Abstract
Clinical trials are vital for advancing care. However, a systematic approach to tracking trial participation across different facilities and sponsors has been lacking. We developed natural language processing (NLP) methods to extract study enrollment history, including enrollment status, consent date, and study title from information on clinical trial participation recorded in clinical notes in the electronic health record based on national Veterans Affairs electronic health record data. The method exhibited high test-set precision for enrollment status (0.94), consent date (0.97), and study title (0.87) and acceptably high recall (0.76, 0.70, and 0.84, respectively). From a single center, the classifier correctly identified 111 of 125 trial participants (88.8%) across 12 distinct trials. Our study demonstrates the feasibility of using NLP to capture trial enrollment from a nationwide healthcare system. This algorithm creates a novel data resource for analyzing and tracking trial enrollment at the population level.
Keywords
Introduction
Electronic health records (EHR) are increasingly used to facilitate clinical trial recruitment and follow-up. 1 Specifically, in the case of follow-up of trial participants, clinicians and study personnel can obtain event data in real-time from the EHR with or without need for manual review. Thus, various methods have been developed to mark trial participant data in the EHR for long-term follow-up in clinical research. 1 When systematically integrated into the EHR on a large scale, these electronic markers of trial participation provide a unique opportunity to study and monitor trial participation at a population level. For example, population-level monitoring of patient demographics (e.g., geographic location and rurality) and trends (e.g., over time and location) can inform efforts to increase clinical trial access and participation.
To facilitate EHR use for clinical research, the United States Veterans Affairs (VA) healthcare system encourages use of “research” notes for documentation of clinical trial participation.2,3 These notes often have research-specific note titles to distinguish them from other patient chart documentation. Research note titles are metadata labels (e.g., “RESEARCH ENROLLMENT NOTE,” “RESEARCH CONSENT”) assigned by VA clinical staff to distinguish research-related documentation from routine clinical care notes. The VA system is the largest integrated healthcare system in the United States, providing care for over 9 million Veterans each year at over 1,255 facilities across the nation. 4 As part of required or recommended documentation for trial participation, the “research” trial enrollment consent note contains information on both the study title for which the patient was consented and the date they were consented for the study. 5 The study title can provide insights into the study’s intent, and when the titles are matched to trials known to have recruited at the VA, additional details can be obtained such as intervention, sponsor, start and completion dates.
The VA’s use of a nationwide shared EHR and shared clinical trial documentation processes enables tracking trial enrollments on a large scale, all linked to the rich clinical data contained within the EHR. This approach will enable in-depth analysis of multifaceted barriers to trial enrollment at various levels: trial, facility, provider, and patient. 6 Performing this detailed, multi-level analysis is currently not feasible because existing methods for identifying comprehensive trial enrollment do not link to the EHR, 6 and other large-scale resources that cover multiple facilities lack patient-level or facility-level data. 7 In this study, we create a generalizable EHR-linked trial enrollment classifier. This EHR-based classifier will systematically identify and categorize trial enrollment, enabling a comprehensive and systematic understanding of enrollment gaps at various levels.
The purpose of the current study is to develop and evaluate a method to extract research study enrollment as recorded in unstructured clinical notes, using data from the VA’s national EHR database. We use independently recorded trial enrollments from a medical oncology at a VA facility to determine the completeness of our trial enrollment capture. We focus specifically on cancer because oncology trials are a leading specialty in clinical trial development and execution, representing over 20% of interventional clinical studies registered on ClinicalTrials.gov, a comprehensive and robust trial registry whose use is mandated by US regulations. 8
Methods
Data source and primary dataset
This study is based on EHR data from the nationwide VA healthcare system, which is collated in the VA Corporate Data Warehouse (
System architecture
We developed a rule-based system with three components. First, we created a rule-based NLP algorithm to identify the subset of notes in the Primary Dataset that are most likely to record study enrollment, called the Enrollment Classifier. This step helped us distinguish enrollment notes from other notes, such as those that primarily described research follow-up appointments. Application of the Enrollment Classifier produces the Refined Dataset containing notes classified as containing information about a patient’s enrollment in a study. Second, we created two rule-based NLP algorithms to extract information the Refined Dataset about each study a patient enrolled in: (a) the Study Title NLP, which extracts the title of the study the patient enrolled in, and (b) the Consent Date NLP, which extracts the date of the patient’s consent to participate in the study. These three algorithms (the Enrollment Classifier, the Study Title NLP, and the Consent Date NLP) are described in the following sections. Source code embodying the algorithms is described in Code Availability below.
