Abstract
Background:
Gastric cancer (GC) is a common malignancy with high morbidity. Long non-coding RNAs (LncRNAs) have been demonstrated to be critical post-transcriptional regulators in tumorigenesis. This study aimed to investigate the effect of LncRNA NEAT1 on the proliferation and metastasis of GC.
Material and Methods:
The expression of LncRNA NEAT1 was examined in clinical samples and GC cell lines. GC cell lines (SGC-7901 and BGC-823) and human normal gastric epithelial cell line (GES-1) were employed. The correlation between NEAT1, miR-103a and STAMBPL1 was determined by luciferase reporter assay. Cell viability was determined by CCK8 assay. Cell invasion capacity was examined by Transwell assay. The protein level of STAMBPL1 was analyzed by western blotting.
Results:
LncRNA NEAT1 was found to be up-regulated in GC cell lines. Further studies identified LncRNA NEAT1 as a direct target of miR-103a. Moreover, NEAT1 knockdown and miR-103a overexpression inhibited cell proliferation and cell invasion. NEAT1 knockdown and miR-103a overexpression also decreased STAMBPL1 levels.
Conclusion:
Our study indicated that LncRNA NEAT1 was up-regulated in GC cells and tissues. NEAT1 was targeted and inhibited by miR-103a and acted as an oncogene, which promoted the malignant behavior of GC cells. This regulatory effect of NEAT1 may be associated with STAMBPL1. Therefore, NEAT1 could be used as a biomarker for predicting the progression of GC.
Gastric cancer (GC), with an increasing incidence in recent years, is the most common type of digestive malignancies. 1,2 In China, the distribution of gastric cancer is widespread, and the mortality rate in each region is significantly different, and there is a trend of regional concentration. About 400,000 new cases of gastric cancer occur each year, accounting for 42% of the total number of cases in the world. By 2005, gastric cancer has become the leading malignant tumor in China with cancer morbidity and mortality. 3 Although novel biomarker and therapy strategies have been developed rapidly, people with advanced gastric cancer have a poor survival rate. 4 Therefore, it is essential to identify more effective therapeutic strategies and targets.
Recently, people have paid more attention on long non-coding RNAs (lncRNAs). Long non-coding RNAs (lncRNAs) are transcripts of more than 200 nucleotides in length and do not have protein coding potential. 5 LncRNAs are involved in a variety of cell behaviors, including cell proliferation, apoptosis, invasion and drug resistance. 4,6 In addition, numerous studies have uncovered that lncRNAs can act as oncogenes or tumor suppressors through interactions with tumor-related genes in tumorigenesis. 7,8 STAM binding protein-like 1 (STAMBPL1) is a major factor in JAMM family members. 9 Studies have shown that STAMBPL1 is closely related to tumor development. 10 Li et al found that in HeLa cells of cervical cancer, methylation of STAMBPL1 leads to cell proliferation and anti-apoptosis through NFκB, TGF-β and PI3 K signaling pathways. 11 However, there is limited research on the role of STAMBPL1 in GC. Therefore, the present study aims to explore the potential role of STAMBPL1 in GC and to highlight the emerging roles of ceRNAs in the biological regulation of GC cells.
MicroRNAs (miRNAs) are a series of small non-coding RNA molecules (18-25 nucleotides) that can endogenously inhibit mRNA translation by binding to the 3′-untranslated region (3′-UTR) of the target gene mRNA. 12 A large number of reports show that miRNAs are widely involved in the development of various cancers, including GC. 13 Recent studies have shown that lncRNAs have the ability to act as miRNA sponges and regulate miRNA target derepression at the post-transcriptional level. 14 -16 Based on this theory, lncRNAs can antagonieze miRNA’s inhibitory effect by competitively binding to it.
In our present study, we investigated the expression level of NEAT1 in GC tissues and cell lines. The expression level of NEAT1 was found to be upregulated in both GC tissues and cells. Then, we explored the role of NEAT1 in GC progression through function gain & loss experiments. Results suggested that NEAT1 was negatively correlated to the expression of miR-103a. In addition, miR-103a was proved to directly bind to NEAT1. Moreover, results showed that miR-103a functioned as a tumor suppressor in GC. Finally, our study uncovered that NEAT1 promote GC progression by competitively binding to miR-103a. Collectively, our study suggests that NEAT1/ miR-103a/ STAMBPL1 axis has a potential to be used as therapeutic targets for GC therapy.
