Abstract
Introduction
Colorectal cancer is the third most common cause of cancer-related deaths worldwide. 1 Currently, standard treatments of colorectal cancer include surgery, chemotherapy, and targeted therapy.2,3 However, 50% to 60% of patients diagnosed with colorectal cancer will develop distant metastases, by which point when treatments have a very limited impact on the progression of diseases and become very challenging.4,5 In addition, approximately 40% of the surgically cured patients will experience cancer recurrence within 5 years.6,7 Therefore, identifying novel diagnostic and prognostic biomarkers is of significant importance for improving survival.
An increasing number of studies have demonstrated that long noncoding RNAs (lncRNAs) play key roles in various kinds of cancers, which can participate in multiple gene regulatory networks and tumor gene transcription.8,9 Nuclear paraspeckle assembly transcript 1 (NEAT1) is a lncRNA that is located on chromosome 11 and is widely expressed in a variety of mammalian cell types. 10 It has been found that NEAT1 has abnormally high expression in prostate cancer, colorectal cancer, nonsmall cell lung cancer, and promotes tumorigenesis.9–12 NEAT1 is a noncoding RNA related to cancer susceptibility. 13 Zhang et al. 14 tested NEAT1 in colorectal cancer and noncancerous cervical tissues of polish colorectal cancer patients and found that NEAT1 level in colorectal cancer tissues was higher than that in noncancerous cervical tissues. However, its role in colorectal cancer remains largely unknown.
ZEB1 has been implicated as a pivotal step for tumor infiltration and distant metastasis in a variety of carcinomas. 15 ZEB1 expression is also dysregulated in colorectal cancer and is related to a high grade of colorectal cancer. 16 It was demonstrated that ZEB1 is the target of miR-448 and negatively correlates with the expression of miR-448. 17 Whether miR-448 is involved in the progression of colorectal cancer through ZEB1 remains to be explored. Thus, we aimed to elucidate the role and precise mechanism of NEAT1 in colorectal cancer.
In the present study, we analyzed the expression of NEAT1 in colorectal cancer tissue and cell lines. Function studies were performed to determine the effect of NEAT1 on the phenotype of colorectal cancer cell lines. It was found that NEAT1 was upregulated in the colorectal cancer tissues and cells. NEAT1 promotes the proliferation and invasion of colorectal cancer cells. Mechanistically, NEAT1 sponges miR-448 and promotes the expression of ZEB1. Taken together, our findings reveal that NEAT1-miR-448-ZEB1 axis regulates malignant progression in colorectal cancer in vitro and in vivo, suggesting that NEAT1 may serve as a potential target for colorectal cancer treatment.
Materials and Methods
Clinical Specimens
The colorectal cancer tissues and the paracancerous tissues were collected from the Hubei Provincial Hospital of Traditional Chinese Medicine. Our study was approved by The Ethic Committee of the Hubei Provincial Hospital of Traditional Chinese Medicine (Approval No. 20200144, ethical approval date: January 7, 2020). All patients provided written informed consent prior to enrollment in the study.
Cell Culture and Transfection
We purchased human colorectal epithelial cell line DLD-1 and colorectal cancer cell lines SW480, SW620, Caco-2, and HCT116 cells from Shanghai cell bank of Chinese Academy of Sciences. These cells were cultured with eagle's minimum essential medium (DMEM, 90%), fetal bovine serum (10%), and double-antibody penicillin streptomycin in 37 °C and 5% CO2 incubators.
The sh-NEAT1, miR-448 mimic, miR-448 inhibitor, pcDNA3.1/NEAT1, and pcDNA3.1ZEB1, and their negative controls were obtained by Genepharma. These plasmids were transfected into colorectal cancer cells using the Lipofectin 3000 (Thermo Fisher Scientific) regent according to the manufacturer's instructions. At 48 h post transfection, cells were harvested for quantitative polymerase chain reaction (qPCR) analysis.
