Abstract
Background
Extraction of medical terms and their corresponding values from semi-structured and unstructured texts of medical reports can be a time-consuming and error-prone process. Methods of natural language processing (NLP) can help define an extraction pipeline for accomplishing a structured format transformation strategy.
Objectives
In this paper, we build an NLP pipeline to extract values of the classification of malignant tumors (TNM) from unstructured and semi-structured pathology reports and import them further to a structured data source for a clinical study. Our research interest is not focused on standard performance metrics like precision, recall, and F-measure on the test and validation data. We discuss how with the help of software programming techniques the readability of rule-based (RB) information extraction (IE) pipelines can be improved, and therefore minimize the time to correct or update the rules, and efficiently import them to another programming language.
Methods
The extract rules were manually programmed with training data of TNM classification and tested in two separate pipelines based on design specifications from domain experts and data curators. Firstly we implemented each rule directly in one line for each extraction item. Secondly, we reprogrammed them in a readable fashion through decomposition and intention-revealing names for the variable declaration. To measure the impact of both methods we measure the time for the fine-tuning and programming of the extractions through test data of semi-structured and unstructured texts.
Results
We analyze the benefits of improving through readability of the writing of rules, through parallel programming with regular expressions (REGEX), and the Apache Uima Ruta language (AURL). The time for correcting the readable rules in AURL and REGEX was significantly reduced. Complicated rules in REGEX are decomposed and intention-revealing declarations were reprogrammed in AURL in 5 min.
Conclusion
We discuss the importance of factor readability and how can it be improved when programming RB text IE pipelines. Independent of the features of the programming language and the tools applied, a readable coding strategy can be proven beneficial for future maintenance and offer an interpretable solution for understanding the extraction and for transferring the rules to other domains and NLP pipelines.
Keywords
Introduction
The HiGHmed consortium combines and integrates the competencies of nine university hospitals. Currently, over 20 partners participate in the project, including medical faculties, research institutions, and industry specialists.
HiGHmed’s fundamental purpose is to develop an open platform for the infrastructural and innovative exchange of information, thus allowing a nationwide health information exchange interoperability that enables the development of sophisticated new solutions in precision medicine, medical data analytics, health sharing, and medical education.1,2
For each location, various diversified information systems (IS) exist that store their data in structured, unstructured, and semi-structured data sources, and extract, transform, load (ETL) processes are required for the data integration task. HiGHmeds goal is to simplify and support the ETL and data mapping processes through sophisticated interoperable information exchange approaches.
In the ETL processes, in the transformation task, the extraction of particular terms, and their respective values from Hospital Information Systems (HIS) that contain information from unstructured or semi-unstructured sources, requires the implementation of an NLP method.
Locke et al., define NLP in medicine as: “
In this paper, we apply the NLP method of RB to computationally extract clinical concepts in documents through the implementation of hard-coded rules 4, therefore to study the manual coding with REGEX and AURL. The main scope is to demonstrate developers to regard programming coding techniques for improving readability and not only to evaluate their extraction pipelines according to precision and recall.
TNM staging system
The TNM staging system is an internationally recognized standard that through the definition of three primary codes along with additional modifiers, describes the amount and spread of cancer in a patient’s body. The T defines the size of the tumor or the spreading of cancer in a near tissue. The N describes the spread of cancer to near lymph nodes. The M stands for metastasis and describes the spread of cancer to other parts of the body. 4
The objectives of the TNM code for the tumor classification are: 5
Aid treatment planning
■ Indicate a prognosis. ■ Assist in the evaluation of treatment results. ■ Facilitate the exchange of information between treatment centers. ■ Contribute to continuing investigations of human malignancies. ■ Support cancer control activities, including through cancer registries.
