Abstract
Patients with rare diseases commonly suffer from severe symptoms as well as chronic and sometimes life-threatening effects. Not only the rarity of the diseases but also the poor documentation of rare diseases often leads to an immense delay in diagnosis. One of the main problems here is the inadequate coding with common classifications such as the International Statistical Classification of Diseases and Related Health Problems. Instead, the ORPHAcode enables precise naming of the diseases. So far, just few approaches report in detail how the technical implementation of the ORPHAcode is done in clinical practice and for research. We present a concept and implementation of storing and mapping of ORPHAcodes. The Transition Database for Rare Diseases contains all the information of the Orphanet catalog and serves as the basis for documentation in the clinical information system as well as for monitoring Key Performance Indicators for rare diseases at the hospital. The five-step process (especially using open source tools and the DataVault 2.0 logic) for set-up the Transition Database allows the approach to be adapted to local conditions as well as to be extended for additional terminologies and ontologies.
Introduction
Rare diseases are defined with an incidence of 1 in 2,000 individuals in the European Union and 1 in 2,500 in the USA. 1 Because there are about 6,000 to 8,000 different rare diseases, probably 263 to 446 million people worldwide are affected. 2 Rare diseases are diverse, highly variable and thus a “global public health burden” 3 : They all have in common that they have chronic effects on the quality of life of the patients and are sometimes life-threatening. 4 Due to a lack of expertise and geographically dispersed experts, false diagnoses or long waiting periods – on average of about 7 years until the disease is correctly diagnosed – are common.3,5 50 % of patients remain undiagnosed. 6
In addition to the difficult diagnostics and the diagnostic delay, the documentation of the diagnoses is a huge hurdle. Rare diseases are currently only insufficiently described by classifications, such as by the International Statistical Classification of Diseases and Related Health Problems 10th Version (ICD-10). Aymé et al
The Orphanet approach (https://www.orpha.net) represents an alternative to coding with ICD-10. 11 It is a multilingual portal containing information about rare diseases and orphan drugs. An important component is the ORPHAcode, which is a unique and stable identifier for a single rare disease. In this way, significantly more diseases can be provided with a unique code than with ICD-10. The resulting code system enables mapping to other terminologies, such as ICD-10-WHO, Systematized Nomenclature of Medicine Clinical Terms (SNOMED CT), HUGO Gene Nomenclature (HGNC) or Online Mendelian Inheritance in Man (OMIM). There are also references to the Human Phenotype Ontology (HPO) to describe symptoms and phenotypes. 12 Using specific coding system for rare diseases significantly increases the documentation quality and representation of rare diseases.11,13 They thus form the basis for both quality-assured secondary data use and specific additional documentation, for example through registers.
In Germany, diagnoses have so far been documented exclusively with ICD-10 of the German Modification (ICD-10-GM) and coding with ORPHAcodes is done on a voluntary basis. An obligation to document rare diseases with ORPHAcode and the German Alpha-ID-SE is mandatory since 2023.14,15 However, few approaches have been published on how to integrate the specific coding systems into clinical practice and IT infrastructure.
The University Center for Rare Diseases at the University Hospital Carl Gustav Carus in Dresden, developed a parametrized form within the clinical information system to enable the obligatory coding of rare diseases using ORPHAcodes for all inpatients. 16 As part of the project “Collaboration on Rare Diseases” of the German Medical Informatics Initiative (CORD-MI), the underlying data basis of the form was refactored. This resulted in the Transition Database for Rare Diseases. The aim of this work is to review current approaches for documentation of diagnoses for rare diseases and to explain how the information in the Orphanet catalog can be used to support clinical documentation of rare diseases by using a database with semantic mappings. Necessary steps are described to store and version ORPHAcodes and their mappings as well as to structure them according to local specifics. This results in lessons learned that allow the adaptation of the approach to other clinical IT infrastructures.
State of the art
For an overview of methods, which are already in use, for rare disease documentation support, a rapid scoping review was conducted by two authors. The methodology based on “PRISMA Extension for Scoping Reviews”,
17
which consists of the following steps: (1) Identifying the research questions (2) Identifying relevant studies (search strategy) (3) Study selection (4) Charting the data (5) Collating, summarizing and reporting the results.
Research questions
The primary research question was: What approaches are there to support documentation of rare diseases and how is the support provided? This led to secondary research questions to detail the approaches: • Which terminologies or ontologies are used for the documentation? • In which stage is the approach (e.g., proof-of-concept or routine)? • Which systems or applications are supported (e.g., in the clinical information system, in the documentation system, as an external application)?