Real-Time Quantitative Polymerase Chain Reaction
Total RNA was extracted from tissues and cells by using Trizolregent (Invitrogen) according to the instructions. The total RNA was reverse transcribed into complementary DNA by using a PrimeScrpt RT reagent Kit (TaKaRa). The SYBR@Premix Ex Taq™ (TaKaRa) was used to perform quantitative RT-PCR in a 20 μL reaction which was subsequently proceeded in a Real-Time PCR detection system (Applied Biosystems™). The above mixture was put into the PCR reaction instrument. The real-time quantitative polymerase chain reaction (qRT-PCR) analysis of miRNA was performed using the All-in-One™ miRNA qRT-PCR detection kit (Genecopeia). The ΔΔCt was calculated by subtracting the ΔCt of the control cells from the ΔCt of the experimental cells. Fold change was generated using the 2−ΔΔCt equation. The qRT-PCR primer sequences used are as follows:
sh-NEAT1 1-forward: 5′-CTTCCTCCCTTTAACTTATCCATTCAC-3′; sh-NEAT1 1-reverse: 5′-CTCTTCCTCCACCATTACCAACAATAC-3′; sh-NC-forward: 5′-ATCATGTTTGAGACCTTCAACA-3′; sh-NC-reverse: 5′-CATCTCTTGCTCGAAGTCCA-3′.
Cell Counting Kit-8 Test
Cell Counting Kit-8 (CCK-8) test was performed on SW480 and HCT116 cells after transfection. The logarithmic cells were collected and cultured in the incubator for tested 10 μL CCK-8 (5 mg/mL) (Sigma) was added into each pore and cultured the cells for 4 h; 150 μL DMSO was added into each well to dissolve formazan. After that, the absorbance value of each well at 490 nm was measured by a Bio-Rad model 550 microplate reader (Bio-Rad).
Transwell Test
Cells after transfection were washed with PBS for 3 times, and then 10 μL cell suspension with the density of 1.25 × 105/mL was seeded into the 24-well plate (BD company), the medium containing 10% serum was added into each well. The inoculated cells were put into the incubator and cultured for 24 h under the condition of 37 °C and 5% CO2. After that, cells were stained with Giemsa kit (Abcam). The cells were taken photos and observed.
Dual-Luciferase Reporter Assay
We used the TargetScan (http://www.targetscan.org/) and DIANA (http://mirdb.org/) to predict the targets of NEAT1. The sequence of NEAT1 and the 3′-UTR of ZEB1containing the predicted binding sites (wt) or mutant binding sites (mut) were synthesized by Genepharma and cloned into pmirGLO vector (Promega). miR-493 mimics or mimics control, along with the plasmids containing wild or mutant type of NEAT1-3′-UTR, were cotransfected into SW480 and HCT116 cells using Lipofectamine3000 (ThermoFisher Scientific). The intensity of luciferase was measured.
Nude Mice Xenograft Assay
Male BALB/c nude mice, 6-week-old, were purchased by Vital River Laboratory Animal Technology Company and housed in a specific pathogen-free environment with a temperature of 25 °C and 60% relative humidity. 5 × 106 SW480 cells stably expressing pSilencer vector or pSilencer/NEAT1 were inoculated subcutaneously into the flanks of nude mice. After 7 days, the width and length measurement of tumors was measured every 2 days and the volume was calculated (Tumor volume = 1/2 × [width]2 × length).
Immunohistochemistry
Paraffin-embedded tumor tissues were sectioned at 4um thickness. These sections were deparaffinized in xylene and rehydrated. Antigen retrieval was done by a pressure cooker for 30 min in 10 mM citrate buffer. 0.5% hydrogen peroxide was used to block endogenous peroxidase activity. The slides were incubated with a specific primary antibody against ZEB1 and plexin A4 overnight at 4 °C. Immunostaining was performed using DAB according to the manufacturer's instructions. The dyed slides were photographed with a microscope.
Statistical Analysis
Each experiment was performed at least 3 times, and take the average value of multiple test results was the final data. When P value <.05, the result was considered significantly different. For comparisons between 3 or more groups, one-way analysis of variance was used, followed by Tukey's multiple comparison tests. Statistical analysis was performed using the SPSS25.0 (SPSS Inc.) and GraphPad Prism software, version 5.0 (GraphPad Software, Inc.).