Results
LncRNA NEAT1 Is Up-Regulated in GC Cell Lines and Tissue
The relative expression levels of NEAT1 in GC cell lines (SGC-7901 and BGC-823) and human normal gastric epithelial cell line (GES-1) were also measured. NEAT1 expression was significantly up-regulated in GC cell lines than that in GES-1 cells (Figure 1A). The relative expression level of lncRNA NEAT1 in 32 GC tissues and paired normal adjacent tissues was measured using qRT-PCR and was normalized to GAPDH. NEAT1 expression was significantly up-regulated in GC tissues compared with that in normals (P < 0.05, Figure 1B). Moreover, the subcellular distribution assay was performed and showed that NEAT1 was predominantly located in the nucleus rather than cytoplasm (Figure 1C and D). Collectively, the results showed that NEAT1 was up-regulated in GC.

LncRNA NEAT1 is up-regulated in GC cell lines and tissue. (A) NEAT1 expression in GC cells (SGC-7901 and BGC-823) and human normal gastric epithelial cell line (GES-1) was detected by qRT-PCR. (B) NEAT1 expression in 32 GC tissues and paired normal adjacent tissues was measured using qRT-PCR. (C and D) The expression levels of NEAT1 in the subcellular fractions of SGC-7901 and BGC-823cells was determined by qRT-PCR.U6 and GAPDH were used as nuclear and cytoplasmic markers, respectively. U6 was used as a loading control in qRT-PCR (*P < 0.05).
NEAT1 Knockdown Inhibits Cell Proliferation and Cell Invasion in GC
Because NEAT1 was up-regulated in GC, we investigated the impact of NEAT1 knockdown in GC cell lines to explore its biological functions. After transfection with siRNAs, NEAT1 was significantly reduced in SGC-7901 and BGC-823 cells (Figure 2A and B). The CCK-8 assay showed that NEAT1 knockdown remarkably inhibited proliferation in SGC-7901 and BGC-823 cells (Figure 2C and D). Cell invasion ability was determined by Transwell invasion assay. The number of invaded cells in the si-NEAT1 group was markedly reduced when compared to that in NC group for both SGC-7901 and BGC-823 cells (Figure 2E). In summary, these results showed that knockdown of NEAT1 can repress both proliferation and invasion capacity in GC cells.

NEAT1 knockdown inhibits cell proliferation and cell invasion in GC. (A and B) qRT-PCR results showed si-NEAT1 were transfected into SGC-7901 and BGC-823 cells successfully. (C and D) CCK-8 assay was performed and the result revealed that the viability of SGC-7901 and BGC-823 cells transfected with si-NEAT1 were remarkably inhibited when compared with si-NC group. (E) Transwell invasion assay showed that the number of invaded cells in the si-NEAT1 group was markedly reduced compared to the si-NC group in SGC-7901 and BGC-823 cells (*P < 0.05).
NEAT1 Negatively Correlated to miR-103a in GC Cells
The putative miRNA candidates targeting NEAT1 were predicted via StarBase3.0. 17 The predicted sites of miR-103a binding to the NEAT1 sequence are depicted in Figure 3A. We then explored that miR-103a expression was down-regulated in the GC cell lines when compared with that in the normal cells (Figure 3B). The expression of miR-103a in clinical samples and found it was significantly down-regulated in GC tissue samples (Figure 3C). Then, a luciferase reporter assay was adopted to verify whether miR-103a directly binds to NEAT1. Cells were co-transfected with miR-103a mimics and the NEAT1-Wt or NEAT1-Mut vector. Data showed that miR-103a significantly suppressed the luciferase activity of NEAT1-Wt when compared with that of the negative control, but miR-103a did not affect the luciferase activity of NEAT1-Mut in SGC-7901 and BGC-823 cells (Figure 3D and 3E). In sum, above results suggested that miR-103a can directly bind to NEAT1 in GC cells.

NEAT1 interacts with miR-103a in GC cells. (A) Putative binding site of miR-103a on NEAT1. (B) MiR-103a expression in GC cells (SGC-7901 and BGC-823) and human normal gastric epithelial cell line (GES-1) was detected by qRT-PCR. (B) MiR-103a expression in 32 GC tissues and paired normal adjacent tissues was measured using qRT-PCR. (D and E) Dual-luciferase reporter assay was performed to determine luciferase activity in SGC-7901 and BGC-823 cells co-transfected with miR-103a mimics and NEAT1-WT or NEAT1-MUT (*P < 0.05).
