Abstract
Objectives
Gastroesophageal reflux-related chronic cough (GERC), an extraesophageal manifestation of gastroesophageal reflux disease (GERD). Although 24h MII-pH monitoring is the gold standard for diagnosing GERC, its invasiveness, high cost, and limited accessibility hinder widespread use in many clinical settings. This study aimed to develop a non-invasive machine learning model incorporating Peptest™ and GerdQ scores to facilitate GERC detection, particularly in primary care and resource-limited environments where MII-pH testing is not readily available.
Methods
210 chronic cough patients were enrolled between September 2022 and June 2024. GERC diagnosis followed established guidelines, and salivary pepsin levels were measured via Peptest™. Feature selection was performed using the Boruta algorithm (hereafter referred to as Boruta), a method based on random forest (RF), designed to identify relevant variables by comparing them to random shadow features. The selected optimal features were then evaluated using nine ML models, including logistic regression (LR), RF and others. Model performance was assessed through area under the curve (AUC), decision curve analysis (DCA), and calibration curves.
Results
73 (34.76%) patients had GERC. Peptest™ and GerdQ scores were key predictors. Logistic regression was selected for its balance of accuracy (AUC: 0.876) and clinical utility. The nomogram model showed excellent discrimination and calibration. DCA indicated high net benefit at prediction thresholds of 0.10–0.90. RCS analysis revealed non-linear relationships: GERC risk increased with GerdQ >8.66 and Peptest™ >54.791 ng/ml.
Conclusion
The nomogram model provides a reliable, non-invasive tool for GERC diagnosis, aiding timely clinical intervention, especially for patients unsuitable for pH testing.
Keywords
Introduction
GERD is recognized as a condition caused by the reflux of stomach contents, as defined by the Montreal consensus. 1 GERC is a common extraesophageal manifestation of GERD, presenting as a persistent chronic cough,2,3 should be distinguished from laryngopharyngeal reflux (LPR), which is primarily characterized by laryngopharyngeal symptoms. While GERC is associated with acid reflux, LPR often presents with non-acidic or weakly acidic reflux. Additionally, the symptomatology of GERC predominantly involves chronic cough, whereas LPR is marked by symptoms such as hoarseness, chronic throat clearing, and globus pharyngeus. 4
Diagnosing GERC remains a significant challenge. Common diagnostic methods include multichannel intraluminal impedance and pH monitoring (MII-pH), endoscopy, barium meal, empirical anti-reflux therapy, and questionnaires. Historically, 24h esophageal pH monitoring was the primary diagnostic tool for GERC, 2 but it showed limited accuracy in detecting non-acid reflux. 5 Currently, 24h MII-pH is regarded as the gold standard for diagnosing GERC with greater sensitivity and reliability. However, technical difficulties, poor patient tolerance, and limited use in primary care settings remain significant barriers to its widespread application.6,7 Furthermore, traditional methods such as endoscopy and barium meal exams demonstrate low sensitivity. 8
The Gastroesophageal Reflux Diagnostic Questionnaire (GerdQ) was developed from high-quality clinical trials to assist in diagnosing GERC, assessing its impact on patient quality of life, and guiding treatment selection.9–11 While GerdQ is practical for clinical use, its poor sensitivity for non-acid reflux cases remains a limitation. 12
Recently, pepsin, a protease produced by gastric chief cells, has emerged as a potential biomarker for reflux, as its presence in laryngeal mucosa, paranasal sinuses, saliva, and bronchoalveolar lavage fluid has been linked to GERD.13–16 PeptestTM, a diagnostic tool that detects pepsin, has shown promise, with studies indicating positive results in 94% of non-erosive reflux disease (NERD) patients. 17 However, its utility as a standalone test has been questioned, as some studies showed it struggles to reliably distinguish GERD patients from controls. 18
Machine learning (ML) offers a novel approach to improving GERC diagnosis by analyzing complex multivariate data, capturing non-linear relationships, and handling noisy or missing data. 19 ML algorithms can potentially enhance diagnostic accuracy by identifying patterns and valuable factors that traditional methods might overlook. In this study, we aim to utilize ML techniques to identify key predictors of GERC and develop more accurate diagnostic models. Through the Boruta, we ultimately selected GerdQ and Peptest™ as key factors, and therefore, these two factors were primarily included in our ML learning model. Both GerdQ scores and PeptestTM results are clinically significant in the diagnosis and management of GERC, offering non-invasive yet effective assessment of reflux-related symptoms and pepsin detection.