Enrollment classifier
To develop the Enrollment Classifier, we randomly selected 300 notes from the Primary Dataset and allocated 50% of the notes (150 documents) as the training set for Enrollment Classifier development and the remaining 50% (150 documents) as the held-out test set used for final performance evaluation. All model development work, including all preliminary performance evaluations conducted as part of model development, were conducted in the training set; it was not necessary to create a separate “tuning” or “development” set due to the use of a manual rule-development process. These 300 notes were split into 3 sets of 100 notes which were each annotated by one of our 3 pairs of annotators. Each member of the pair independently labeled their set of 100 notes so that all notes were annotated twice. To measure annotator reliability while accounting for expected chance agreements in the annotations, we calculated a note-level Cohen’s Kappa as the measure for inter-annotator agreement. The note-level Cohen’s Kappa averaged across the three pairs of annotators was κ = 0.61 on the initial annotation, demonstrating moderate-to-substantial agreement. Informal review of conflicting annotations revealed disagreement about how to handle notes that mixed information on study enrollment with information on screening for eligibility (before enrollment) and/or study follow-up (after enrollment). Conflicting annotations were adjudicated by a third reviewer, who included all notes that recorded study enrollment, regardless of whether they also included other information. Clinical note annotation was performed using Label Studio, an open-source data labeling tool that provides an easy-to-use interface for annotating data.
Annotators were instructed to label the notes as “Enrollment” or “Not Enrollment” and to annotate the text phrases relevant to making their decision. The annotators used the following annotation guidelines. (a) If the note wording indicated that a patient had enrolled in a study, e.g., “a consent form was signed”, it was labeled “Enrollment”. (b) Notes detailing patient screening that included phrases such as “patient enrolled” were also labeled “Enrollment”. (c) Notes incidentally describing earlier patient enrollment or a patient’s progress, e.g., containing phrases such as “visit #2”, “3 weeks visit”, “6 months visit”, were classified as “Not Enrollment”. (d) If the note text only referenced a scanned consent document stored elsewhere but did not provide further details, then it was also labeled “Not Enrollment”.
We used the training set of annotated notes to build a rule-based enrollment classifier. The training set was reviewed for the presence of repeating patterns consistent with either the note being an Enrollment note (inclusion patterns) or being a Not Enrollment note (exclusion patterns). Inclusion patterns included phrases indicating a signature for informed consent or patient agreement to participate in the study. Exclusion patterns included phrases indicating the note was related to a follow-up visit, a patient progress note, or a patient’s withdrawal from a study. For each note, the algorithm first searched for relevant exclusion patterns in the note text and assigned the “Not Enrollment” label to matching notes. Next, the algorithm searched for inclusion patterns and assigned the note “Enrollment” as applicable. Finally, all residual unassigned notes were labeled “Not Enrollment”. We iteratively refined the algorithm using a
Study title NLP and consent date NLP
The Enrollment Classifier was executed on the full Primary Dataset and classified approximately 24% (1,068,879 notes) of the notes as Enrollment. This set of clinical notes was identified as the Refined Dataset. We randomly selected 750 notes from the Refined Dataset and allocated 50% of them (375 notes) as the training set for the Study Title NLP and Consent Date NLP development and the remaining 50% (375 notes) as the test set. Three pairs of annotators were assigned a set of 250 notes for which each member labeled the set independently. The 750 notes were split into 3 sets of 250 notes which were each annotated by one of our 3 pairs of annotators. As with the previous annotation scheme, each set of 250 notes was independently annotated by each member of the assigned pair so that all notes were annotated twice. The token-level Cohen’s Kappa was κ = 0.89 for the consent date annotation and κ = 0.93 for the study title annotation. Conflicting annotations were adjudicated by a third reviewer. Figure 1 presents examples of annotated enrollment notes. Enrollment notes with study title and consent date annotation: The general structure of clinical notes can vary significantly. (A-C) Three de-identified notes are shown with their respective study title (yellow) and consent date (pink) annotated.
Annotators used the following annotation guidelines for Study Title: (a) Annotate the first occurrence of the fully written-out, human-readable study title, if such a title is present. (b) If title is not present, annotate the first occurrence of study acronym, for example, “EVALUATE-HF”. (c) If neither is present, then annotate the first occurrence of trial number, IRB number or other study identifier.
