Abstract
Lifestyle modification, including diet, exercise, and tobacco cessation, is the first-line treatment of many disorders including hypertension, obesity, and diabetes. Lifestyle modification data are not easily retrieved or used in research due to their textual nature. This study addresses this knowledge gap using natural language processing to automatically identify lifestyle modification documentation from electronic health records. Electronic health record notes from hypertension patients were analyzed using an open-source natural language processing tool to retrieve assessment and advice regarding lifestyle modification. These data were classified as lifestyle modification assessment or advice and mapped to a coded standard ontology. Combined lifestyle modification (advice and assessment) recall was 99.27 percent, precision 94.44 percent, and correct classification 88.15 percent. Through extraction and transformation of narrative lifestyle modification data to coded data, this critical information can be used in research, metric development, and quality improvement efforts regarding care delivery for multiple medical conditions that benefit from lifestyle modification.
Keywords
Background and significance
National guidelines have recommended the use of lifestyle modification in the treatment and prevention of prevalent disorders plaguing the US today including hypertension, obesity, coronary artery disease, diabetes, peripheral vascular disease, and cancer.1–9 The top seven US health risks for combined disability and death identified in the Global Burden of Disease Study 2016 were tobacco, dietary risks, high body mass index, alcohol and drug abuse, high blood pressure, high fasting plasma glucose, and high cholesterol.10,11 These risks can be minimized through the implementation of lifestyle modifications (i.e. tobacco cessation, dietary changes, alcohol moderation, illicit drug use abstinence, increased aerobic exercise, and weight loss to achieve a healthy weight). The efficacy of addressing lifestyle in just one counseling session has been shown. 12 For example, behavior change can be elicited by making patients aware of their elevated blood pressure readings after a provider encounter. 13 Despite the effectiveness and importance of lifestyle modification in treating hypertension and many chronic illnesses, lifestyle modification is under-utilized and not easily measured, due to it being buried in narrative text rather than in consistent coded form (e.g. diagnosis or procedure codes).14–21 This limits accurate evaluation of the efficacy of these interventions, tracking of quality care metrics, and reimbursement. Evaluation of lifestyle modification includes both the assessment of lifestyle modification activities as reported by a patient or observed by a provider (e.g. “patient started running and weight is down”) and advice on lifestyle modification activities given by a provider to a patient (e.g. “recommend patient lose 30 pounds”). Systematic identification of lifestyle modification is the first step toward improving its use in clinical practice and establishing it as a quality metric. While some clinical information systems code limited individual behaviors (e.g. smoking history), much of this information continues to be recorded primarily in narrative form.
There is a need for automated methods that can facilitate the extraction and integration of lifestyle behavior factors for use in research. Historically, manual chart review has been used to abstract information from patient records, but this has proven to be a time and labor-intensive process, making large-scale chart abstractions nearly impossible. In order to accomplish this task more efficiently, this study used natural language processing (NLP) software tools and processes that can automatically extract text-based information. NLP tools can process many thousands of notes per hour. 22 This technology makes larger chart abstractions feasible and allows a more comprehensive evaluation of documentation of lifestyle modification.
NLP has been used to successfully extract data from electronic clinical records and applied in many fields for efficient and accurate chart abstraction.23–27 Some studies have explored the automated extraction of information on smoking status as an isolated finding.28–31 One study looked at NLP tool augmentation to extract cardiovascular risk identification. 32 Another study used the MediClass NLP tool for extraction of information on weight management counseling in postpartum visits and showed extraction capabilities similar to human abstractors. 33 A separate study extracted limited lifestyle modification documentation in evaluation of diabetes management. 34 And multiple prior studies of lifestyle modification have used survey data with potential bias and/or generalizability concerns. To our knowledge, this work is the first to evaluate lifestyle modification for hypertension in a large population using automated methods.
The primary objective of this study was to use an existing open-source natural language processing tool, cTAKES, along with rules and regular expressions on existing electronic health records to make previously invisible data on lifestyle modification documentation visible and ready for analysis. We used an existing data set that had been used in prior evaluations of hypertension treatment and clinical inertia. 35 This data set was chosen because it could accomplish two goals: (1) evaluate the feasibility of automatic extraction of lifestyle modification (LM) using an open-source NLP tool allowing use and application to multiple chronic disease evaluations at different institutions, and (2) facilitate extension of prior work in this patient population to improve care of hypertension patients early in their disease process. Methods used in this study are designed so that they can be applied to many other patient populations including those with other chronic diseases such as obesity, coronary artery disease, diabetes, and peripheral vascular disease.
Methods and materials
Institutional and clinical settings: UW Health is the academic health system for the University of Wisconsin–Madison. UW Health adopted the Epic Systems Corporation’s electronic health record in 2004 and has created a Health Information Management Center, which is devoted to the integrity of system-wide electronic health record data and facilitating its use for improved patient care and health. All relevant full-text documents from patients meeting inclusion criteria were retrieved from the UW Health electronic health record. An overview diagram of this study’s steps is provided in Figure 1 with more details of the steps and methods following.
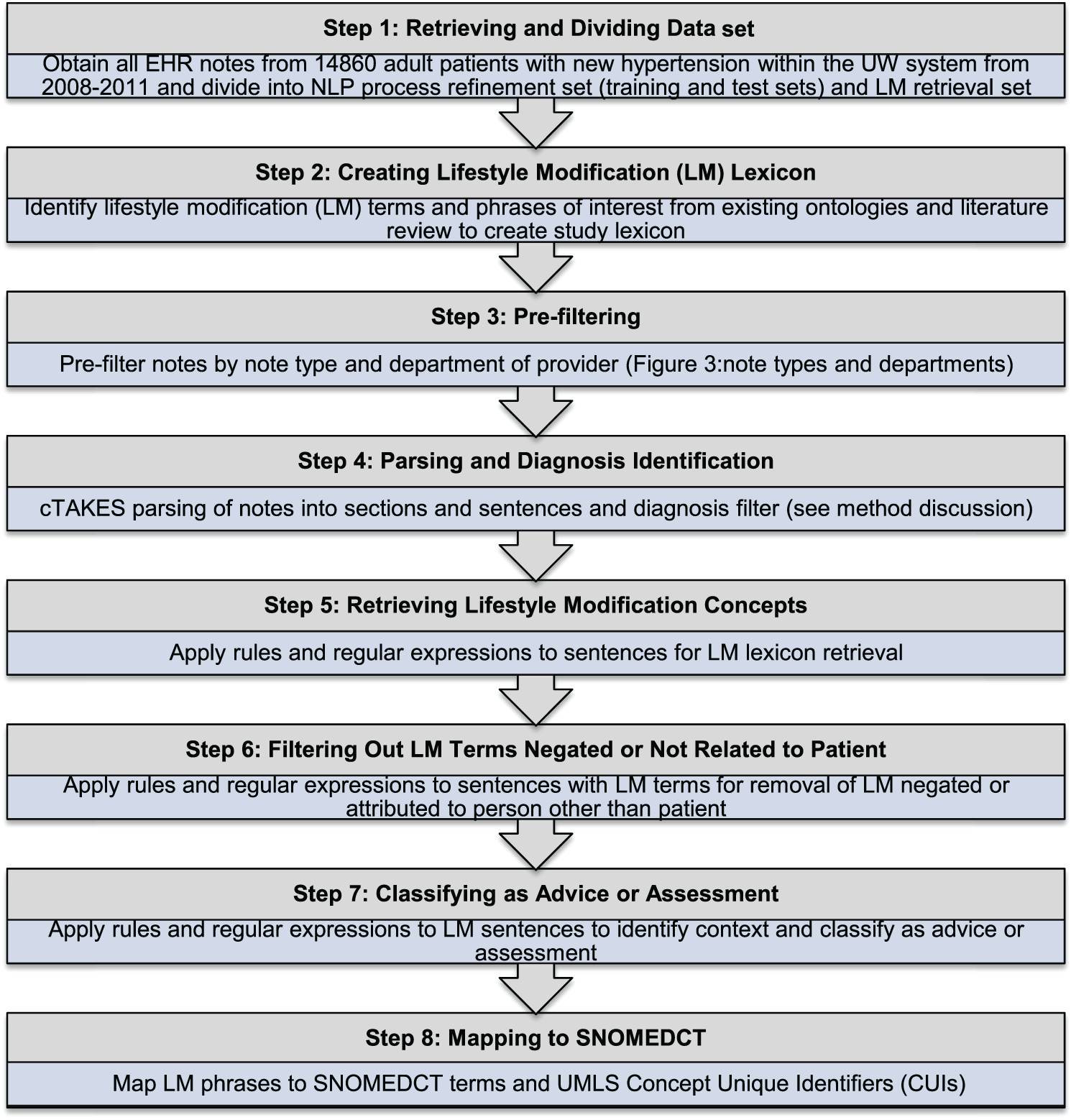
Overview of study steps.