Methods
Study setting and participants
This study was conducted in the Outpatient Clinic in the Department of Respiratory and Critical Care Medicine at Tongji Hospital of Tongji University (Shanghai, China). 210 consecutive patients with cough lasting for ≥ 8 weeks were included, and those who met the following criteria were assigned to the GERC group.20,21 The diagnosis of GERC was determined according to the criteria established by the American College of Chest Physicians and the Chinese Medical Association.
22
The diagnosis of GERC was established based on the following criteria: (1) a chronic cough (lasting >8 weeks) that occurred mainly during the day, with normal chest radiography or computed tomography (CT) images and with or without co-existing typical reflux-related symptoms such as regurgitation and heartburn; (2) positive results from MII-pH, such as AET >6.0%, SI ⩾50%, and/or SAP ⩾95%; (3) significant improvement in cough symptoms (a reduction of >50% in cough symptom score) following stepwise anti-reflux therapy, even in the absence of positive MII-pH findings; (4) potential GERC patients were considered when the other common causes of chronic cough including cough variant asthma, upper airway cough syndrome, eosinophilic bronchitis were excluded after a sequential laboratory workup according to an established algorithm.
23
GERC was diagnosed when patients met either the (1)+(2)+(4) or (1)+(3)+(4) combination of the diagnostic criteria described above. If the above conditions were not met, patients would be assigned to non-GERC group. The study has received approval from the Ethics Committee of Tongji Hospital (K-2015-007) and was registered with the Chinese Clinical Trials Register (ChiCTR1800020221). Informed consent was obtained from all participants. Figure 1(a) illustrates the study’s flowchart. Clinical characteristics collected for all enrolled patients included demographic variables (age, sex), anthropometric measures (height, weight and body mass index (BMI)). These data were obtained through standardized patient interviews and medical record review at the time of enrollment. A:Study flowchart. GERC: Gastroesophageal reflux-induced chronic cough; ML: Machine Learning. B: Boruta Algorithm Variable Selection Plot. Feature selection for the relationship between various clinical characteristics and risk of GERC analyzed by the Boruta algorithm. The horizontal axis shows the name of each variable, while the vertical axis represents the Z-value of each variable. The box plot depicts the Z-value of each variable in the model calculation, with green boxes representing important variables, yellow representing tentative attributes, and red representing unimportant variables. C: Receiver operating characteristic curves assessing the predictive ability of 9 models for GERC. D. Decision curves analysis (DCA) of the 9 models.
Laboratory investigation
GerdQ
The GerdQ is a simple, self-administered questionnaire including six items, developed from three questionnaires (Gastrointestina Symptom Rating Scale – GSRS, 24 Reflux Disease Questionnaire – RDQ 25 and the GERD Impact Scale – GIS 26 ). (Supplemental Table 1). The scores are added together to calculate the total GerdQ score, which ranges from 0 to 18. GERD should be considered when the GerdQ score is 8 or higher.
HARQ (Hull Airway Reflux Questionnaire)
HARQ was created by Morice et al. and is accessible for clinical use in 38 different languages at no cost on the website https://www.issc.info/. 27 The HARQ comprised 14 items related to cough triggers or aggravating factors and associated symptoms. Patients were asked to recall how these items affected their lives over the past month and rate them on a 6-point scale from 0 to 5. The total score can range from 0 to 70, with higher scores indicating greater cough sensitivity. A HARQ score of 13 or higher was considered indicative of reflux-related symptoms. Additionally, the HARQ was used in conjunction with other assessments. (Supplemental Table 2).
