Abstract
The incidence of colorectal cancer (CRC), as well as subsequent patient mortality, has increased in the last decade; an unhealthy diet is considered to be the leading cause. Previous studies have shown the potential of the bromodomain containing 1 (BRD1) gene as a therapeutic target for CRC based on its specificity; however, the genetic mode of action and expression in CRC cells are yet to be investigated. In this study, target genes were screened from single-cell transcriptome sequencing data, and the collected clinical specimens were subjected to immunohistochemistry (IHC) to identify the protein expression of target genes; the results were verified in the GSE17536 array set. Receiver operating characteristic curves (ROC) and overall survival (OS) were used to test target genes as biomarkers and independent predictive markers for CRC. Based on these results, BRD1 was screened as a target gene, and IHC results showed that BRD1 protein expression in CRC was higher than that in normal tissues and was significantly upregulated in poorly differentiated (PD) CRC. ROC analysis showed that the area under the curve in the collected clinical specimens and GSE17536 were 0.6062 and 0.6094, respectively. OS analysis showed that higher BRD1 protein expression was associated with a significantly shorter survival time. In conclusion, BRD1 expression was positively correlated with PD CRC and negatively correlated with OS, indicating that BRD1 could predict the differentiation state of CRC and may be a novel predictive biomarker.
Keywords
Introduction
Colorectal cancer (CRC) is one of the most common cancers worldwide, accounting for the third highest number of novel cases diagnosed per year. The prevalence of CRC cases was 38.2 per 100,000 men and women per year. The death rate was 13.7% per 100,000 men and women per year. These trends are age-adjusted and are based on cases from 2013 to 2017 and deaths from 2014 to 2018. 1 In 2018, China's age-standardized CRC incidence and mortality rates were 23.7% and 10.9%, respectively. 2 It is possible to locate CRC using colonoscopy. The most prevalent types of CRC are polyp-type and ulcer infiltration-type; serrated polyps are the most significant type. 3 Inspired by morphological observations, current molecular studies have provided strong evidence for the development of serrated polyps as a CRC pathway.4,5 Molecular biological studies have provided strong evidence for the development of serrated polyps in polyp-type CRC. On this basis, many new treatment options have emerged, such as biochemical treatment with trifluoropyridine/tepraxilate and treatment of the intestinal microbiota.6,7 However, there is no specific indicator to predict the degree of differentiation and survival risk of CRC. Once CRC patients are found to have reached a stage of relatively high tumor malignancy or when accompanied by metastasis involving other organs, it is challenging to increase their survival rates.
Single-cell RNA sequencing (scRNA-seq) offers the possibility of characterizing transcriptional dynamics throughout differentiation and determining the final differentiation product. 8 In this study, the Bromodomain containing 1 (BRD1) gene was selected from the raw LI ScRNA-seq data, 9 to further explore its expression in CRC by analyzing its expression in a single cell. The scope of observation was expanded to the entire tumor tissue. This provides us with the possibility of exploring the changes in the same substance at different research levels.
BRD1 encodes a bromodomain-containing protein that acts as a regulator of hematopoiesis by interacting with DNA histone tails to build up the scaffold subunit of various histone acetyltransferase complexes. 10 Immunohistochemistry (IHC) detection has revealed the expression of BRD1 protein in the nucleus, perikaryal cytosol, and proximal dendrites of the neurons in the adult rat, rabbit, and human central nervous systems, which corresponds to the widespread expression of BRD1 mRNA in the brain tissue. 11 Moreover, BRD1 has been associated with nodular prostate diseases and schizophrenia BRD1 also shows single-nucleotide variant enrichment in estrogen-deprived breast cancer. 12
To date, no research has focused on the relationship between BRD1 and CRC. In this study, the expression characteristics of BRD1 in CRC were evaluated to provide a further understanding of the occurrence and development of CRC.
Materials and Methods
Samples and clinical information of 42 CRC cases with various grades of differentiation were collected at the First Affiliated Hospital of Chongqing Medical University (Chongqing, China) from September 5, 2020, to November 28, 2020. Of these, 15 cases were well differentiated (WD), 17 cases were moderately differentiated (MD), and 10 cases were poorly differentiated (PD). All patients were diagnosed by pathologists according to world health organization guidelines. 13 This study was approved by the Ethics Committee of Chongqing Medical University. The study was conducted from December 3, 2020, to February 9, 2021. It is worth noting that the specimen collector could not obtain the participants’ information during or after specimen collection.