Related work
Methods of NLP provide the means to extract various information from clinical texts and for information exchange in general while their efficiency is proven in the scientific community when parsing unstructured texts.6–9
Current state-of-the-art NLP techniques for text extraction are either RB, machine learning-based (MLB), or deep learning (DL) methods. Even though many attempts are made to create sophisticated hybrid methods that combine RB with MLB or DL, with promising results, still 60% of the studies and solutions refer to RB solutions in which the rules are manually written. 10
In the study from Michael et al.,
11
in which they focused their research on a more practical approach regarding personal opinions and difficulties that developers face when using REGEX, the following critical points were mentioned: ■ The crypticness of the syntax. ■ Difficulty to validate the syntax during the design time. ■ A little cheat sheet is required that outlines what each symbol does. ■ Hard to validate and document.
The concept of readability and maintainability with RΒ IE pipelines are considered important for the life cycle of the rules. 12
Heitlager et al. 13 relate the amount of effort to maintain any software solution with the coding quality of the source code.
Some examples of tools that support the programming of RB extraction pipelines are: ■ Unstructured Information Management Architecture (UIMA) ■ Apache OpenNLP library ■ spaCy library ■ CoreNLP
The UIMA is an architecture platform designed for Java applications to define custom annotators with inbuild custom expressions and embed them through the included analysis engine in external java projects for IE pipelines. 14
The Apache OpenNLP is a Java-based library for ML learning tasks. The library supports the definition of manual annotated terms and training of data for building Named entity recognition (NER) classification models. Li’s et al. definition for NER is, “
The spaCy library for the Python programming language is built for general NLP extraction tasks; like the pre-processing of large text volumes and feature extraction. Also, the inbuild token-based matching with REGEX through spaCy’s RB matcher engine, allows the definition of string matching specific commands. 17
The CoreNLP library is a Java-based collection of various NLP processing tools, that use RB, ML, and DL components. The UIMA analysis engine can be also embedded for RB feature extraction. 18
Objectives
For using MLB methods, the expertise of ML specialists for maintaining and retraining the domain adaption is required at all times together. Also, the lack of enough training data especially in medical domains and particularly in German requires non-MLB approaches.
In our project, we decided to design a framework independent of ML specialists and to apply manual RB extraction approaches for extracting TNM codes in text-based unstructured pathological reports that are manually entered by medical practitioners. Our main research is focused on the two following questions: ■ What is the time to correct wrong rules or include any exceptions since a TNM code can be written in various ways? ■ If we decide to change the programming language, can we implement the rules efficiently?
Research tools
Manual writing of extraction rules with REGEX has been the dominant approach for text extraction of unstructured and semi-structured texts. 19 However, the AURL is an all-in-one solution that includes a workbench for the writing and unit-testing 20 of the extraction scripts and closes the gap between code readability and maintainability. 21
Regular expressions
Briefly, REGEX is a formal language that contains various lookup expressions for string pattern matching. REGEX is applied in various NLP tasks like morphology, text analytics, speech recognition, and IE. REGEX is included in many software tools and programming languages. 22
In the preparation part, usually, a developer reads the specification documents and annotates and identifies the rules needed. In the composition part, the rules are written and tested through for example an external online tool like regex101.com 23 or through a programmed unit testing, in which each extraction rule is required to be exclusively tested separately.
In the implementation part, the REGEX is then encapsulated in a programming language that supports REGEX. Wrongly written scripts, that do not satisfy the extraction results, are re-entering the composing part for correction until the extraction is satisfied.
For our research with REGEX, we used the Python programming language and the spaCy library for programming the extraction pipeline.
Apache UIMA ruta language
The AURL is an imperative RB language that extends the UIMA framework for the mapping of expressions and annotations and enables the rapid creation, editing, and debugging of extraction scripts while REGEX can be also included.
An Eclipse plugin called the UIMA Ruta Workbench is included that validates the script commands in design time, enables the import of files for testing, and can also in the same environment visualize the extraction results of multiple rules and significantly help reduce the time for writing and validating the rules.
The feature of immediate visualization helps to solve the problem of meaningfulness of the annotations through visual cues results so that users can understand the function of the coding components required to improve and optimize existing rules and to add efficiently new rules to an unseen set of documents in which the current extraction rule will fail.21,24
In AURL, the composition and implementation parts are processed together since the included Java Eclipse workbench allows the parallel editing, correction, and visualization of the results from multiple rules.