PeptestTM
PeptestTM was used to detect pepsin in saliva samples as a non-invasive diagnostic marker of extraesophageal reflux. According to the manufacturer’s instructions, patients were instructed to provide unstimulated saliva samples under standardized conditions. Samples were centrifuged, and the supernatant was analyzed using a lateral flow device with monoclonal anti-pepsin antibodies. A test result was considered positive when the pepsin concentration exceeded 75 ng/mL, based on the manufacturer’s threshold. 28
The principle of Peptest™ is based on the detection of pepsin in saliva, which reflects reflux into the upper aerodigestive tract. Pepsin is commonly regarded as a biomarker for LPR, a condition pathophysiologically distinct from GERD. While GERD is typically acid-mediated, LPR often involves weakly acidic or non-acidic reflux. It is important to note that the diagnostic sensitivity of Peptest™ may be influenced by the timing of sample collection, with postprandial and nocturnal samples potentially increasing detection rates. In this study, only fasting saliva samples were collected for Peptest™ analysis. While standardized, this approach may have limited the ability to capture reflux events occurring during peak periods. Future studies would incorporate multi-timepoint sampling protocols to better capture peak reflux episodes, particularly in patients with non-acidic reflux, thereby enhancing the diagnostic sensitivity of Peptest™.
Feature extraction and selection
Feature selection was a vital step in narrowing down the number of features within a dataset, following the identification of relevant study variables. A key feature selection method used was the Boruta. Boruta is a wrapper-based feature selection method built on the random forest classifier that ranks feature importance using Z-scores. Boruta works by generating shadow features—duplicates of the original dataset’s features—and then comparing the Z-scores of these shadow features with those of the real features during each iteration of model development. If a real feature’s Z-score exceeded the maximum Z-score of the shadow features, it was considered important and retained; Conversely, if a real feature’s Z-score was lower than the highest shadow feature score, it was considered uninformative and discarded. This process helped to identify the most relevant features for further analysis. 29
Model development and performance
We use 9 machine learning techniques to develop diagnosis model: LR, 30 K-nearest neighbors (KNN), 31 Naive Bayes (NB), 32 decision tree (DT), 33 support vector machine (SVM), 34 RF, 35 Xtreme Gradient Boosting (XGB), 36 Light Gradient Boosting Machine (LGBM), 37 and artificial neural network (ANN). 38 Machine learning models were trained with selected viables as input to clarify patients. The data were randomly split into a training set (n = 147) and a test set (n = 63) at a ratio of 7:3. The training set was used to fit the model, while the test set was used to evaluate the model’s performance. Although there is no universally established sample size threshold for ML modeling, prior studies suggest that datasets with over 150 samples are generally adequate for binary classification tasks involving a limited number of predictors, particularly when paired with cross-validation techniques. In our study, the sample size and feature-to-sample ratio were deemed acceptable for the selected ML methods.39,40
Sample size and statistical analysis
The sample size was calculated based on the anticipated diagnostic performance of the combined model using GerdQ and Peptest™. Preliminary data suggested a sensitivity of 0.79 and a specificity of 0.87 for identifying GERC. Assuming a GERC prevalence of 40% among patients with chronic cough, and setting the allowable margin of error (L) at 0.12 with a two-sided α of 0.05 (Z = 1.96), the required sample size was estimated using Buderer’s method.41,42
This calculation specifically targeted the model’s discriminative capacity (sensitivity/specificity) for GERC diagnosis, not pepsin concentration thresholds. This aligns with guideline recommendations for diagnostic model development, 43 where sample size determination focuses on classification accuracy rather than biomarker levels. A total of 210 patients were ultimately included to ensure robustness.. Based on these calculations, the minimum required sample size was 160 patients. A total of 210 patients were ultimately included in the final analysis.
Continuous variables following a normal distribution are presented as mean (standard deviation), and comparisons between two groups were conducted using independent samples t-test. For continuous variables not following a normal distribution, data are presented as median (lower quartile, upper quartile), and comparisons between groups were made using the Wilcoxon rank-sum test. Categorical variables are expressed as frequency and percentage, and comparisons between groups were performed using the chi-square test. The area under the receiver operating characteristic (ROC) curve (AUC) was used as the primary indicator in this study, and the 95% confidence interval (CI) for the AUC was calculated using the Bootstrap method with 1000 repetitions. The ROC curve was plotted, and accuracy, sensitivity, specificity, and F1 score were selected to evaluate the model’s discriminative performance. Calibration of the model was assessed by plotting a calibration curve and calculating the Brier Score. 44 Decision Curve Analysis (DCA) was employed to reflect the clinical net benefit of the model. 45 Statistical analyses were conducted using R 4.2.3 and Python 3.8.10 software, with all tests being two-sided, and p-values <0.05 considered statistically significant.
Results
Patient characteristics
General clinical characteristics of patients.