MiR-103a Inhibits GC Cell Proliferation and Invasion
We then studied the biological function of miR-103a through in vitro experiments. We transfected mimics-NC or miR-103a-mimics into GC cells. After transfection, miR-103a expression in SGC-7901 and BGC-823 cells increased significantly (Figure 4A and B). Results of CCK-8 assay showed that miR-103a overexpression significantly inhibited cell proliferation in SGC-7901 and BGC-823 cells (Figure 4C and D). Subsequently, cell invasion assays were performed to determine the effect of miR-103a on GC cell invasion. The number of invaded cells was markedly reduced after transfection with miR-103a mimics compared with NC group in both SGC-7901 and BGC-823 cells (Figure 4E). Collectively, these data indicate that miR-103a can suppress GC cells proliferation and invasion, which inversely correlates with the effects of NEAT1 in GC cells.
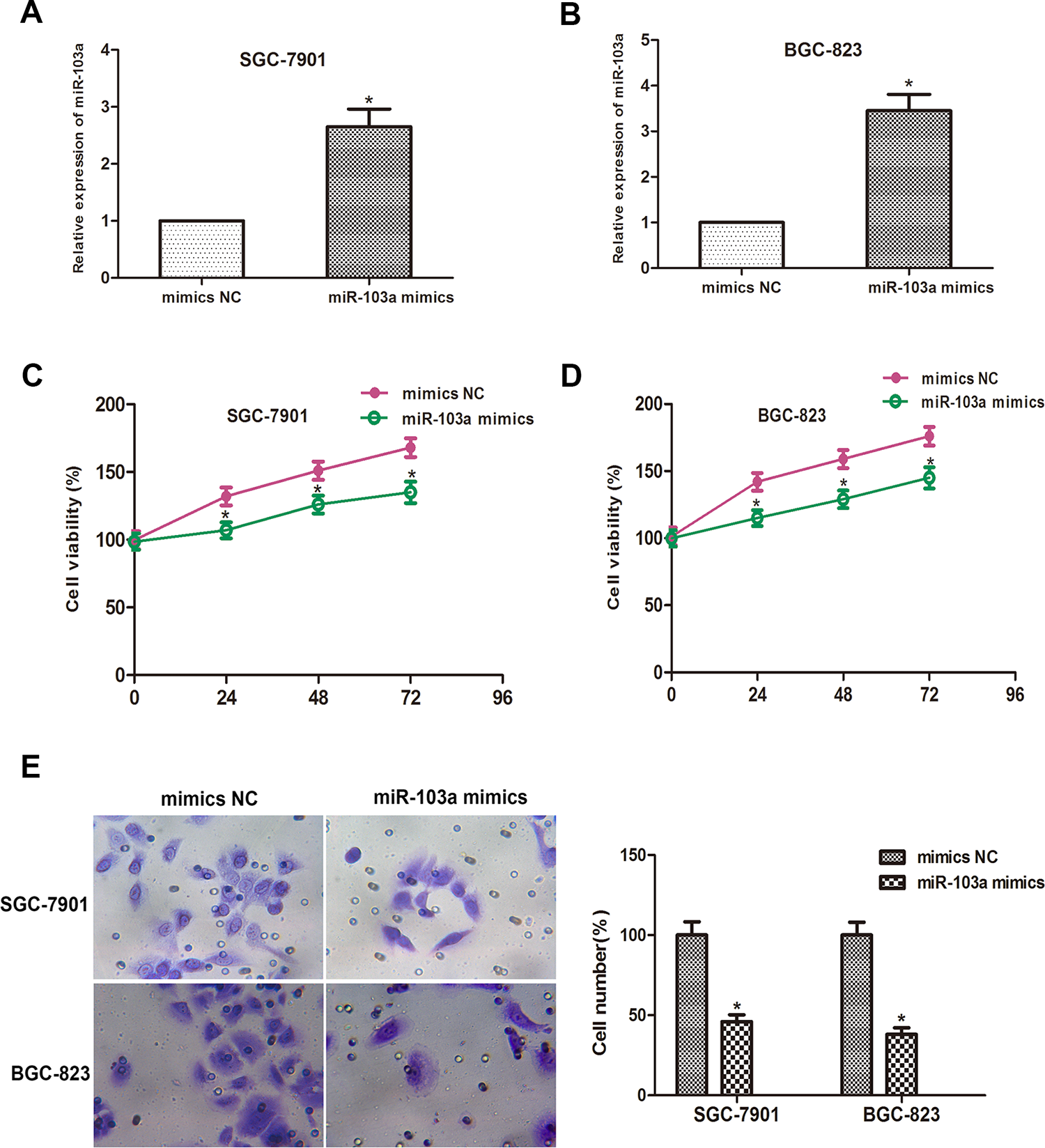
MiR-103a inhibits GC cell progression. (A and B) qRT-PCR results revealed that miR-103a mimics were transfected into SGC-7901 and BGC-823 cells successfully. (C and D) CCK-8 assay was measured and the result showed that the viability of SGC-7901 and BGC-823 cells transfected with miR-103a mimics were remarkably inhibited when compared with NC mimics group. (E) Transwell invasion assay showed that the number of invaded cells in the miR-103a mimics group was reduced compared to the NC mimics group in SGC-7901 and BGC-823 cells (*P < 0.05).