IHC was performed on the 42 samples to detect BRD1 protein expression. The fresh tumor tissue obtained after the patients’ operations was directly frozen and stored in liquid nitrogen after being labeled, followed by dehydration, paraffin embedding, and sectioning. The anti-BRD1 antibody (ab181060, from Abcam (China), diluted with 2% bovine serum albumin (BSA)) was used at a dilution of 1:200. Image-Pro Plus 6.0 (Media Cybernetics, America)was used to calculate the integral optical density (IOD) of gray analysis in IHC images.
CancerSEA (http://biocc.hrbmu.edu.cn/CancerSEA/home.jsp) is the first database dedicated to analyzing cancer single-cell state maps, involving 14 functional states of 41,900 cancer cells from 25 cancer types. Our single-cell data were obtained from the Li H. Nat Genet. 2017 (Colon), with accession number GSE81861 and includes 290 cells from 11 cell groups. Simultaneously, the array sets used for verification were obtained from GSE17536 and contains 232 CRC samples.
Genes significantly related to BRD1 expression were identified using co-expression analysis. Then, on the Database for Annotation, Visualization and Integrated Discovery (DAVID) (http://david.ncifcrf.gov/), the signaling pathways involved in co-expression of these genes and their biological functions were analyzed by Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG).
The student's t-test was performed to explore the expression distribution in different differentiation groups. Receiver operating characteristic (ROC) curves were constructed to explore the model of differentiation biomarkers. The overall survival (OS) was estimated from the date of diagnosis to the date of death or the last follow-up. The log-rank test was used to verify the differences. Pearson correlation analysis was used to identify the genes related to BRD1 expression. Differences were considered statistically significant at p < 0.05.
Results
BRD1 is Significantly Differentially Expressed in CRC ScRNA-seq
First, 90 cells containing 3 samples were subjected to ScRNA-seq; 583 genes were found to be of interest for single-cell expression. The correlation between the expression data of each gene and biological function was analyzed. Based on its correlation and specificity with CRC, BRD1 was selected as the research target (Figure 1). As shown in Figure 1A, BRD1 was highly positively correlated with angiogenesis, inflammation, apoptosis, and differentiation, and highly negatively correlated with DNA repair. As an integral aspect of diagnosis, we considered the CRC distinction grades. As shown in Figure 1B, the expression of BRD1 was positively correlated with the differentiation grades of CRC, with a correlation of 0.33 and a p-value of 0.01 (p < 0.05).