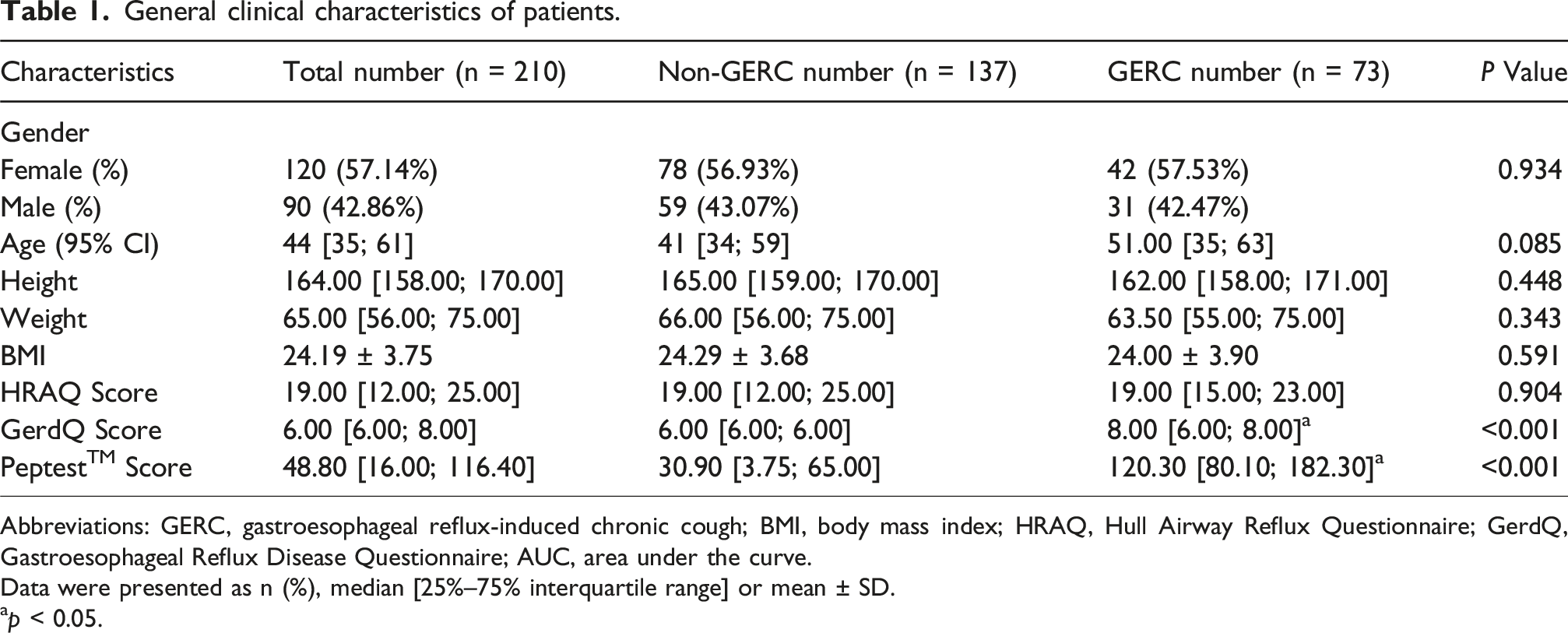
Abbreviations: GERC, gastroesophageal reflux-induced chronic cough; BMI, body mass index; HRAQ, Hull Airway Reflux Questionnaire; GerdQ, Gastroesophageal Reflux Disease Questionnaire; AUC, area under the curve.
Data were presented as n (%), median [25%–75% interquartile range] or mean ± SD.
ap < 0.05.
Feature extraction and selection
The Boruta analysis was conducted to identify the determinants that have significant importance in predicting risk of GERC. In each iteration, the Boruta algorithm compares the Z-score of each original feature with the highest Z-score among the shadow features (randomly permuted duplicates). Features with lower importance than the maximum shadow feature are considered uninformative and are excluded from further modeling. As a result, several characteristics-including age, gender, weight, height, BMI, and HARQ scores-were considered as irrelevant and removed. In contrast, GerdQ and PeptestTM scores emerged as high importance factors. (Figure 1(b)).
Machine learning model performance
The predictive performance of each model.