MiR-103a Directly Targets STAMBPL1
STAMBPL1 was predicted to be one of the potential targets of miR-103a by
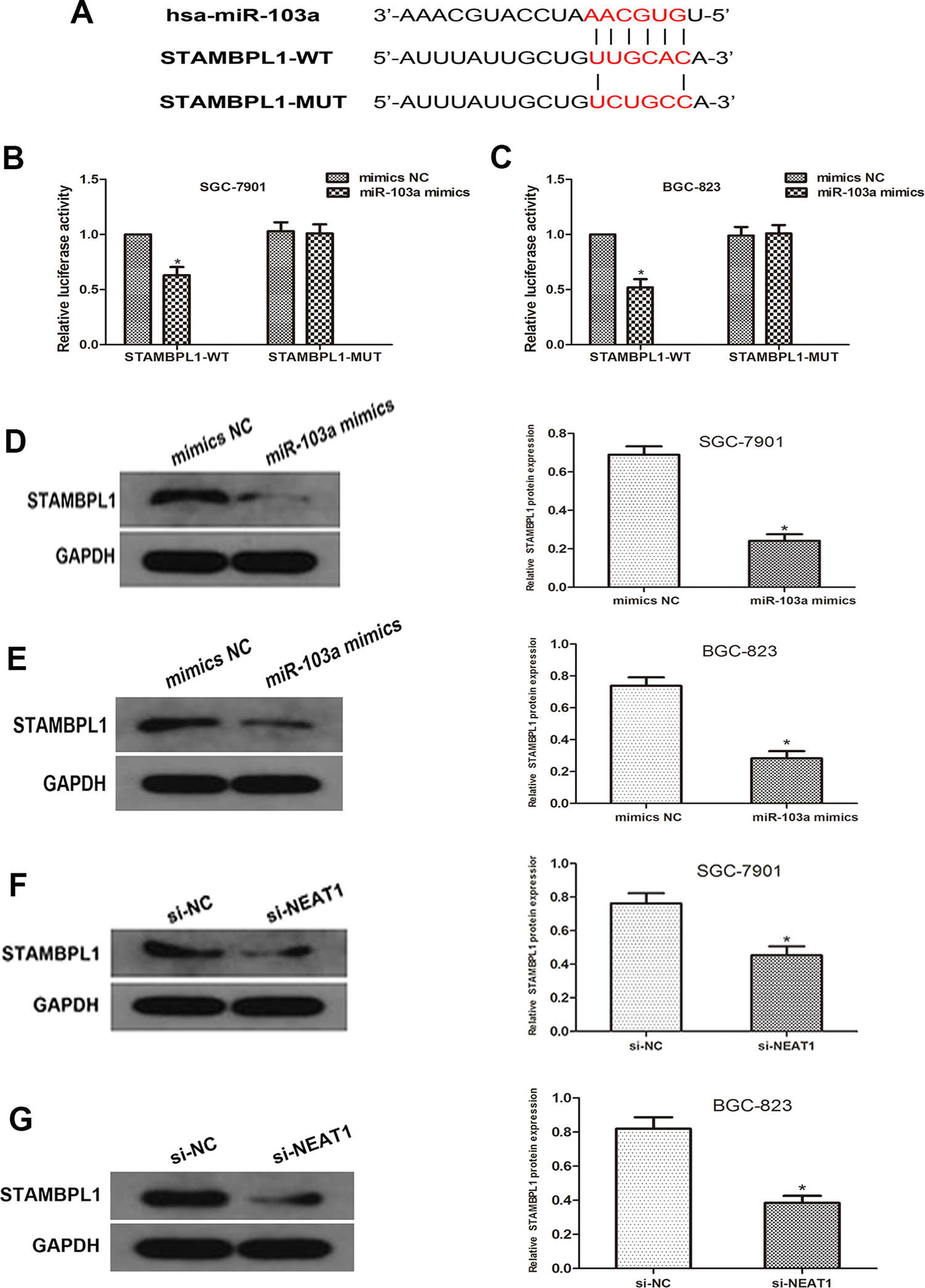
MiR-103a directly targets STAMBPL1. (A) Sequence alignment of predicted miR-103a binding sites with the STAMBPL1 3′UTR and the mutated sequence of miR-103a. (B and C) Luciferase reporter assay was performed in SGC-7901 and BGC-823 cells that were co-transfected with miR-103a mimics and reporter vectors that containing STAMBPL1 3′UTR or mutated STAMBPL1 3′UTR. Relative luciferase activities are shown. (D and E) Western blot analyses showed that overexpression of miR-103a markedly decrease the STAMBPL1 expression level in SGC-7901 and BGC-823 cells. (F and G) Western blot analyses showed that NEAT1 silence led to an decrease in the STAMBPL1 expression level SGC-7901 and BGC-823 cells (*P < 0.05).