Methods
This section outlines script coding factors for writing readable and maintainable manual RB pipelines. The writing of readable code offers a level of abstraction and provides an understanding of the intended purpose of the extraction script. For our study, we measure how readable and non-readable rules have an impact on importing into another language and especially on maintaining RB IE pipelines.
We program two separate pipelines, in REGEX with spaCY and AURL. Each rule is programmed then once in out-of-the-box and also in a readable style. We measure the total time that is required to update faulty rules in both pipelines and each style. Record of the time is through a handheld start-stop timer in which we log the data on an excel table.
Computation performance, portability, and accuracy extraction are outside the scope of this paper.21,25
Code readability
Code readability as a software quality metric has an immediate effect that can positively or negatively influence maintainability and reusability. 26
Reusability requires software design and testing principles 27 and is also an achievement concept that can be accomplished through readable code. 26
Code readability can not be easily quantified and measured by a deterministic function, nonetheless, maintainability can be measured by summarizing either separately or simultaneously the duration for editing, modifying, testing, and validating the results of the extraction rules. 28
When writing RB extraction scripts, we relate readability with the understanding of the extraction logic by reading without the presence of a knowledge engineer or developer. As script extraction logic, we refer to the steps for parsing and allocating portions of a search string from a given sequence of symbols.
Simple REGEX and AURL script for an easy extraction rule.

The extract logic for both scripts in Table 1 is readable: in the search text “13. pT1b, pNx, L0, V0, Pn0, R0; G2” allocate portions that qualify the conditions: a string that starts with an uppercase “T” character and following through a number.
Standard rules of thumb for writing readable programming code are decomposition and redundancy 29 and declaring rules with intention-revealing names. 28
In the context of writing extraction scripts, decomposition is the ability to split complicated rules into more manageable and readable sub-rules. Decomposition relates also to redundancy and helps avoid the repetition of rules by removing duplicate ones. Redundant rules can be reusable and through proper definition, they can be shared and included between various extraction pipelines.
Also according to the survey of Tashtoush et al., 30 intention-revealing names along with the spacing of programming commands have an important impact factor on code readability.
Code maintainability
Akour et al.
31
comment that “
Maintainability is not a design pattern but a software achievement concept and belongs to one of the most important software quality factors in general. 32
Especially for RB systems, in which many rules are to be written, tested, and corrected, if the concept of readability is not considered as a guideline, it can negatively influence the correction or the addition of new rules in unseen data.
For RB coding, maintainability is affected also by the available tools and methods in which a rule programmer can easily edit, control, evaluate, and update extraction scripts.
An RB script can only be maintainable if it is readable through coding design principles and the application of methods/tools that allow the editing, evaluation of the test rules, and visualization of the results for correction.
Research dataset
10 examples selected in the specifications document between the data curator and domain expert for building the rules.