Search strategy
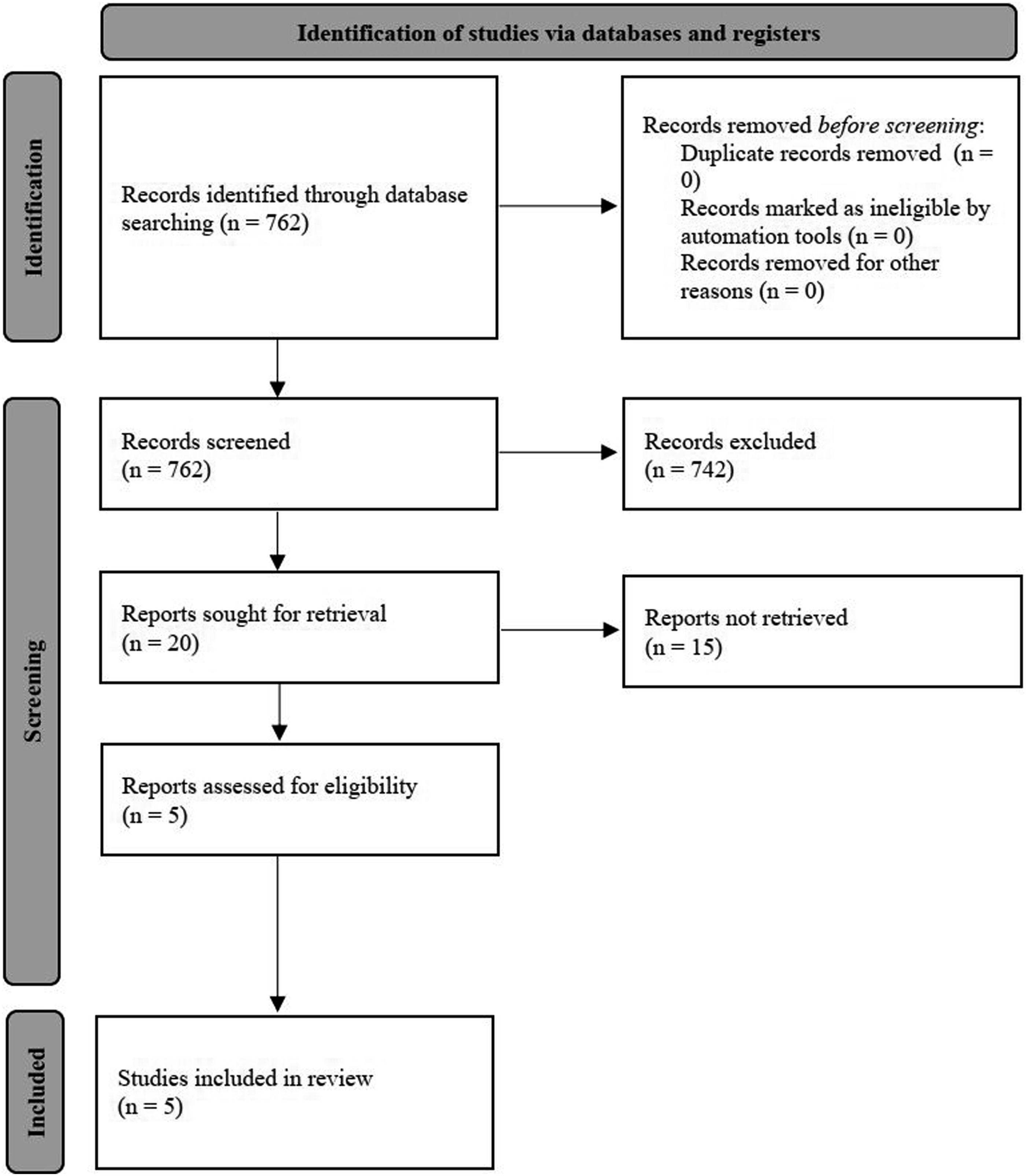
A search (on 10th on January 2023) in PubMed of the last 50 years for the search term (Documentation [Title/Abstract] OR Code* [Title/Abstract] OR Coding* [Title/Abstract]) AND (Rare Disease* [Title/Abstract] OR Orphan Disease* [Title/Abstract]) resulted in 762 papers (see Figure 1). Two authors screened these papers and discussed in case of disagreement. PRISMA flowchart to identify approaches for the state of the art (model from
18
).
Restriction to rare diseases is important because, on the one hand, they require specific terminologies and, on the other hand, multiple diagnosis codes are also needed, e.g. for billing purposes as well as for more precise designation, for the exact indication of suspected and excluded diagnoses.
Study selection
Inclusion and exclusion criteria of the rapid scoping review.