Characteristics of bromodomain containing 1 (BRD1) in single-cell sequencing.
BRD1 Protein Expression is Upregulated in CRC Compared With That in Normal Tissues
IHC staining was performed on 42 CRC specimens and the corresponding controls (normal tissues). To evaluate the expression of the BRD1 protein, the average optical density (AOD = IOD / Area) was introduced as the evaluation standard. The results showed that compared with expression levels in normal tissues, BRD1 levels in CRC samples increased significantly, as can be observed in the IHC images in Figure 2A. After digital evaluation, the AOD of BRD1 protein expression was significantly higher than that in normal tissues (p < 0.05) (Figure 2B).
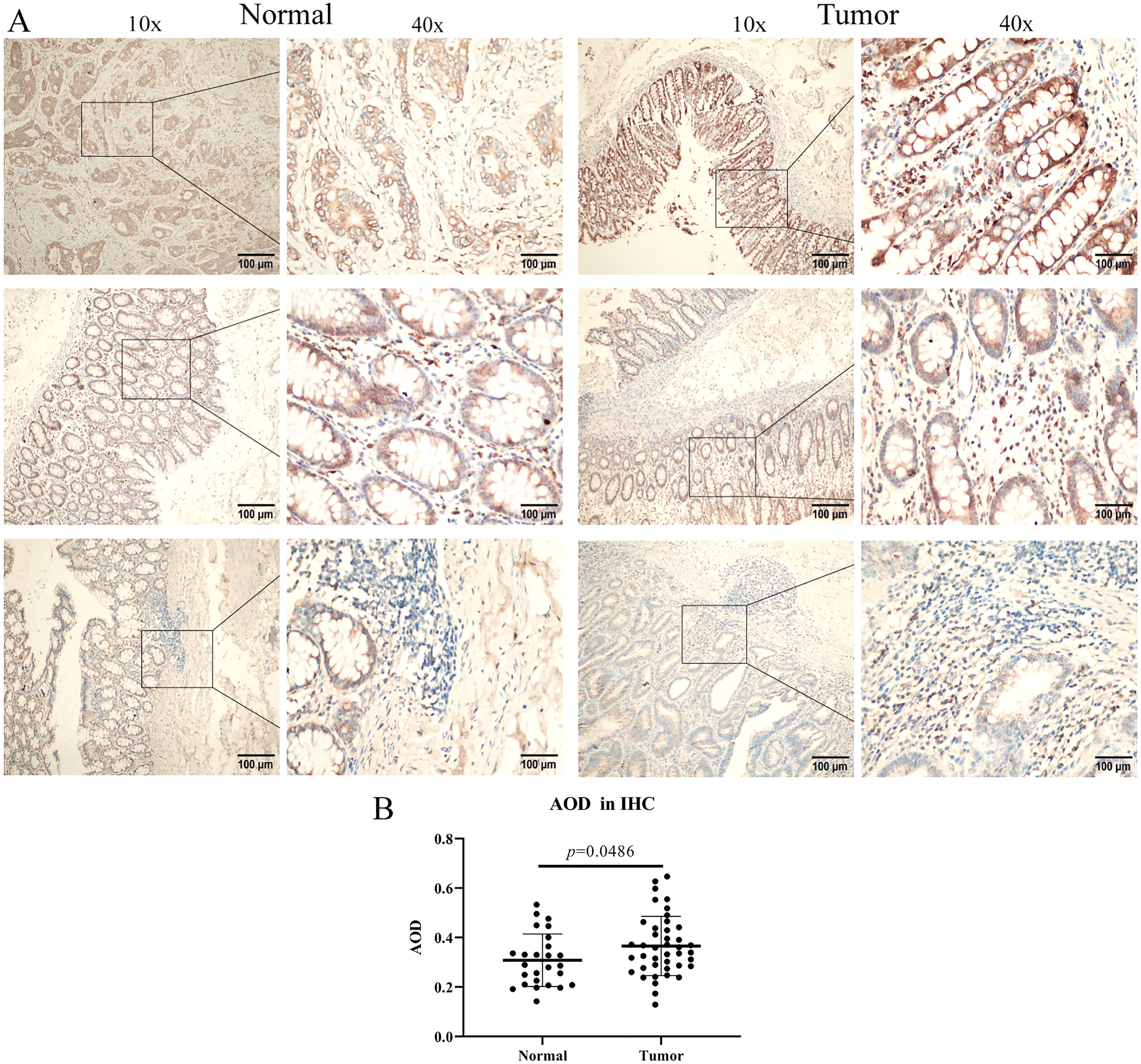
IHC staining of BRD1 in normal and tumor tissues.
BRD1 Protein Expression Is Upregulated in PD CRC
Clinically, the grades of differentiation are usually used to evaluate CRC deterioration and predict prognosis. After IHC staining of 15 WD cases, 17 cases of MD, and 10 cases of PD, IHC images showed significantly more positive results for BRD1 protein in PD than in MD and WD (Figure 3A). Moreover, after calculating the AOD of IHC images, the results (Figure 3B) show that AOD increased with the deterioration of differentiation (p < 0.05). To further validate this result, GSE17536, with 232 samples, was found in Gene Expression Omnibus (GEO) and used for verification. Fortunately, this result is also applicable to this array set (Figure 3C).

Bromodomain containing 1 (BRD1) protein expression in different differentiations.
BRD1 Is a Predictive Biomarker of PD CRC
The ROC is typically used to test the validity of a model. If the area under the ROC curve (AUC) was greater than 0.5, the model was considered valid. In this study, we used BRD1 expression to predict whether a CRC is PD. The results with an AUC of 0.6094 in the collected samples indicated that BRD1 could be an effective predictive biomarker for PD CRC (Figure 4A). Interestingly, the AUC for GSE17536 was 0.6062 (Figure 4B). We also assessed patients’ survival time at each level of BRD1 expression (We sorted the BRD1 expression data of 177 samples from small to large, and then equally divided them into 3 parts, and set >8.784506 as the high expression group, set <8.520746 as the low expression group, and the rest as the medium expression group), and found that patients with high expression of BRD1 had remarkably shorter survival times than others (Figure 4C and D).