Step 1: retrieving and dividing data set
Study inclusion and exclusion criteria are detailed in Table 1. 35
Inclusion and exclusion criteria.
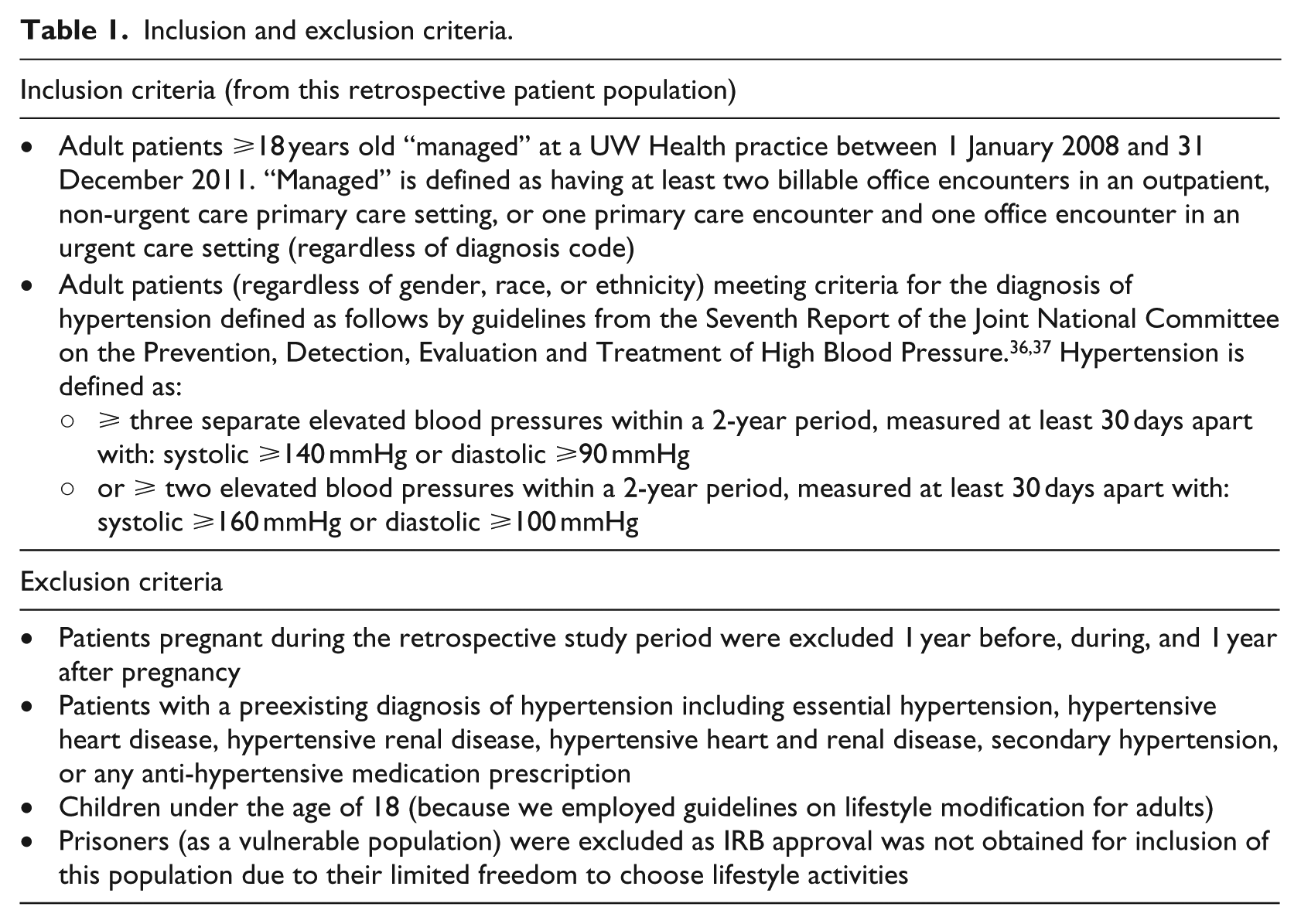

The hypertension diagnosis criteria were based on JNC 7 criteria, reflecting the guidelines available during the time of the hypertension data set creation. However, since this analysis focuses on lifestyle modifications, more recent guidelines including, JNC 836–39 and the 2017 American Heart Association’s guidelines reflect similar lifestyle modification recommendations. LM concepts reflect all three sources. Blood pressure measurements were extracted from discrete fields within the EHR. Preexisting conditions were identified using ICD9 codes. The 14,860 patient data set was divided into a 500 patient NLP tool refinement set and a 14,360 patient lifestyle modification retrieval set (Figure 2).
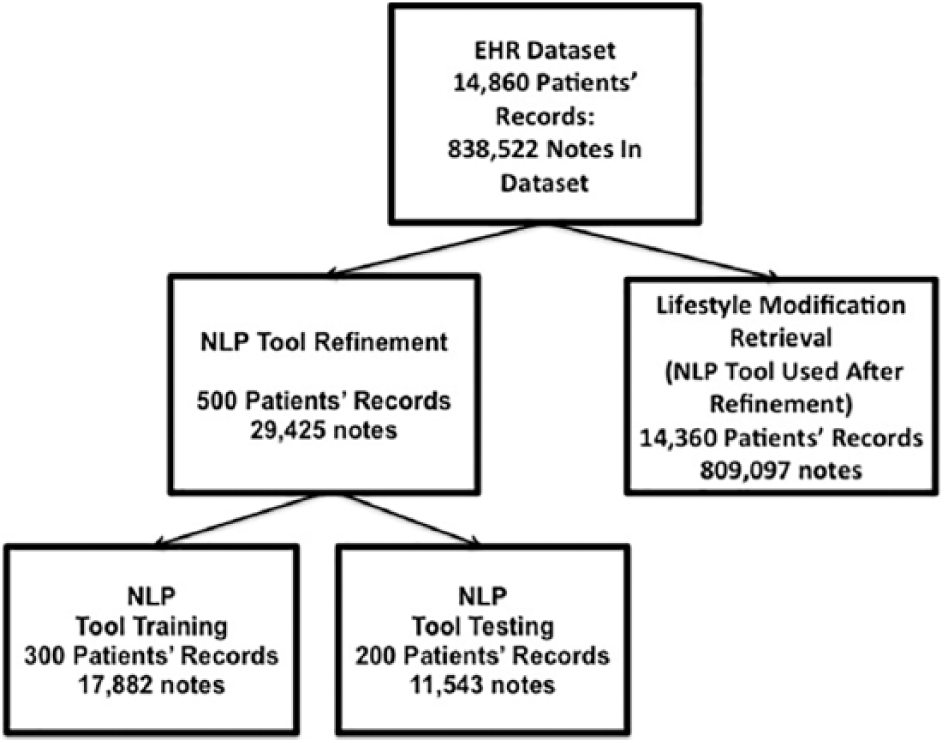
Data set divisions with initial note counts.
The subset of patient notes for NLP tool refinement was randomly selected using Python’s random module: “random sample.”40,41 This subset was composed of notes from throughout the outpatient clinical encounter including nursing notes, provider notes, patient instructions, nutrition consultation, and exercise consultation. The University of Wisconsin IRB approved this study. Baseline characteristics of the study population are listed in Table 2 with the 500 patient subset characteristics listed beside the 14,860 entire data set characteristics.
Study subjects’ baseline characteristics.