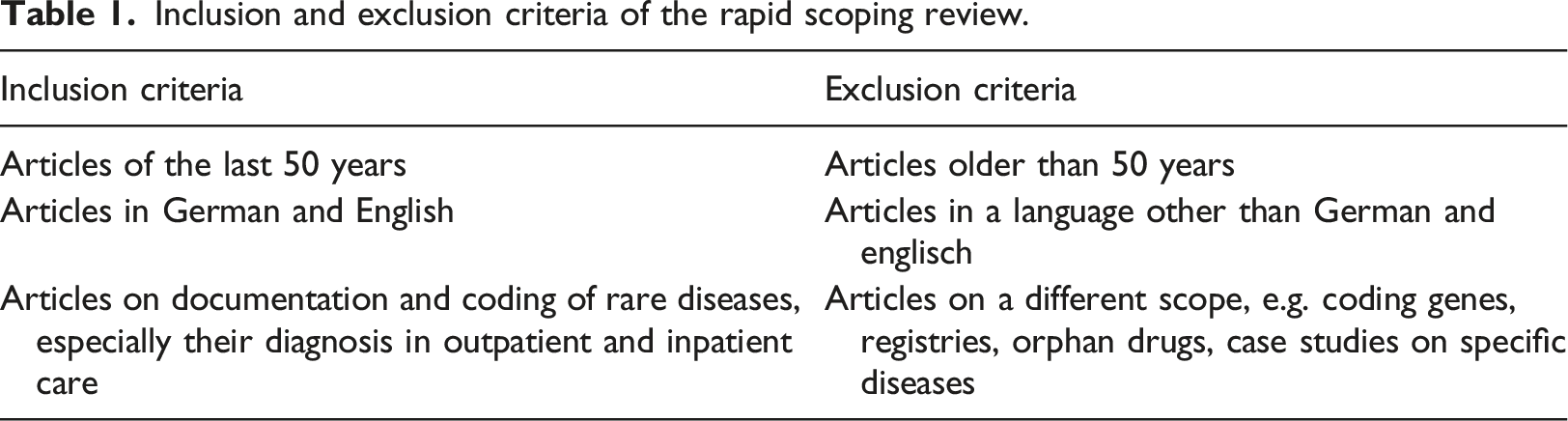

Of the 762 articles 20 papers were included through title-abstract-screening. Five studies were included and 15 studies were excluded by the full-text screening in accordance with the inclusion and exclusion criteria (see Table 1). Only three papers discuss explicit approaches to documentation support; two additional papers discuss coding support by extra applications.
A large number of papers found by the search string were excluded because they dealt with genes encoding a specific phenotype or problems in public health and epidemiology but did not show any relation to rare disease coding or documentation.
Charting the data
Extracted information according to research questions.

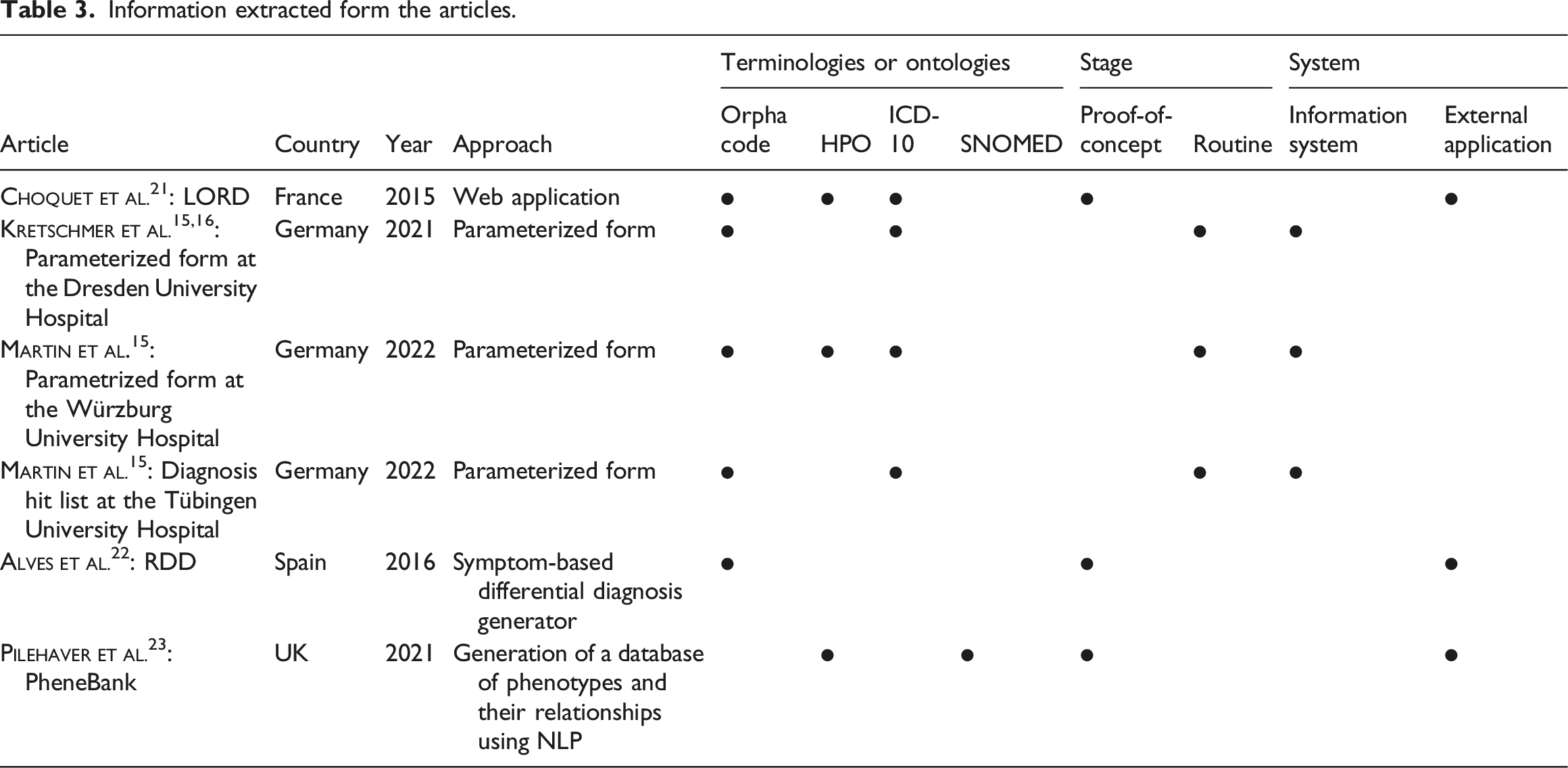
Information extracted form the articles.
Reporting the results
Three papers specifically describe how rare diseases can be documented.
The two additional papers were about IT systems for correct coding as part of diagnostic support.
Thus, six separate approaches to documenting rare disease diagnoses were identified: 1. Choquet et al.
21
: LORD, 2. Kretschmer et al.15,16: Parameterized form at the Dresden University Hospital 3. Martin et al 4. Martin et al 5. Alves et al.
22
: RDD 6. Pilehaver et al.
23
: PheneBank
All approaches have in common that they use international terminologies and ontologies for coding diagnoses (see Table 3). The majority (n = 5) use ORPHAcodes, half (n = 3) also use HPO, and the approaches developed and used in Germany additionally use German terminologies, such as ICD-10-GM (n = 3) or Alpha-ID-SE (n = 1). However, SNOMED (n = 1) and OMIM (n = 1) are rarely used.
The identified approaches are already in routine use (n = 4) or are being evaluated as proof-of-concepts (n = 2) (see Table 3). The approaches used in Germany are integrated into the clinical information system or the documentation system (n = 3). The other approaches are external applications (n = 3) (see Table 3).
Despite the commonalities, a uniform approach to documenting rare disease diagnoses is not evident.
Concept
In a cooperation between the University Center for Rare Diseases and the IT division of the University Hospital Carl Gustav Carus in Dresden, a form for the clinical documentation of rare diseases was designed. In the following, the functionality of the form is explained briefly, but the focus is on the Transition Database for Rare Diseases used for the underlying data catalog as well as their interaction.