Annotators used the following annotation guidelines for Consent Date: (a) Annotate the first occurrence of the consent or enrollment date. (b) If there is both a generic visit date and a more specific date in regard to consent or enrollment, then use the more specific date. (c) Do not annotate testing date as a consent date. (d) Exclude the time part of the date, if present (e.g., “3/14/15 @ 9:26am”).
We used the training set of annotated clinical notes to build a rule-based NLP algorithm. The set was reviewed for the presence of repeating word patterns associated with the protocol title and consent date. These patterns were then used to generate regular expressions to facilitate automated rule-based annotation. The rule-based NLP algorithm was designed to extract information consistent with the annotation guidelines described above.
More specifically, we used two strategies to identify research protocol titles. The first approach employed two regular expression sets, one to detect the start of a protocol title, e.g., “Research study name:”, “Name of protocol:”, and another to detect the end of a title, e.g., line break, paragraph end or subsequent section title. The word context between the start and end of the title was annotated as the research protocol title.
The second strategy utilized visual exploration of word order and syntax within high frequency clinical notes, i.e., those likely to be the same study, to create more accurate regular expression sets. More specifically, this entailed minimizing extraction of unwanted context flanking the title as well as that surrounding consent dates.
We iteratively refined our rule-based NLP algorithm using the training note set and a
Performance evaluation
Performance of the Enrollment Classifier, Consent Date NLP, and Study Title NLP were evaluated with reference to their respective test sets. After tabulating the confusion matrix (the number of true positives, true negatives, false positives, and false negatives), standard evaluation measures were evaluated, including precision, recall, accuracy, and the F1 score.
The Enrollment Classifier is a binary classifier, so tabulation of the confusion matrix was straightforward. For Consent Date NLP, we converted all annotated dates into a normalized form (YYYY-MM-DD) and compared the results by value. For Study Title NLP, we used two different methods to produce the confusion matrix. In the Offset Overlap Method, we considered any non-zero length overlap between the NLP annotation and the human annotation as a match. In the Normalized Levenshtein Distance Method, we used the normalized Levenshtein distance metric with a threshold of 0.7 to evaluate the results. Cases where both the NLP algorithm and the annotator produced a value but these results did not match under the chosen criterion were classified as false positives.
Validation relative to external clinical trial enrollment data
In addition to validating the NLP algorithm’s performance relative to the labels our annotators assigned in the held-out test set, we aimed also to assess its performance in identifying individuals confirmed to have enrolled in trials based on gold-standard enrollment data that is external to our EHR data source. We randomly selected patients known to have consented to specific cancer clinical trials from a single VA facility (Durham VA Medical Center) medical oncology department, encompassing various trial types, enrollment periods, and sponsors. There was no overlap between these patients and those with notes annotated in the training and test sets described in prior subsections. In addition to the trial name, we obtained different possible trial identifiers that may be used instead of the study title, such as protocol number and IRB number. We used NLP to identify trial enrollment status in the validation set as follows: We obtained patient notes from the period of potential trial enrollment, during which a patient could have been enrolled in more than one trial. If no enrollment note was detected by the Enrollment Classifier in the patient’s notes during this period, the patient was labeled as “not found”. If at least one NLP-derived trial title matched the recorded trial identifier (title, IRB number, or protocol number), the patient was labeled as “found-match”. If the NLP study title did not match the recorded study identifier, the patient was labeled as “found-nonmatch”.
Code availability
Code for the complete system, including the Enrollment Classifier, the Consent Date NLP, and the Study Title NLP, is available at https://github.com/bostoninformatics/.