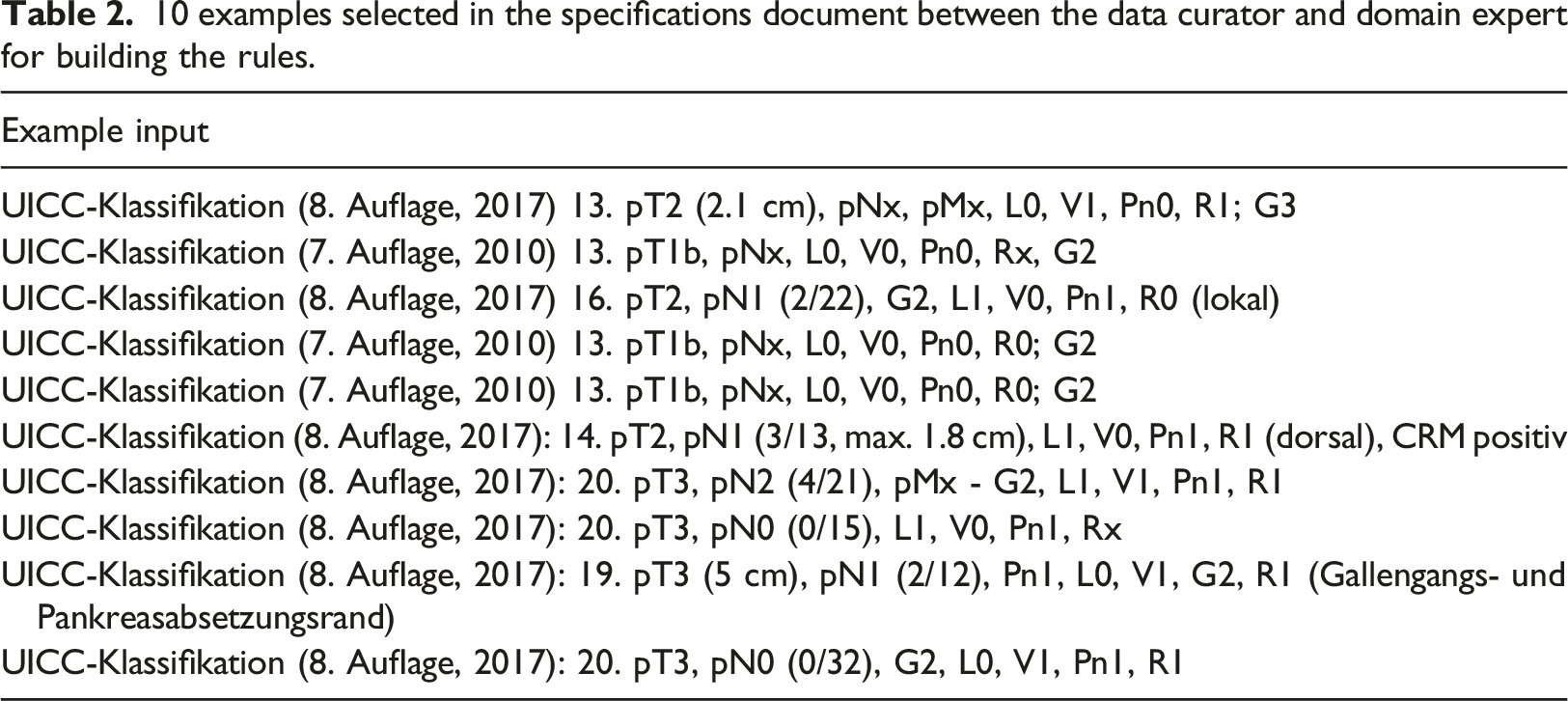

A total of 113 semi-structured cases containing TNM strings from the Oncology department of XXX1 were available for optimizing the rules in the German language.
Example of the table of the tags for the manual assignment of the documents and for writing the export rules. The third column is to determine the various extraction strings for each term. The as underlined marked values represent the required string to be extracted.
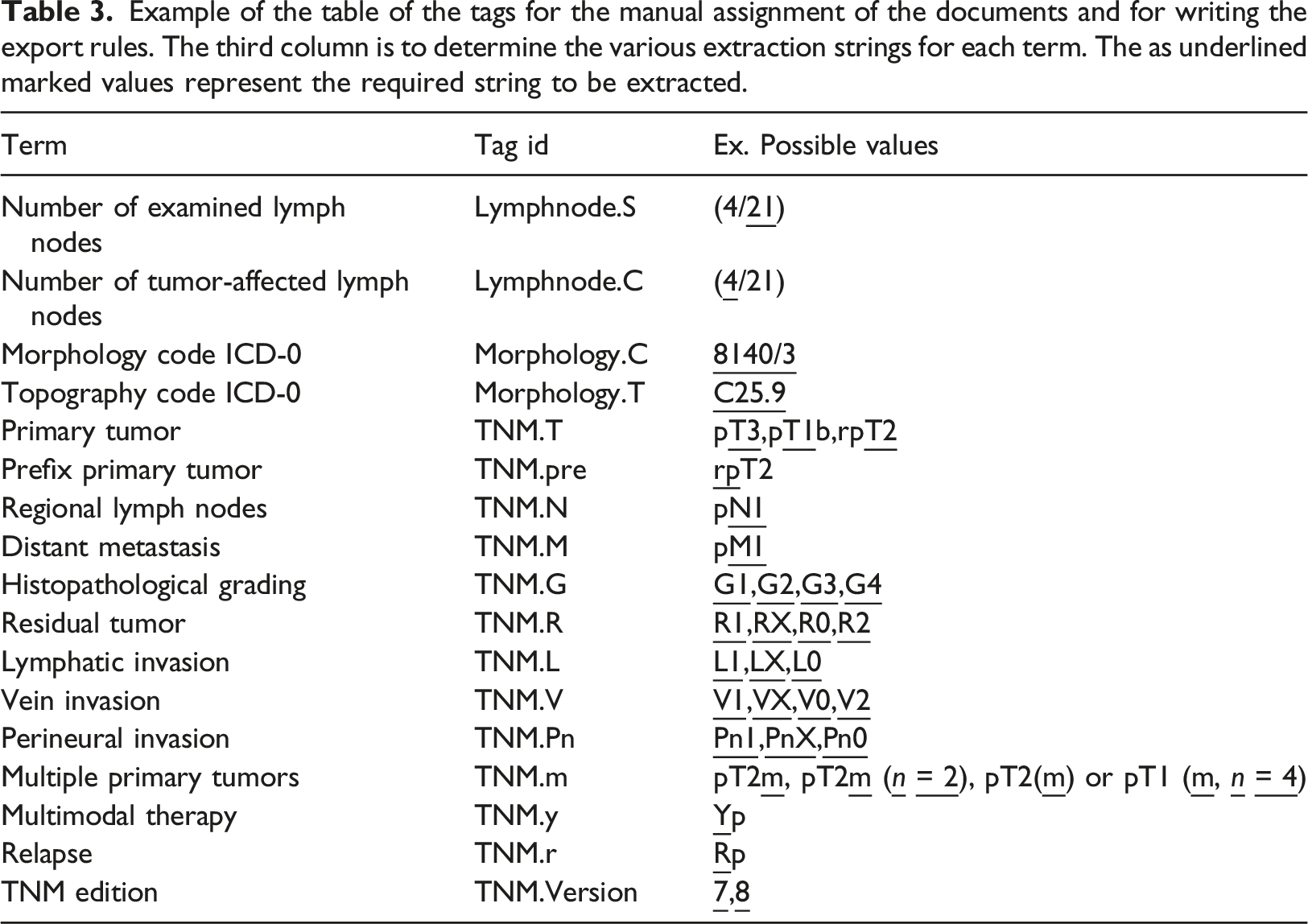
Table containing example texts in the German language from semi-structured and unstructured documents and for writing the export rules for the TNM.
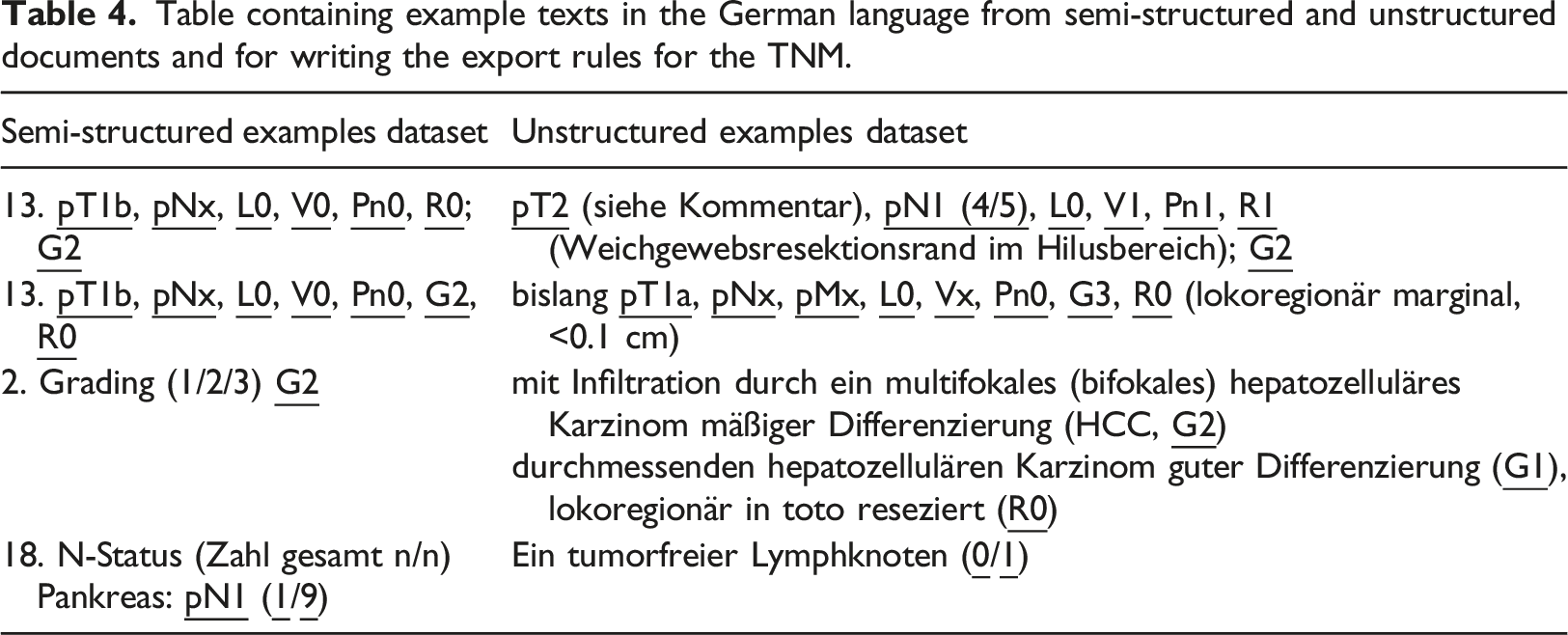

Through the unstructured examples, we were able to exemplarily study the difference in factors of readability and maintainability when writing the extraction scripts.
Results
Two NLP developers implemented the pipelines in AURL and REGEX. The experience level of the AURL developer was intermediate, while the level of the REGEX developer was advanced.