Parametrized form for clinical documentation of rare diseases
The parametrized form allows coding of rare diseases using ORPHAcode in the clinical information system ORBIS of the Dedalus Healthcare Group. The form allows searching by ORPHAcode, disease name, ICD-10 code and genes based on the Orphanet catalog. The search result can be selected and stored in the electronic patient record. Since the majority of rare diseases are mainly chronic, documentation is per patient and not per medical case. This is unusual for the German healthcare system, as documentation is strongly based on case-related billing of medical services.
On the one hand, the form is used for more accurate medical coding of rare diseases and is mandatory for inpatient care at the hospital. On the other hand, the results are used for statistical analysis to create greater awareness of rare diseases and their incidence. For this purpose, statistical parameters are displayed within a dashboard. 25
The clinical information system expects one comma-separated values (CSV) file per searchable field (i.e. ORPHAcode, name, ICD-10 code, genes). Previously, the CSV files were updated manually by an Excel form every half year. The refactoring presented here enables an automated update every month.
Transition database for rare diseases
In 2022, the form was refactored. The aim was to update the underlying data catalog on a monthly basis in order to always use the most recent status of the Orphanet catalog. New rare diseases are constantly being researched and included in the Orphanet catalog. The update of the source data is determined along a strict decision path by Orphanet Nomenclature Manager and a medical and scientific validating authority (see Procedural document: Orphanet nomenclature and classification of rare diseases). This process includes both the collection of information through literature and expert knowledge, as well as discussion in a selected committee. For the provision of the updated information, there is a release cycle in which the files required for the database are updated monthly. This high variability requires constant updates. The Transition Database combines all monthly data as well as changes of the Orphanet catalog since 2020.