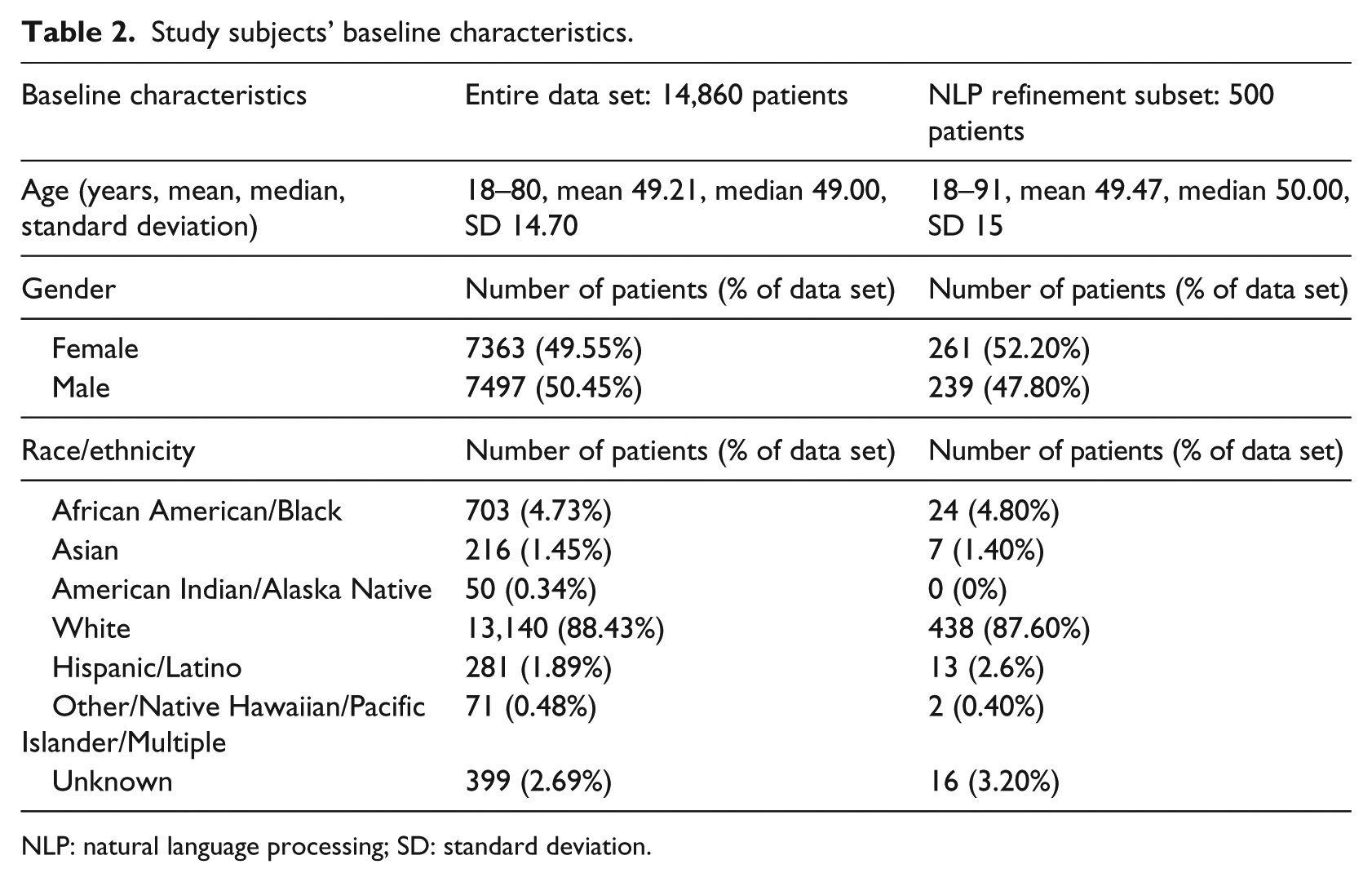
NLP: natural language processing; SD: standard deviation.
Step 2: creating lifestyle modification (LM) lexicon
An empirical method was used to create and iteratively enrich and refine the lifestyle modification terminology using four approaches: (1) literature review to identify related terms, including terms and concepts discussed in JNC 7 and 8; (2) existing ontology review (terms and their interconnections) to identify relevant terms including acronyms and abbreviations such as those in the SNOMED CT ontology, the National Center for Biomedical Ontology and the Consumer Health Vocabulary 42 ; (3) domain expert collaboration to identify words, acronyms, abbreviations, and phrases relevant to hypertension and lifestyle modification; and (4) electronic health record note training. Identified terms and phrases from this fourfold process generated the initial list of relevant terms and phrases for extraction (Table 3).
Example of terms and phrases in lifestyle modification lexicon.
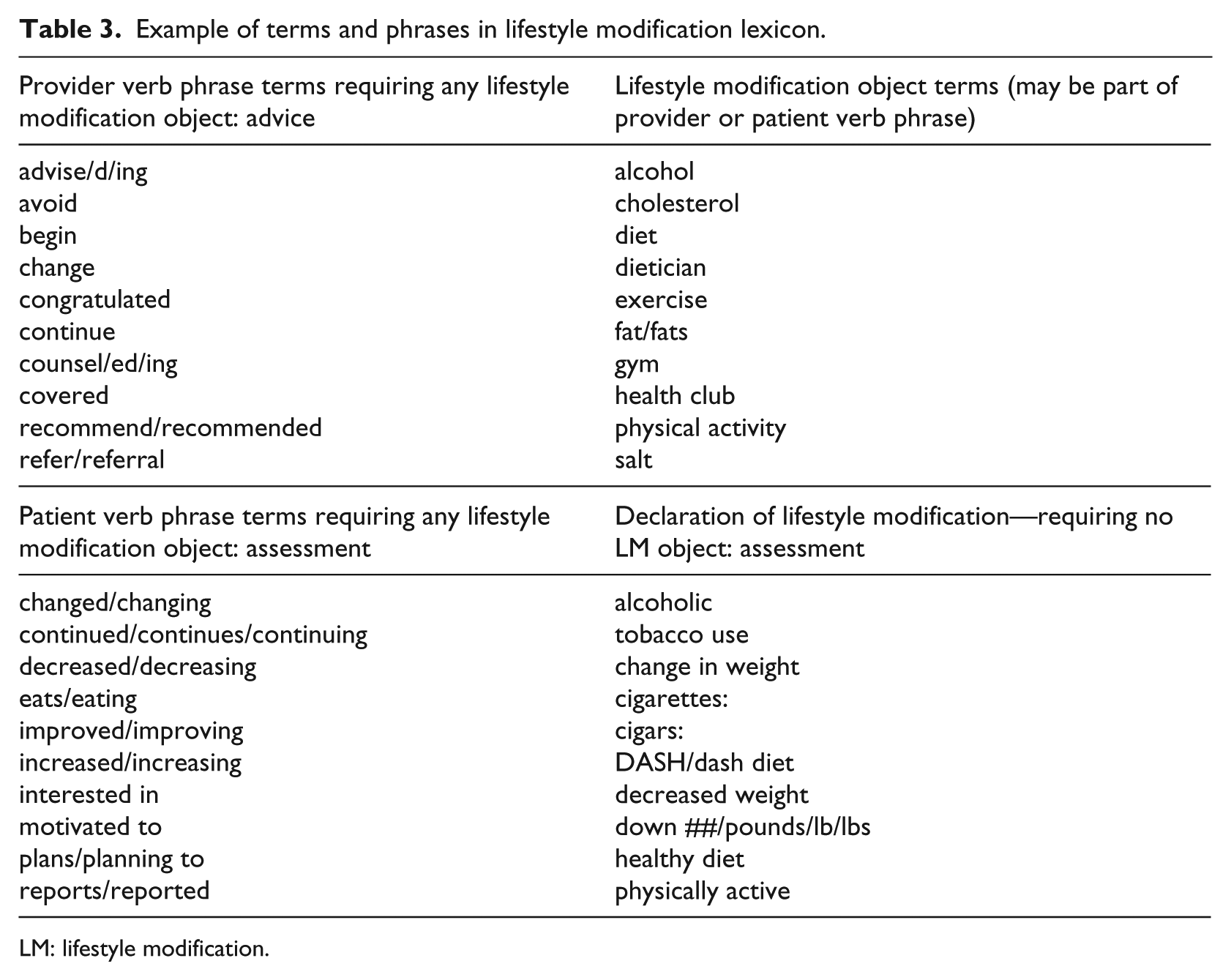
LM: lifestyle modification.
Our goal was to iteratively extend the NLP tool to have additional capabilities to handle discourse. Terms and phrases were mapped to codes in the Unified Medical Language System (UMLS) and semantic types based on the UMLS semantic net. Details available as Supplemental Appendices A and B.
Step 3: pre-filtering to identify notes of interest
The total data set was composed of 14,860 patients’ EHR notes with an average of 56 notes per patient. Each note was composed of multiple sentences, some with multiple concepts of interest (lifestyle modification, family history). Many notes were not relevant to lifestyle modification, so pre-filtering for note types and departments likely to have documented lifestyle modification was performed (Figure 3 shows LM-relevant note types and departments agreed upon by three study physicians).
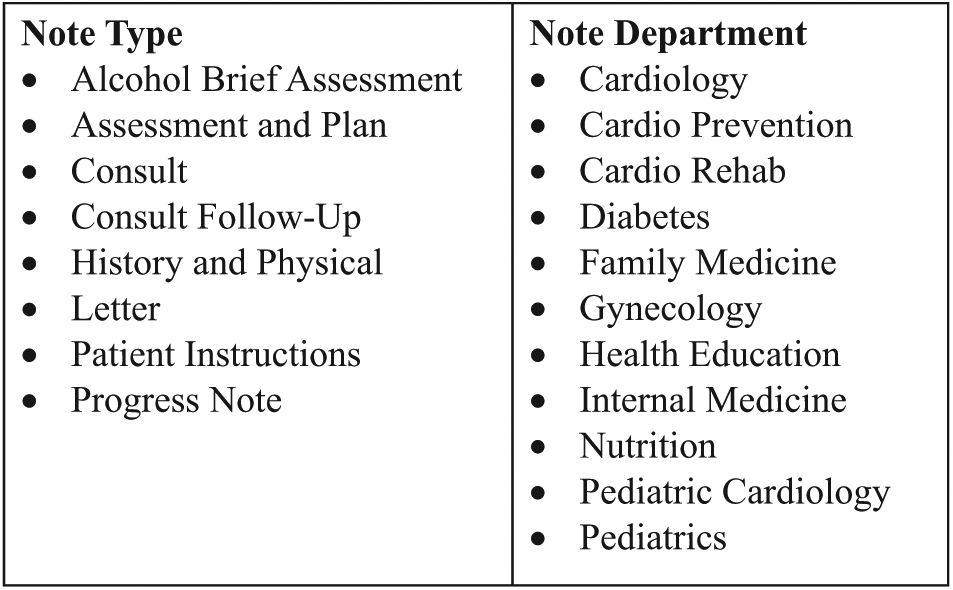
Note types and departments evaluated.