Discussion
Recent evidence suggests that the role of non-coding RNAs in a variety of cellular processes has improved our understanding of cancer biology. 18 Therefore, the role of dysregulated non-coding RNAs may provide new insights into the identification of potential therapeutic targets in clinical applications. In our present study, we found that NEAT1 was markedly up-regulated in GC tissues and cell lines. Knockdown of NEAT1 significantly retarded cell proliferation and invasion in GC cells. Moreover, NEAT1 was identified as a direct target of miR-103a. miR-103a was significantly downregulated in GC and acted as a tumor suppressor in GC cells through inhibiting STAMBPL1. Taken together, NEAT1 induced cell proliferation and invasion by sponging miR-103a in GC.
Increasing studies has revealed that several lncRNAs are dysregulated in GC. Some of them have been proved to act as oncogenic lncRNAs. LncRNA CTC-497E21.4 promotes cell proliferation through modulating miR-22/NET1 axis through RhoA signaling pathway. 19 LncRNA UCA1 promotes tumor metastasis by inducing miR-203/ZEB2 axis in gastric cancer. 20 Some lncRNAs act as tumor suppressors in GC. LINC01939 inhibits the metastasis of gastric cancer by acting as a molecular sponge of miR-17-5p to regulate EGR2 expression. 21 LINC01133 inhibits gastric cancer progression by sponging miR-106a-3p to regulate APC expression and the Wnt/β-catenin pathway. 22 In the present study, we found that NEAT1 was up-regulated in GC cells and tissues, and its knockdown significantly inhibited cell proliferation and invasion, indicating that NEAT1 acts as an oncogene in GC progression.
Accumulating evidence has shown that miRNAs play critical roles in tumor progression and development. MiR-103a-3p situated in 5q34, and was associate with tumor, Parkinson disease and pregnancy-related complications. 23 -25 It has been reported that microRNA-103a increases the oncogenic effects of macrophages by targeting PTEN in lung cancer. 26 MiR-103a can also mediate tumor suppressive effect in glioma by regulating FEZF1/CDC25A pathway. 27 In our study, we found that miR-103a could inhibit cell proliferation and invasion by targeting STAMBPL1, which is consistent with previous reports.
In summary, increased expression of NEAT1 was observed in GC patients. NEAT1 was targeted and inhibited by miR-103a and acted as an oncogene, which promoted the malignant behavior of GC cells. Our study facilitates the understanding of NEAT1 function in GC tumorigenesis and provides a novel therapeutic target.
Materials and Methods
Clinical Samples Collection
This research was approved by the Animal Ethics Committee of Bengbu Medical College and the informed consent was obtained from all patients diagnosed with GC. GC tissues and adjacent normal tissues were obtained from patients by surgery and immediately frozen in liquid nitrogen at the first affiliated hospital of Bengbu Medical College.
Cell Culture and Reagents
The human normal gastric epithelial cell line GES-1 and the human gastric cancer cell lines SGC-7901 and BGC-823 were obtained from Bengbu Medical College (Anhui, China). The cells were cultured in RPMI 1640 medium supplemented with 10% fetal bovine serum (FBS, Invitrogen, Carlsbad, CA, USA), 100 U/mL penicillin, and 100 μg/mL streptomycin (Invitrogen) at 37°C under 5% CO2 and 1% O2. All experiments were repeated 3 times independently. Hsa-miR-103a mimics, pcDNA3.1/NEAT1 plasmid, siRNA-NEAT1 and negative control miRNA (miR-nc) were chemically synthesized by Shanghai Integrated Biotech Solutions Co., Ltd. (Shanghai, China). Transfection was performed by using Lipofectamine 2000 (Invitrogen, USA) according to the manufacturer’s instructions.