Abbreviations: KNN, K-Nearest Neighbors; NB, Naive Bayes; SVM, Support Vector Machine; RF, Random Forest; XGB, Xtreme Gradient Boosting; LGBM, Light Gradient Boosting Machine; ANN, artificial neural network; LR, Logistic Regression; DT, Decision Tree. AUC, area under the curve; AUC. low/up, lower/upper bound of the AUC (95% CI).

A Calibration Curve analysis of the 9 models. B. Restricted cubic spline curves for risk of GERC in chronic cough by GerdQ (A) and PeptestTM (B). The cutoff values in the plot of GerdQ were 8.663, and the cutoff values for PeptestTM were 54.791 ng/ml. C. Nomogram to estimate the risk of GERC in chronic cough patients. Total points were sum of GerdQ and PeptestTM scores.
Pairwise comparison of AUC differences using DeLong’s test among each model.

Abbreviations: KNN, K-Nearest Neighbors; LGBM, Light Gradient Boosting Machine; LR, Logistic Regression; ANN, artificial neural network; NB, Naive Bayes; RF, Random Forest; SVM, Support Vector Machine; DT, Decision Tree; XGB, Xtreme Gradient Boosting.
*p < 0.05.
Comparison of the diagnostic value of Peptest™ and GerdQ score alone versus in combination for diagnosing GERC
Comparison of the diagnostic value of Peptest™ and GerdQ score alone versus in combination for diagnosing GERC.
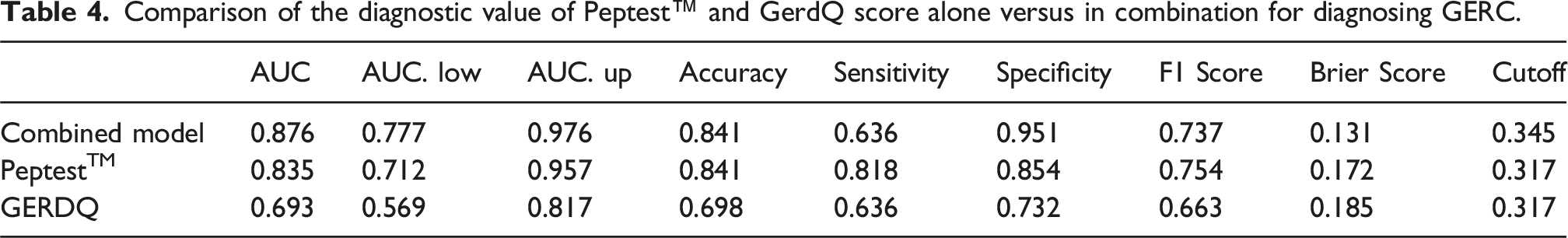
Restricted cubic spline (RCS)
The RCS curve analysis indicated the GerdQ (non-linear p = 0.014) and PeptestTM (non-linear p = 0.010) were non-linearly related to the risk of GERC. The cutoff values in the plot of GerdQ were 8.663, and the turning point for PeptestTM were 54.791 ng/ml, with a significant increase in risk after crossing cutoff values rexpectively. (Figure 2(b)).
Nomogram construction
A predictive nomogram was developed based on the logistic regression analysis, incorporating two significant risk factors for predicting GERC (Figure 2(c)). Total points were sum of GerdQ and PeptestTM score. This total score could then be projected onto the lower total point scale to estimate the probability of GERC.
Discussion
This study provided evidence about the diagnostic accuracy of the GerdQ and Peptest™ scores in a chronic cough population with GERC. We compared 8 viables and screened out GerdQ and PeptestTM scores as 2 key factors via Boruta algorithm.