We included pediatric departments because the National Institutes of Health (NIH) defines children as those persons 18–21 years old and some patients in this age range may continue care in pediatric clinics. Gynecology was also included as a primary care clinic because many women see gynecologists as their primary care provider.
Step 4: parsing and diagnosis identification
After pre-filtering, 14,331 notes were in the NLP tool refinement set and 403,018 notes were in the LM retrieval set. These notes were processed using the Clinical Text Analysis and Knowledge Extraction System (cTAKES). This is an open-source NLP pipeline that processes clinical notes and identifies types of clinical named entities—drugs, diseases/disorders, signs/symptoms, anatomical sites and procedures. 43 Each named entity has attributes for the text span, the ontology mapping code, context (e.g. family history of, current, unrelated to patient), and negated/not negated. Figure 4 shows components of a typical NLP pipeline (with study enhancements to improve relevant term and phrase identification depicted by the down-arrow boxes). The typical process starts with detection of each section of the text and then each sentence, followed by the identification of sentence tokens (e.g. words, dates, numbers) in the sentence. A part-of-speech is assigned to tokens (e.g. noun, preposition, noun-phrase). The Named Entity Recognition component implements a dictionary lookup, so that each entity is mapped to a concept from the tool dictionary. Post-coordination then combines multiple concepts into a single one (e.g. DASH + diet).
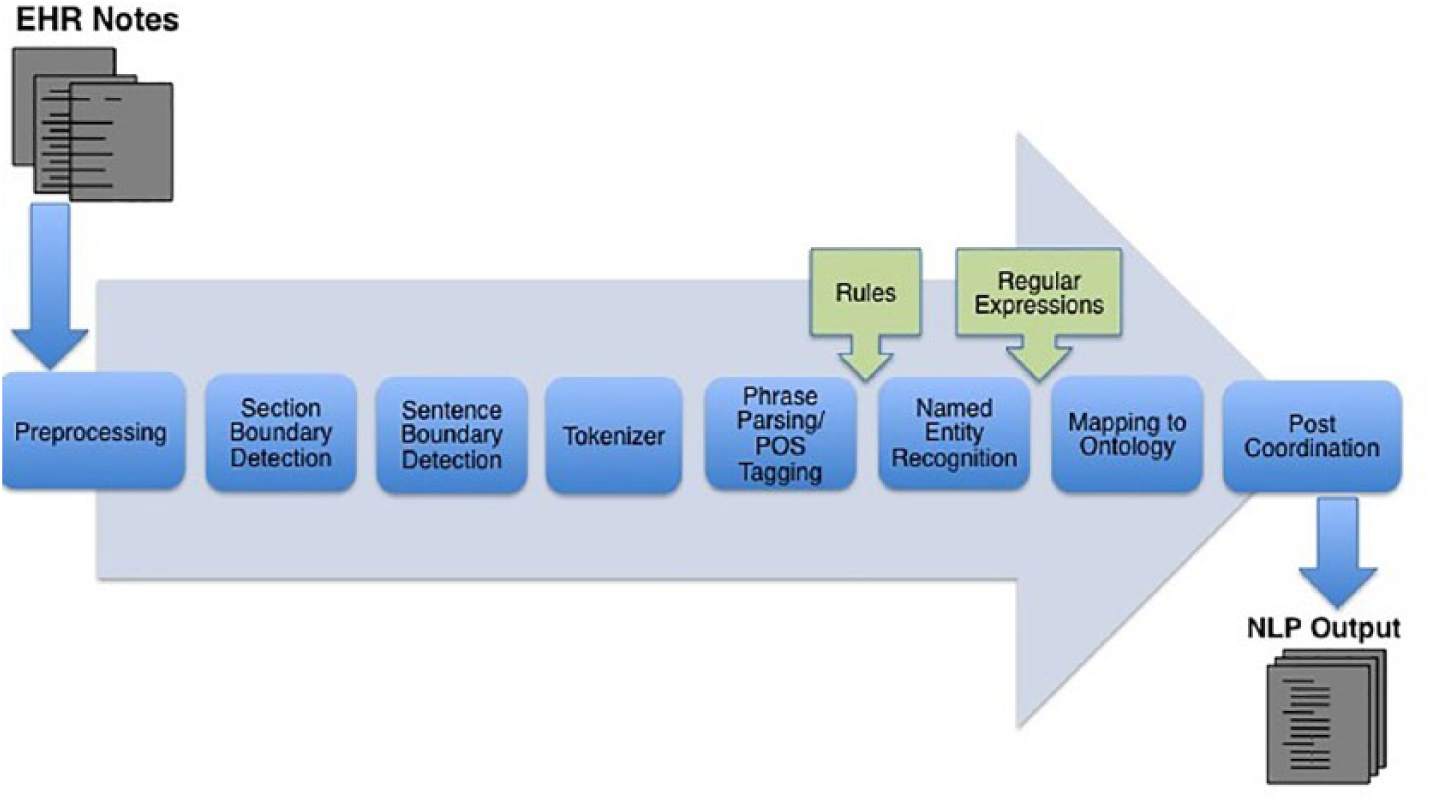
Augmented cTAKES processing steps.
Notes were further processed using Python code to retrieve notes with diagnoses likely to have LM (e.g. obesity, diabetes—details of diagnosis filter terms available by request as Supplemental Appendix C).
Steps 5-8: retrieving LM, filtering out notes with negated or LM terms not related to patient, and mapping
This study initially tried to use cTAKES as the sole method to identify relevant LM, but this approach resulted in poor precision (67.15%) and poor recall (62.48%) because:
Searching for semantic types of interest (TUIs) was too broad and extracted many irrelevant items (e.g. “dietary history” = TUI T033 = “Finding.” Many irrelevant terms are “Findings” such as “sweaty palms” which was pulled as a relevant retrieval when searching based on TUI alone).
Using concept unique identifiers (CUIs) was too restrictive due to identifying only a specific term and inability to identify phrases with LM (e.g. “exercise” = CUI C0015259. If searching on this CUI, relevant terms such as “swimming” with CUI C0039003 were missed).
Using cTAKES’ retrieval and dictionary lookup effectively identified nouns, but missed verbs and verb phrases of LM documentation (e.g. retrieved “diet,” missed “counseled on diet changes”).
Therefore, this study used a combined approach to processing the notes. cTAKES was used for parsing the notes into sections and sentences, but instead of using the parsed parts of speech identified by cTAKES for ontology mapping, Python code with regular expressions and rules was used to process each sentence to identify lifestyle modification terms and phrases from the created lexicon. This enhanced the natural language processing workflow to facilitate extraction of data related to lifestyle modification, including social and behavioral factors that historically have been difficult to extract and verbs. These data were mapped to corresponding terms within the SNOMED CT ontology using the Unified Medical Language System (UMLS).
For example, a provider recommendation of “cut down on alcohol intake” was retrieved using the study’s enhanced NLP methods resulting in lifestyle modification advice retrieval: “alcohol consumption counseling” in SNOMED CT UMLS CUI = C1531491 and TUI = Health Car Activity T058. Without enhancements to NLP, no match for this phrase was retrieved or mapped by cTAKES to the SNOMED CT ontology.
In order to capture context-sensitive phrases, we used semantic and syntactic rules, as well as regular expressions. Due to many notes containing unusual syntactic expressions impeding retrieval of context-sensitive lifestyle modification, existing note punctuation was modified. Specifically, when a sentence about LM contained a colon (:), the NLP processor read this as a hard stop and classified concepts after it as belonging to the next sentence. The colon mark was replaced with a comma to allow capture of concepts after the colon mark as contextually related to the first phrase. This improved retrieval of the entire lifestyle modification context. For example, prior to modification, cTAKES identified the negated history of “Patient smoking: no.” as a positive history of patient tobacco use: “Patient smoking. No.” After modification, the intended meaning was retrieved: “patient history negative for smoking.” Tokenization per cTAKES was not modified, but cTAKES only identifies nouns and noun phrases and this study also needed to identify verbs and verb phrases. Therefore, the algorithm searched the entire sentence using regular expressions for LM object terms and phrases (Figure 5 and details of LM lexicon available as Supplemental Appendix A).