Results
The Expression Level of NEAT1 Was Higher in Colorectal Cancer Tissues and Cell Lines
We first evaluated the mRNA expression level of NEAT1 in colorectal cancer tissue and paracancerous tissue by qPCR. And the expression level of NEAT1 gene in colorectal cancer was significantly higher than that in paracancerous tissues. In addition, we determined the expression of NEAT1 in human colorectal epithelial cell line DLD-1 and colorectal cancer cell lines including SW480, SW620, Caco-2, and HCT116 by qPCR. The expression of NEAT1 gene was higher in SW480 and HCT116 and was the highest in SW480 cells (Figure 1).

Upregulation of nuclear paraspeckle assembly transcript 1 (NEAT1) expression and downregulation of miR-448 expression in colorectal cancer. (A) The levels of NEAT1 in adjacent normal tissues and colorectal cancer tissues (n = 30). (B) The levels of NEAT1 in human colorectal cancer cell lines and normal cells. (C) The levels of miR-448 in normal cells and human colorectal cancer cell lines. (D) Correlation between NEAT1 and miR-448.
Knockdown of NEAT1 Inhibited Cell Proliferation and Invasion of Colorectal Cancer Cells
The expression level of NEAT1 in sh-NEAT1 transfected cells was decreased significantly in SW480 and HCT116 cells (Figure 2A). The results of CCK-8 assay showed that compared with the control group, the proliferation of SW480 and HCT116 cells transfected with sh-NEAT1 was significantly reduced (Figure 2B and C). The invasion ability of SW480 cells transfected with sh-NEAT1 was significantly impaired in comparison to the control group (Figure 2D and E).
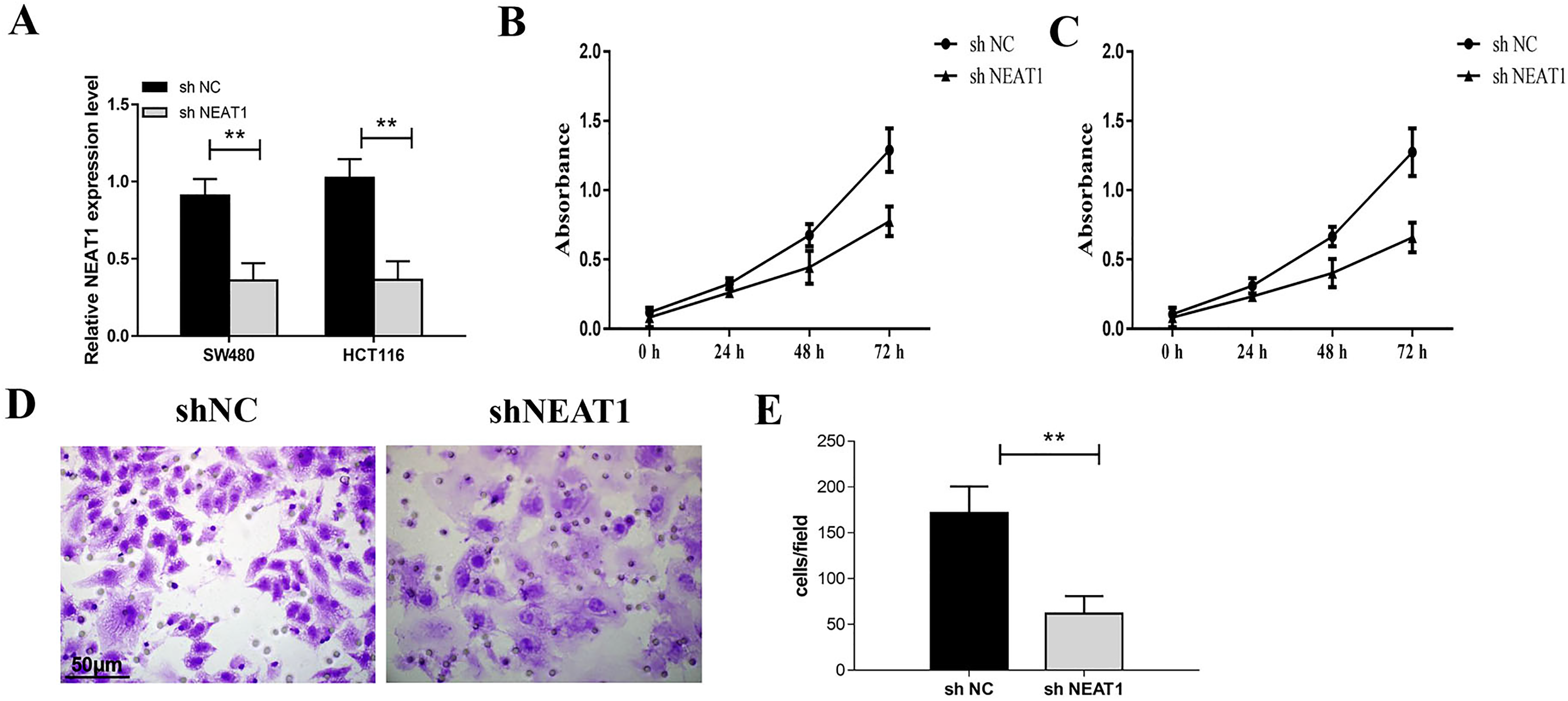
Knockdown of nuclear paraspeckle assembly transcript 1 (NEAT1) inhibited the cell proliferation and invasion of colorectal cancer cells. (A) The expression of NEAT1 in SW480 and HCT116 cells was detected by PCR. The results showed that the expression level of NEAT1 in sh-NEAT1 transfected cells was decreased significantly. (B) Cell Counting Kit-8 (CCK-8) assay was used to detect the proliferation of SW480 cell. (C) CCK-8 assay was used to detect the proliferation of HCT116 cell. (D) Transwell test was used to detect the invasion of SW480 cells after knockdown of NEAT1. *P < .05, **P < .01.
NEAT1 Sponges miR-448 in Colorectal Cancer Cells
The luciferase reporter assay showed that miR-448 overexpression largely decreased the luciferase activity of the reporter plasmid carrying NEAT1-wt but not the NEAT1-mut (Figure 3C and D). The expression of NEAT1 was regulated negatively by miR-448 in SW480 and HCT116 cells. Compared with the control group, the expression level of NEAT1 transfected with miR-448 mimic was significantly lower, while transfected with miR-448 inhibitor was significantly higher (Figure 3E and F).