Results
Performance evaluation results
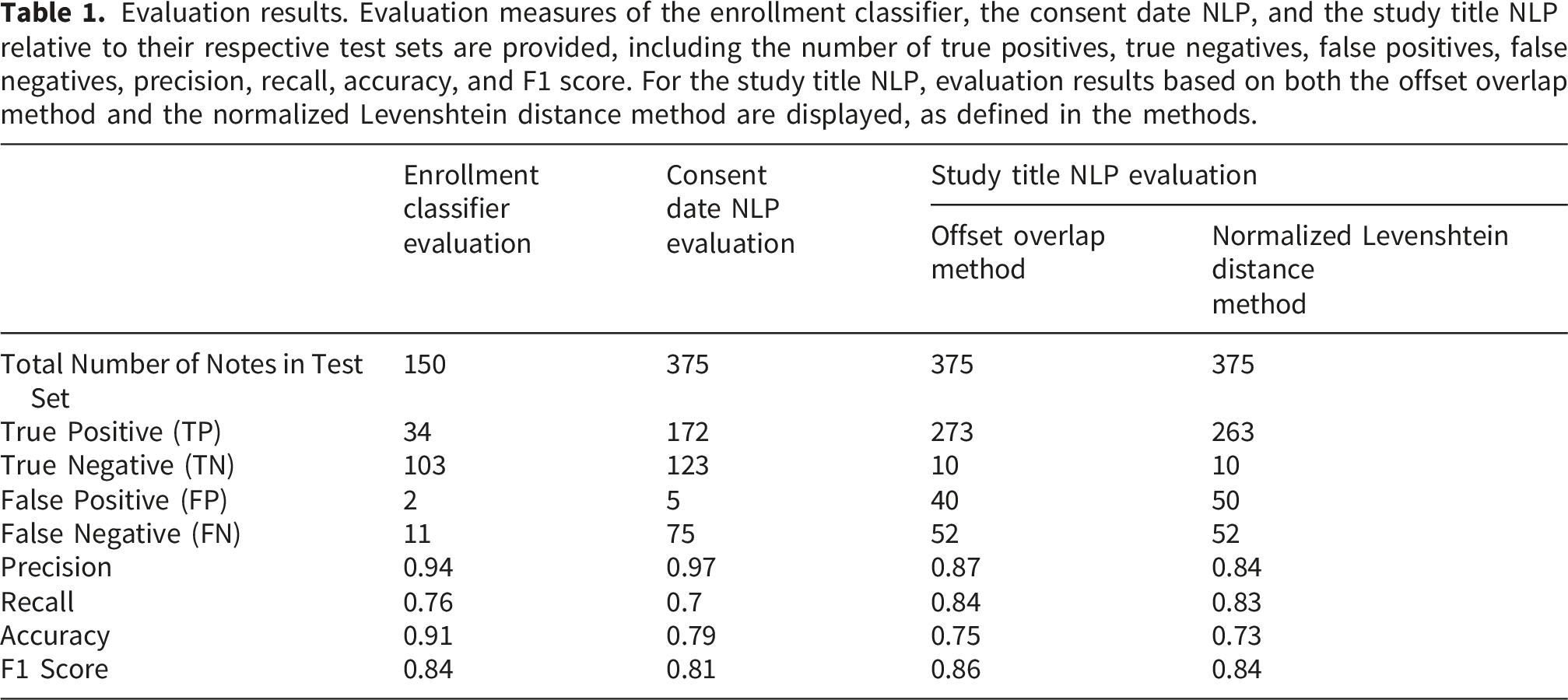
Evaluation results. Evaluation measures of the enrollment classifier, the consent date NLP, and the study title NLP relative to their respective test sets are provided, including the number of true positives, true negatives, false positives, false negatives, precision, recall, accuracy, and F1 score. For the study title NLP, evaluation results based on both the offset overlap method and the normalized Levenshtein distance method are displayed, as defined in the methods.
Error analysis
Error analysis. False positive error analysis for the study title and consent date NLP are shown. Each false positive in the test set was reviewed and the error was classified as to its type. For study title NLP, error analysis is shown only relative to evaluation with the offset overlap method, since performance was similar relative to evaluation with the normalized Levenshtein distance method.

For the Study Title NLP false positives, the majority (29 out of 40, i.e., 73% of the false positives) are only technically false positives, meaning that while the extracted span did not exactly match the annotated target used for evaluation, the algorithm still correctly captured a valid study identifier. This includes 15 errors where the study number or IRB number was matched instead of title, 7 errors where the correct/partial title was matched in a different part of the note from the annotation, 4 errors where the study title abbreviation was matched but the study title is present along with the abbreviation in the same sentence, and 3 errors where the correct study title was captured in the wrong note type (e.g., in termination note or progress note). If these errors were reclassified as true positives, the precision for Study Title NLP would be 0.96.