The use of more flexible approaches, such as terminology servers or API is unsuitable for the present case, as the clinical information system requires only CSV files. Thus, a more static approach was deliberately chosen: The Transition Database is the basis for the rare disease parametrized form for clinical documentation of rare diseases.
The automated workflow of how the database is loaded and used as the basis for the data catalog of the parametrized form is shown in Figure 2. The extent to which the individual steps have been implemented is explained in the following chapter. Workflow of updating the transition database for rare diseases representing the DAGs from Airflow (without step (E) “Manual input of the CSV files into the clinical information system”; dv = DataVault, h = hub, s = satellite).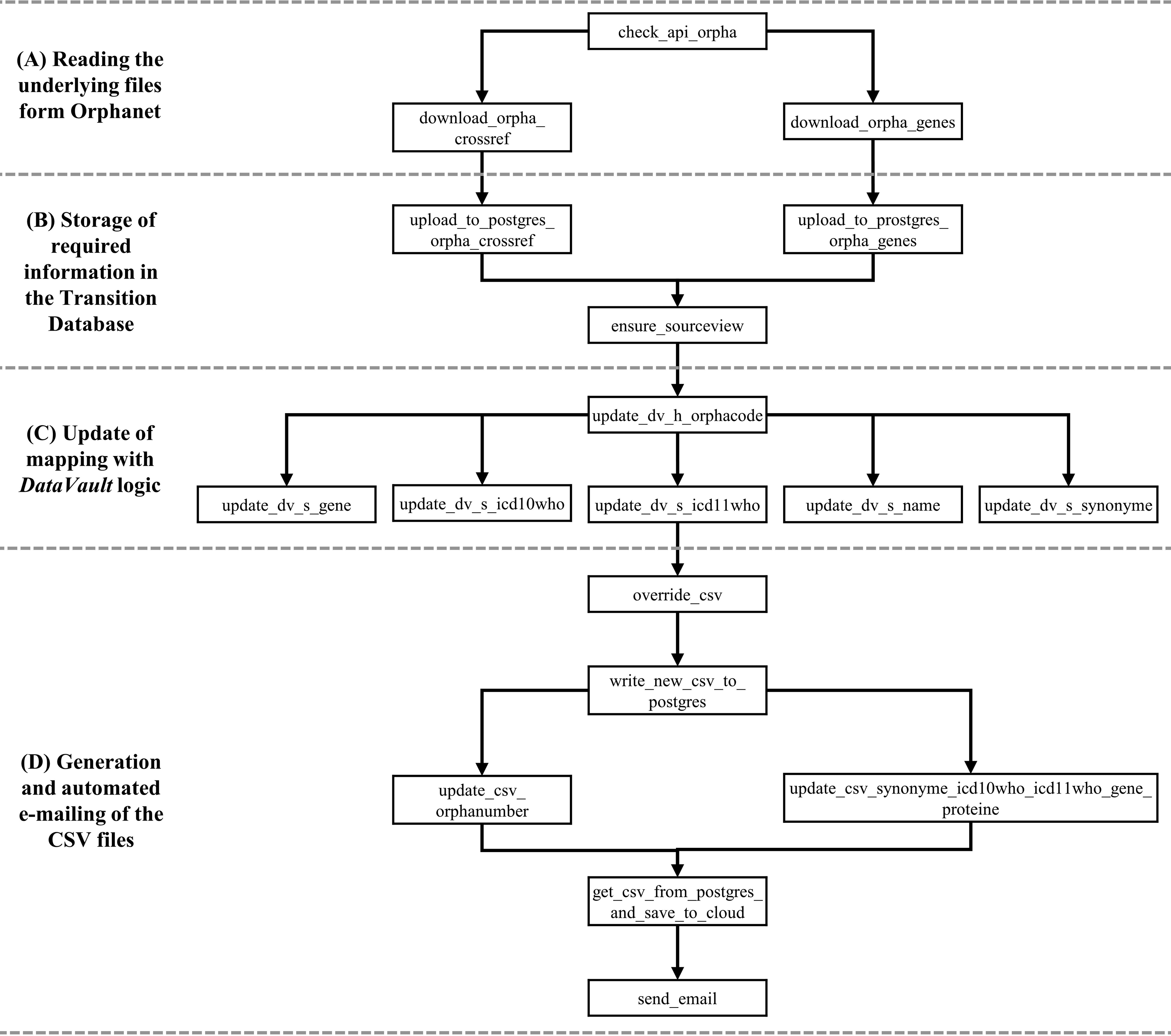
Implementation
The following section focuses solely on the implementation of the Transition Database for Rare Diseases following the steps of the automated workflow (see Figure 2.). To manage the workflow orchestration we used Apache Airflow (https://github.com/apache/airflow). It is an open source tool for workflow management. Directed acyclic graphs (DAGs) with tasks and dependencies can be programmed via Python and are scheduled by Apache Airflow. 26
Reading the underlying files form Orphanet
Orphadata (https://www.orphadata.com) provides aggregated information from Orphanet in the form of predefined datasets (referred to as the Orphanet catalog).