Quantitative Real-Time PCR (qRT-PCR) Analysis
Total RNA was extracted from fresh frozen tissue or cultured cells using Trizol reagent (Life Technologies, USA). The absorbance ratio of nucleic acids at 260 and 280 nm (A260/280) was measured with a Beckman-DU800 spectrophotometer to evaluate the purity of nucleic acids. First-strand cDNA was synthesized using reverse transcriptase superscript III (Invitrogen, USA) and approximately 2 µg of total RNA. The ABI Prism 7500 system (American Applied Biosystems) was used to quantitatively analyze the expression level of NEAT1 in gas chromatography tissue or cultured cells. GAPDH was used as the internal standard. The expression level of mature miR-103a was quantified by TaqMan miRNA probe and ABI Prism 7500 system (American Applied Biosystems). The relative levels of miR-103a in cultured cells or tissue samples normalized to U6. All reactions are in triplicate.
Cell Proliferation Assay
Cells were treated with different transfection conditions and plated on 96-well plates. According to the manufacturer’s protocol, cell proliferation was assessed daily using the CCK-8 test kit (Beyotime, China) for 3 consecutive days. Cell viability was determined by measuring the absorbance at 450 nm wavelength on the Elx800 system (BioTek, USA). All experiments were performed in 6-fold solution and repeated at least 3 times.
Cell Invasion Assay
Cell invasion analysis uses matrix gel-coated transwell plates (Biosciences, USA). Cells were taken 24 hours after transfection and inoculated into the upper chamber with serum-free medium. Add fresh medium containing 10% serum to the lower chamber as a chemical attractant. After incubation for 48 h, fix and stain with 0.1% crystal violet, and take out the non-invasive cells with a cotton swab. Under the IX71 microscope (Olympus Japan), ImagePro Insight software (Olympus Japan) was used to count 6 random fields in each well. The average number of invasive cells is expressed as a percentage compared to the control group. Data are expressed as the mean ± standard deviation of at least 3 independent experiments
Dual-Luciferase Assay
Cells (2.0*104) grown in 96-well plates were co-transfected with 150 ng NC-miRNA or miR-103a mimics, 50 ng psi-check luciferase expression reporter (Promega), including 3′UTR of NEAT1 (Wild type or mutant type) to detect the relationship between miR-103a mimic and NEAT1 plasmid. 150 ng miR-103a mimic and 50 ng pEZX-MT01 luciferase miRNA expression report (genecopia) were used to detect the relationship between miR-103a and STAMBPL1. In addition, cells were co-cultured with 150 ng miR-103a mimic, 25 ng pEZX-MT01 luciferase miRNA expression reporter (genecopia) consisting of 3′UTR of wild-type STAMBPL1, 75 ng Gas5 plasmid or empty vector to determine miR-103a, The relationship between STAMBPL1 and NEAT1. According to the manufacturer’s instructions, 48 hours after transfection, cells were collected using a luciferase detection kit (Promega) on a Synergy 2 microplate reader fluorometer (BioTek) for luciferase detection. The relative luciferase activity was calculated as the ratio of firefly luciferase activity to renin luciferase activity.
Western Blot
Western blot analysis was performed according to standard protocols. Briefly, whole cell lysates were prepared, and the protein concentration was colorimetrically quantified using the BCA protein detection kit (Beijing Solarbio). The samples were separated in 10% SDS polyacrylamide gel (Solarbio, Beijing) and blotted on PVDF membrane (Millipore). Immunoblotting was performed using commercially available antibodies (anti-STAMBPL1 antibody, 1:5000, Santa Cruz; anti-GAPDH antibody, 1:5000, Santa Cruz.). Capillary electrophoresis (ECL) was used for chemiluminescence detection. Image J software was used to quantify the immunoblot signal.
Statistical Analysis
All data were expressed as mean ± standard from 3 independent experiments. The independent experiments need to be at least 3 times. Statistical analysis was performed using ANOVA with SPSS 19.0 software (IBM Corporation, Armonk, NY, USA). The P < 0.05 were considered statistically significant.
Footnotes
Authors’ Note
Ethical approval number: BBFY2019-K131.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by The Natural Science Research Project of Education Office of Anhui Province (No. KJ2019A0387).