For Consent Date NLP false positives, all but 1 (4 out of 5 false positives, or 80%), were captures of the correct consent date but in a context that did not align with our guidelines. If these errors were reclassified as true positives, the precision of the Consent Date NLP would be 0.99. We also analyzed Consent Date NLP false negatives. The most common cause of missed consent dates is the case where consent date appears in a context not seen in the training set. The NLP algorithm relies on an extensive set of specific patterns that capture the majority of consent dates seen in the training set data, but it limits its use of wild card matching to reduce the number of false positives. The second most common cause of the missed consent dates is unresolved coreference. For example, the date of initial study participation discussion is mentioned in one part of the document, while the other part says that the patient has consented on that date. Other causes of the missed consent dates include misspelled dates (e.g., with year being specified as 3 digits) and dates in obscure formats that are not recognized by NLP as valid dates.
Validation relative to external clinical trial enrollment data
Missingness analysis of participants in cancer clinical trials from a single center. Comparison of recorded enrollments from a single VA center from the medical oncology department from years 2000 to 2023, representing 12 distinct trials.

§Includes all trials that enrolled within the given time period. Trials do not have to recruit for the entire duration of the time period listed. Trials that span multiple time periods are split by the years of enrollment of the individual trial participants. Specific dates of enrollment were not recorded externally and so the “total patients enrolled” and “enrollment not found” column has been purposefully left blank for those rows.
Application to full dataset
Enrollment note count based on information content. The number of notes identified in the primary dataset, the refined dataset, and those with a study title, a study title and consent date, and a study title but not consent date are presented. The number of distinct patients represented in these notes is also shown.
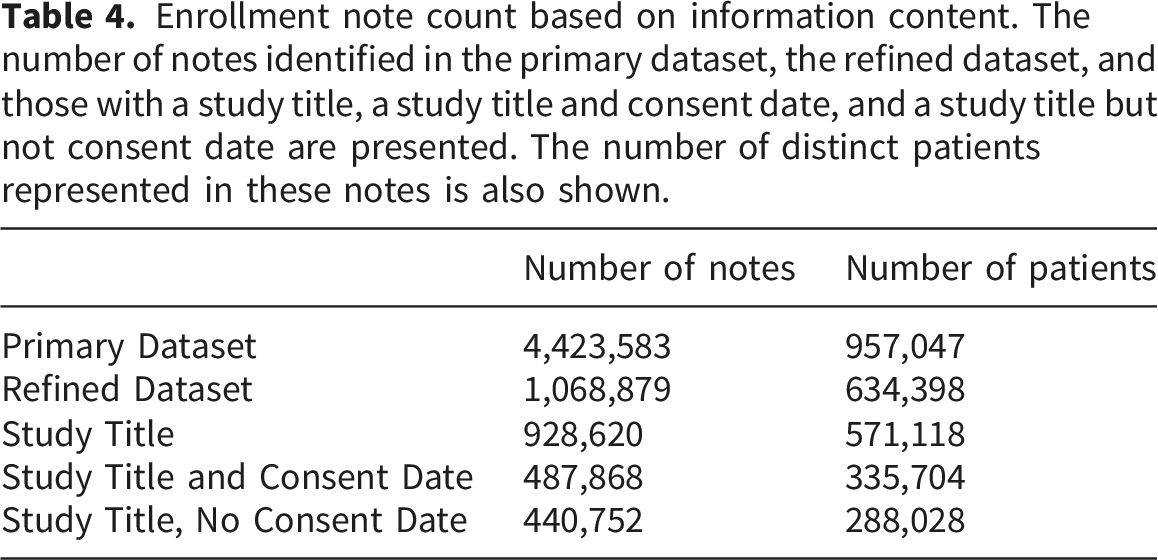
Among the 335,704 patients with both a Study Title and Consent Date, the vast majority (271,842; 81%) had only one record of enrollment, 42,140 (13%) had a record of enrollment in two studies, and 21,722 (6%) had a record of enrollment in three or more studies. For patients with multiple enrollments, there were a median of 796 days (mean, 1226 days) between enrollments.
Discussion
In this study, we developed and evaluated an automated and accurate approach to extract clinical study enrollment information from unstructured clinical notes using NLP methods. We chose a rule-based approach over other approaches such as large language models because a rule-based approach is more interpretable and was feasible to deploy in our environment. We found that rule-based NLP methods performed very well in identification of study title and consent dates. In fact, even most of the errors identified through our pre-designed evaluation measures actually still captured useful information such as the second rather than first mention of a study title or capture of an abbreviation of a title rather than the full title. Using data from a single center, we also determined that the NLP captured known trial enrollments across a variety of trial types, sponsors, and enrollment periods.