In the programming phase of the rules, the visualization of the results that the AURL workbench provided offered a valuable feature that helped us to immediately identify missed or wrong annotated parameters, the total line of commands for the 20 extraction rules of the source code was 73.
The multi-extraction of the TNM parameters including the test/maintenance in AURL was programmed in 8 h.
With REGEX and spaCY, the programming and testing of the scripts were programmed in 16 h. The total programming lines for 20 rules of the source code were 205.
Comparison of correcting out-of-the-box programmed rules with readable ones.

The examples following were chosen from the development phase of the NLP pipeline to discuss the efficiency of both methods from a developer and non-developer perspective and to serve as a paradigm for a better practical understanding of the factor’s readability and maintainability. A rule in REGEX which can be seen in the following coding example 1, was directly implemented with AURL within 10 min. After decomposing the rule, and declaring intention-revealing names, the rule was reprogrammed in AURL in 5 min.
Examples in readability
The next code in example 1 demonstrates the extraction of the primary tumor string from an unstructured pathological medical report through a REGEX script.
For this example, the density, non-intuitive syntax, and crypticness of the REGEX are demonstrated. Non-REGEX experienced developers require a cheat sheet to understand the intent parallel to the extra process of inputting the text and testing with a REGEX parser. The expression is also non-redundant. Decomposition and redundancy are only possible in the programming language, like in the following pseudo-code example in 2.
In the AURL in the following coding example 3, a demonstration of a script for extracting the direct extent of the primary tumor from a TNM classification is coded.
Furthermore, the script can be decomposed, like in code example 4.
The previous extraction rule is now decomposed into two rules: the prefix of the primary tumor that is declared as a variable and is now redundant and the primary tumor. Because of the decomposition and redundancy, the long rule is now in smaller parts and the extraction logic can be better read and understood.