Example of regular expression to retrieve exercise.
Negation identification was performed using negated regular expressions already employed to extract LM terms and phrases. This approach allowed for more efficient and thorough negation identification of verbs and verb phrases that were not identified by cTAKES’ negation module. Attribution of a concept, as pertaining to the patient or other person for LM terms/phrases and diagnoses, was expanded to include, in addition to mother and father, grandparents, grandmother, grandfather, roommate, spouse, husband, wife, son, daughter, partner, roommate, parent, friend, significant other, coworker, brother, sister, aunt, and uncle. Family history retrieval using name of relation (e.g. mom, sister) within rules and regular expressions, along with mapping schemes to SNOMED CT, was created specifically for this project. Mapping of terms to the SNOMED CT hierarchy was restricted to parent and child concepts, and their CUIs, to reduce granularity and facilitate effective use of this coded data in future machine learning projects that required less dimensionality. For example, “brother with non-insulin-dependent diabetes mellitus” was mapped to “FH: diabetes mellitus,” a child of the parent concept “FH: metabolic disorder,” and not mapped to the grandchild concept “FH: diabetes mellitus type 2.” This allowed all diabetics to be binned under one code. Details of family history mapping available as Supplemental Appendix D.
This study also classified lifestyle modification words and phrases as “assessment” and “advice” as was done in a prior analysis of videotaped encounters involving lifestyle modification. 44 Classification as assessment or advice was based on logic rules that included key words. This allowed for evaluation of LM documentation as either provider documentation of LM activities that the patient reported (e.g. patient reported exercising = assessment) or provider documentation of advice that she or he offered to the patient (e.g. recommended exercise = advice). Matching of LM terms to SNOMED CT was further refined with specific LM terms and phrases grouped under overarching concepts such as exercise education (advice) or exercise history (assessment). Details of advice/assessment phrases available as Supplemental Appendix B. This process continued until performance was close to reproducing the manually coded set of terms and phrases, and additional modifications to the system minimally altered NLP tool performance. The training/testing tasks and iterative process details are shown in Table 4.
Annotation task iteration details.
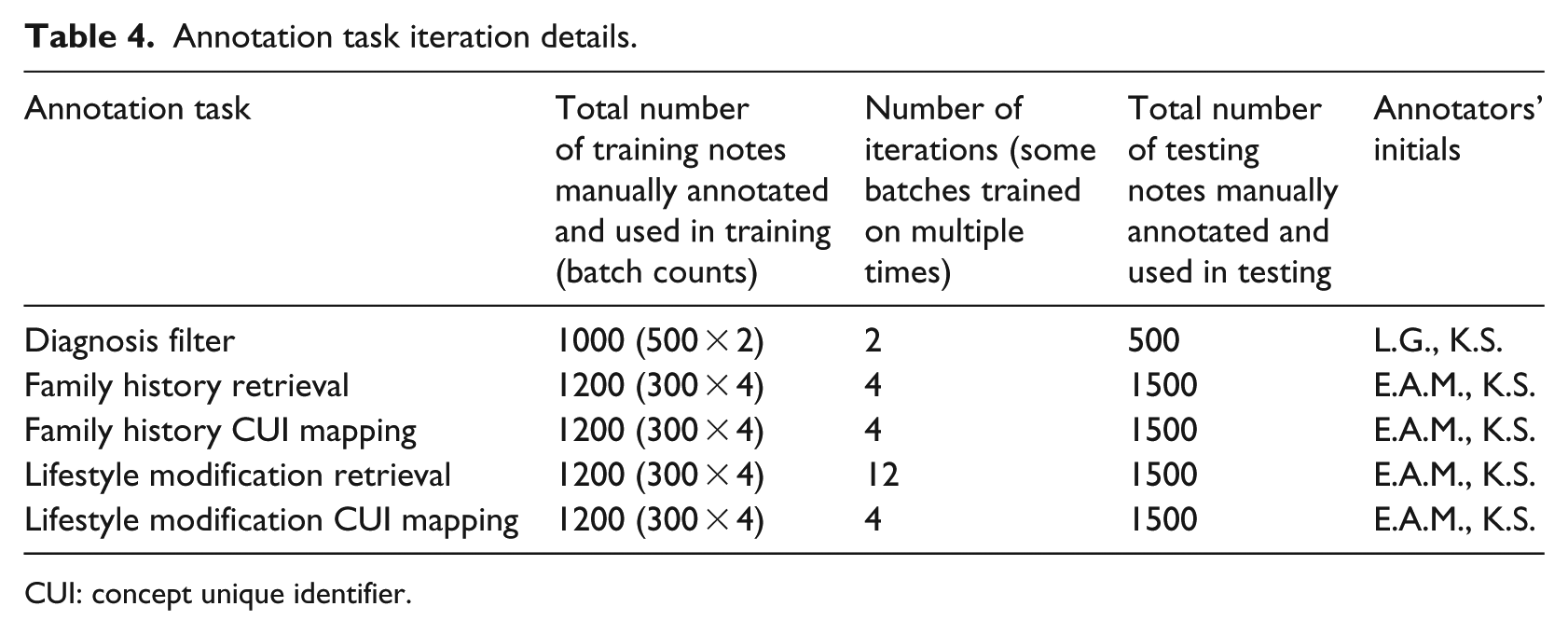
CUI: concept unique identifier.
Overfitting was discussed by researchers (K.S., E.A.M., Y.S.) after each iteration to determine whether the retrieved terms/phrases from that iteration reflected relevant and generalizable results or included concepts or terms reflective only of this patient sample. All three members reached consensus in determining whether changes made to the tool were to be kept or represented overfitting and should not be retained in the tool development process. This iterative process resulted in retrieval of textual LM documentation and transformation into coded data ready for future statistical and machine learning analyses (Figure 6).
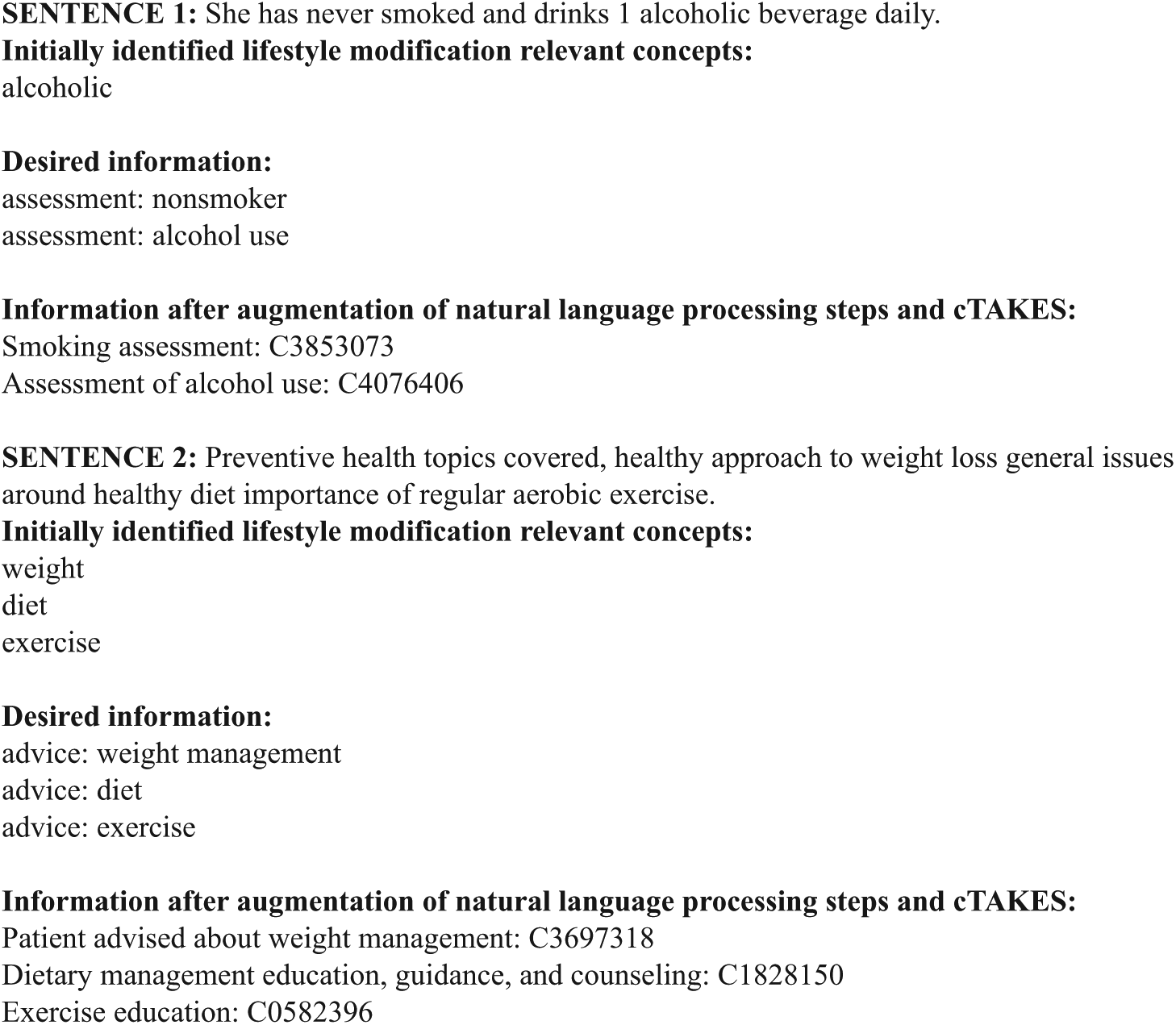

Example of LM retrieval and transformation to coded data.