Our work complements prior work on information extraction related to clinical trials. First, a large body of literature investigates using information extraction techniques to determine patient eligibility for clinical trials. This contrasts with our work, which identifies patients already enrolled in past trials rather than determining eligibility for future ones. For example, numerous methods have been developed to parse unstructured trial documentation into structured eligibility rules,10,11 while others use structured eligibility criteria to automatically extract information from the EHR to identify potentially eligible patients.12,13 More recently, machine learning and large language models have been used to combine these steps,14,15 and dedicated user-facing applications like MatchMiner and the VA’s Matching Patients to Accelerate Clinical Trials (MPACT) system have been developed to integrate these processes into easy-to-use prescreening interfaces.16,17 A second major area of work involves extracting information about clinical trial design or characteristics from publications of trial results. For example, methods have been developed for information extraction of study characteristics such as eligibility criteria, sample size, treatments, and outcomes from publications reporting clinical trial results, 18 while other work examines published trials to evaluate empirical barriers to clinical trial enrollment. 19 This line of research differs from our own, as it focuses on extracting summary data from publications, whereas our method extracts individual-level data from the EHR.
LLM approaches have recently shown strong performance on information-extraction tasks, and some of the errors we observed, such as mismatches arising from ambiguous context, might occur less often with LLMs.20,21 However, rule-based methods retain important operational advantages, including minimal computational requirements, easier deployment at national VA scale, fewer dependencies, and full auditability. Future work can compare this system with LLM-based extraction to assess potential accuracy gains and explore hybrid approaches that balance performance with operational constraints.
Limitations of the study include the following. First, we only attempt to extract study enrollment information in EHR notes with a title that includes the keyword “Research”, since this is by far the most common location where such data is captured in the VA’s EHR. Expansion to different note types such as oncology notes could potentially capture additional trial enrollment data from non-standard locations and improve sensitivity but would also likely capture more spurious information such as consent to non-research procedures. Continued standardization efforts in trial enrollment reporting can improve data accuracy. In order to apply our approach in another healthcare system, similar pre-selection of notes to those that are likely to include some trial enrollment information will be an important factor for generalizability of our results, and further refinement of the rules may be necessary to optimize performance.
Conclusion
We developed and evaluated a method to extract research study enrollment history as recorded in unstructured clinical notes. The method exhibited strong performance in a held-out test set and an external validation set. This method establishes a unique data source for studying nationwide, population-level barriers to clinical trial enrollment. Unlike existing resources, it encompasses clinical trial participation across multiple sponsors, including industry and federal entities, and incorporates patient- and facility-level data. This resource enables comprehensive analysis of multi-level barriers to trial enrollment, which can yield insights to guide interventions to boost trial accrual and enhance access.
Footnotes
Ethical considerations
The analyses for this report were conducted as part of a VA project to support clinical operations that was classified as non-research by the VA Boston Healthcare System Research and Development Committee.
Consent to participate
Informed consent was not applicable as the project was classified as non-research by the VA Boston Healthcare System Research and Development Committee.
Author contributions
DCE, NVD, and NRF conceived the study. SG, EL, JL, CY, and DC annotated data. SG, EL, RZ, JL, and JC conducted data analysis. SG, JTW, EL, DRF, RD, DCE, NVD, MTB, and NRF interpreted data. SG, RD, EL, JTW, and NRF drafted the manuscript. All authors critically edited the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the U.S. Veterans Affairs (VA) Cooperative Studies Program (MTB, NVD, NRF) and the VA Boston Medical Informatics Fellowship (EL). The views expressed are those of the authors and do not necessarily reflect the position or policy of the Department of Veterans Affairs nor the United States government.
Declaration of conflicting interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Research funding from Bayer and Merck unrelated to the present work (JL, NRF; research funds to institution).
Data Availability Statement
The United States Department of Veterans Affairs (VA) places legal restrictions on access to veteran’s health care data, which includes both identifying data and sensitive patient information. The analytic data sets used for this study are not permitted to leave the VA firewall without a Data Use Agreement. This limitation is consistent with other studies based on VA data. However, VA data are made freely available to researchers behind the VA firewall with an approved VA study protocol. For more information, please visit ![]() or contact the VA Information Resource Center (VIReC) at
or contact the VA Information Resource Center (VIReC) at