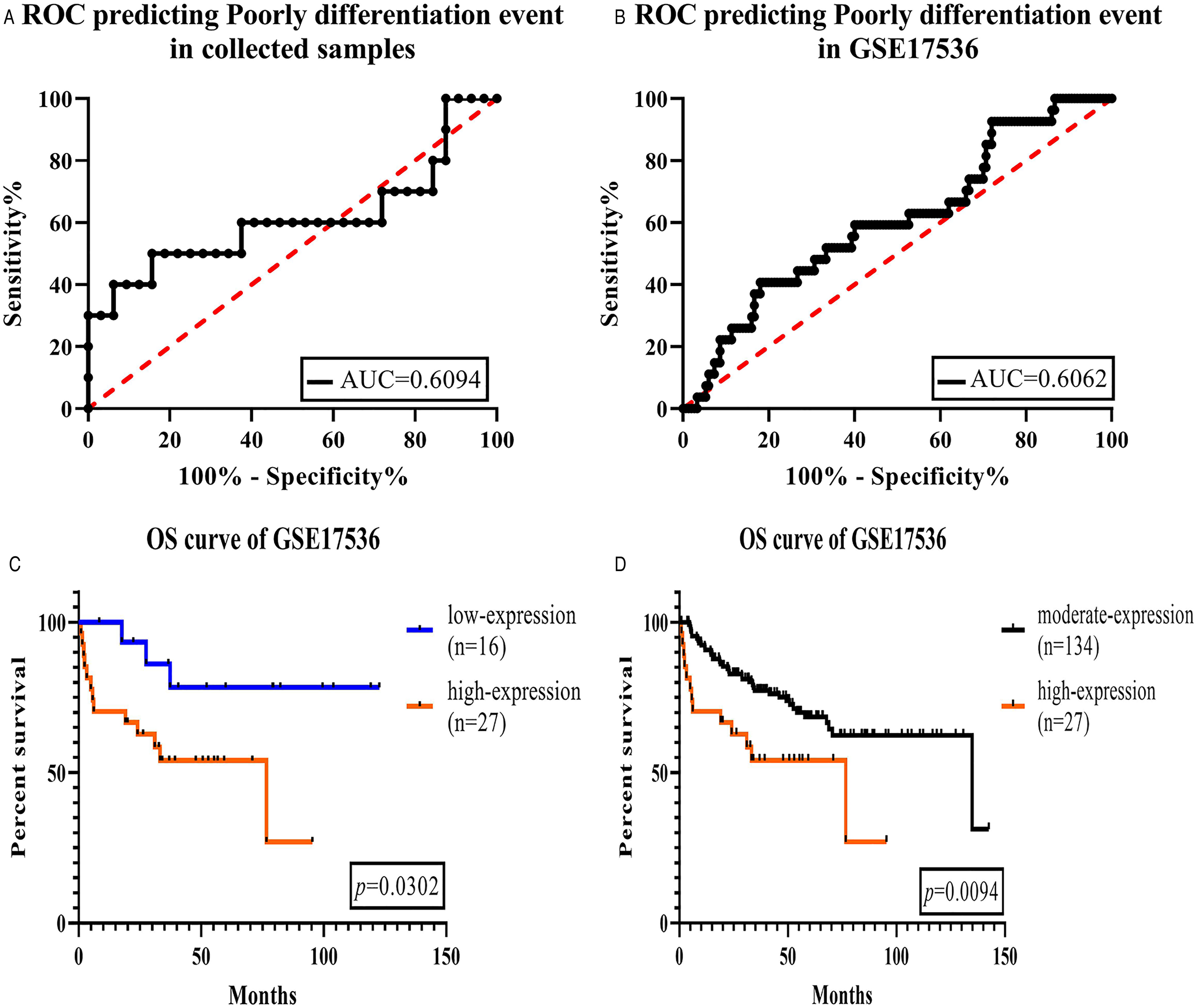
BRD1 is a predictive biomarker of the patients with poorly differentiated CRC.
BRD1 is Associated With Transcriptional Regulation of the Nucleus
To investigate the relationship between BRD1 and genes co-expressed with BRD1, co-expression analysis was performed. Based on the supportability of BRD1, the top 50 co-expression genes were identified and are shown in Figure 5A. The results revealed that BRD1 is highly associated with ZBED4, FAM193A, and EP300, which are closer to the center of the circle, with a higher correlation with BRD1. To explore the biological process of co-expressed genes, GO enrichment and KEGG pathway analyses were performed using the DAVID website. The results are listed in Figure 5B, wherein 6 BPs for the top 50 genes were observed, including DNA-templated transcription; regulation of transcription from RNA polymerase II promoter; positive regulation of transcription from RNA polymerase II promoter; negative regulation of transcription; histone H3-K9 demethylation; and histone H2A monoubiquitination. In Cellular components (CC) and molecular function (MF) of the cell, BRD1 is the primary CC in the nucleus. Protein binding and DNA binding are mainly included in MF. In addition, the Notch signaling pathway is the main KEGG pathway involved in BRD1 regulation.

Co-expression analysis of bromodomain containing 1 (BRD1).