Here there are different packages, so the used German Orphanet Nomenclature Pack contains different files. The Orphanet Nomenclature File is the main source for the concept of the Transition Database: Regular updated moderated XML files containing every valid ORPHAcode, name of the disease, status (active or inactive), synonyms, disorder type (disease or syndrome or histopathological subtype), classification level (disease or group) can be downloaded from their Github Repository (https://github.com/Orphanet/Orphadata_aggregated).
To track changes the Github Repository is checked daily. A SHA256 hash is created for the data and compared with the hash value of the previous data exports. If the hash differs, the new XML files are automatically downloaded, read, transformed, written to the database and saved as a backup.
Storage of required information in the transition database
The Transition Database contains all the information of the Orphanet catalog (i.e. ORPHAcode, name, status, synonyms, ICD-10-WHO code and since recently ICD-11-WHO codes with associated mapping relationships, codes from OMIM, UMLS, MeSh, GARD, Meddra, link to Orphanet website) as well as an ID, the date of the underlying file and a generated unique job ID for the executed update (see Figure 3). Sample excerpt from the database (tool: DbVisualizer; mapping relations are hidden).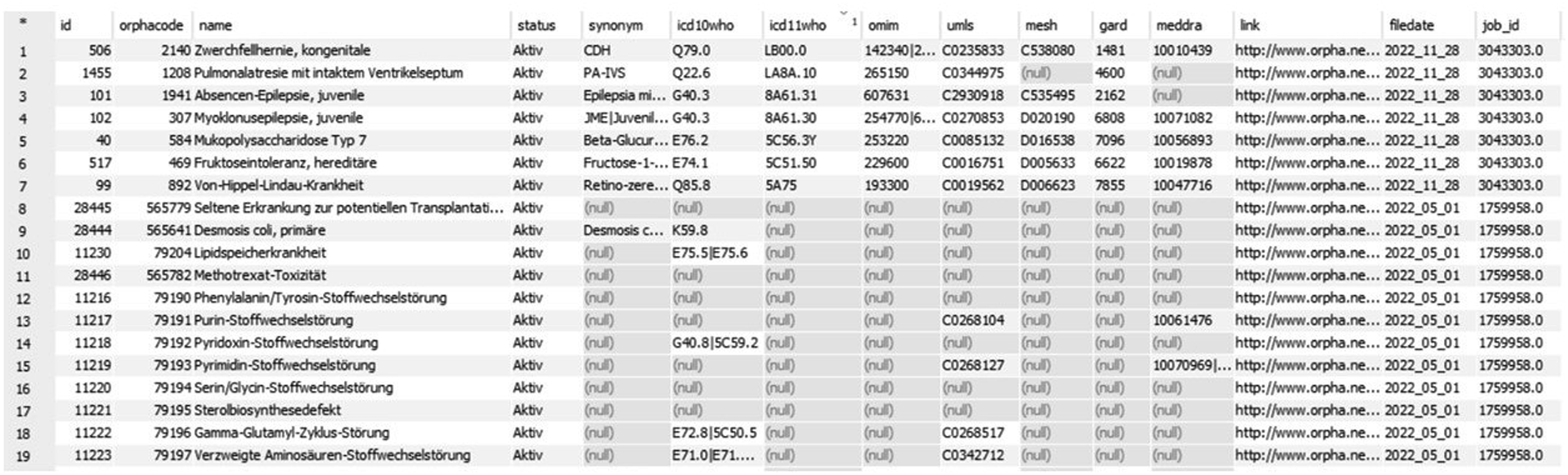
When a new version of the data is available, the associated XML file is parsed into a flat structure for later processing in a relational database. This function requires multiple transformation and pivoting steps in order to receive the target structure containing the unique ID, ORPHAcode, link to the resource, name, synonyms as well as the status and the external references. Using Apache Airflow this functionalities are orchestrated to stage the data into the database.
Every downloaded XML file contains the complete and valid ORPHAcode catalog. Each ORPHAcode and its information is written line by line to the Transition Database and then displayed in whole in its most recent version.
U pdate of mappings with DataVault 2.0 logic
The mappings are updated and versioned using the DataVault 2.0 logic. DataVault is a logic of data warehousing, which is based on a two-layer architecture (stage and data warehouse) with the goal to have an exact copy of the original data.