The extraction logic is as follows: allocate portions of text that contain the string of prefixes with a “
The decomposition of the extraction pattern, through the declaration of rules, actions, and conditions, in AURL, offers a more intuitive declaration that helps to eliminate the gap of syntax crypticness and understand the intent of the rule.
Long rules are decomposed into shorter ones, and declaring intention-revealing names can simplify the process of reading and understanding the extraction logic. In reusability, repeated rules or complicated ones that contain many symbols can be included in one file as a library or class object.
Examples in maintainability
AURL offers a workbench in which multiple rules can be written and in one batch tested directly with test texts while the results of the extracted rules are visualized and each rule can be selected and the annotated text is marked visually.
In three cases of the unstructured data, the histopathological grading was inputted as “G2-3,” this input of the parameter was not specified on the data modeling document. The batch visualization of the results provided helpful to identify the missed annotation and easily define and change the rule.
Because of the ambiguous tokenization of TNM strings, when implementing the multiple variations found after testing the pipeline that refers to the same extraction rule with spaCy and REGEX in our pipeline, the adaption proved not optimal.
In five cases of unstructured data, the value of the Primary tumor was entered with space in between.
The maintenance of the previous extraction rule required the writing of an extra on spaCy’s rule matching parser of an empty character following after a “T”. In code example 5, this issue can be viewed, in which an extra command was added.
The first matching command searches for the string that does contain a space in between, while the second is for numbers following the T character.
In programming with REGEX, each rule was tested separately and for each document, applied rules cannot be visualized in a batch. This factor can significantly negatively influence the general aspect of maintainability.
Discussion and Conclusion
The structured approach of composing and decomposing the rules together with the visualization of the extraction results can help to provide a clear intent and understanding of the written syntax. For the programming of new rules or editing, the time factor can be reduced.
Readable composing of the commands allows the exploitation of domain knowledge on a programming and readable level that help developers immediately identify portions of the commands and the intention of the extraction code while offering a level of interpretability.
In this research study, while seeking a reliable method for programming a readable and maintainable RB IE pipeline, the demonstration of the application of the AURL closed the main gap when programming with REGEX which is the general problem with the lack of abstraction. 33
However since that REGEX is used in various programming languages for IE pipelines, a readable code strategy can help to achieve a level of abstraction and also make it maintainable.
The combination of readable and structured writing of extraction strings parallel to an immediate visualization of multiple rule results can accomplish a maintainable and portable pipeline and should be considered independent of the development platform in every IE framework.
When developing rules, a challenging aspect is the various ways to tokenize an input expression due to the morphological variability in real-world clinical text. For instance, the expression “cT2
Such exceptions when decomposed in smaller tokenizations, can be therefore easily changed, without having to debug and update a more complex rule. This programming approach can be also applied to other RB IE applications in which items from other domains need to be extracted. In the case that medical departments allow the practitioners to freely write without considering the standards of annotating a coding system for example, they can ente spaces, commas, or other characters, it would be of advance if such rules are from the beginning in such a way designed that can be easily changed.
In the case of reconfiguring IE pipelines after some time to add new rules on unseen data or error correction, the constructive composing of rules through the principles of readability can provide the means to easily reprogram the pipeline and save development time for production environments, and reduce the overall cost.
Coding Examples
T“ NUM,
Footnotes
Acknowledgements
The project within this work was done is funded by the German Ministry of Education and Research.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: This publication is funded by the Deutsche Forschungsgemeinschaft (DFG) as part of the “Open Access Publikationskosten” program.
Availability
The source codes both in AURL and spaCy with REGEX associated with our research for extracting TNM staging strings in medical german texts are freely available at https://gitlab.plri.de/NektariosLadas/tnm-extraction-with-apache-uima-ruta-language/-/blob/master/uima_ruta/tnmextract_nodes.ruta and ![]() .
.
Approval statement
The complete content and data on this study is approved from Hannover Medical School in Hannover Germany for publishing. Approval code: 8411_BO_K_2019.