Validation of the extended tool
The augmented NLP process was applied to the testing data set as the manually annotated gold standard. Data set divisions and uses are detailed in Figure 7 and Table 4 describes researchers and numbers of notes employed in each test set evaluation.
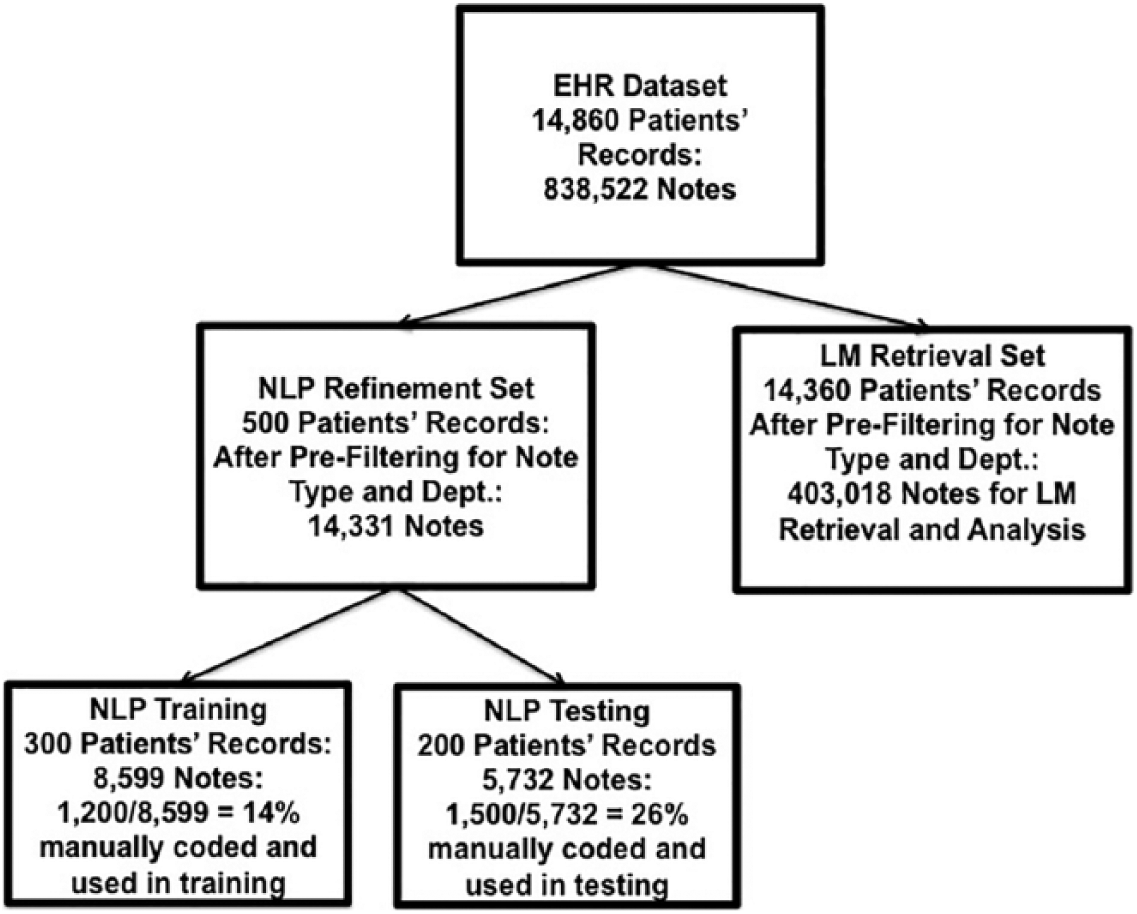
After pre-filtering for note type and department—data divisions and uses.
Two sets of domain experts were employed to establish validity in each test task. One team worked on validation of the diagnosis filter and consisted of one clinical data analyst trained in linguistics and healthcare terminologies and one physician (L.G., K.S.). The other team consisted of two physicians who worked on validation of the LM retrieval and CUI mapping, and FH retrieval and CUI mapping, and LM classification as assessment or advice (E.A.M., K.S.). Each validation task used the randomly selected test set of patients’ records that each team’s members manually abstracted independently. From these manual abstractions, inter-annotator agreements were calculated using percent agreement and kappa coefficient calculations. Testing results of the enhanced NLP process are reported using precision, recall, and F measurements in Table 5. For this study, precision is defined as the number of terms correctly extracted divided by the sum of the number of terms correctly extracted and the number of terms incorrectly extracted. Formally this is: Precision = TP/(TP + FP) where TP denotes true positive (a term that was correctly extracted) and FP denotes false positive (a term that was extracted as relevant but should not have been). Recall is the number of terms correctly extracted divided by the sum of the number of terms correctly extracted and the number of terms that should have been extracted but were missed. Formally this is: Recall = TP/TP + FN where FN denotes false negative (a term that was not extracted but should have been). The F-measure is the harmonic mean between precision and recall with F = 2[(precision × recall)/(precision + recall)]. In addition to lifestyle modification retrieval and CUI mapping, our process was able to identify attribution and mapping to family history CUIs, and LM documentation type (i.e. assessment or advice). Success of classification of LM documentation as advice or assessment was evaluated using percent of LM terms and phrases correctly. Unit of measure for classification as advice or assessment was each LM concept (e.g. if a sentence contained two LM concepts with one being “advised weight loss” and classified correctly identified and the other being “patient reports gaining weight” and not correctly classified, then one concept would be counted as correct and one counted as incorrect classification identified).
NLP augmentation validation performed on 1500 manually annotated notes from the 200 patient testing subset.
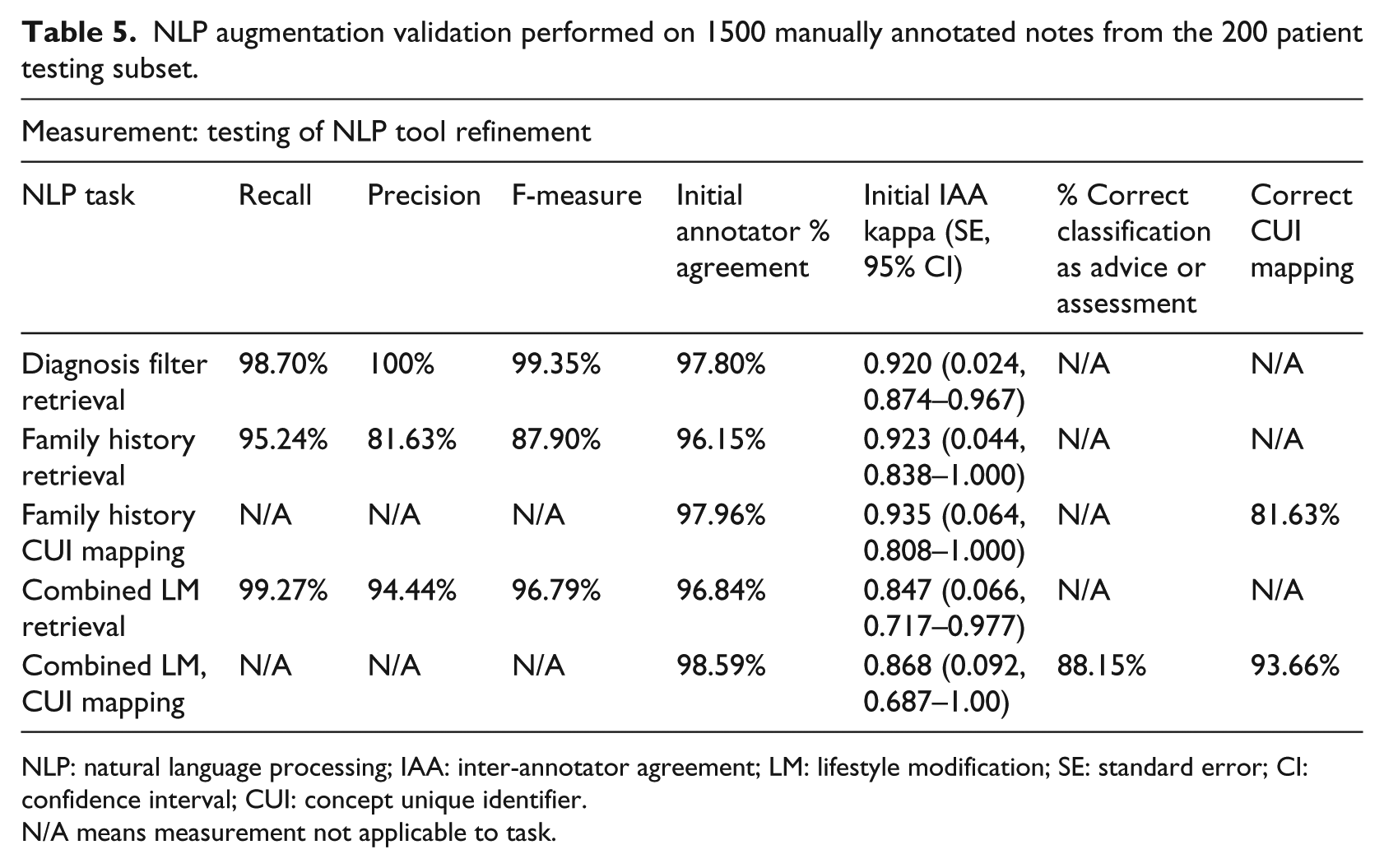
NLP: natural language processing; IAA: inter-annotator agreement; LM: lifestyle modification; SE: standard error; CI: confidence interval; CUI: concept unique identifier.