27
The model of DataVault consists of a single hub table and multiple satellite tables, which are linked by unique business keys. For the Transition Database we created a hub containing the unique keys and five satellites containing all the describing attributes (see Figure 4). Database model of DataVault 2.0 logic with hub (h) and satellites (s).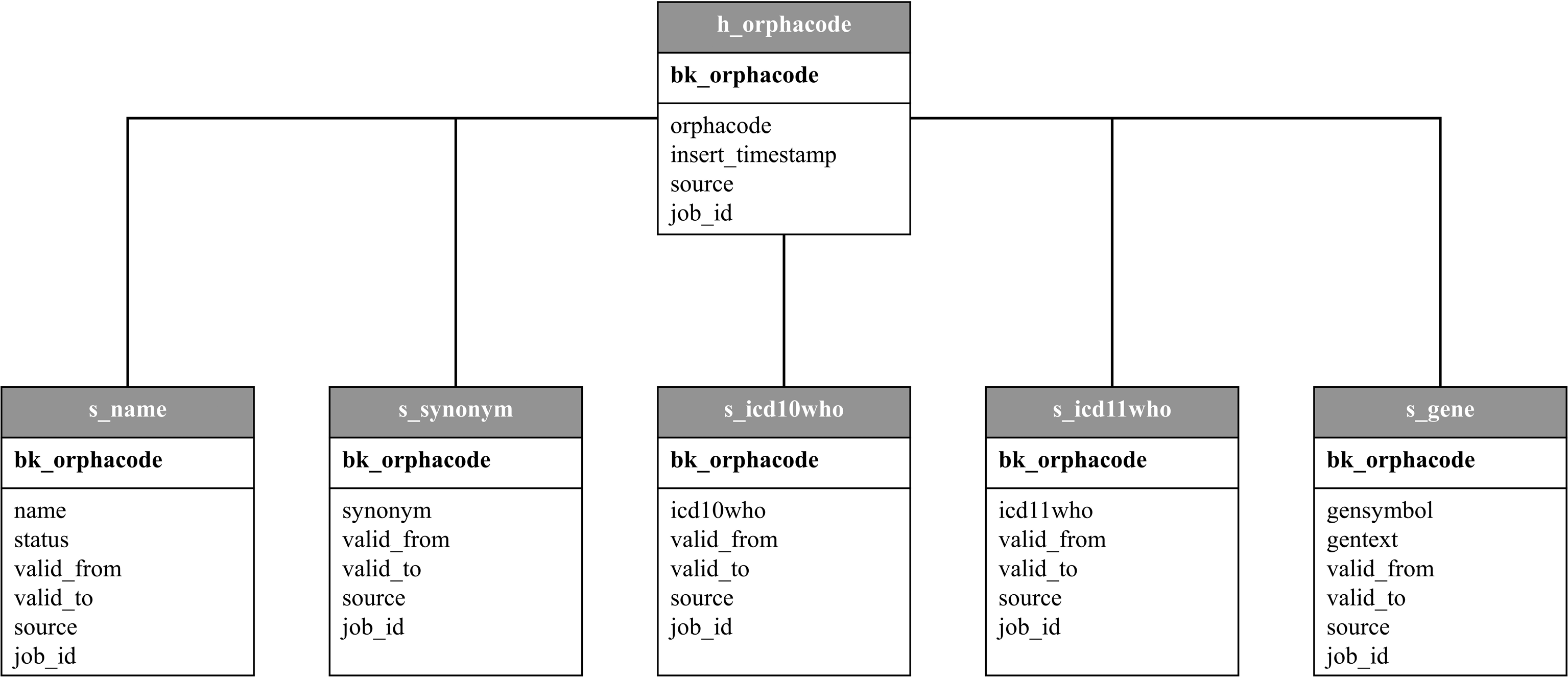
The hub contains all ORPHAcodes ever written to the Transition Database, a MD5 Hash as business key for each one and valid_from and valid_to dates. This is the main table for the DataVault. The five satellites contain available information about the name, status, synonyms, ICD-10 (and since recently ICD-11 codes) and associated genes for each ORPHAcode as well as valid_from and valid_to dates for each satellite. The entries are connected by the business key to ensure fast transactions. The hub and each satellite can be displayed as a most recent valid view or a historicized view with all recorded changes since DataVault creation. The DataVault is created manually with a self written Airflow Operator and uses Structured Query Language (SQL) queries. This allows to generate and customize a DataVault for different use cases independently form commercial software. In our case, the most recent views of each satellite are used for the creation and update of five CSV files (ORPHAcode and name, synonym, ICD-10, genes, names of genes).
Generation and automated e-mailing of the CSV files
The required CSV files correspond to the satellites and contain additional information that the clinical information system expects.
Each CSV file is written to a table in the Transition Database and from there updated with new information and again saved to CSV files. A task in the Airflow DAG compares the CSV table to the most recent view of the satellite associated with it. No longer valid entries get the valid_to date from today and new entries are created with valid_from today in the CSV Table. In the next step, all these CSV tables are written to separate CSV files, compressed into a ZIP archive and saved to our local cloud storage. After that, the last task sends an email with a link to the archive to the recipient list responsible for updating the clinical information system.
