillary nodal burden.
The clinical model contributed patient-level risk factors, including tumor size, peritumoral edema, and MRI-ALN status, while C-radiomics provided quantitative descriptors of tumor texture beyond visual assessment. Habitat radiomics, in turn, captured biologically active subregions that better represented intratumoral heterogeneity, thereby enhancing model performance.23–26
It is noteworthy that the relatively low PPV observed in the validation cohort was largely attributable to class imbalance, as patients with high nodal burden (≥3 ALNMs) comprised a smaller proportion of the dataset. Under such conditions, PPV naturally decreases despite satisfactory model discrimination and calibration. Importantly, the nomogram achieved high sensitivity and NPV, underscoring its potential clinical role in identifying patients unlikely to harbor extensive nodal metastasis—those who may benefit from less invasive axillary management. Thus, the model's predictive focus aligns with its intended clinical purpose: risk stratification and safe exclusion of high-burden disease, rather than definitive classification of positive cases.
Despite these encouraging results, several limitations should be acknowledged. First, its retrospective and single-center design may have introduced selection bias. Large-scale, multicenter prospective studies are warranted to validate the present findings and improve their generalizability. Second, the dataset exhibited an inherent imbalance between patients with high and limited nodal burden, which may have affected certain evaluation metrics such as specificity and PPV. To avoid introducing synthetic bias, no resampling or weighting techniques were applied. Future research should explore appropriate class-balancing methods to further improve model robustness. Third, several pathological factors associated with ALNM—including molecular subtypes (eg, HER2 status), tumor grade, and LVI—were excluded due to incomplete records, limiting the ability to fully explore the correlation between habitat radiomics and histopathologic heterogeneity. Future studies should incorporate these variables to enhance analytical robustness. Fourth, a direct performance comparison between single-sequence and multiparametric MRI models was not performed. Although this study focused on the integration of complementary multiparametric features, future research should further investigate the individual contribution of each MRI sequence to optimize feature selection and model performance. Fifth, while the model's accuracy and discriminatory power are promising, its clinical utility is further validated by its ability to incorporate subregional information, offering a deeper understanding of the tumor's heterogeneity. This subregional analysis could be particularly useful in identifying biologically aggressive areas within the tumor, which might otherwise be overlooked in traditional clinical assessments. Finally, our methodology sets itself apart from automated systems such as mAIstro, which provides end-to-end automation for model development. 40 Our approach hinges on manual feature extraction and model construction. In contrast, mAIstro automates the entire process, including preprocessing, feature selection, and model building, with a strong emphasis on efficiency and consistency. Future research could delve into integrating such automation techniques to bolster the scalability and reproducibility of our methodology, particularly when dealing with large datasets and multi-center studies.
Conclusions
We developed and validated a nomogram integrating habitat radiomics, C-radiomics, and MRI morphological features to predict high-burden ALNM in breast cancer. The model demonstrated excellent discrimination, calibration, and clinical utility, suggesting potential for reducing unnecessary ALND and identifying patients likely to benefit from neoadjuvant chemotherapy. Future large-scale, prospective multicenter studies are warranted to confirm these findings and to further explore the incremental value of filtered-image radiomics features and comprehensive pathological variables in improving predictive accuracy.
Supplemental Material
sj-docx-1-tct-10.1177_15330338261416806 - Supplemental material for Multiparametric MRI-Derived Habitat Radiomics in Subregional Analysis for Predicting Axillary Lymph Node Metastatic Burden in Breast Cancer
Supplemental material, sj-docx-1-tct-10.1177_15330338261416806 for Multiparametric MRI-Derived Habitat Radiomics in Subregional Analysis for Predicting Axillary Lymph Node Metastatic Burden in Breast Cancer by Yaoqi Han, Fei Gao, Aimei Ouyang, Jing Wang, Chunling Zhang, Guoyue Chen, Xue Bing and Zhen Gao in Technology in Cancer Research & Treatment
Footnotes
Acknowledgments
The authors wish to thank Li Li for critically reviewing the study proposal, providing technical support, or assisting with patient care. Written permission has been obtained from all acknowledged individuals. This placeholder ensures that all non-author contributions can be properly documented and consented for, in line with journal policy.
Ethical Considerations
The study protocol received approval from the Institutional Review Board of the Central Hospital affiliated with Shandong First Medical University (Approval No. 20250120011; Date: February 8, 2025).
All patient data were fully de-identified prior to analysis, ensuring that no personal identifiers, medical record numbers, or traceable imaging information were retained. Given the retrospective nature of this study and the anonymization of patient data, the requirement for written informed consent was waived by the Institutional Review Board.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Author Contributions
[Yaoqi Han]: Conceptualization, Methodology, Formal analysis, Writing – original draft.
[Fei Gao]: Investigation, Data curation, Writing – review & editing.
[Aimei Ouyang]: Resources, Visualization.
[Jing Wang, Chunling Zhang, and Guoyue Chen]: Supervision, Project administration.
[Xue Bing, and Zhen Gao]: Funding acquisition, Supervision.
All authors read and approved the final manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Science and Technology Development Program of the –Jinan Municipal Health Commission of China (grant nos. 2023-2-29 and 202134045).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The data that support the findings of this study will be made available by the corresponding author upon reasonable request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.