In the exploratory analysis of Diamond study, cutoff 8 of GerdQ provided the highest specificity (71.4%) and sensitivity (64.6%) in diagnosing GERD.46,47 C. Jonasson et al. 47 indicated that a GerdQ cutoff of 9 might provide more optimal balance between sensitivity, at 66% (95% CI: 58–74), and specificity, at 64% (95% CI: 41–83), for diagnosing GERD. While because of high incidence of reflux esophagitis in study, the GerdQ demonstrated a high positive predictive value of 92% (95% CI: 86–97), but a low negative predictive value of 22% (95% CI: 13–34) for GERD. It is noteworthy that the primary care patients involved in the two studies sought consultation for a diverse array of common upper gastrointestinal symptoms, such as reflux and/or dyspeptic symptoms, or were part of a more specifically selected reflux population. GerdQ includes six main items: reflux, heartburn, nausea, insomnia, epigastric pain and medication use. It is evident that the questions are based on typical symptoms of gastroesophageal reflux, which are more commonly observed in patients with acid GERC. 48 In our previous studies found that optimal GerdQ cutoff for predicting GERC was identified as 8.0. At this cutoff, the sensitivity was 66.7% and specificity was 91.7%. When focusing solely on acid GERC in a subanalysis, the GerdQ demonstrated even higher predictive accuracy, with a sensitivity of 90.9%, specificity of 78.6%, a positive predictive value of 71.4%, and a negative predictive value of 96.4%. However, it was not possible to determine a reliable GerdQ cutoff for predicting non-acid GERC. 49 Thus, GerdQ has limited diagnostic value in non-acid GERC. The GerdQ cutoff in this study was 8.663, which was almost consistent with those previously reported. Our combined diagnostic model models presented value with a sensitivity of 63.6% and specificity of 95.1%. Consistent with previous studies, our model showed a significant improvement in specificity, with a similar sensitivity, compared with single GerdQ in diagnosis. This was also confirmed in the comparison with our standalone GerdQ model, where the AUC was 0.693, significantly lower than that of the combined diagnostic model. Notably, the GerdQ is primarily designed to assess typical acid reflux symptoms such as heartburn and regurgitation, and it does not capture otolaryngological symptoms commonly associated with LPR, including hoarseness, chronic throat clearing, and globus pharyngeus. 50 This limits its utility in detecting non-acid GERC or LPR-related phenotypes. Therefore, to improve screening sensitivity for patients presenting with extraesophageal symptoms, combining the GerdQ with additional instruments such as the Reflux Symptom Index (RSI) may be beneficial. Further incorporation of supplementary tools like the RSI will be considered in future studies to enhance the diagnostic performance in atypical reflux cases.
Pepsin consists of a family of isoenzymes, with pepsin 3b being the most predominant. It remains active across a broad pH range and is not irreversibly denatured until the pH exceeds 7.5. This characteristic makes pepsin an ideal biomarker for detecting reflux in clinical samples, particularly for identifying reflux above the upper esophageal sphincter in conditions like EER, LPR, and respiratory reflux. 51 PeptestTM was a noninvasive diagnostic device via utilizing a proprietary monoclonal antibody to detect pepsin in salivary. Although pepsin is recognized for its role in GERD at low pH due to its proteolytic activity, refluxed pepsin, even in nonacidic or weakly acidic reflux, could cause mucosal inflammation in the laryngopharynx, which is not addressed by PPIs. This may help explain why some reflux-related symptoms and findings remain resistant to PPI treatment. The receptor-mediated uptake of nonacidic pepsin, as seen in nonacid LPR, and the resulting inflammatory and neoplastic changes, are not prevented by PPIs.52,53 This may also be the main reasons why PeptestTM is useful in detecting both acid and non-acid reflux patients.
Rongrong Chen et al. 54 compared postprandial, post-symptom and random salivary samples. Results indicated that the post-symptom PeptestTM had the highest area under the curve among the three time points, with a diagnostic specificity of 81.0% and a sensitivity of 53.3% at a diagnostic threshold of 86 ng/ml. Jinyuan Gan et al. 55 also observed a significant correlation between positive endoscopy findings of erosive esophagitis (EE) and salivary pepsin concentrations. The predictive probabilities derived from binary regression of fasting and post-prandial pepsin concentrations resulted in an AUC of 0.955 ± 0.044 (95% CI 0.868 to 1.000, p < 0.001). Another research of our group is also suggested salivary pepsin concentration >76.10 ng·mL−1 had high diagnostic value both for acid and non-acid GERC. 56 The cutoff value for PeptestTM in our study was 54.791 ng/ml. The variability in diagnostic values across different studies suggests that further studies need to determine a more appropriate diagnostic cutoff value. Although the comparison between our combined diagnostic model and the single Peptest™ model showed no significant difference in p-value, it may be necessary to increase the sample size and further classify GERC for more detailed comparisons.