Nuclear paraspeckle assembly transcript 1 (NEAT1) sponges miR-448 in colorectal cancer cells. (A) The putative target sequence of miR-448 on the 3′-UTR in NEAT1. (B) miR-448 mRNA levels in SW480 cell. (C, D) Luciferase assay was performed to investigate whether miR-448 binds with NEAT1. (E) The expression of NEAT1 in SW480 cells was detected by PCR after miR-448 inhibitor or mimic. (F) The expression of NEAT1 in HCT116 cells was detected by PCR after miR-448 inhibitor or mimic. *P < .05, **P < .01.
miR-448 Inhibitor Transfection Reversed the Effect of NEAT1 Silencing
The results of CCK-8 tests indicated that compared with the control group, the proliferation of SW480 and HCT116 cells was significantly reduced after sh NEAT1 + inhibitor nc transfected. And the proliferation of the cells transfected with sh-NEAT1 + miR-448 inhibitor was promoted in comparison with the sh-NEAT1 groups (Figure 4A and B). Knockdown of NEAT1 inhibited the invasion of colorectal cancer cells while miR-448 inhibitor reversed the effect of NEAT1 silencing (Figure 4C and D).

Inhibition of miR-448 reversed the effect of nuclear paraspeckle assembly transcript 1 (NEAT1) knockdown. (A) Cell Counting Kit-8 (CCK-8) assay was used to detect the proliferation of SW480 cell. (B) CCK-8 assay was used to detect the proliferation of HCT116 cell. (C, D) Transwell test was used to detect the invasion of SW480 cell. *P < .05, **P < .01.
miR-448 Directly Targets ZEB1 in Colorectal Cancer
We predicted the targets of miR-448 by using Targetscan 7.2 and DIANA software and got 535 genes predicted by both the 2 software. Luciferase assay indicated that miR-448 mimic transfection largely decreased the activity of the reporter plasmid carrying ZEB1-3′UTR-wt but not the ZEB1-3′UTR-mut (Figure 5A). In order to clarify the effect of miR-448 on ZEB1expression, we detected the level of ZEB1 in colorectal cancer cells under the overexpression or knockdown of miR-448. The results showed that both the mRNA and protein level of ZEB1 was increased in colorectal cancer cells were transfected with miR-448 inhibitor, while miR-448 mimic group decreased that significantly (Figure 5C and D). Meanwhile, ZEB1 expression was decreased in colorectal cancer cells after NEAT1 silencing (Figure 5E). The expression levels of miR-448 gene and ZEB1 were negatively correlated with clinical colorectal cancer tissues (Figure 5F).