Manual input of the CSV files into the clinical information system
The sent CSV files are then manually inserted into the information system by an employee of the IT division. After that, the new information is available to all users.
Lessons learned and conclusion
The correct and clear documentation of rare diseases diagnoses is essential for patient care and research. However, there are only few published approaches like the scoping review shows. This is in line with the statement of Choquet et al. who described that a processing is necessary because, although the data is publicly available, it has a high heterogeneity and update frequency, and thus it can be poorly used and integrated into existing systems. 21 Therefore, we propose the Transition Database for Rare Diseases containing information from the Orphanet catalog as well as mappings of the ORPHAcode. We explained this approach using the example of the parameterized form at the at the University Hospital Carl Gustav Carus in Dresden. It can also be adapted to local specifics due to the explanation of the single steps.
The lessons learned during development were diverse: (1) Automation for increased timeliness, (2) database as a solid basis, (3) DataVault 2.0 logic for flexible updates, (4) use of open source tools for easy embedding and (5) transferability and expandability. (1) Manual updating of the database for the parameterized form is very complex, which is why automating the process is important. Apache Airflow is used to automate and orchestrate the workflows for downloading, saving, updating, transforming and sending the data. The automation increases the timeliness of the data and creates an overview of the frequency and content of the updates of both the data source (Orphadata) and the prepared data (Transition Database for Rare Diseases). (2) Although terminology servers are often used to store and update terminologies and classifications, this approach is not suitable for local requirements. Since the clinical information system requires CSV files as input, the authors decided to store the data in a database and transform it according to the required data structure. This also corresponds to the described state of the art.
24
(3) The rapid frequency of change in the source data set, which contains highly linked information, requires a dynamic method to update the data. Therefore, the DataVault 2.0 logic was used. The hub-satellite relationship corresponds to the logic of the required CSV files, making programming easier. In addition, this structure allows for quick expansion. Thus, a new satellite could be programmed for ICD-11 codes already present in the source without changing the original program code. Therefore, we assume that a transfer to other terminologies and classifications is easily and quickly possible. So far, the focus is on the ORPHAcodes, which corresponds to the state of the art.15,21,22 (4) The use of open source tools enables independence from commercial tools. Thus, the database could be easily and quickly embedded into the existing IT infrastructure. Apache Airflow is already used for different dashboards
25
and the DataVault 2.0 logic is an essential part of the clinical data warehouse
28
(even if the commercial tool DataVaultBuilder is used). The use of other tools is conceivable; for example, the prototype of the Transition Database was created with Pentaho Data Integration (https://www.hitachivantara.com/en-us/products/dataops-software/data-integration-analytics/download-pentaho.html). (5) The concept can also be applied to other use cases, for example, it is successfully used in the clinical data warehouse for mappings to an international research data repository
29
and for mappings of medication data. In general, the step-by-step approach allows for adaption to local conditions (e.g. use of other tools, different structure of the output).
For the future, we plan to expand the database. In particular, it will be expanded to include terminologies and ontologies for genetic rare diseases, i.e. HPO and HGVS. In addition, the database will be used as a basis to store mappings to other rare disease documentation approaches: The focus here is on minimal data sets21,30 and registries31–33. By accurately documenting diagnoses of rare diseases, patients can be included in registries in a more precise and targeted manner. This in turn forms the basis for research on quality-assured data and enables access to improved care. Examples of the successful implementation of registries include the German registry for Cystic Fibrosis (https://www.muko.info/), LORIS MyeliNeuroGene rare disease database 34 , the United Kingdom Primary Immune Deficiency registry 36 as well as the GAIN Registry 37 . In addition, registry software tools can also use the described approach to update the underlying terminologies.
The Transition Database for Rare Diseases is routinely used as a basis for the documentation of rare diseases at the University Hospital Carl Gustav Carus in Dresden. The data obtained from this serves both patient care and the monitoring of key performance indicators for rare diseases within a dashboard. 25
The structured storage and continuous updating of terminologies and ontologies as well as their mappings supports the documentation of rare diseases in patient care and hence the research of these conditions. Accurate naming of diseases will increase the chance of personalized medicine for patients with rare diseases and can shorten the odyssey of diagnosis.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported in part by the German Federal Ministry of Education and Research within the project „Collaboration on Rare Diseases“ (01ZZ1911I) of the German Medical Informatics Initiative.
Open Access funding enabled and organized by Projekt DEAL. We gratefully acknowledge support from the SLUB / TU Dresden Open Access Publication Fund. The funders for the publication had no influence on the study design, data collection, analysis, decision to publish, or preparation of the manuscript.