N/A means measurement not applicable to task.
Results
Results from testing the NLP tool refinement process for combined lifestyle modification retrieval were excellent with 99.27 percent recall and 94.44 percent precision and an F-measure of 96.79 percent. CUI mapping for lifestyle modification was also very good with phrases correctly classified as advice or assessment 88.15 percent. These results along with the excellent diagnosis filter testing results and family history retrieval and mapping results are shown in detail in Table 5. Inter-annotator agreement was also excellent with initial agreement percentages (range 96.15%–98.59%) and kappa scores (range 0.847–0.935) (Table 5). In the rare occurrence of reviewers extracting a different number of terms, the consensus-agreed-upon terms were used as the gold standard to calculate inter-annotator agreement. After consensus discussions, 100 percent agreement was reached between annotators for each task.
Each specific type of lifestyle modification retrieval recall and precision is detailed in Table 6. Overall retrieval testing results for each type of lifestyle modification were very good.
Recall and precision of specific types of LM concepts.
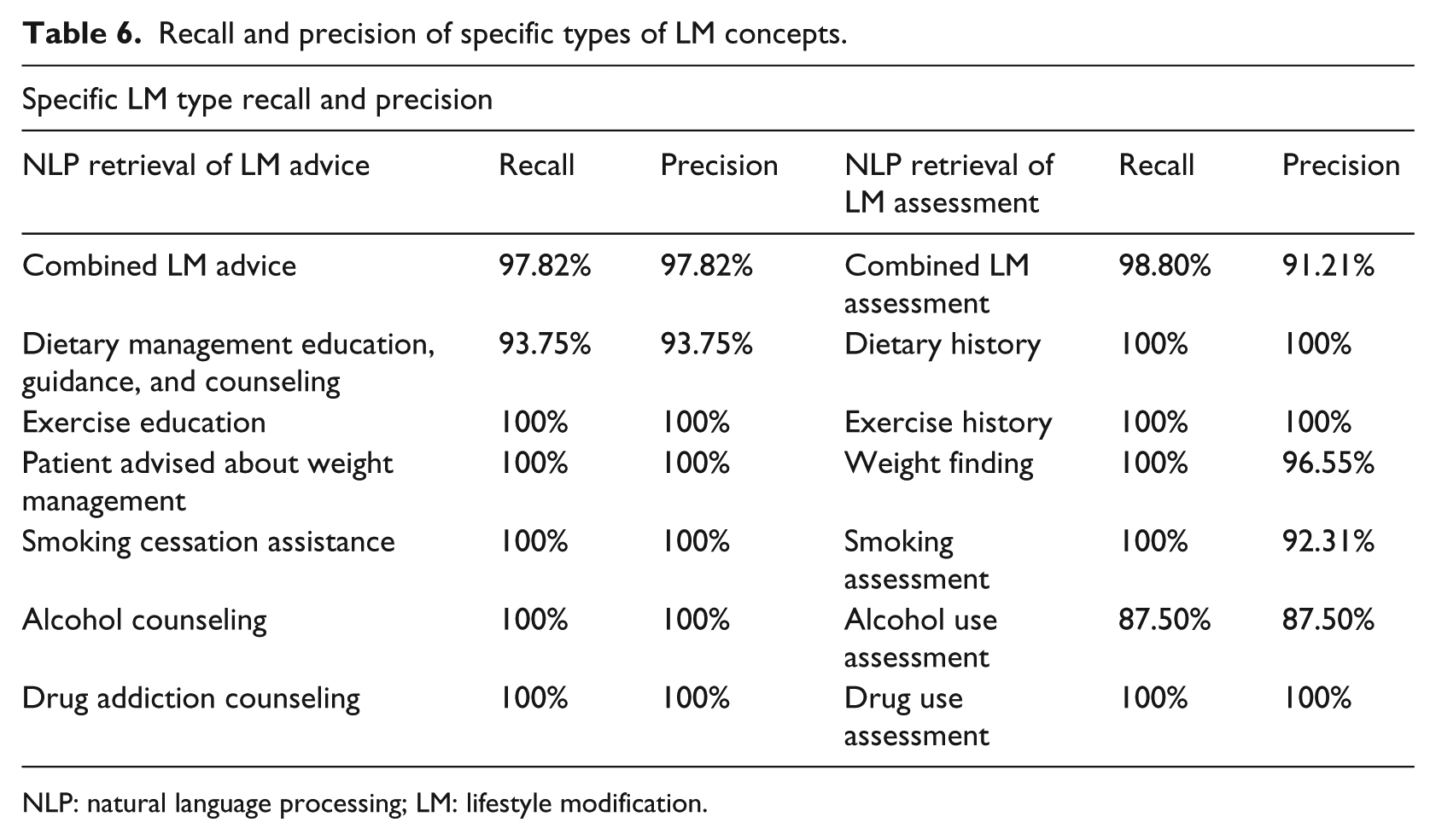
NLP: natural language processing; LM: lifestyle modification.
Of the 14,360 patients in the LM retrieval set, 11,252 patients (78.36%) had notes documenting lifestyle modification. Each patient had an average of 56 notes in the initial data set. From the total 809,097 notes in the LM retrieval set (NLP refinement set removed), after filters and processes described in Figure 1 were applied, 47,838 notes had at least one documentation of lifestyle modification. Many notes contained more than one documented LM activity. Specific lifestyle modification activities and their CUIs are detailed in Table 7 with patient and note counts for each type of LM.
Lifestyle modifications, concept unique identifiers (UMLS CUIs), and retrieval counts.