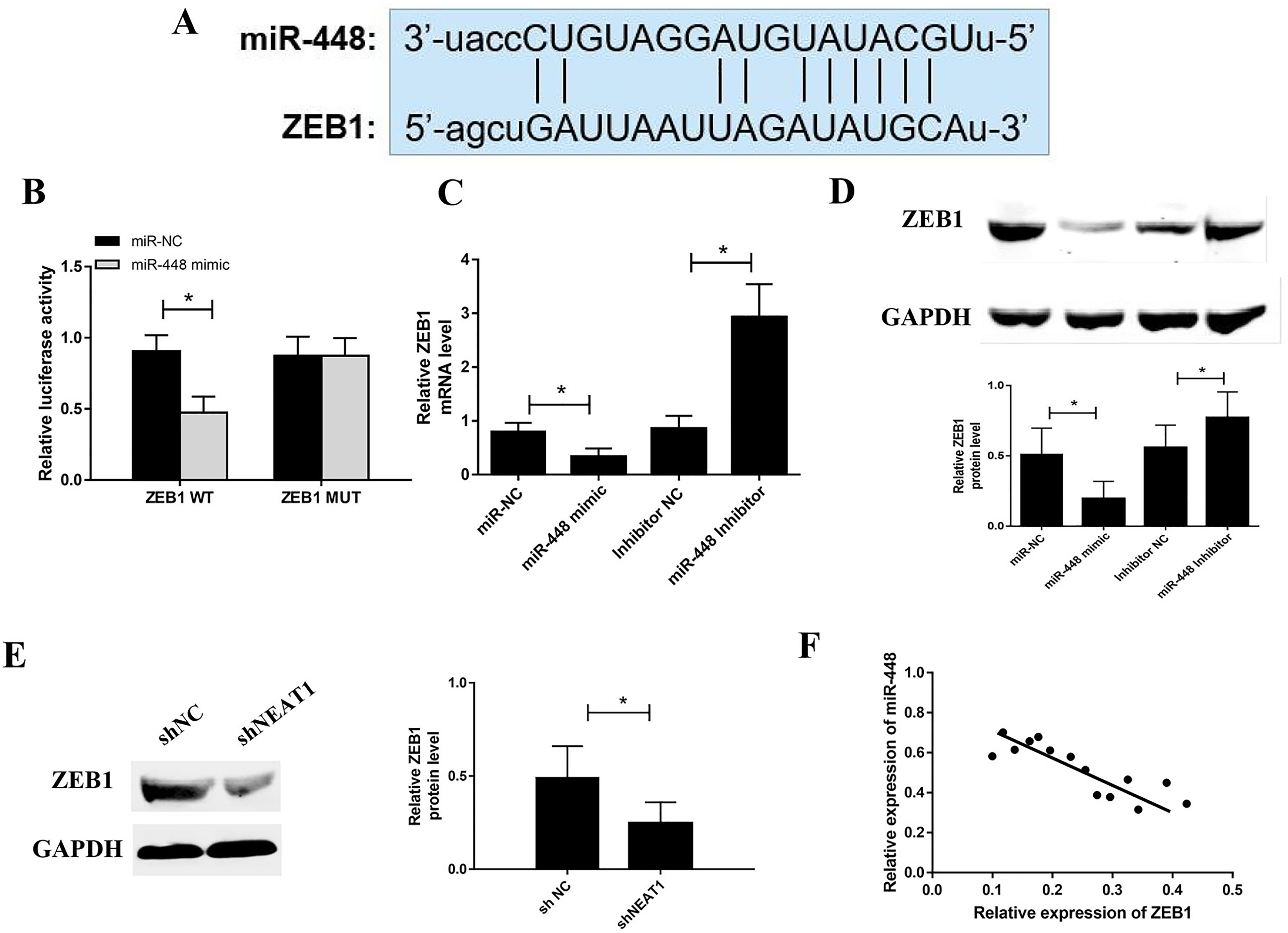
miR-448 targets ZEB1 in colorectal cancer cells. (A) The putative target sequence of miR-448 on the 5′-UTR in ZEB1. (B) Luciferase assay was performed to investigate whether miR-448 bind with ZEB1. (C) The expression of ZEB1 in SW480 cell was detected by PCR after miR-448 overexpression or knockdown. (D) Effect of miR-448 on ZEB1 protein levels. (E,) Effect of nuclear paraspeckle assembly transcript 1 (NEAT1) on ZEB1 protein levels. (F) Correlation between miR-448 and ZEB1.*P < .05, **P < .01.