Discussion
The third most common disease and the fourth most common cause of cancer-related deaths are CRC. 14 Currently, the most widely used pathological diagnostic markers in the diagnosis of CRC are: microsatellite instability, APC WNT signaling pathway regulator, kirsten rat sarcoma viral proto-oncogene (KRAS), v-raf murine sarcoma viral homolog B1 proto-oncogene (B-Raf), and serine/threonine kinase.15–17 Nevertheless, the diagnostic accuracy is still not sufficiently high. Furthermore, the anguished treatment experiences of CRC patients and the high death rates of those with PD CRC highlight the urgent need for its effective diagnosis and treatment.
There are many methods employed to study CRC; scRNA-seq is currently one of the most in-depth and accurate of them. 18 In this study, we retrospectively analyzed scRNA-seq data GSE81861. BRD1 was filtered as our target due to its correlation with CRC differentiation of 0.33 and a p-value of 0.01, which suggested that BRD1 may play an important role in the development of PD CRC.
In clinical samples, IHC is a useful approach for detecting protein expression in situ. 19 We tested 42 pairs of samples and normal control tissues. Convincingly, the results of more positive staining in CRC indicated that BRD1 also significantly promoted the occurrence of CRC. Differentiation is an essential indicator of CRC to assess its severity and predict outcomes. 20 We separately calculated BRD1 protein expression in CRCs of different grades of differentiation. We found clear results that the worse the differentiation of CRC, the higher the expression of the BRD1 protein. It is possible that the evolution of CRC from PD to WD was promoted by BRD1. We obtained the same results after expanding the sample size. At the same time, there is no research showing that BRD1 has the same effect on other cancers.
By showing the limits of a test's ability to distinguish between alternative health states across the entire range of operational environments, ROC curves offer a pure index of precision. 21 ROC curves were generated to explore the ability of all grades of CRC to predict PD CRC. The results of AUC for BRD1 expression and GSE17536 analysis, which were 0.6094 and 0.6062, respectively, indicate that the model of BRD1 predicting PD CRC is entirely feasible. Furthermore, PD CRC patients had a lower OS time.
In previous studies, BRD1 was usually associated with mental diseases. 22 In this study, genes co-expressed with BRD1 actively participated in the transcriptional regulation of nuclear DNA. This also confirms that BRD1 is located in the nucleus. Hence, further studies are needed to determine whether the function of BRD1 in brain cells is consistent with that in intestinal cells.
Conclusions
To the best of our knowledge, this is the first study to show an association between CRC and BRD1 expression. These results illustrated that BRD1 may be considered as a novel predictive biomarker for CRC.
Supplemental Material
sj-odt-1-tct-10.1177_15330338211039678 - Supplemental material for Expression Characteristics and Clinical Correlations of BRD1 in Colorectal Cancer Samples
Supplemental material, sj-odt-1-tct-10.1177_15330338211039678 for Expression Characteristics and Clinical Correlations of BRD1 in Colorectal Cancer Samples by Zhou Li, Junjie Wang, Yuzhu Ji and Fangzhou Song in Technology in Cancer Research & Treatment
Supplemental Material
sj-xlsx-2-tct-10.1177_15330338211039678 - Supplemental material for Expression Characteristics and Clinical Correlations of BRD1 in Colorectal Cancer Samples
Supplemental material, sj-xlsx-2-tct-10.1177_15330338211039678 for Expression Characteristics and Clinical Correlations of BRD1 in Colorectal Cancer Samples by Zhou Li, Junjie Wang, Yuzhu Ji and Fangzhou Song in Technology in Cancer Research & Treatment
Supplemental Material
sj-xlsx-3-tct-10.1177_15330338211039678 - Supplemental material for Expression Characteristics and Clinical Correlations of BRD1 in Colorectal Cancer Samples
Supplemental material, sj-xlsx-3-tct-10.1177_15330338211039678 for Expression Characteristics and Clinical Correlations of BRD1 in Colorectal Cancer Samples by Zhou Li, Junjie Wang, Yuzhu Ji and Fangzhou Song in Technology in Cancer Research & Treatment
Footnotes
Acknowledgments
This study sincerely thanks LI for the raw single-cell sequencing data, and also thanks to the English advanced editing provided by editage.
Authors’ Note
Zhou Li contributed to conceptualization, methodology, writing—original draft; Fangzhou Song contributed to conceptualization, writing—review and editing, supervision, funding acquisition; Junjie Wang contributed to the investigation, validation, visualization; Yuzhu Ji contributed to resources; and all authors read and approved the final manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Key Research and Development of Social and People’s Livelihood (grant number cstc2018jscx-mszdX0031).
Supplemental Material
Supplemental material for this article is available online.
Abbreviations
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