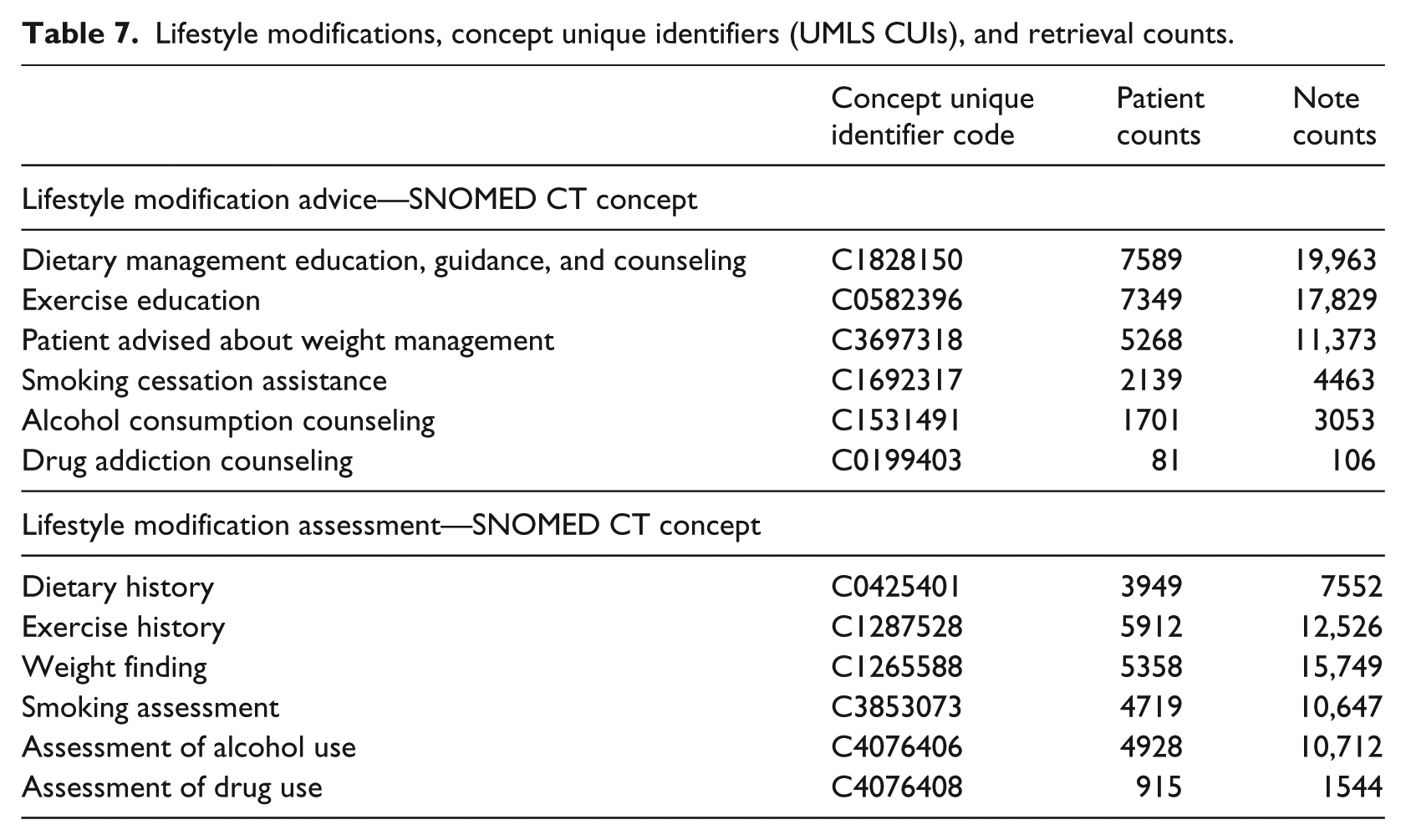
For advice, diet and exercise advice were most frequently documented. For assessment, exercise and weight assessments were most frequently documented. Counts for advice on exercise were greater than counts for assessment, suggesting that providers are offering advice on exercise regardless of patients’ current exercise status. Counts for drug abuse counseling and assessment were low. This was a hypertension population and drug use assessment and counseling is not a recommended lifestyle modification intervention for hypertension. Drug abuse assessment and counseling were included in the design of this NLP tool enhancement for comprehensiveness and facilitation of future training and testing in populations with higher numbers of drug abuse assessment and counseling.
Outcome
Our modified process using an augmented open-source natural language processing tool was successful in identifying lifestyle modification documentation in electronic health records of hypertension patients at an academic medical center. Given the importance of lifestyle modification interventions for multiple medical issues, this is an important and innovative step in care transparency for future comparative effectiveness studies, outcome analyses, and efforts in care improvement. To date, most studies evaluating lifestyle modification as a medical treatment rely on surveys and self-reports which are inherently vulnerable to reporting and recall bias.45,46 With increasing emphasis placed on the need for LM to treat multiple chronic diseases, our study offers more objective and comprehensive measurement of LM care delivery via EHR analysis than previous studies. It is encouraging that 78 percent of patients in this study had LM addressed at least once during the study period, but there is room for improvement in how often LM is addressed with each patient. The percentage of notes containing LM was low at 5.9 percent, suggesting that although providers are addressing LM with many patients, providers are not repeatedly reviewing LM with patients despite the need for behavior modification for treatment of many diseases. As more efforts are made to improve lifestyle modification interventions, this study has shown that LM documentation can be automatically extracted from EHRs, thus offering increased identification of actual use of lifestyle modification in the care of hypertension and multiple chronic disorders in the future (Figure 8). 47
Chronic conditions that can be evaluated using these methods.
Challenges and limitations
Limitations of this study included the secondary use of EHR data that can present data quality issues (e.g. missing data, reporting bias, recording bias). Caveats and challenges in secondary use of EHR data have been well documented.48–51 One model of data quality defined completeness as, “the extent to which data are of sufficient breadth, depth, and scope for the task at hand.” 52 This study’s ability to accurately retrieve and classify LM from this EHR data set confirms that this data set is sufficient for this task. A second challenge was this study’s use of an open-source NLP tool with requirements for customization. Initial lifestyle modification data retrieval attempts using only cTAKES were unsuccessful with poor recall and precision but, after iterative development, the final testing results were successful in LM retrieval and classification as assessment or advice. This study overcame a key challenge in NLP evaluation of care delivery: the inability to retrieve verbs and verb phrases. cTAKES alone could not accurately identify verbs, verb phrases, or verb-tense-specific classification of a term as advice or assessment. For example, “patient started walking” is assessment and documentation of a patient reported LM. This is different from “recommended patient start walking 30 minutes per day” which is provider LM advice. cTAKES alone could not retrieve or distinguish these two different concepts, but using a combination of regular expressions, rules, and key words, these critical concepts centered on verb phrases were accurately retrieved and classified. However, inherent limitations in generalizability and scalability are present with this current approach.
Future development
Future research will attempt to improve and augment cTAKES and its dictionary to extract and map verb phrases directly. This approach will minimize use of rules and regular expressions and make this work more generalizable and scalable. We also plan to extend lifestyle modification mapping to ICD10 and other dictionaries of interest. This work will be made available to researchers in related medical areas that use LM as a treatment and could be used to support evaluation of current and future initiatives such as “Exercise Is Medicine” and “Healthy People 2030.”53–55 Another area for future development could be the assessment of quality of counseling for lifestyle modification with a more granular extraction of lifestyle modification concepts and counseling details to better understand best practices. 56
Conclusion
This study successfully extracted lifestyle modification documentation from EHR notes. Its methods and future planned expansion of these methods could be used in studies involving multiple chronic medical conditions. This is an important step in better understanding and quantifying the use of lifestyle modification as a prevention and treatment modality for many disorders. This information can be used in future outcome and comparative effectiveness research and inform metric development for lifestyle modification documentation and counseling. The automatic identification and mapping of terms, especially verbs, related to care delivery is a major innovation that can allow further evaluation and improvement in care delivery models and treatment approaches to multiple chronic illnesses.
Supplemental Material
Shoenbill_HTN-NLP-HIJ-AppendixA – Supplemental material for Natural language processing of lifestyle modification documentation
Supplemental material, Shoenbill_HTN-NLP-HIJ-AppendixA for Natural language processing of lifestyle modification documentation by Kimberly Shoenbill, Yiqiang Song, Lisa Gress, Heather Johnson, Maureen Smith and Eneida A Mendonca in Health Informatics Journal
Supplemental Material
Shoenbill_HTN-NLP-HIJ-AppendixB – Supplemental material for Natural language processing of lifestyle modification documentation
Supplemental material, Shoenbill_HTN-NLP-HIJ-AppendixB for Natural language processing of lifestyle modification documentation by Kimberly Shoenbill, Yiqiang Song, Lisa Gress, Heather Johnson, Maureen Smith and Eneida A Mendonca in Health Informatics Journal
Supplemental Material
Shoenbill_HTN-NLP-HIJ-AppendixC – Supplemental material for Natural language processing of lifestyle modification documentation
Supplemental material, Shoenbill_HTN-NLP-HIJ-AppendixC for Natural language processing of lifestyle modification documentation by Kimberly Shoenbill, Yiqiang Song, Lisa Gress, Heather Johnson, Maureen Smith and Eneida A Mendonca in Health Informatics Journal
Supplemental Material
Shoenbill_HTN-NLP-HIJ-AppendixD – Supplemental material for Natural language processing of lifestyle modification documentation
Supplemental material, Shoenbill_HTN-NLP-HIJ-AppendixD for Natural language processing of lifestyle modification documentation by Kimberly Shoenbill, Yiqiang Song, Lisa Gress, Heather Johnson, Maureen Smith and Eneida A Mendonca in Health Informatics Journal
Footnotes
Acknowledgements
The authors are grateful to the Health Innovation Program at the University of Wisconsin–Madison for assistance with data acquisition. K.S. and Y.S. contributed equally to this work and should be listed as first authors.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Funding for this project for investigators Shoenbill, Song, and Mendonca provided by: The Clinical and Translational Science Award (CTSA) program, through the NIH National Center for Advancing Translational Sciences (NCATS), grant UL1TR002373 (A. Brazier, PI) UW-Madison Office of the Vice Chancellor for Research and Graduate Education Research-2014 Fall Research Competition Award. “Predictors of Lifestyle Modification in Hypertension: A Computational Analysis”. (E. Mendonca, PI) Additional funding for investigator Shoenbill provided by: University of North Carolina Chapel Hill: start up funding. (K. Shoenbill, PI) NLM Grant 5T15LM007359 to the Computation and Informatics in Biology and Medicine Training Program. (M. Craven, PI).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