NEAT1 Inhibits the Growth of Colorectal Cancer Cells in Vivo
In the in vivo study, as shown in Figure 6A and B, SW480 cells transfected with pSilencer/NEAT1 grow more slowly than the pSilencer transfection group. The tumor volume in the pSilencer/NEAT1 was also lower than that of the vector control group. We performed IHC test to confirm if the expression of ZEB1 was modulated. The expression of ZEB1 in the NEAT1 knockdown group was significantly lower than that in the control group (Figure 7D and E).

Nuclear paraspeckle assembly transcript 1 (NEAT1) can inhibit the growth of colorectal cancer in vivo. SW480 cells transfected with pSilencer and pSilencer/NEAT1 were injected into 6-week-old nude mice separately. (A) The tumors in the 2 groups were showed. (B) The survival curve of tumors was made. (C) The volumes of the tumors were detected. (D) The expression of ZEB1 and Ki67 in the tumors was detected by IHC. (E) The expression of ZEB1 in the tumors was assessed by western blot. (F) The model summarized the mechanisms of long noncoding RNA (lncRNA) NEAT1 promoting the malignant progression of colorectal cancer by targeting ZEB1 via miR-448. *P < .05, **P < .01.
Discussion
LncRNA NEAT1 is a gene located at the human chromosome 11 and is correlated to the progression of cancer. It was demonstrated that NEAT1 was significantly increased and was associated with tumor–node–metastasis and maybe a potential diagnostic biomarker for colorectal cancer. It is involved in the development of colon cancer by regulating MYC and WNT. 18 Increasing evidence has confirmed that NEAT1 plays critical role in the progression of other human cancers. Ectopic expression of NEAT1 has been reported in many types of cancer including breast cancer, hepatocellular carcinoma, thyroid cancer, ovarian cancer, pancreatic cancer, and osteosarcoma. 19 It was reported that NEAT1 was significantly upregulated in glioma tissues and correlated with the advanced tumor stage (III/IV). NEAT1 promoted breast cancer growth and metastasis by regulating TGF-β signaling pathway. Silencing of NEAT1 attenuates the proliferation, cell cycle, and migration of glioma cells. Yu et al. verified that NEAT1 promoted EC progression via acting as a molecular sponge of miR-181a-5p. 20 NEAT1 overexpression promoted tumorigenesis in clear cell renal cell carcinoma through regulating Wnt/beta-catenin and plays the role of oncogenes. 21
As in colorectal cancer, it was reported that NEAT1 was up-regulated in colorectal cancer tissues and high expression of lncRNA NEAT1 is related to the prognosis of cervical squamous cell cancer. 22 Our results also indicated that the expression level of NEAT1 gene in 4 colorectal cancer cell lines was significantly higher than that in colorectal epithelial cells DLD-1. NEAT1 silencing can inhibit cell proliferation and trigger the cells apoptosis of colorectal cancer cells. Accordingly, in our study, we confirmed that lncRNA NEAT1 gene is highly expressed in colorectal cancer. Inhibition of NEAT1 can suppress the proliferation, migration, and invasion of colorectal cancer cells.