Rena Yadlapati et al. 57 hold that it was not appreciable differences between healthy volunteers and subjects with a combination of laryngeal and reflux symptoms via single salivary pepsin analysis. Studies also pointed those detectable differences existed in mean salivary pepsin concentration, and future studies would benefit more from use of a quantitative reader in replace of visual semi-quantitative assessment. Gastroesophageal reflux involves acid and non-acid components of gastric contents. 58 It is the burden of refluxate rather than acidic burden which drives laryngeal complaints. 59 Therefore, patients with atypical symptoms or non-acid reflux, acid-suppressing medications may be less effective, requiring more comprehensive diagnostic and treatment approaches. In this study, we focused on assessing the diagnostic value of Peptest™ for detecting reflux-related chronic cough. However, for suspected LPR patients, it is crucial to note that combining Peptest™ results with laryngoscopic findings could enhance the detection rate of non-acidic reflux. Laryngoscopy allows direct visualization of reflux-induced changes in the larynx and pharynx, which are typical in LPR but often overlooked in conventional GERD diagnostics. Therefore, integrating these findings with Peptest™ results may provide a more comprehensive assessment, improving the sensitivity and accuracy of LPR diagnosis.
HARQ is also an important questionnaire developed from Reflux Symptom Index (RSI) in UK. 60 It seems to be more valuable to distinguish among different kinds of cough. 61 While our previous model combined with HARQ and GerdQ exhibiting effective value, HARQ did not express a similar valuable importance in this study. 62 This discrepancy may be attributed to differences in symptom distribution and sample size across study populations. It is also worth noting that HARQ includes several items reflecting typical LPR symptoms, such as throat clearing, hoarseness, and globus pharyngeus, which are not always present in patients with acid-dominant GERC. Therefore, its diagnostic value may be more relevant in LPR-specific subgroups. Future studies are warranted to explore the role of HARQ in such patient populations through dedicated subgroup analyses.
This may be the first time to explore the combined diagnostic value of GerdQ and PeptestTM in GERC, which may help to further improve the diagnosis of GERC. However, the study had some limitation. First, it is a single-center study and sample size needs to be increased for further validation. Second, we opted for a clinically practical approach by collecting a single fasting sample, but the absence of salivary pepsin testing at multiple time intervals limited our ability to identify the optimal time-point for diagnostic value. Third, the sample sizes were relatively small, and study lacked validation for the diagnostic value of the combined model for acidic versus non-acidic GERC. Additional promotion and validation of these models are necessary. In the future, we will refine the data collection methods and design a prospective study to further validate the findings of this research.
Conclusion
In conclusion, this single-center study developed and internally validated a novel, non-invasive nomogram that combines Peptest™ and GerdQ scores to predict the risk of GERC. The model demonstrates high accuracy, excellent calibration, and user-friendly interpretability. Importantly, it holds strong potential as a practical tool for the early screening and triage of patients with suspected GERC, particularly in primary care and resource-limited settings where access to 24h MII-pH monitoring is restricted. By enabling personalized risk assessment, this model may assist clinicians in making more informed and timely treatment decisions.
Supplemental Material
Supplemental Material - Diagnostic value of Peptest™ combined with gastroesophageal reflux disease questionnaire in identifying patients with gastroesophageal reflux-induced chronic cough
Supplemental Material for Diagnostic value of Peptest™ combined with gastroesophageal reflux disease questionnaire in identifying patients with gastroesophageal reflux-induced chronic cough by Jiaying Yuan, Xiao Luo, Lina Huang, Yaxing Zhou, Bingxian Sha, Tongyangzi Zhang, Shengyuan Wang, Li Yu, Xianghuai Xu in Chronic Respiratory Disease
Footnotes
Ethical considerations
The study protocol was approved by the Ethics Committee of Shanghai Tongji Hospital (approval number: 2018-LCYJ-013). Written informed consent was obtained from all participants prior to their involvement in the study.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Natural Science Foundation of China (No. 82270114 and 82070102), the Project of Science and Technology Commission of Shanghai Municipality (No. 22Y11901300, 21Y11901400 and 20ZR1451500), the Program of Shanghai Academic Research Leader (No. 22XD1422700).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The data and related materials from this study can be obtained from the corresponding author upon reasonable request.
Study Registration
This study is registered with the Chinese Clinical Trials Register under the identifier ChiCTR1800020221.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.