The mechanism underlying the effect of NEAT1 in cancer development is complicated. It was demonstrated by the previous studies that NEAT1 can regulate cancer metabolism in an allele-specific manner by binding the Cleavage Factor I complex. 23 NEAT1 modulates the expression of Wnt/β-catenin signaling in nonsmall cell lung cancer and esophageal cancer. It can be activated by E2F1 and interacts with PTTG1 to promote the progression of pituitary adenomas. 24 Competing endogenous RNA (ceRNA) mechanism is the most well-identified function of lncRNAs in regulating gene expression. Zhang et al. study found that NEAT1 contributes to paclitaxel resistance of ovarian cancer cells by targeting miR-194. 25 In this study, Bioinformatics analysis also discovered that lncRNA NEAT1 directly sponges miR-448. In addition, using a dual-luciferase reporter and RIP assays, we demonstrated that lncRNA NEAT1, as a molecular sponge for miRNA, targeted miR-448 by ceRNA way. To further confirm whether the regulation of NEAT1 occurs via miR-448, we silenced NEAT1 in colorectal cancer cells with miR-448 inhibitor and found that the miR-448 inhibitor reversed the effect of NEAT1 silencing.
It is well known that lncRNAs can work as the sponges of miRNAs to block the regulatory effect of them. NEAT1 was demonstrated to bind with miR-145 in colon cancer and miR-424 in glioma.26,27 Here, we predicted and verified that NEAT1 sponges miR-448 in colorectal cancer. miR-448 has been reported to be downregulated in colorectal cancer which is consistent with what we found in the present study.28–30 Furthermore, we searched the downstream target genes of miR-448 and ZEB1was found.
The downregulation of miR-448 has been found to be function as a tumor suppressor in colorectal cancer. However, the regulation mechanisms of miR-448 in CRC are still unknown. A previous study reported that miR-448 might serve as a tumor suppressor in CRC partly through targeting IGF1R and that it may serve as a tumor suppressor 31 Another recent study demonstrated that miR-448 was decreased in colorectal tissues and cell lines by targeting Rictor mRNA to negatively regulate the growth and colony formation of colorectal cancer. 32 In our study, bioinformatics analysis also discovered that miR-448 directly targets ZEB1. In addition, using a dual-luciferase reporter assay, we found that miR-448 targeted ZEB1. Further study showed that both the mRNA and protein level of ZEB1was increased in colorectal cancer cells when transfected with miR-448 inhibitor, while miR-448 mimic group decreased that significantly.
ZEB1 can induce EMT process and promote cancer progression and play an oncogenic role in various cancers including bladder, lung, and breast cancer. ZEB1 is also upregulated in colorectal cancer. ZEB1-driven expression of miR-409-3p promotes the proliferation of breast cancer.33,34 We found that NEAT1 works as a ceRNA of NEAT1 and also plays as an oncogene in colorectal cancer. ZEB1can regulate the expression of numerous downstream genes involved in the proliferation and invasion of cancers such as Bcl-2, Bax, cyclinA1, cyclinB1, cyclinD2, and MMP9.
This study has some limitations. First, we determined the NEAT1/miR-448/ ZEB1signal axis in colorectal cancer, but whether there are other molecules and links among these 3 molecules, whether they are affected by other factors remain unknown. Second, the colorectal cancer cell types we used were SW480 and HCT116 cells, more cell lines, and even primary colorectal cancer cells need to be used to confirm what we found in colorectal cancer.
In conclusion, since the literature about the relationship between NEAT1, miR-448, and ZEB1 is rare, our research has a certain degree of innovation, and we have found the possible signal pathway for the development of colorectal cancer. We have found the relevant targets of colorectal cancer, such as NEAT1, miR-448, and ZEB1. We have also proved that NEAT1 can inhibit miR-448 so as to promote the expression of ZEB1, promote the development of colorectal cancer. These findings are likely to provide new targets and ideas for the research and development of drugs for the treatment of colorectal cancer.
Footnotes
Abbreviations
Authors’ Note
Our study was approved by The Ethic Committee of the Hubei Provincial Hospital of Traditional Chinese Medicine (approval no. 20200144). All patients provided written informed consent prior to enrollment in the study. Hanquan Wu and Dengwen Dong contributed equally to the article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
